1. Introduction
More than 60% of new drugs with small molecules are derived from natural products, indicating that natural products are important sources for the discovery and development of new drugs [
1]. Marine actinomycetes are one of the richest producers of bioactive natural products [
2,
3,
4,
5]. For example, salinosporamide A from the marine
Salinospora strain CNB-392 entered clinical trials against multiple myeloma only three years after its discovery [
6], and abyssomicin C from marine
Verrucosispora strain AB 18-032 is the first natural product that inhibits
p-aminobenzoate biosynthesis, a pathway used only by microorganisms [
7].
Bagremycins are bacterial secondary metabolites and belong to phenol esters formed from
p-hydroxystyrene and
p-hydroxybenzoic acid. The first two bagremycins A and B, from the soil actinomycete
Streptomyces sp. Tü 4128, were reported in 2001 [
8]. Since then, no new bagremycins have been reported until our recent publication, which reported three new bagremycins C–E from a mangrove-derived actinomycete
Streptomyces sp. Q22 [
9]. Both bagremycins A and B showed antibacterial activity against
Arthrobacter aurescens DSM20166 and
Streptomyces viridochromogenes Tü 57 [
8]. Bagremycin C inhibited the proliferation of different glioma cell lines; induced apoptosis in human glioma U87-MG cells in a dose- and time-dependent manner; and arrested the U87-MG cell cycle at the G
0/G
1 phase [
9]. The biosynthesis of bagremycins was also investigated, and several genes including bagA, bagB, bagC, and bagI are involved in bagremycin biosynthesis [
10,
11,
12].
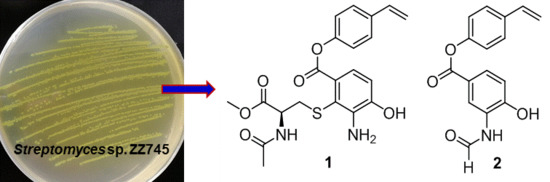
As part of our research for discovering novel bioactive compounds from marine-derived microorganisms [
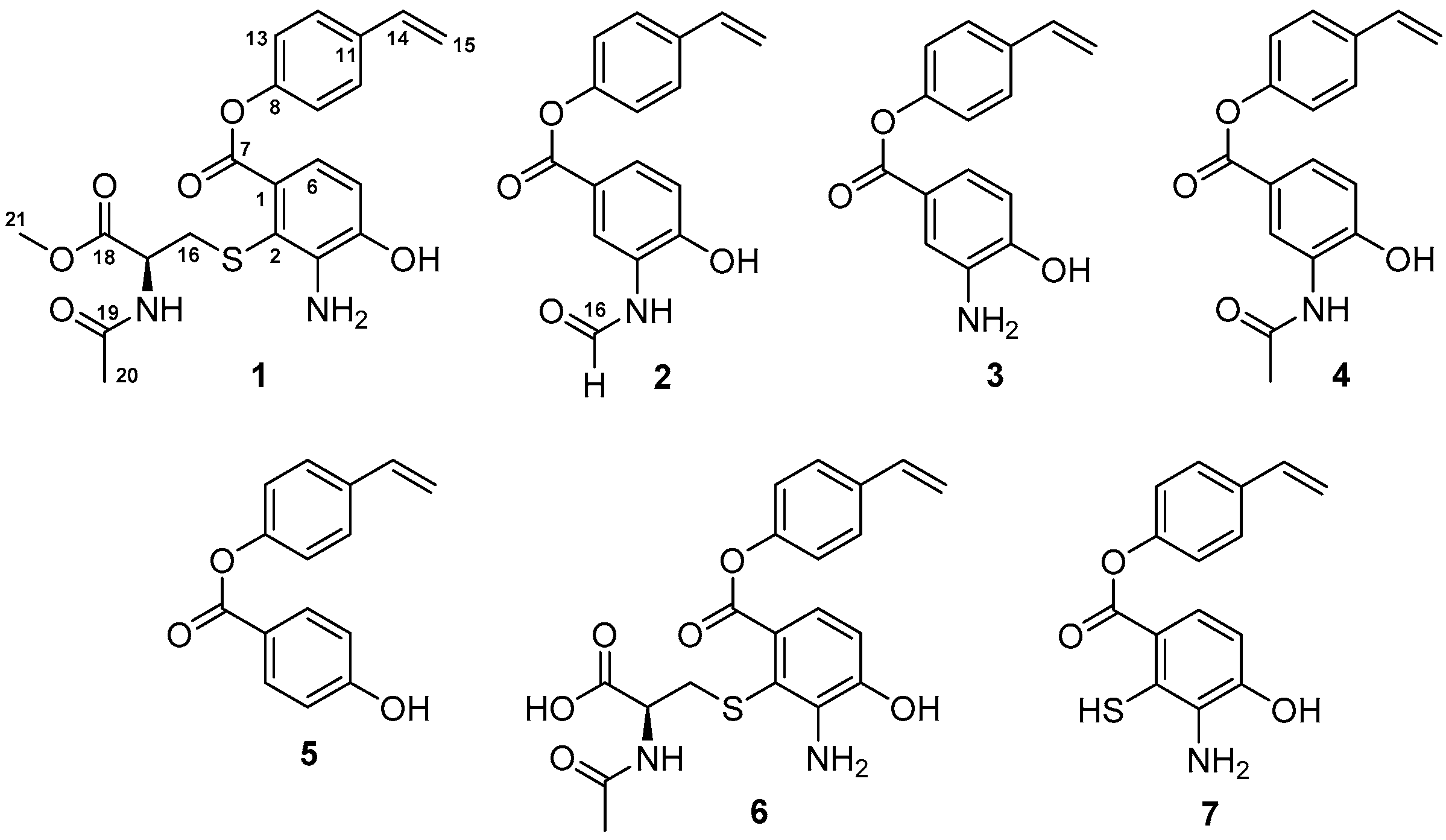
9], a marine strain ZZ745 was isolated from a coastal mud. A scale up culture of the strain ZZ745 in Gause’s liquid medium produced two new members of bagremycins, named bagremycins F (
1) and G (
2), together with the known bagremycins A (
3), B (
4), and E (
5) (
Figure 1). Herein, we reported the isolation and culture of the strain ZZ745, as well as the isolation, structural elucidation, and antibacterial activity of the isolated compounds.
2. Results and Discussion
The strain ZZ745 (
Supplementary materials, Figure S1) isolated from a sample of coastal mud was identified as
Streptomyces sp. ZZ745, according to the results from its 16S rDNA sequence analysis (
Supplementary materials, Table S1 and Figure S2). A crude extract prepared from a 50 L culture of the strain ZZ745 in Gause′s liquid medium was fractionated by repeated column chromatography, followed by purification with high performance liquid chromatography (HPLC) to give
1–
5. On the basis of the NMR spectral analysis and comparison of their NMR data with those reported from the literatures,
3–
5 were identified as bagremycins A (
3), B (
4), and E (
5) [
8,
9]. Their
13C and
1H NMR data are shown in the
Supplementary materials (Supplementary materials, Table S2).
Compound
1 was obtained as a colorless amorphous powder and gave ion peaks at
m/
z 431.1270 [M + H]
+ and 453.1089 [M + Na]
+ in the (+)-HRESIMS, corresponding to a molecular formula of C
21H
22N
2O
6S. The
13C NMR spectrum (
Supplementary materials, Figure S4) of
1 showed 21 carbon signals, fifteen of which were attributed to the scaffold of
1, resonating at
δC 126.5 (C-1), 113.1 (C-2), 140.4 (C-3), 147.7 (C-4), 113.2 (C-5), 119.1 (C-6), 165.7 (C-7), 150.5 (C-8), 122.0 (C-9, C-13), 127.1 (C-10, C-12), 134.7 (C-11), 135.8 (C-14), and 114.3 (C-15). The
1H NMR spectrum (
Supplementary materials, Figure S3) displayed the signal at
δH 6.78 (1H, d,
J = 8.2 Hz, H-5); 7.10 (1H, d,
J = 8.2 Hz, H-6); 7.21 (2H, d,
J = 8.6 Hz, H-9, H-13); 7.55 (2H, d,
J = 8.6 Hz, H-10, H-12); 6.74 (1H, dd,
J = 17.7, 10.9 Hz, H-14); 5.29 (1H, dd,
J = 10.9, 0.6 Hz, Ha-15); and 5.82 (1H, dd,
J = 17.7, 0.6 Hz, Hb-15) (
Table 1). The
1H and
13C NMR data for the scaffold of
1 were very similar to those of bagremycins C (
6) and D (
7) [
9], suggesting that the core structures of
1,
6, and
7 had the same substitution pattern. The remaining six carbons were assigned to an
N-acetylcysteine methyl ester moiety, comprising of the following
13C and
1H NMR signals:
δC 35.5 (
δH 2.98, dd,
J = 13.1, 8.4 Hz and 3.11, dd,
J = 13.1, 5.4 Hz, CH
2-16);
δC 52.2 (
δH 4.25, m, NH-CH-17);
δC 171.0 and 169.3 (C-18 and C-19);
δC 22.2 (
δH 1.81, s, CH
3-20);
δC 51.9 (
δH 3.54, s, CH
3-21) and (
δH 8.41, d,
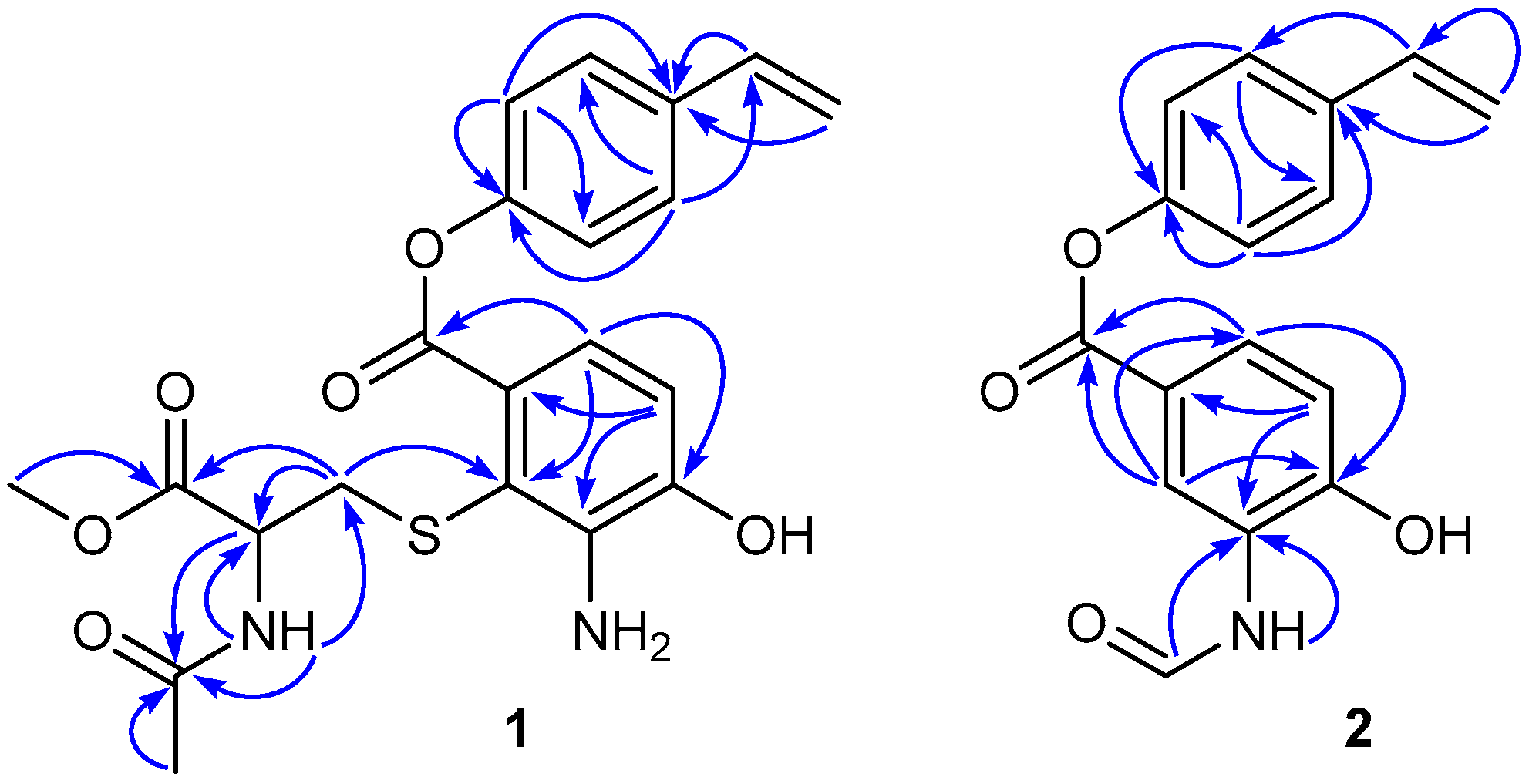
J = 7.5 Hz, NH). The significant HMBC correlations (
Figure 2) from H
2-16 to C-2 (
δ 113.1), placed the
N-acetylcysteine methyl ester on C-2. Since both
1 and
6 are not only levorotatory, but also have nearly identical structures, it was legitimate to assume that both compounds had the same absolute configuration at C-17 of the amino acid moiety. Since the absolute configuration of C-17 of
6 was determined as 17
S [
9], it was concluded that the configuration of C-17 of
1 was also 17
S. The structure of
1 was a new member of bagremycins, named bagremycin F. Its
13C and
1H NMR data were fully assigned by the HSQC and HMBC correlations (
Table 1 and
Figure 2).
Since bagremycin F (
1) was a methyl ester of bagremycin C (
6), it was possible that
1 could be an artifact formed by esterification of the carboxylic acid of
6 by MeOH, which was used as the solvent for extraction. To prove this possibility, an ethanol extract prepared from the mycelia of the strain ZZ745 was analyzed by HPLC-HRESIMS. The results (
Supplementary materials, Figures S23 and S24) showed that bagremycin F (
1) was detected in the ethanol crude extract. Therefore, it was clear that
1 was not an artifact, but a new naturally occurring compound.
The molecular formula of C
16H
13NO
4 for
2 was deduced from the (+)-HRESIMS at
m/
z 284.0929 [M + H]
+ and 306.0734 [M + Na]
+, as well as its
13C data. The
13C and
1H NMR spectra (
Supplementary materials, Figures S10 and S11) of
2 showed the signals for one
p-hydroxybenzoic acid ester carbonyl (
δC 164.3), one formamide unit (
δC 160.2;
δH 8.33, 9.78), twelve aromatic carbons and seven aromatic protons, including:
δC 118.1 (C-1), 121.9 (C-2), 126.2 (C-3), 152.6 (C-4), 114.8 (C-5), 126.9 (C-6), 150.4 (C-8), 122.1 (C-9, C-13), 127.1 (C-10, C-12), 134.7 (C-11), and
δH 8.90 (d,
J = 2.2 Hz, H-2); 6.99 (d,
J = 8.5 Hz, H-5); 7.74 (dd,
J = 8.5, 2.2 Hz, H-6); 7.20 (d,
J = 8.6 Hz, H-9, H-13); 7.54 (d,
J = 8.6 Hz, H-10, H-12); and an exocyclic double bond (
δC 135.8, 114.3;
δH 6.74, 1H, dd,
J = 17.6, 10.9 Hz, 5.82, 1H, dd,
J = 17.6, 0.7 Hz, 5.27, 1H, dd,
J = 10.9, 0.7 Hz) (
Table 1). These NMR data indicated that
2 was an analogue of bagremycins. The molecular formula, together with the
1H and
13C NMR data of
2, revealed that its structure was nearly identical to that of
4, except for the acetyl group in
4, which was replaced by the formyl group in
2. Literature search revealed that
2 was a new member of bagremycins, which was named bagremycin G.
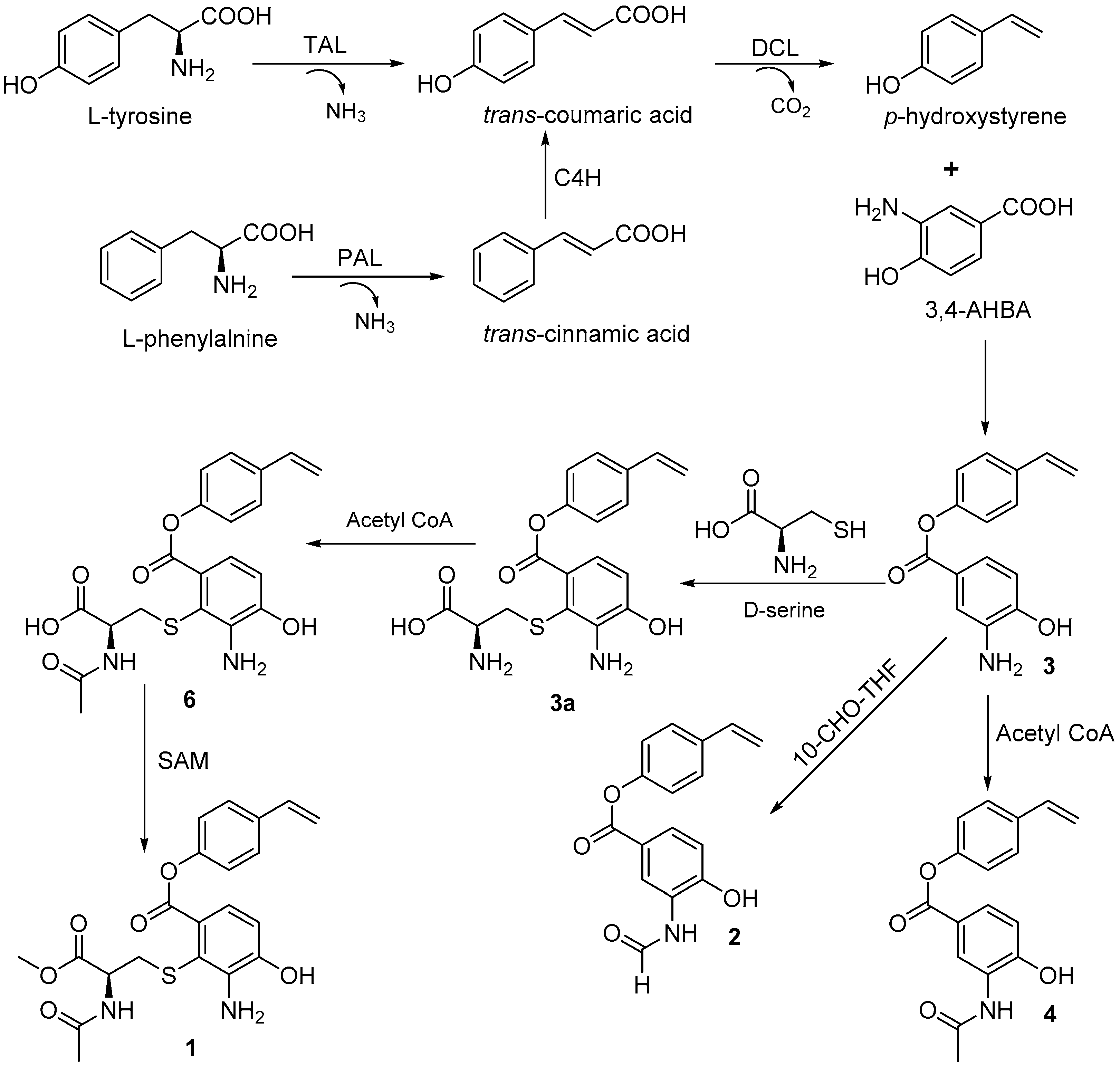
The biosynthetic pathway for bagremycins A (
3) and B (
4) was previously proposed, as described in Reference [
10]. They are the condensation products of 3-amino-4-hydroxybenzoic acid (3,4-AHBA) and
trans-coumaric acid, which is derived from
l-tyrosine or
l-phenylalanine. Accordingly, bagremycins F (
1) and G (
2) should share the same biosynthetic pathway as that of bagremycins A (
3) and B (
4) and may be derived from bagremycin A (
3) (
Figure 3).
The antibacterial activity of
1–
5 against methicillin-resistant
Staphylococcus aureus (MRSA),
Escherichia coli, and
Candida albicans was evaluated using the micro broth dilution method [
15]. Gentamicin (an antibiotic against both Gram-positive and Gram-negative bacteria) and amphotericin B (an antifungal drug) were used as positive controls. As shown in
Table 2, bagremycins F (
1) and G (
2) showed antibacterial activity against
E. coli, with MIC values of 41.8 and 67.1 μM, respectively, but were inactive against MRSA and
C. albicans in a concentration of 116.2 μM for
1 and 176.5 μM for
2.
Bagremycin C (
6) was previously reported to have strong growth inhibitory activity against glioma cells [
9]. Therefore, bagremycins F (
1) and G (
2) were also evaluated for their activity against human glioma U87-MG and U251 cell lines by sulforhodamine B (SRB) assay [
9,
16]. Doxorubicin (DOX) was used as a positive control. The results showed that both bagremycins F (
1) and G (
2) were not active against both cell lines, even at a concentration as high as 100 μM.
3. Materials and Methods
3.1. General Experimental Procedures
Ultraviolet-visible (UV) and infrared radiation (IR) spectra were recorded on a METASH UV-8000 spectrometer (Shanghai METASH Instruments Co. Ltd., Shanghai, China) and a Bruker TENSOR II high performance FT-IR spectrometer (Bruker, Karlsruhe, Germany), respectively. Optical rotation was measured on an Autopol I polarimeter (Rudolph Research Analytical, Hackettstown, NJ, USA). HRESIMS data was obtained from an Agilent 6230 time-of-flight liquid chromatography–mass spectrometry (TOF LC-MS) (Agilent, CA, USA). NMR spectra were acquired on a Bruker 500 spectrometer (Fällanden, Switzerland), and chemical shifts were expressed in δ (ppm). Sephadex LH-20 (GE Healthcare, Stockholm, Sweden) and octadecyl-functionalized silica gel (ODS, Cosmosil 75C18-Prep, Nacalai Tesque Inc., Kyoto, Japan) were used for column chromatography. HPLC separation was conducted on an Agilent 1260 HPLC system with a diode array detector (DAD), using a Zorbax SB-C18 column (250 × 9.4 mm, 5 μm, Agilent Technologies, Palo Alto, CA, USA). All solvents were purchased from the Shanghai Lingfeng Co., Ltd. (Shanghai, China). Amphotericin B (>95.0%) and gentamicin (99.6%) were obtained from Meilune Biotechnology Co. Ltd. (Dalian, China), and doxorubicin (DOX, >98.0%) from Sigma-Aldrich. Human glioma U87-MG and U251 cells were obtained from the Cell Bank of the Chinese Academy of Sciences. Gause’s medium was bought from Guangdong Huankai Microbial Science and Technology Co. Ltd. (Guangzhou, China). Methicillin-resistant Staphylococcus aureus ATCC 43300 (MRSA), Escherichia coli ATCC 25922, and Candida albicans were gifts from Professors Zhongjun Ma, Pinmei Wang, and Bin Wu, respectively.
3.2. Isolation and Identification of Strain ZZ745
Strain ZZ745 was isolated from a sample of marine mud, which was collected from a coastal area located at the Jintang Island of Zhoushan (Zhejiang, China), in September 2016. Summarily, soil (1.0 g) was transferred into a sterile tube containing 9 mL sterile sea water to make a 1 × 10−1 g/mL suspension, after shaking for 10 min on a rotary shaker at 180 rpm. A diluted suspension (1 × 10−3 g/mL, 200 μL) from the 1 × 10−1 g/mL suspension was dispersed evenly on the surface of Gause’s agar medium in a plate, and then incubated for seven days at room temperature. The single colony (strain ZZ745) was shifted onto the surface of Gause’s agar medium in another plate, and then incubated for another six days for securing pure strain. The pure strain ZZ745 was finally transferred onto slant with Gause’s agar medium, and then stored at 4 °C for further use.
The strain ZZ745 was identified by 16S rDNA sequence analysis, which was conducted by Legenomics (Hangzhou, China). Its 16S rDNA sequence has been deposited in GenBank (accession number: MH734530). A voucher strain (Streptomyces sp. ZZ745) was preserved at the Laboratory of the Institute of Marine Biology, Ocean College, Zhejiang University, China.
3.3. Large Scale Culture of Strain ZZ745
The pure colony of strain ZZ745 from slant was refreshed and inoculated into 250 mL Gause’s liquid medium in a 500 mL Erlenmeyer flask, which was cultured at 28 °C for five days in a shaker at 180 rpm, to make seed broth. The seed broth of 5 mL was transferred into a 500 mL Erlenmeyer flask containing 250 mL Gause’s liquid medium, and then cultured for 21 days at the same conditions for the culture of seed broth. A total of 50 L culture was prepared for this study.
3.4. Isolation of Compounds 1–5
The total 50 L culture was separated into two parts of mycelia and supernatant by centrifugation. The mycelia were extracted with MeOH (3 × 300 mL). The methanolic solutions were combined and evaporated under reduced pressure to give a crude extract A (3.3 g). The supernatant was partitioned with EtOAc (3 × 25 L), and the EtOAc solutions were combined and evaporated under reduced pressure to afford a crude extract B (5.5 g). A combination of the extracts A and B (8.8 g) was fractionated by an ODS column eluting with 20%, 40%, 60%, 80%, and 100% MeOH, to give five fractions A–E. The fraction C was further separated by a Sephadex LH-20 column eluting with 50% MeOH to furnish subfractions C1 and C2, based on the results from thin layer chromatography (TLC) analysis. Compounds 1 (1.2 mg, tR 30.2 min, acetonitrile:H2O, 48:52) and 2 (1.3 mg, tR 34.4 min, MeOH:H2O, 60:40) were obtained from the subfractions C1 and C2, respectively, by HPLC purification through a Zorbax SB-C18 column (250 × 9.4 mm, 5 μm). Similarly, the fraction D was separated by an ODS column eluting with 75% MeOH to give subfractions D1 and D2, based on the results from TLC analysis. By HPLC purification using the same Zorbax SB-C18 column, 3 (8.2 mg, tR 18.5 min, MeOH:H2O, 75:25) and 4 (22.1 mg, tR 24.3 min, MeOH:H2O, 75: 25) were purified from the subfraction D1, and 5 (19.8 mg, tR 29.2 min, MeOH:H2O, 80:20) from the subfraction D2.
Bagremycin F (1): colorless amorphous powder; molecular formula C
21H
22N
2O
6S; [α]
−14.0 (
c 0.04, MeOH); UV (MeOH) λ
max (log ε) 201 (3.49), 250 (2.92), 331 (2.34) nm; IR (KBr)
νmax 3317, 2921, 2853, 1714, 1461, 1254, 1171, 1033 cm
−1;
13C and
1H NMR data (in DMSO-
d6), see
Table 1, HRESIMS
m/
z 431.1270 [M + H]
+ (calcd. for C
21H
23N
2O
6S, 431.1277) and 453.1089 [M + Na]
+ (calcd. for C
21H
22N
2NaO
6S, 453.1096).
Bagremycin G (2): colorless amorphous powder; molecular formula C
16H
13NO
4; UV (MeOH) λ
max (log ε) 201 (3.14), 247 (2.97), 321 (1.92) nm; IR (KBr)
νmax 3380, 2918, 2853, 1714, 1540, 1464, 1256, 1198, 1050 cm
−1;
13C and
1H NMR data (in DMSO-
d6), see
Table 1, HRESIMS
m/
z 284.0929 [M + H]
+ (calcd. for C
16H
14NO
4, 284.0923) and 306.0734 [M + Na]
+ (calcd. for C
16H
13NNaO
4, 306.0742).
3.5. Antimicrobial Activity Assay
The antimicrobial activity of
1–
5 against MRSA,
E. coli, and
C. albicans was determined by the micro broth dilution method, as described in the previous study in Reference [
15].
3.6. Antitumor Activity Assay
The activity of
1–
5 against human glioma U87-MG and U251 cells was evaluated by SRB assay and DOX was used as a positive control. U87-MG and U251 cells were cultured in MEM (Minimum Essential Medium, Gibco, Grand Island, NY, USA) and DMEM (Dulbecco’s Modified Eagle Medium, Gibco, Grand Island, NY, USA), both adding 10% FBS (Fetal Bovine Serum, PAA Laboratories Inc., Toronto, ON, Canada), respectively. All glioma cells were incubated at 37 °C in a humidified incubator with 5% CO
2. Cells after the third generation were used for the experiment. The detailed procedure for SRB assay was described in previous publications in References [
9,
16].