Chitin Oligosaccharide Modulates Gut Microbiota and Attenuates High-Fat-Diet-Induced Metabolic Syndrome in Mice
Abstract
1. Introduction
2. Results
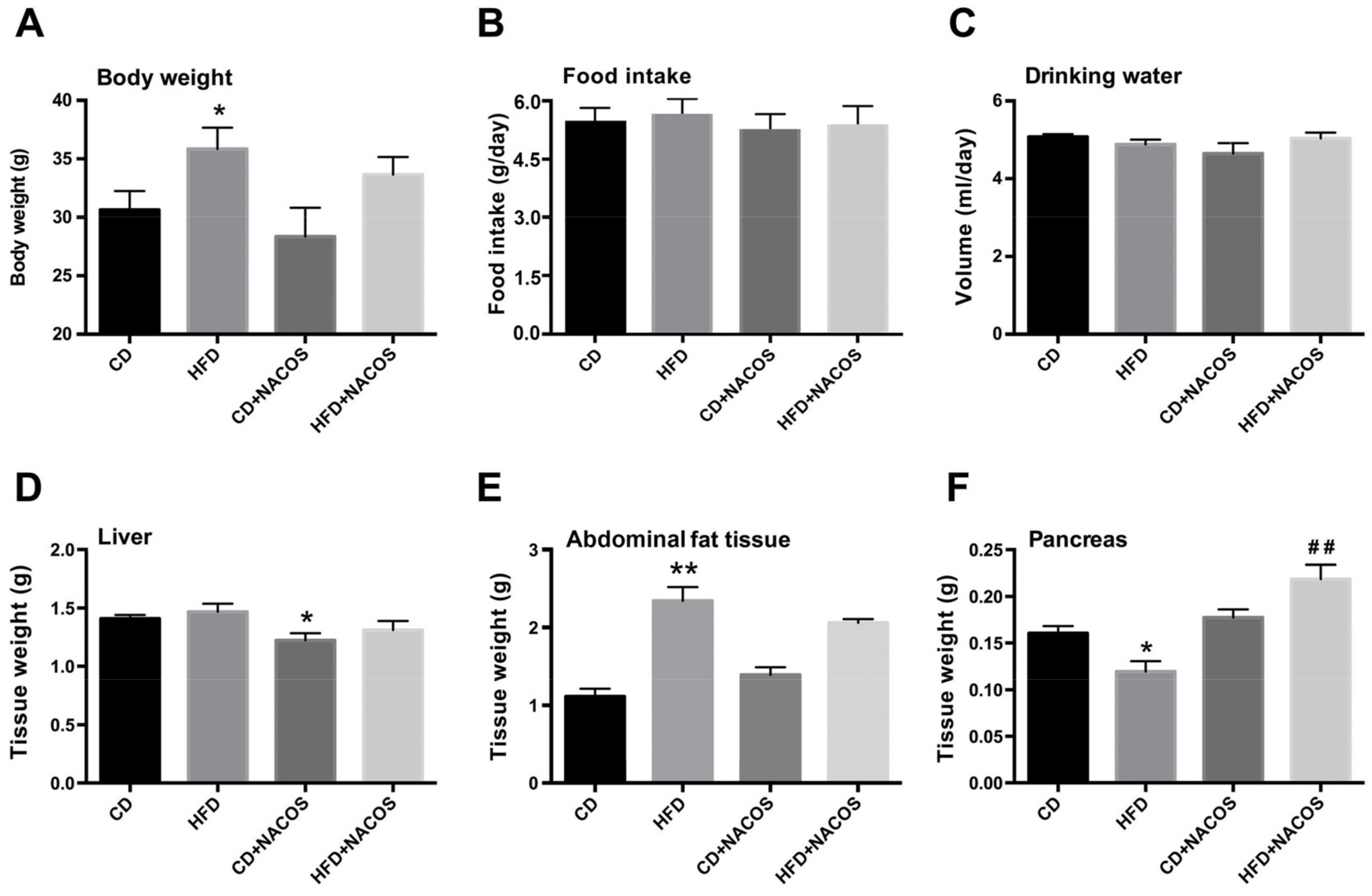
2.1. Effect of NACOS on Food Intake, Drinking Water, and Body Composition of HFD-Fed Mice
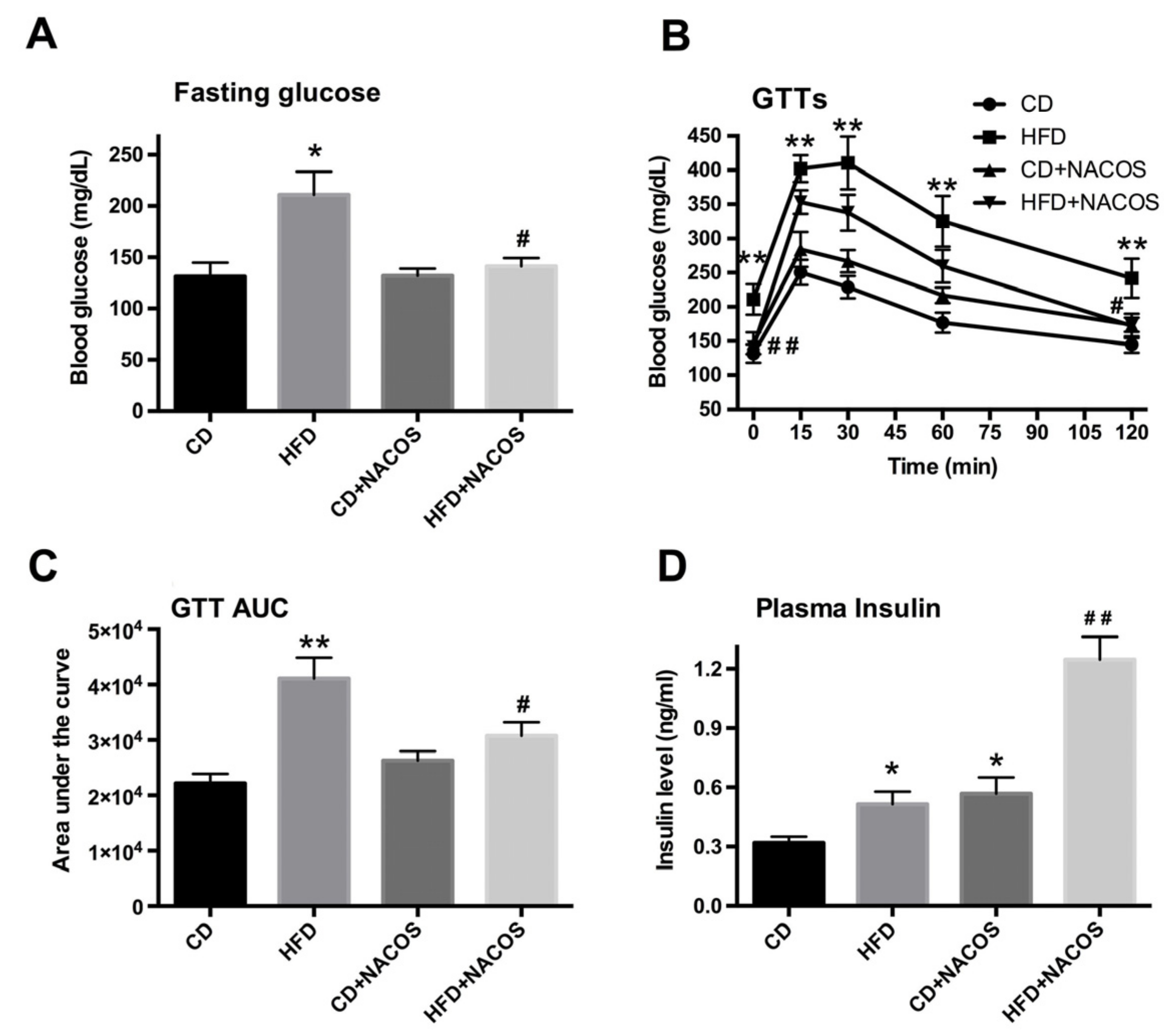
2.2. NACOS Ameliorated Glucose Intolerance and Increased Insulin Secretion of HFD-Treated Mice
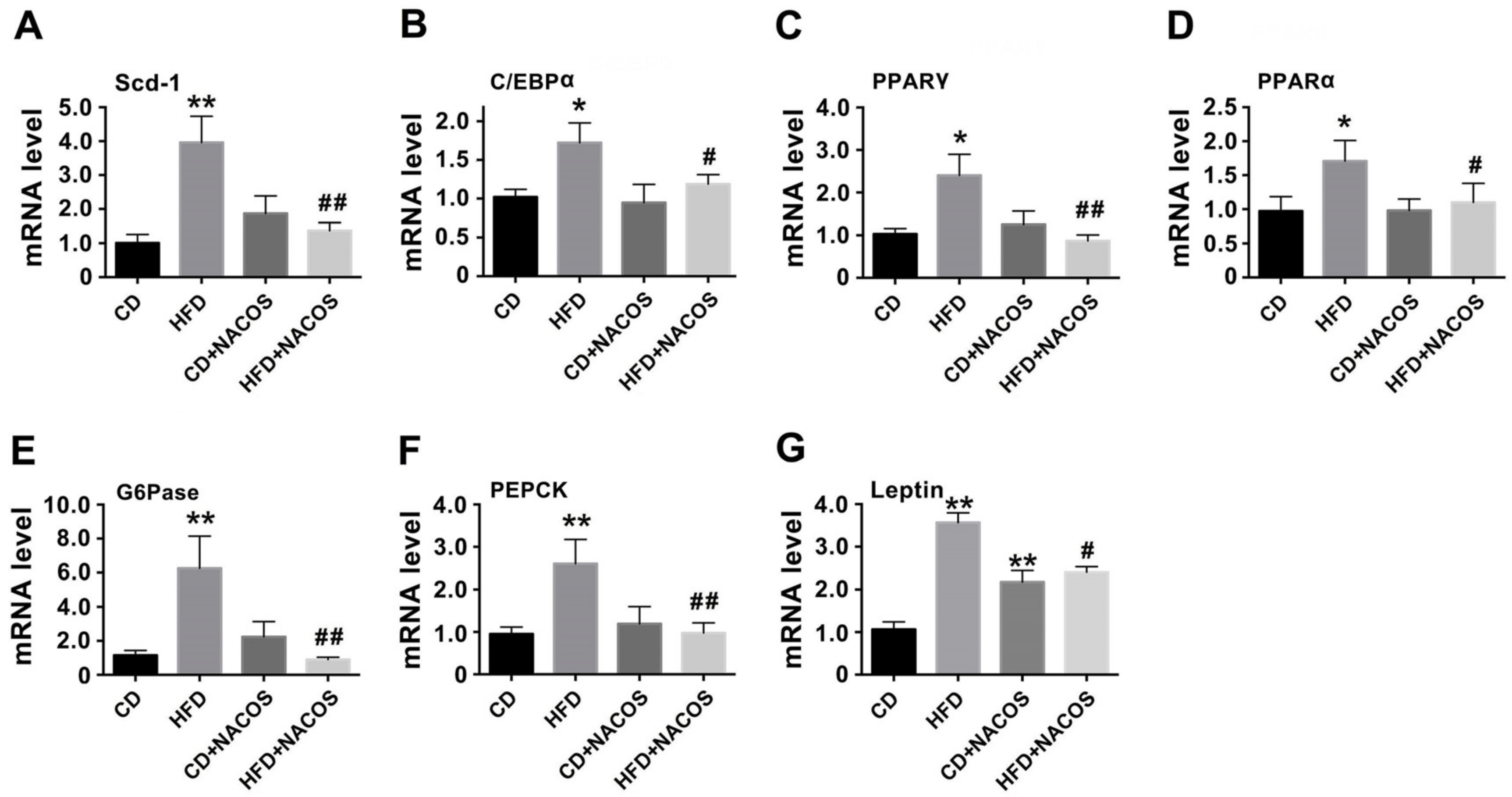
2.3. NACOS Alleviated Dyslipidemia in HFD-Fed Mice
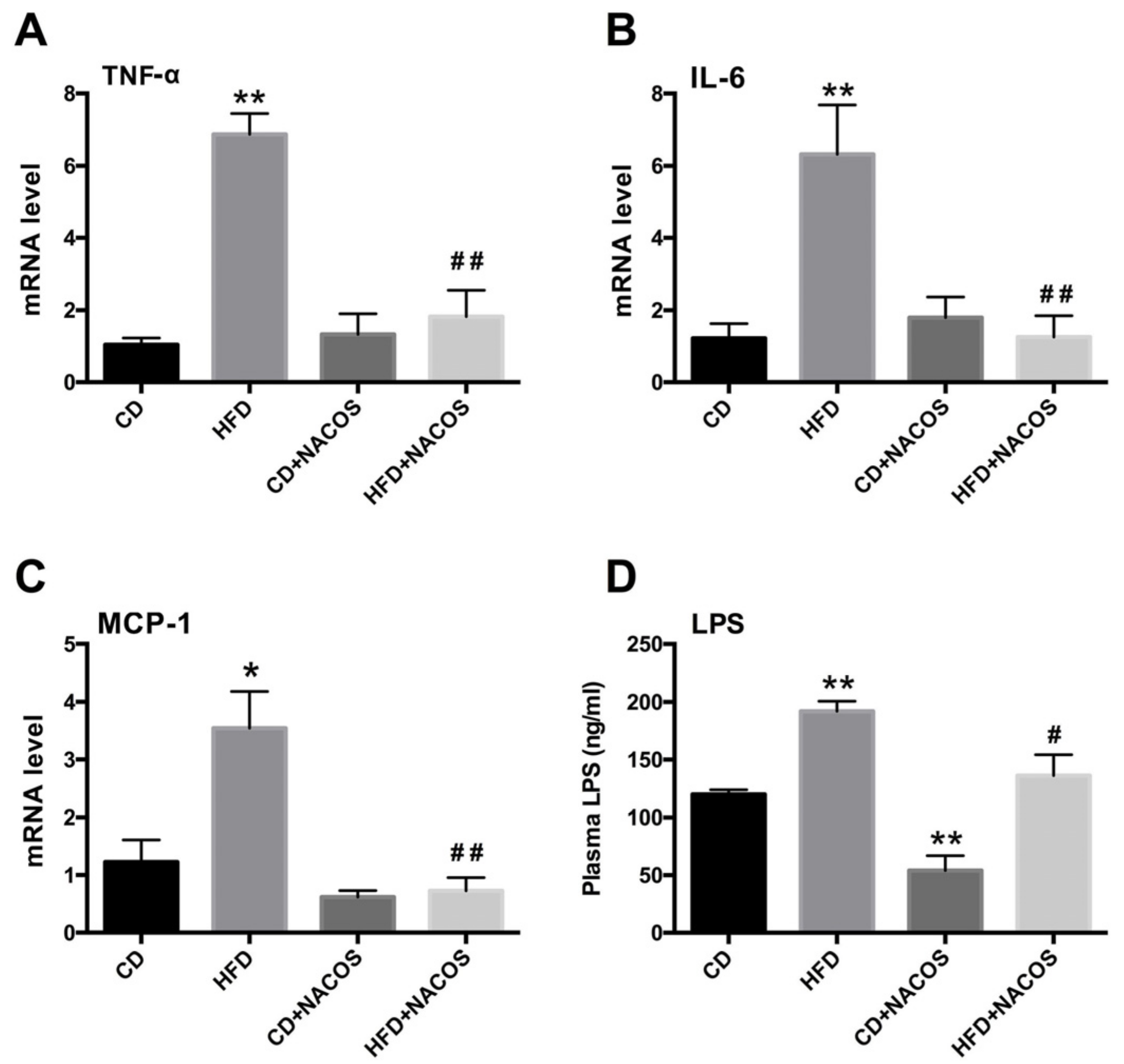
2.4. NACOS Attenuated Inflammation in HFD-Fed Mice
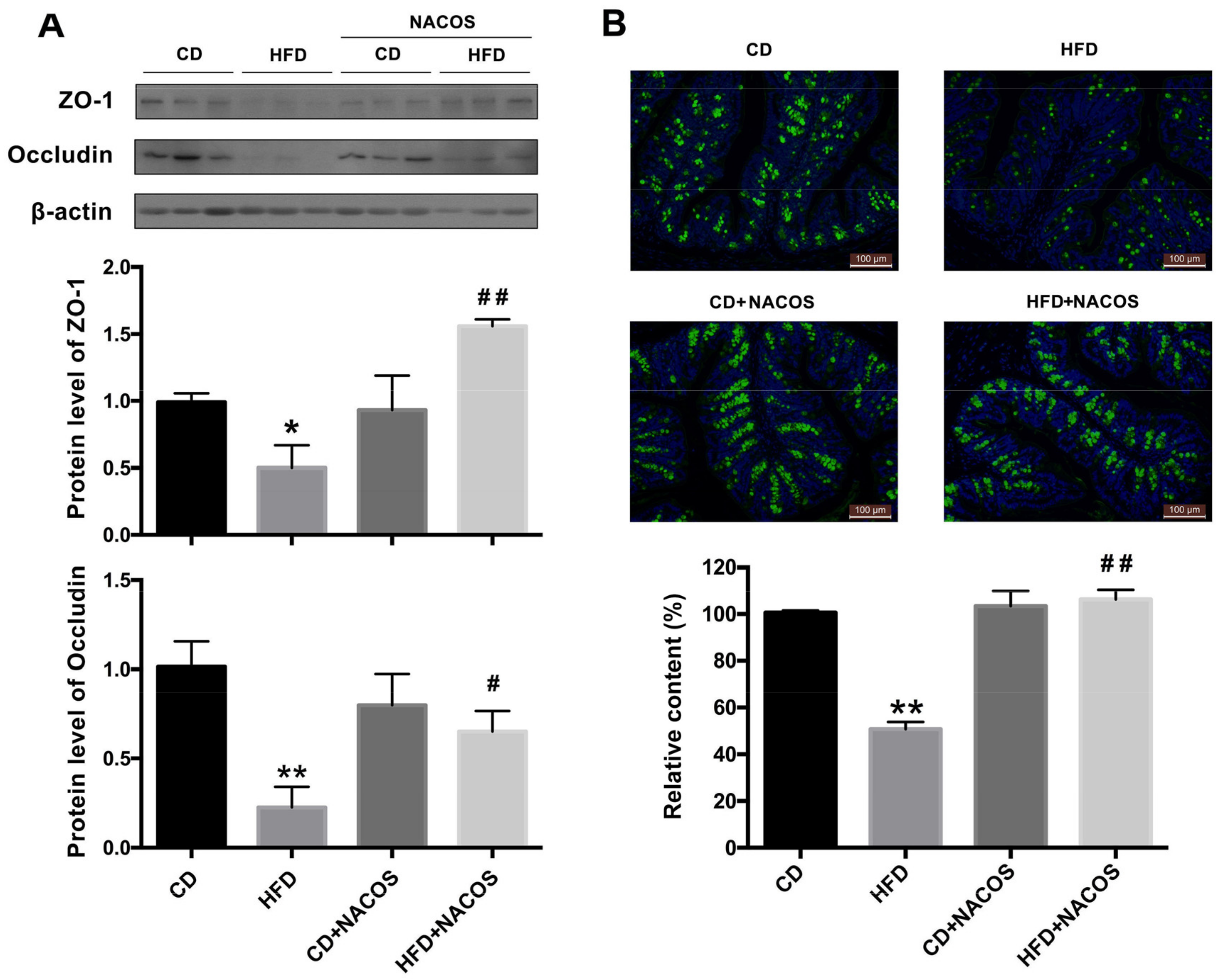
2.5. NACOS Protected Intestinal Integrity in HFD-Fed Mice
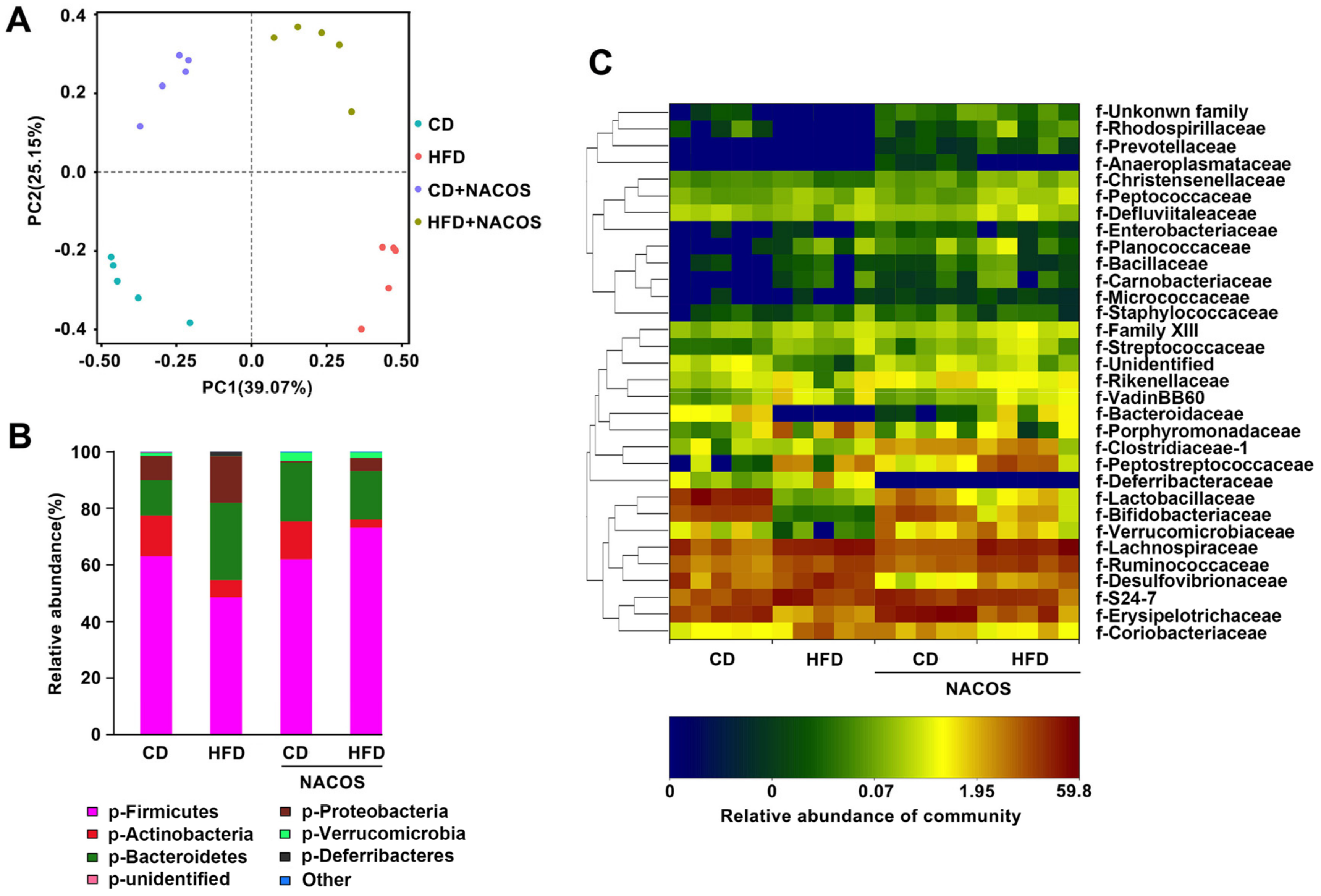
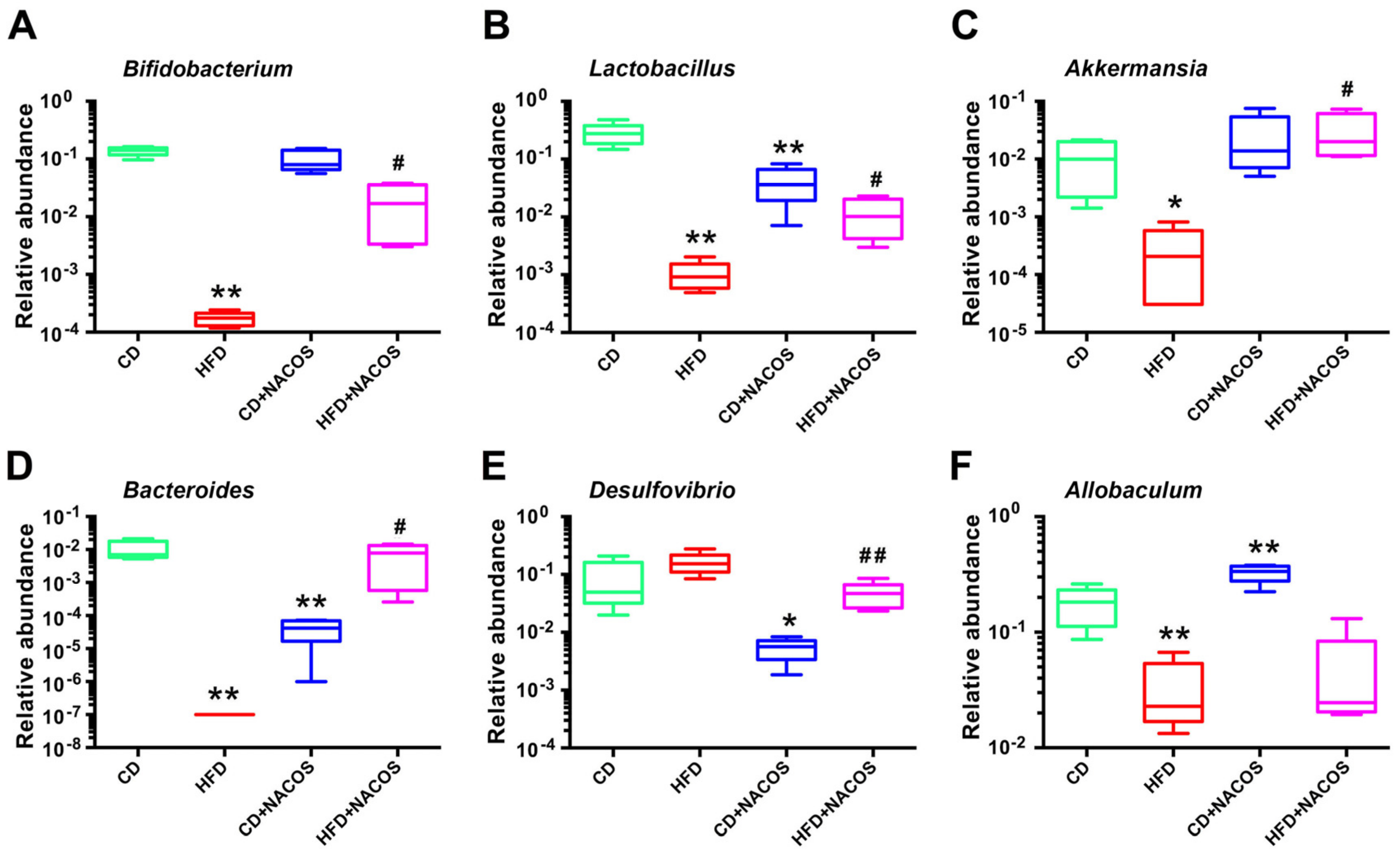
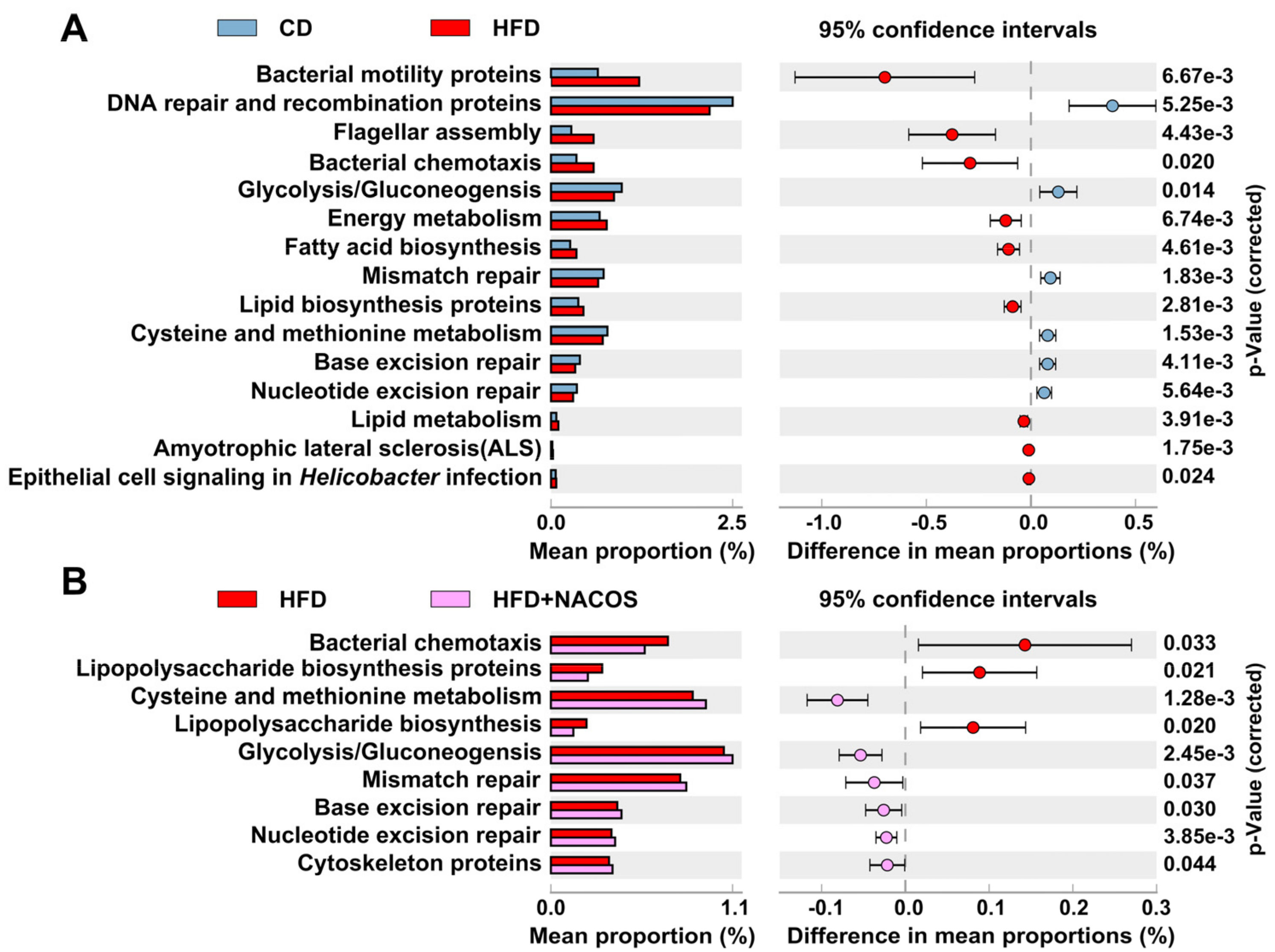
2.6. NACOS Protected Gut Microbiota from Destruction in HFD-Fed Mice
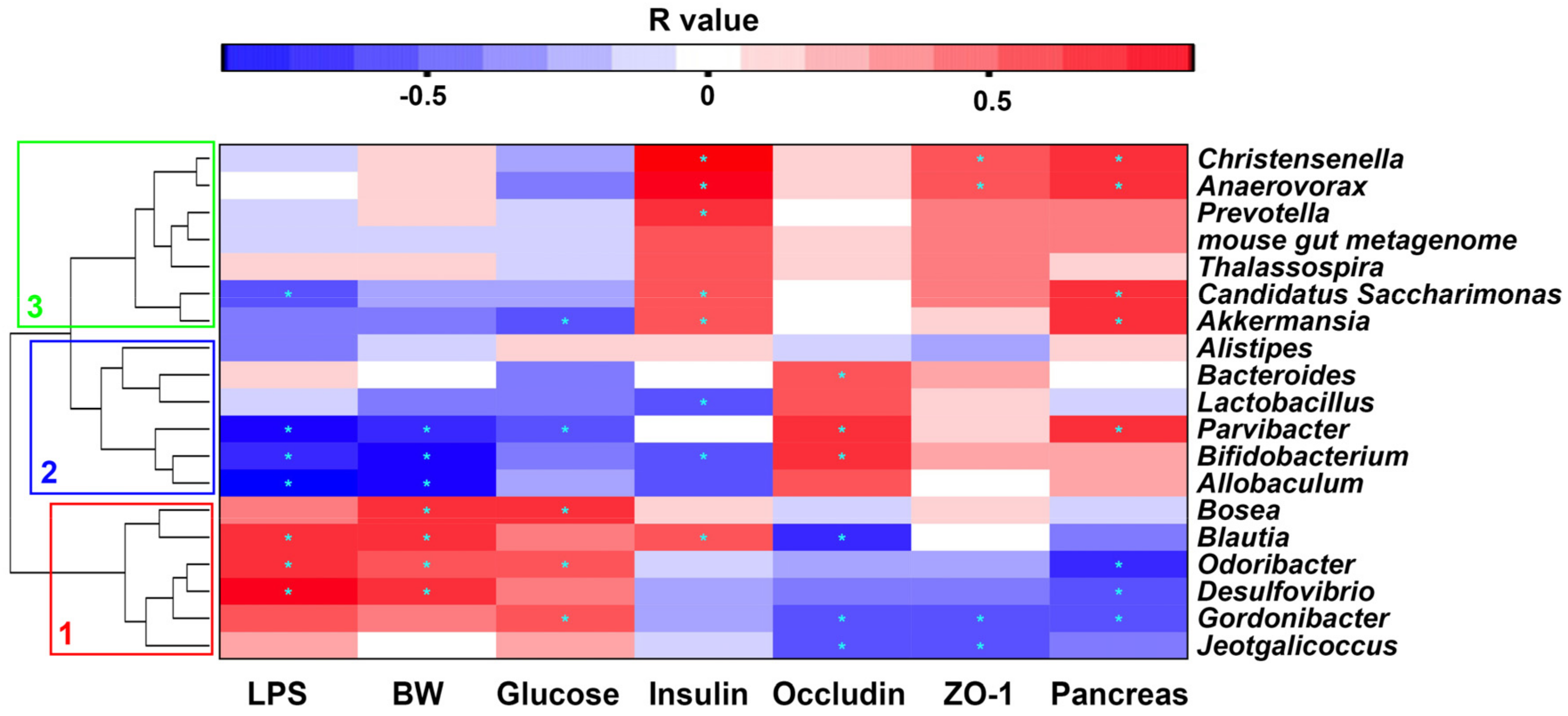
2.7. Correlation between Gut Microbiota and Metabolic Parameters
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Mouse Treatment
4.3. Glucose Tolerance Test and ELISA Assay
4.4. RNA Extraction and Quantitative Real-Time PCR (qRT-PCR)
4.5. Western Blotting
4.6. Immunofluorescent Staining
4.7. 16S rRNA Microbial Community Analysis
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Goldberg, R.B.; Mather, K. Targeting the consequences of the metabolic syndrome in the Diabetes Prevention Program. Arterioscler. Thromb. Vasc. Biol. 2012, 32, 2077–2090. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Neyrinck, A.M.; Fava, F.; Knauf, C.; Burcelin, R.G.; Tuohy, K.M.; Gibson, G.R.; Delzenne, N.M. Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia 2007, 50, 2374–2383. [Google Scholar] [CrossRef] [PubMed]
- Suarez-Zamorano, N.; Fabbiano, S.; Chevalier, C.; Stojanovic, O.; Colin, D.J.; Stevanovic, A.; Veyrat-Durebex, C.; Tarallo, V.; Rigo, D.; Germain, S.; et al. Microbiota depletion promotes browning of white adipose tissue and reduces obesity. Nat. Med. 2015, 21, 1497–1501. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Backhed, F.; Fulton, L.; Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 2008, 3, 213–223. [Google Scholar]
- Ravussin, Y.; Koren, O.; Spor, A.; LeDuc, C.; Gutman, R.; Stombaugh, J.; Knight, R.; Ley, R.E.; Leibel, R.L. Responses of gut microbiota to diet composition and weight loss in lean and obese mice. Obesity (Silver Spring) 2012, 20, 738–747. [Google Scholar] [CrossRef] [PubMed]
- Serino, M.; Luche, E.; Gres, S.; Baylac, A.; Berge, M.; Cenac, C.; Waget, A.; Klopp, P.; Iacovoni, J.; Klopp, C.; et al. Metabolic adaptation to a high-fat diet is associated with a change in the gut microbiota. Gut 2012, 61, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Clavel, T.; Desmarchelier, C.; Haller, D.; Gerard, P.; Rohn, S.; Lepage, P.; Daniel, H. Intestinal microbiota in metabolic diseases: from bacterial community structure and functions to species of pathophysiological relevance. Gut Microbes 2014, 5, 544–551. [Google Scholar] [CrossRef] [PubMed]
- Andersson, U.; Branning, C.; Ahrne, S.; Molin, G.; Alenfall, J.; Onning, G.; Nyman, M.; Holm, C. Probiotics lower plasma glucose in the high-fat fed C57BL/6J mouse. Benef. Microbes 2010, 1, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Everard, A.; Belzer, C.; Geurts, L.; Ouwerkerk, J.P.; Druart, C.; Bindels, L.B.; Guiot, Y.; Derrien, M.; Muccioli, G.G.; Delzenne, N.M.; et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. USA 2013, 110, 9066–9071. [Google Scholar] [CrossRef] [PubMed]
- Shin, N.R.; Lee, J.C.; Lee, H.Y.; Kim, M.S.; Whon, T.W.; Lee, M.S.; Bae, J.W. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 2014, 63, 727–735. [Google Scholar] [CrossRef] [PubMed]
- Stenman, L.K.; Waget, A.; Garret, C.; Klopp, P.; Burcelin, R.; Lahtinen, S. Potential probiotic Bifidobacterium animalis ssp. lactis 420 prevents weight gain and glucose intolerance in diet-induced obese mice. Benef. Microbes 2014, 5, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Ottman, N.; Reunanen, J.; Meijerink, M.; Pietila, T.E.; Kainulainen, V.; Klievink, J.; Huuskonen, L.; Aalvink, S.; Skurnik, M.; Boeren, S.; et al. Pili-like proteins of Akkermansia muciniphila modulate host immune responses and gut barrier function. PLoS One 2017, 12, e0173004. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.E.; Larsson, J.M.; Hansson, G.C. The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 4659–4665. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Possemiers, S.; Van de Wiele, T.; Guiot, Y.; Everard, A.; Rottier, O.; Geurts, L.; Naslain, D.; Neyrinck, A.; Lambert, D.M.; et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 2009, 58, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Khare, P.; Jagtap, S.; Jain, Y.; Baboota, R.K.; Mangal, P.; Boparai, R.K.; Bhutani, K.K.; Sharma, S.S.; Premkumar, L.S.; Kondepudi, K.K.; et al. Cinnamaldehyde supplementation prevents fasting-induced hyperphagia, lipid accumulation, and inflammation in high-fat diet-fed mice. Biofactors 2016, 42, 201–211. [Google Scholar] [PubMed]
- Cani, P.D.; Amar, J.; Iglesias, M.A.; Poggi, M.; Knauf, C.; Bastelica, D.; Neyrinck, A.M.; Fava, F.; Tuohy, K.M.; Chabo, C.; et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 2007, 56, 1761–1772. [Google Scholar] [CrossRef] [PubMed]
- Blasco-Baque, V.; Serino, M.; Burcelin, R. Metabolic therapy at the edge between human hosts and gut microbes. Ann. Pharm. Fr. 2013, 71, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Childs, C.E.; Roytio, H.; Alhoniemi, E.; Fekete, A.A.; Forssten, S.D.; Hudjec, N.; Lim, Y.N.; Steger, C.J.; Yaqoob, P.; Tuohy, K.M.; et al. Xylo-oligosaccharides alone or in synbiotic combination with Bifidobacterium animalis subsp. lactis induce bifidogenesis and modulate markers of immune function in healthy adults: A double-blind, placebo-controlled, randomised, factorial cross-over study. Br. J. Nutr. 2014, 111, 1945–1956. [Google Scholar] [CrossRef] [PubMed]
- Goh, Y.J.; Klaenhammer, T.R. Genetic mechanisms of prebiotic oligosaccharide metabolism in probiotic microbes. Annu. Rev. Food Sci. Technol. 2015, 6, 137–156. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.; Lyu, W.; Xie, M.; Yuan, Q.; Ye, H.; Hu, B.; Zhou, L.; Zeng, X. Effects of alpha-Galactooligosaccharides from Chickpeas on High-Fat-Diet-Induced Metabolic Syndrome in Mice. J. Agric. Food Chem. 2017, 65, 3160–3166. [Google Scholar] [CrossRef] [PubMed]
- Aranda-Martinez, A.; Lopez-Moya, F.; Lopez-Llorca, L.V. Cell wall composition plays a key role on sensitivity of filamentous fungi to chitosan. J. Basic Microbiol. 2016, 56, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Mhamdi, S.; Ktari, N.; Hajji, S.; Nasri, M.; Sellami Kamoun, A. Alkaline proteases from a newly isolated Micromonospora chaiyaphumensis S103: Characterization and application as a detergent additive and for chitin extraction from shrimp shell waste. Int. J. Biol. Macromol. 2017, 94, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Ohnuma, T.; Numata, T.; Osawa, T.; Inanaga, H.; Okazaki, Y.; Shinya, S.; Kondo, K.; Fukuda, T.; Fukamizo, T. Crystal structure and chitin oligosaccharide-binding mode of a ‘loopful’ family GH19 chitinase from rye, Secale cereale, seeds. FEBS J. 2012, 279, 3639–3651. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.H.; Ha, K.S.; Moon, K.S.; Kim, J.G.; Oh, C.G.; Kim, Y.C.; Apostolidis, E.; Kwon, Y.I. Molecular weight dependent glucose lowering effect of low molecular weight Chitosan Oligosaccharide (GO2KA1) on postprandial blood glucose level in SD rats model. Int. J. Mol. Sci. 2013, 14, 14214–14224. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Ahn, H.Y.; Kwak, J.H.; Shin, D.Y.; Kwon, Y.I.; Oh, C.G.; Lee, J.H. The effects of chitosan oligosaccharide (GO2KA1) supplementation on glucose control in subjects with prediabetes. Food Funct. 2014, 5, 2662–2669. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.W.; Park, Y.S.; Choi, J.W.; Yi, S.Y.; Shin, W.S. Antidiabetic effects of chitosan oligosaccharides in neonatal streptozotocin-induced noninsulin-dependent diabetes mellitus in rats. Biol. Pharm. Bull. 2003, 26, 1100–1103. [Google Scholar] [CrossRef] [PubMed]
- Yuan, W.P.; Liu, B.; Liu, C.H.; Wang, X.J.; Zhang, M.S.; Meng, X.M.; Xia, X.K. Antioxidant activity of chito-oligosaccharides on pancreatic islet cells in streptozotocin-induced diabetes in rats. World J. Gastroenterol. 2009, 15, 1339–1345. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.G.; Jo, S.H.; Ha, K.S.; Kim, S.C.; Kim, Y.C.; Apostolidis, E.; Kwon, Y.I. Effect of long-term supplementation of low molecular weight chitosan oligosaccharide (GO2KA1) on fasting blood glucose and HbA1c in db/db mice model and elucidation of mechanism of action. BMC Complement. Altern. Med. 2014, 14, 272. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.K.; Park, P.J.; Yang, H.P.; Han, S.S. Subacute toxicity of chitosan oligosaccharide in Sprague-Dawley rats. Arzneimittelforschung 2001, 51, 769–774. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, Q.Q.; Zhao, S.; Li, S.D.; Song, C. Application of Chitosan, Chitooligosaccharide, and Their Derivatives in the Treatment of Alzheimer’s Disease. Mar. Drugs 2017, 15, 322. [Google Scholar] [CrossRef] [PubMed]
- Park, B.K.; Kim, M.M. Applications of chitin and its derivatives in biological medicine. Int. J. Mol. Sci. 2010, 11, 5152–5164. [Google Scholar] [CrossRef] [PubMed]
- Azuma, K.; Osaki, T.; Minami, S.; Okamoto, Y. Anticancer and anti-inflammatory properties of chitin and chitosan oligosaccharides. J. Funct. Biomater. 2015, 6, 33–49. [Google Scholar] [CrossRef] [PubMed]
- Neyrinck, A.M.; Possemiers, S.; Verstraete, W.; De Backer, F.; Cani, P.D.; Delzenne, N.M. Dietary modulation of clostridial cluster XIVa gut bacteria (Roseburia spp.) by chitin-glucan fiber improves host metabolic alterations induced by high-fat diet in mice. J. Nutr. Biochem. 2012, 23, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Mateos-Aparicio, I.; Mengibar, M.; Heras, A. Effect of chito-oligosaccharides over human faecal microbiota during fermentation in batch cultures. Carbohydr. Polym. 2016, 137, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Masuda, S.; Azuma, K.; Kurozumi, S.; Kiyose, M.; Osaki, T.; Tsuka, T.; Itoh, N.; Imagawa, T.; Minami, S.; Sato, K.; et al. Anti-tumor properties of orally administered glucosamine and N-acetyl-d-glucosamine oligomers in a mouse model. Carbohydr. Polym. 2014, 111, 783–787. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Chen, J.; Cao, P.; Pan, H.; Ding, C.; Xiao, T.; Zhang, P.; Guo, J.; Su, Z. Anti-obese effect of glucosamine and chitosan oligosaccharide in high-fat diet-induced obese rats. Mar. Drugs 2015, 13, 2732–2756. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, E.; Fuentes, F.; Vilahur, G.; Badimon, L.; Palomo, I. Mechanisms of chronic state of inflammation as mediators that link obese adipose tissue and metabolic syndrome. Mediators Inflamm. 2013, 2013, 136584. [Google Scholar] [CrossRef] [PubMed]
- Musso, G.; Gambino, R.; Cassader, M. Obesity, diabetes, and gut microbiota: The hygiene hypothesis expanded? Diabetes Care 2010, 33, 2277–2284. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.L.; Lin, H.T.; Liang, T.W.; Chen, Y.J.; Yen, Y.H.; Guo, S.P. Reclamation of chitinous materials by bromelain for the preparation of antitumor and antifungal materials. Bioresour. Technol. 2008, 99, 4386–4393. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Zhang, Y.; Liu, W.; Xu, L.; Yang, Y.; Zhou, Z. Effects of chito-oligosaccharides supplementation on growth performance, intestinal cytokine expression, autochthonous gut bacteria and disease resistance in hybrid tilapia Oreochromis niloticus female symbol x Oreochromis aureus male symbol. Fish Shellfish Immunol 2014, 40, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Liu, W.S.; Han, B.Q.; Sun, Y.Y. Antidiabetic effects of chitooligosaccharides on pancreatic islet cells in streptozotocin-induced diabetic rats. World J. Gastroenterol. 2007, 13, 725–731. [Google Scholar] [CrossRef] [PubMed]
- Collins, K.H.; Paul, H.A.; Hart, D.A.; Reimer, R.A.; Smith, I.C.; Rios, J.L.; Seerattan, R.A.; Herzog, W. A High-Fat High-Sucrose Diet Rapidly Alters Muscle Integrity, Inflammation and Gut Microbiota in Male Rats. Sci. Rep. 2016, 6, 37278. [Google Scholar] [CrossRef] [PubMed]
- Flemer, B.; Gaci, N.; Borrel, G.; Sanderson, I.R.; Chaudhary, P.P.; Tottey, W.; O’Toole, P.W.; Brugere, J.F. Fecal microbiota variation across the lifespan of the healthy laboratory rat. Gut Microbes 2017, 8, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Chassaing, B.; Etienne-Mesmin, L.; Gewirtz, A.T. Microbiota-liver axis in hepatic disease. Hepatology 2014, 59, 328–339. [Google Scholar] [CrossRef] [PubMed]
- Koropatkin, N.M.; Cameron, E.A.; Martens, E.C. How glycan metabolism shapes the human gut microbiota. Nat. Rev. Microbiol. 2012, 10, 323–335. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Vogensen, F.K.; van den Berg, F.W.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sorensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One 2010, 5, e9085. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Lu, J.; Wang, Y.; Gu, W.; Yang, X.; Yu, J. Intake of total saponins and polysaccharides from Polygonatum kingianum affects the gut microbiota in diabetic rats. Phytomedicine 2017, 26, 45–54. [Google Scholar] [CrossRef] [PubMed]
- Everard, A.; Lazarevic, V.; Derrien, M.; Girard, M.; Muccioli, G.G.; Neyrinck, A.M.; Possemiers, S.; Van Holle, A.; Francois, P.; de Vos, W.M.; et al. Responses of gut microbiota and glucose and lipid metabolism to prebiotics in genetic obese and diet-induced leptin-resistant mice. Diabetes 2011, 60, 2775–2786. [Google Scholar] [CrossRef] [PubMed]
- Foley, M.H.; Cockburn, D.W.; Koropatkin, N.M. The Sus operon: A model system for starch uptake by the human gut Bacteroidetes. Cell. Mol. Life Sci. 2016, 73, 2603–2617. [Google Scholar] [CrossRef] [PubMed]
- Rabot, S.; Membrez, M.; Blancher, F.; Berger, B.; Moine, D.; Krause, L.; Bibiloni, R.; Bruneau, A.; Gerard, P.; Siddharth, J.; et al. High fat diet drives obesity regardless the composition of gut microbiota in mice. Sci. Rep. 2016, 6, 32484. [Google Scholar] [CrossRef] [PubMed]
- Arora, T.; Seyfried, F.; Docherty, N.G.; Tremaroli, V.; le Roux, C.W.; Perkins, R.; Backhed, F. Diabetes-associated microbiota in fa/fa rats is modified by Roux-en-Y gastric bypass. ISME J. 2017, 11, 2035–2046. [Google Scholar] [CrossRef] [PubMed]
- Bordalo Tonucci, L.; Dos Santos, K.M.; De Luces Fortes Ferreira, C.L.; Ribeiro, S.M.; De Oliveira, L.L.; Martino, H.S. Gut microbiota and probiotics: Focus on diabetes mellitus. Crit. Rev. Food Sci. Nutr. 2017, 57, 2296–2309. [Google Scholar] [CrossRef] [PubMed]
- Marcobal, A.; Sonnenburg, J.L. Human milk oligosaccharide consumption by intestinal microbiota. Clin. Microbiol. Infect. 2012, 18, 12–15. [Google Scholar] [CrossRef] [PubMed]
- Hale, V.L.; Chen, J.; Johnson, S.; Harrington, S.C.; Yab, T.C.; Smyrk, T.C.; Nelson, H.; Boardman, L.A.; Druliner, B.R.; Levin, T.R.; et al. Shifts in the Fecal Microbiota Associated with Adenomatous Polyps. Cancer Epidemiol. Biomarkers Prev. 2017, 26, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Zhang-Sun, W.; Augusto, L.A.; Zhao, L.; Caroff, M. Desulfovibrio desulfuricans isolates from the gut of a single individual: Structural and biological lipid A characterization. FEBS Lett. 2015, 589, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.S.; Xu, J.; Zhu, H.; Wu, J.; Xu, J.D.; Yan, R.; Li, X.Y.; Liu, H.H.; Duan, S.M.; Wang, Z.; et al. Gut microbiota-involved mechanisms in enhancing systemic exposure of ginsenosides by coexisting polysaccharides in ginseng decoction. Sci. Rep. 2016, 6, 22474. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Chen, H.; Faas, M.M.; de Haan, B.J.; Li, J.; Xiao, P.; Zhang, H.; Diana, J.; de Vos, P.; Sun, J. Specific inulin-type fructan fibers protect against autoimmune diabetes by modulating gut immunity, barrier function, and microbiota homeostasis. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zheng, J.; Cheng, G.; Li, Q.; Jiao, S.; Feng, C.; Zhao, X.; Yin, H.; Du, Y.; Liu, H. Chitin Oligosaccharide Modulates Gut Microbiota and Attenuates High-Fat-Diet-Induced Metabolic Syndrome in Mice. Mar. Drugs 2018, 16, 66. https://doi.org/10.3390/md16020066
Zheng J, Cheng G, Li Q, Jiao S, Feng C, Zhao X, Yin H, Du Y, Liu H. Chitin Oligosaccharide Modulates Gut Microbiota and Attenuates High-Fat-Diet-Induced Metabolic Syndrome in Mice. Marine Drugs. 2018; 16(2):66. https://doi.org/10.3390/md16020066
Chicago/Turabian StyleZheng, Junping, Gong Cheng, Qiongyu Li, Siming Jiao, Cui Feng, Xiaoming Zhao, Heng Yin, Yuguang Du, and Hongtao Liu. 2018. "Chitin Oligosaccharide Modulates Gut Microbiota and Attenuates High-Fat-Diet-Induced Metabolic Syndrome in Mice" Marine Drugs 16, no. 2: 66. https://doi.org/10.3390/md16020066
APA StyleZheng, J., Cheng, G., Li, Q., Jiao, S., Feng, C., Zhao, X., Yin, H., Du, Y., & Liu, H. (2018). Chitin Oligosaccharide Modulates Gut Microbiota and Attenuates High-Fat-Diet-Induced Metabolic Syndrome in Mice. Marine Drugs, 16(2), 66. https://doi.org/10.3390/md16020066




