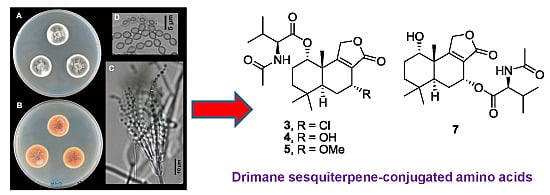
Drimane Sesquiterpene-Conjugated Amino Acids from a Marine Isolate of the Fungus Talaromyces minioluteus (Penicillium Minioluteum)
Abstract
:1. Introduction
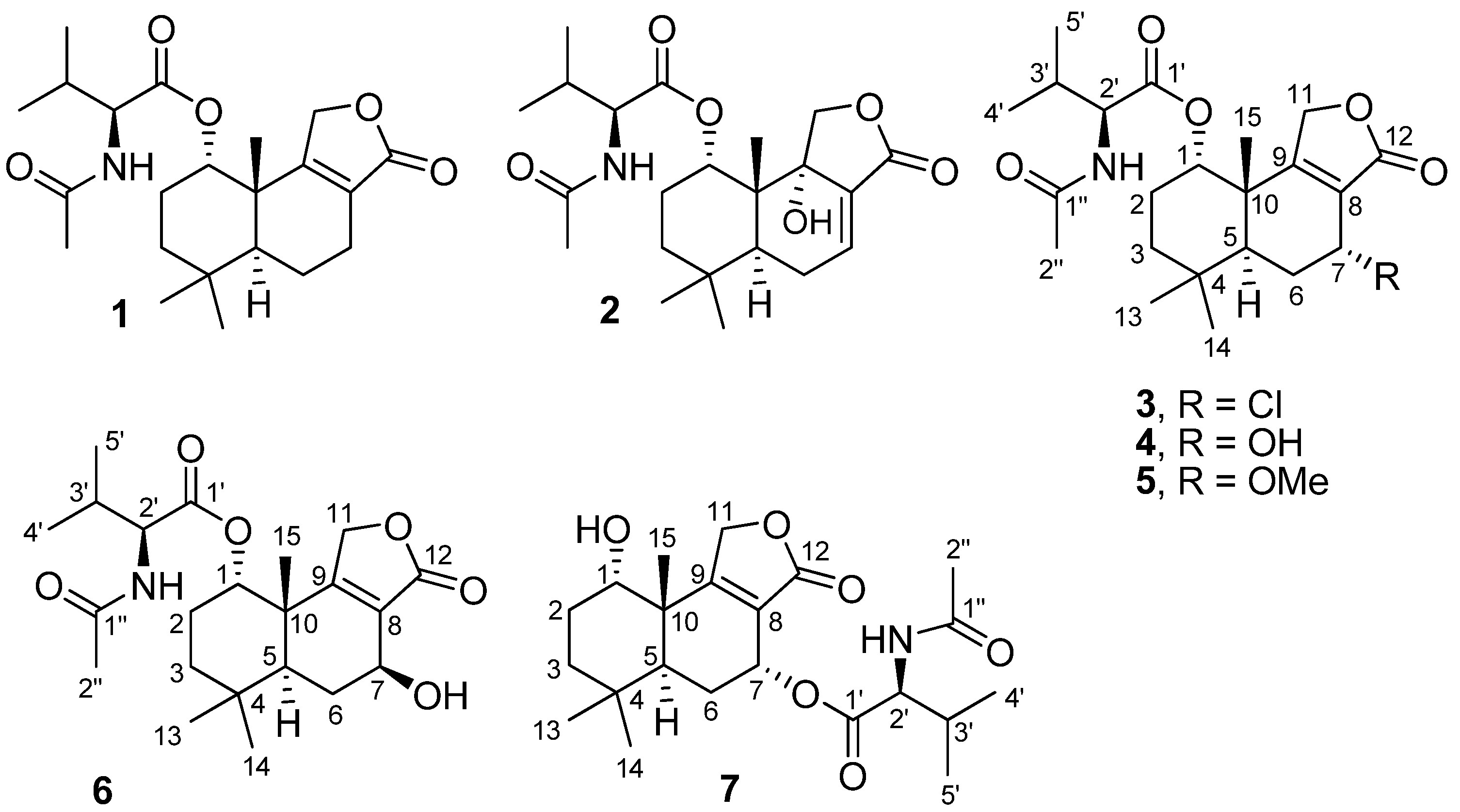
2. Results and Discussion
2.1. Structure Elucidation of New Fungal Metabolites 3, 4, 6 and 7
| 3 | 4 | |||
|---|---|---|---|---|
| δC | δH, multi (J in Hz) | δC | δH, multi (J in Hz) | |
| 1 | 74.3 | 5.02, brs | 74.4 | 4.97, s |
| 2 | 22.4 | 1.85, dq (2.9, 15.1); 1.95–2.07, m | 22.4 | 1.83, dq (2.7, 15.1); 1.94, m |
| 3 | 34.8 | 1.42, dt (3.8, 13.8); 1.65, td (4.0, 14.0) | 34.9 | 1.38, dt (2.5, 13.7); 1.58, td (4.2,13.9) |
| 4 | 32.2 | - | 32.3 | - |
| 5 | 39.8 | 2.40, d (12.4) | 39.7 | 2.20, dd (1.1,12.4) |
| 6 | 28.4 | 1.95–2.07, m; 2.22, d (14.8) | 27.2 | 1.75, td (4.4, 13.9); 2.01, brd (14.1) |
| 7 | 48.6 | 4.86, t (1.8) | 59.6 | 4.55, brd (1.6) |
| 8 | 126.4 | - | 127.1 | - |
| 9 | 170.8 | - | 170.8 | - |
| 10 | 41.5 | - | 41.4 | - |
| 11 | 67.8 | 4.60, d (17.6); 4.72, dd (1.9,17.6) | 68.2 | 4.60, d (17.4); 4.72, dd (1.6,17.4) |
| 12 | 170.8 | - | 173.4 | - |
| 13 | 32.7 | 1.06, s | 33.0 | 1.04, s |
| 14 | 21.8 | 0.97, s | 21.4 | 0.96, s |
| 15 | 20.2 | 1.24, s | 19.6 | 1.20, s |
| 1′ | 170.2 | - | 170.8 | - |
| 2′ | 58.2 | 4.42, dd (5.3, 8.2) | 58.5 | 4.32, dd (6.0, 8.0) |
| 3′ | 30.8 | 2.06, m | 30.3 | 2.07, oct (6.5) |
| 4′ | 18.0 | 0.89, d (6.6) | 18.0 | 0.92, d (6.6) |
| 5′ | 19.0 | 0.94, d (6.6) | 19.1 | 0.95, d (6.6) |
| 1″ | 170.3 | - | 170.5 | - |
| 2″ | 22.9 | 2.00, s | 22.7 | 1.97, s |
| NH | - | 5.74, d (8.0) | - | 6.16, d (7.9) |
| OMe | - | - | - | - |
| 6 | 7 | |||
|---|---|---|---|---|
| δC | δH, multi (J in Hz) | δC | δH, multi (J in Hz) | |
| 1 | 74.6 | 5.00, brs | 71.0 | 3.89, brs |
| 2 | 22.3 | 1.78, q (3.0); 1.96, m | 25.7 | 1.63, dq (3.1,14.8); 2.06, m |
| 3 | 34.5 | 1.36, dt (2.5, 15.2); 1.53-1.63, m | 34.1 | 1.33, dt (3.7, 12.8); 1.74, td (4.1, 14.1) |
| 4 | 32.6 | - | 32.5 | - |
| 5 | 44.4 | 1.81, brd (12.8) | 39.1 | 2.06, brd (13.2) |
| 6 | 27.1 | 1.53–1.63, m; 2.26, dd (6.9, 12.9) | 25.6 | 1.86, ddd (15.0, 13.2, 4.5); 1.97, brd (15.0) |
| 7 | 64.8 | 4.61, t (8.8) | 62.9 | 5.71, t (2.0) |
| 8 | 127.8 | - | 122.7 | - |
| 9 | 168.5 | - | 175.9 | - |
| 10 | 41.6 | - | 42.2 | - |
| 11 | 68.0 | 4.55, dd (2.7, 17.2); 4.73, d (17.2) | 68.5 | 4.81, dd (2.0, 17.1); 5.11, d (17.1) |
| 12 | 172.9 | - | 171.8 | - |
| 13 | 33.1 | 1.02, s | 32.7 | 0.93, s |
| 14 | 21.4 | 0.97, s | 21.1 | 0.91, s |
| 15 | 21.1 | 1.33, s | 20.1 | 1.13, s |
| 1′ | 171.2 | - | 170.8 | - |
| 2′ | 57.7 | 4.43, dd (4.8, 8.2) | 57.3 | 4.56, dd (4.6, 8.7) |
| 3′ | 30.6 | 1.96, m | 31.2 | 2.17, otc (6.8) |
| 4′ | 17.5 | 0.85, d (6.8) | 17.6 | 0.90, d (6.6) |
| 5′ | 19.0 | 0.92, d (6.8) | 19.0 | 0.95, d (6.6) |
| 1′′ | 170.0 | - | 169.7 | - |
| 2′′ | 23.0 | 2.01, s | 23.1 | 2.02, s |
| NH | - | 5.78, d (8.2) | - | 6.04, d (8.6) |
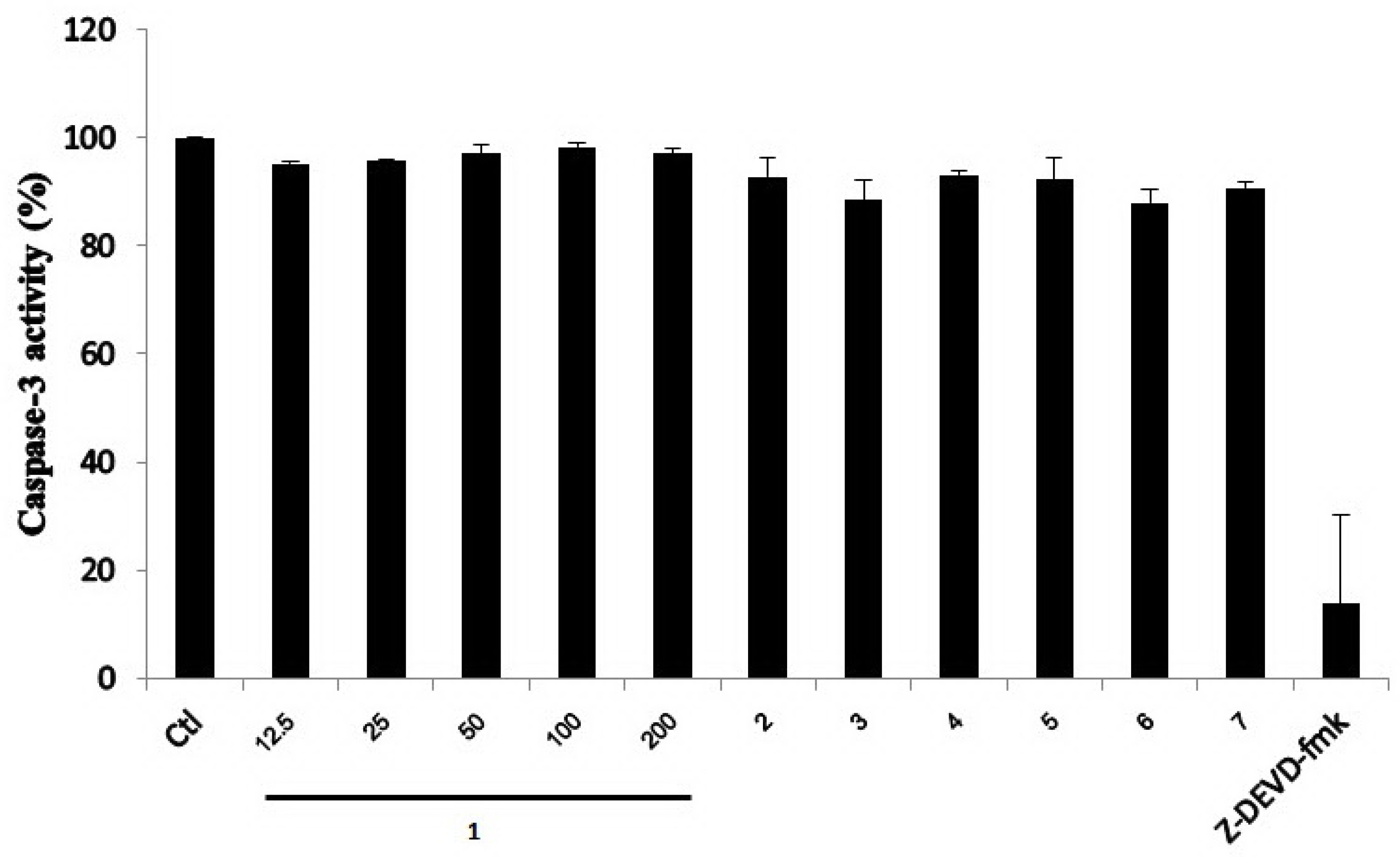
2.2. Biological Activities of the Isolated Compounds 1–7
3. Experimental Section
3.1. General Experimental Procedures
3.2. Fungal Material and Identification of the Fungus PILE 14-5
3.2.1. Morphological Characterization
3.2.2. DNA Sequence-Based Identification
3.3. Fungal Cultivation, Extraction and Isolation of Metabolites
3.4. Spectroscopic Data of Compounds
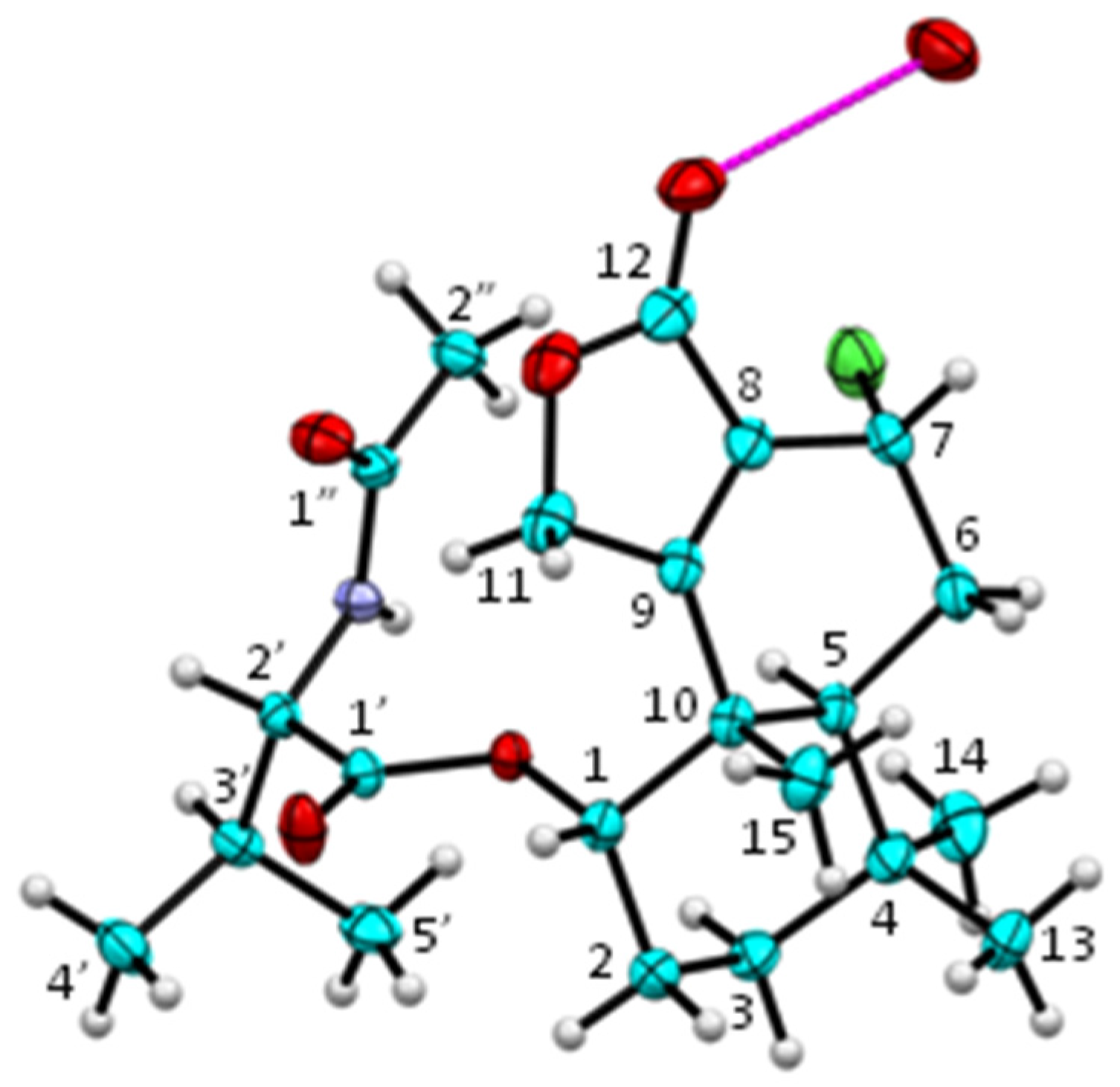
3.5. Single Crystal X-Ray Analysis
3.6. Bioassays
3.6.1. Cell Culture
3.6.2. Cell Viability Assay
3.6.3. Caspase-3 Activity Assay
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bugni, T.S.; Ireland, C.M. Marine-derived fungi: A chemically and biologically diverse group of microorganisms. Nat. Prod. Rep. 2004, 21, 143–163. [Google Scholar] [CrossRef] [PubMed]
- Trisuwan, K.; Rukachaisirikul, V.; Sukpondma, Y.; Preedanon, S.; Phongpaichit, S.; Sakayaroj, J. Pyrone derivatives from the marine-derived fungus Nigrospora sp. PSU-F18. Phytochemistry 2009, 70, 554–557. [Google Scholar] [CrossRef] [PubMed]
- Pejin, B.; Jovanovic, K.K.; Mojovic, M.; Savic, A.G. New and highly potent antitumor natural products from marine-derived fungi: Covering the period from 2003 to 2012. Curr. Top. Med. Chem. 2013, 13, 2745–2766. [Google Scholar] [CrossRef] [PubMed]
- Yuan, W.H.; Wei, Z.W.; Dai, P.; Wu, H.; Zhao, Y.X.; Zhang, M.M.; Jiang, N.; Zheng, W.F. Halogenated metabolites isolated from Penicillium citreonigrum. Chem. Biodivers. 2014, 11, 1078–1087. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.L.; Bao, J.; Liu, K.S.; Zhang, X.Y.; He, F.; Wang, Y.F.; Nong, X.H.; Qi, S.H. Cytotoxic dihydrothiophene-condensed chromones from the marine-derived fungus Penicillium oxalicum. Planta Med. 2013, 79, 1474–1479. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.; Zhang, X.; Du, L.; Wang, W.; Zhu, T.; Gu, Q.; Li, D. Sorbicatechols A and B, antiviral sorbicillinoids from the marine-derived fungus Penicillium chrysogenum PJX-17. J. Nat. Prod. 2014, 77, 424–428. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.W.; Mordhorst, T.F.; Lee, C.; Jensen, P.R.; Fenical, W.; Kock, M. Penilumamide, a novel lumazine peptide isolated from the marine-derived fungus, Penicillium sp. CNL-338. Org. Biomol. Chem. 2010, 8, 2158–2163. [Google Scholar] [CrossRef] [PubMed]
- Guimaraes, D.O.; Borges, W.S.; Vieira, N.J.; de Oliveira, L.F.; da Silva, C.H.; Lopes, N.P.; Dias, L.G.; Duran-Patron, R.; Collado, I.G.; Pupo, M.T. Diketopiperazines produced by endophytic fungi found in association with two asteraceae species. Phytochemistry 2010, 71, 1423–1429. [Google Scholar] [CrossRef] [PubMed]
- Iida, M.; Ooi, T.; Kito, K.; Yoshida, S.; Kanoh, K.; Shizuri, Y.; Kusumi, T. Three new polyketide-terpenoid hybrids from Penicillium sp. Org. Lett. 2008, 10, 845–848. [Google Scholar] [CrossRef] [PubMed]
- Bhandari, R.; Eguchi, T.; Sekine, A.; Ohashi, Y.; Kakinuma, K.; Ito, M.; Mizoue, K. Structure of NG-061, a novel potentiator of nerve growth factor (NGF) isolated from Penicillium minioluteum F-4627. J. Antibiot. 1999, 52, 231–234. [Google Scholar] [CrossRef] [PubMed]
- Okada, H.; Kamiya, S.; Shiina, Y.; Suwa, H.; Nagashima, M.; Nakajima, S.; Shimokawa, H.; Sugiyama, E.; Kondo, H.; Kojiri, K.; et al. BE-31405, a new antifungal antibiotic produced by Penicillium minioluteum. I. Description of producing organism, fermentation, isolation, physico-chemical and biological properties. J. Antibiot. 1998, 51, 1081–1086. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.Y.; Zhang, Q.; Gao, Y.Q.; Zhang, A.L.; Gao, J.M. Miniolins A–C, novel isomeric furanones induced by epigenetic manipulation of Penicillium minioluteum. RSC Adv. 2015, 5, 2185–2190. [Google Scholar] [CrossRef]
- De Souza, G.G.; Oliveira, T.S.; Takahashi, J.A.; Collado, I.G.; Macias-Sanchez, A.J.; Hernandez-Galan, R. Biotransformation of clovane derivatives. Whole cell fungi mediated domino synthesis of rumphellclovane A. Org. Biomol. Chem. 2012, 10, 3315–3320. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, N.; Visagie, C.M.; Houbraken, J.; Frisvad, J.C.; Samson, R.A. Polyphasic taxonomy of the genus Talaromyces. Stud. Mycol. 2014, 78, 175–341. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.P.; Zhu, C.Y.; Zhang, C.P.; Chu, Y.S.; Wang, Y.L.; Zhang, J.X.; Wu, D.K.; Zhang, K.Q.; Niu, X.M. Thermolides, potent nematocidal PKS-NRPS hybrid metabolites from thermophilic fungus Talaromyces thermophilus. J. Am. Chem. Soc. 2012, 134, 20306–20309. [Google Scholar] [CrossRef] [PubMed]
- He, J.W.; Liang, H.X.; Gao, H.; Kuang, R.Q.; Chen, G.D.; Hu, D.; Wang, C.X.; Liu, X.Z.; Li, Y.; Yao, X.S. Talaflavuterpenoid A, a new nardosinane-type sesquiterpene from Talaromyces flavus. J. Asian Nat. Prod. Res. 2014, 16, 1029–1034. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Huang, H.; Shao, C.; Jiang, J.; Zhu, X.; Liu, Y.; Liu, L.; Lu, Y.; Li, M.; Lin, Y.; et al. Cytotoxic norsesquiterpene peroxides from the endophytic fungus Talaromyces flavus isolated from the mangrove plant Sonneratia apetala. J. Nat. Prod. 2011, 74, 1230–1235. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Ohlendorf, B.; Oesker, V.; Wiese, J.; Malien, S.; Schmaljohann, R.; Imhoff, J.F. Acetylcholinesterase inhibitors from a marine fungus Talaromyces sp. Strain LF458. Mar. Biotechnol. 2015, 17, 110–119. [Google Scholar] [CrossRef] [PubMed]
- King, T.J.; Roberts, J.C.; Thompson, D.J. Studies in mycological chemistry. Part XXX and last. Isolation and structure of purpuride, a metabolite of Penicillium purpurogenum Stoll. J. Chem. Soc. Perkin Trans. 1973, 1, 78–80. [Google Scholar] [CrossRef]
- Stierle, D.B.; Stierle, A.A.; Girtsman, T.; McIntyre, K.; Nichols, J. Caspase-1 and -3 inhibiting drimane sesquiterpenoids from the extremophilic fungus Penicillium solitum. J. Nat. Prod. 2012, 75, 262–266. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, Y.; Liu, P.; Wang, W.; Fan, Y.; Zhu, W. Purpurides B and C, two new sesquiterpene esters from the aciduric fungus Penicillium purpurogenum JS03-21. Chem. Biodivers. 2013, 10, 1185–1192. [Google Scholar] [CrossRef] [PubMed]
- Parsons, S.; Flack, H.D.; Wagner, T. Use of intensity quotients and differences in absolute structure refinement. Acta Cryst. 2013, B69, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Maurs, M.; Azerad, R.; Cortes, M.; Aranda, G.; Delahaye, M.B.; Ricard, L. Microbial hydroxylation of natural drimenic lactones. Phytochemistry 1999, 52, 291–296. [Google Scholar] [CrossRef]
- Zang, L.Y.; Wei, W.; Guo, Y.; Wang, T.; Jiao, R.H.; Ng, S.W.; Tan, R.X.; Ge, H.M. Sesquiterpenoids from the mangrove-derived endophytic fungus Diaporthe sp. J. Nat. Prod. 2012, 75, 1744–1749. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; Volume 18, pp. 315–322. [Google Scholar]
- Zhang, Z.; Schwartz, S.; Wagner, L.; Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 2000, 7, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Sheldrick, G.M. A short history of shelx. Acta Cryst. 2008, A64, 112–122. [Google Scholar] [CrossRef] [PubMed]
- The Cambridge Crystallographic Data Centre. Available online: http://www.ccdc.cam.ac.uk/data_request/cif (accessed on 29 May 2015).
- Chiablaem, K.; Lirdprapamongkol, K.; Keeratichamroen, S.; Surarit, R.; Svasti, J. Curcumin suppresses vasculogenic mimicry capacity of hepatocellular carcinoma cells through STAT3 and PI3K/AKT inhibition. Anticancer Res. 2014, 34, 1857–1864. [Google Scholar] [PubMed]
- Zhuravleva, O.I.; Afiyatullov, S.; Denisenko, V.A.; Ermakova, S.P.; Slinkina, N.N.; Dmitrenok, P.S.; Kim, N.Y. Secondary metabolites from a marine-derived fungus Aspergillus carneus Blochwitz. Phytochemistry 2012, 80, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Haga, A.; Tamoto, H.; Ishino, M.; Kimura, E.; Sugita, T.; Kinoshita, K.; Takahashi, K.; Shiro, M.; Koyama, K. Pyridone alkaloids from a marine-derived fungus, Stagonosporopsis cucurbitacearum, and their activities against azole-resistant Candida albicans. J. Nat. Prod. 2013, 76, 750–754. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, Z.; Ju, Z.; Wan, J.; Liao, S.; Lin, X.; Zhang, T.; Zhou, X.; Chen, H.; Tu, Z.; et al. Cytotoxic cytochalasins from marine-derived fungus Arthrinium arundinis. Planta Med. 2015, 81, 160–166. [Google Scholar] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ngokpol, S.; Suwakulsiri, W.; Sureram, S.; Lirdprapamongkol, K.; Aree, T.; Wiyakrutta, S.; Mahidol, C.; Ruchirawat, S.; Kittakoop, P. Drimane Sesquiterpene-Conjugated Amino Acids from a Marine Isolate of the Fungus Talaromyces minioluteus (Penicillium Minioluteum). Mar. Drugs 2015, 13, 3567-3580. https://doi.org/10.3390/md13063567
Ngokpol S, Suwakulsiri W, Sureram S, Lirdprapamongkol K, Aree T, Wiyakrutta S, Mahidol C, Ruchirawat S, Kittakoop P. Drimane Sesquiterpene-Conjugated Amino Acids from a Marine Isolate of the Fungus Talaromyces minioluteus (Penicillium Minioluteum). Marine Drugs. 2015; 13(6):3567-3580. https://doi.org/10.3390/md13063567
Chicago/Turabian StyleNgokpol, Suthatip, Wittaya Suwakulsiri, Sanya Sureram, Kriengsak Lirdprapamongkol, Thammarat Aree, Suthep Wiyakrutta, Chulabhorn Mahidol, Somsak Ruchirawat, and Prasat Kittakoop. 2015. "Drimane Sesquiterpene-Conjugated Amino Acids from a Marine Isolate of the Fungus Talaromyces minioluteus (Penicillium Minioluteum)" Marine Drugs 13, no. 6: 3567-3580. https://doi.org/10.3390/md13063567
APA StyleNgokpol, S., Suwakulsiri, W., Sureram, S., Lirdprapamongkol, K., Aree, T., Wiyakrutta, S., Mahidol, C., Ruchirawat, S., & Kittakoop, P. (2015). Drimane Sesquiterpene-Conjugated Amino Acids from a Marine Isolate of the Fungus Talaromyces minioluteus (Penicillium Minioluteum). Marine Drugs, 13(6), 3567-3580. https://doi.org/10.3390/md13063567