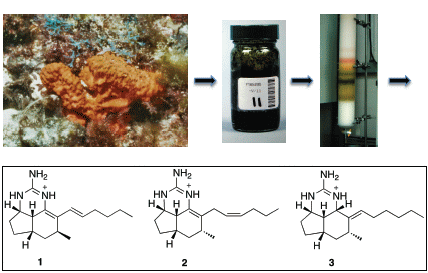
Tricyclic Guanidine Alkaloids from the Marine Sponge Acanthella cavernosa that Stabilize the Tumor Suppressor PDCD4
Abstract
:1. Introduction
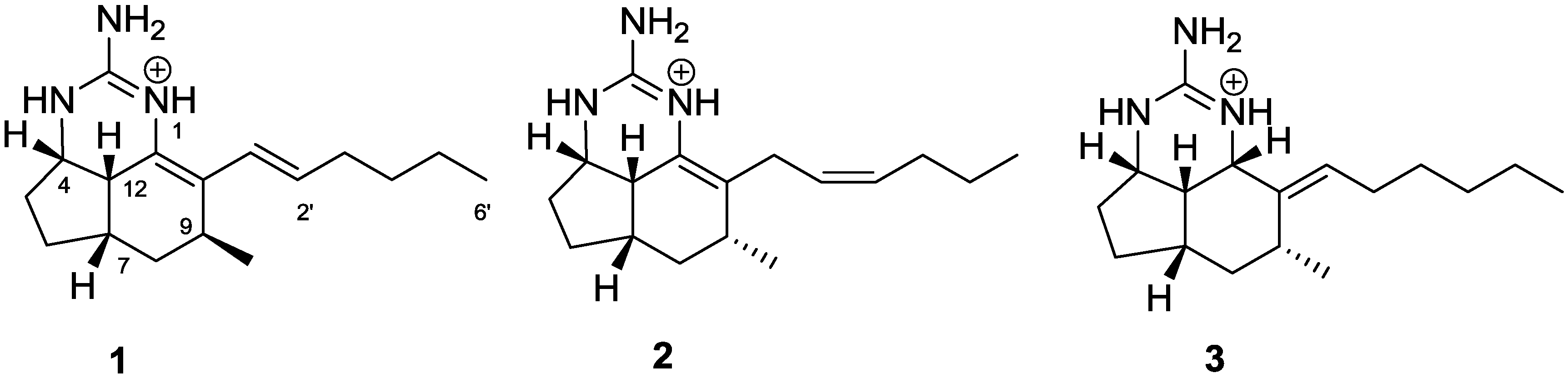
2. Results and Discussion
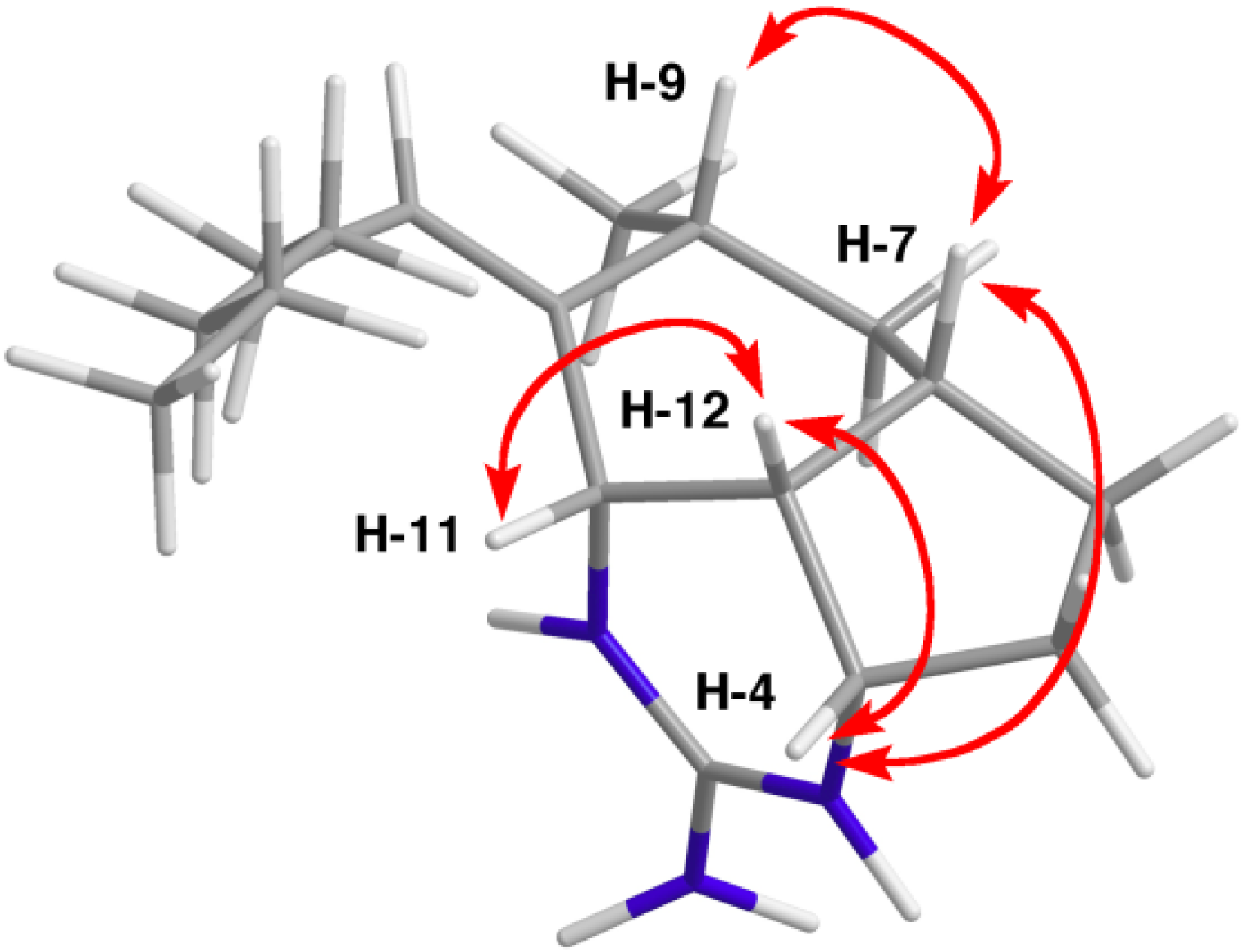
| Position | δC a, Type | δH, Mult. (J in Hz) b | HMBC c | ROESY |
|---|---|---|---|---|
| 1-N | 7.71, br s | |||
| 2 | 154.0, C | |||
| C2-NH2 | 7.00, br s | |||
| 3-N | 7.05, br s | |||
| 4 | 52.9, CH | 3.80, br t (3.9) | 2, 6, 7 | 5b, 7, 12 |
| 5a | 32.9, CH2 | 1.89, m | 4, 7, 12 | |
| 5b | 1.64, m | 4, 7, 12 | 4 | |
| 6a | 29.6, CH2 | 1.99, m | 4, 5, 12 | |
| 6b | 1.43, m | 4, 5, 12 | ||
| 7 | 34.6, CH | 2.05, m | 4,6,12 | 4, 8a, 9 |
| 8a | 37.2, CH2 | 1.84, ddd (13.5, 5.0, 5.0) | 7, 9, 9Me, 12 | 7, 8b, 9 |
| 8b | 1.10, ddd (13.5, 12.6, 12.6) | 6, 9, 9Me, 12 | 8a | |
| 9 | 30.0, CH | 2.50, m | 8, 10, 11, 1′ | 7 |
| 9-Me | 22.4, CH3 | 1.06 3H, d (6.8) | 8, 9, 10 | 8a |
| 10 | 138.1, C | |||
| 11 | 54.9, CH | 3.98, d (6.3) | 7, 9, 10, 12, 1′ | 1′, 12 |
| 12 | 39.6, CH | 2.31, m | 4, 7, 8, 11 | 4, 11 |
| 1′ | 132.2, CH | 5.42, br t (7.1) | 9, 11, 3′ | 11, 2′, 3′ |
| 2′ | 27.8 CH2 | 2.02, m | 1, 10, 3′ | |
| 3′ | 29.6, CH2 | 1.24, m | 4′, 5′ | |
| 4′ | 31.5, CH2 | 1.26, m | 3′, 5′ | |
| 5′ | 22.5, CH2 | 1.28, m | 6′, 4′ | |
| 6′ | 14.0, CH3 | 0.88, t (6.8) | 4′, 5′ |
| Compound | PDCD4 EC50 a (μg/mL) | PDCD4 EC90 a (μg/mL) | Cytotoxicity b (μg/mL) |
|---|---|---|---|
| 1 | 1.8 | - | 4.2 |
| 2 | 2.8 | 5.5 | 17.0 |
| 3 | n.a. c | - | 10.6 |
| Rapamycin d | 0.02 | 0.1 | >1.0 |
3. Experimental Section
3.1. General Experimental Procedures
3.2. Extraction and Isolation
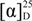 = +170 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.09), 260 (4.10) nm; 1H NMR (CHCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Supplementary Table S1; HRESIMS m/z [M + H]+ 274.2377 (calcd for C17H28N3, 274.2278). Trifluoroacetate salt: yellow oil;
= +170 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.09), 260 (4.10) nm; 1H NMR (CHCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Supplementary Table S1; HRESIMS m/z [M + H]+ 274.2377 (calcd for C17H28N3, 274.2278). Trifluoroacetate salt: yellow oil; 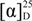 = +64 (c 0.325, CHCl3); HRESIMS m/z [M]+ 274.2270 (calcd for C17H28N3, 274.2278).
= +64 (c 0.325, CHCl3); HRESIMS m/z [M]+ 274.2270 (calcd for C17H28N3, 274.2278).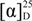 = +60 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.20), 230 (3.89) nm; ECD (MeOH) λmax (Δε) 204 (−2.9), 207 (0), 253 (0), 231 (+8.2), 288 (0) nm; 1H NMR (CHCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Supplementary Table S1; HRESIMS m/z [M]+ 274.2274 (calcd for C17H28N3, 274.2278).
= +60 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.20), 230 (3.89) nm; ECD (MeOH) λmax (Δε) 204 (−2.9), 207 (0), 253 (0), 231 (+8.2), 288 (0) nm; 1H NMR (CHCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Supplementary Table S1; HRESIMS m/z [M]+ 274.2274 (calcd for C17H28N3, 274.2278).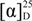 = +40 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.07), 240 (3.54) nm; 1H NMR (CDCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Table 1; HRESIMS m/z [M]+ 276.2439 (calcd for C17H30N3, 276.2434).
= +40 (c 0.1, CHCl3); UV (MeOH) λ (log ε) 202 (4.07), 240 (3.54) nm; 1H NMR (CDCl3, 600 MHz) and 13C NMR (CDCl3, 150 MHz) data, see Table 1; HRESIMS m/z [M]+ 276.2439 (calcd for C17H30N3, 276.2434).3.3. PDCD4 Assay
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Yang, H.S.; Jansen, A.P.; Komar, A.A.; Zheng, X.; Merrick, W.C.; Costes, S.; Lockett, S.J.; Sonenberg, N.; Colburn, N.H. The Transformation Suppressor PDCD4 is a Novel Eukaryotic Translation Initiation Factor 4A Binding Protein that Inhibits Translation. Mol. Cell Biol. 2003, 23, 26–37. [Google Scholar] [CrossRef]
- Yang, H.S.; Jansen, A.P.; Nair, R.; Shibahara, K.; Verma, A.K.; Cmarik, J.L.; Colburn, N.H. A Novel Transformation Suppressor, PDCD4, Inhibits AP-1 Transactivation but not NF-κB or ODC Transactivation. Oncogene 2001, 20, 669–676. [Google Scholar] [CrossRef]
- Leupold, J.H.; Yang, H.S.; Colburn, N.H.; Asangani, I.; Post, S.; Allgayer, H. Tumor Suppressor PDCD4 Inhibits Invasion/Intravasation and Regulates Urokinase Receptor (u-PAR) Gene Expression via Sp-transcription Factors. Oncogene 2007, 26, 4550–4562. [Google Scholar] [CrossRef]
- Nieves-Alicea, R.; Colburn, N.H.; Simeone, A.M.; Tari, A.M. Programmed Cell Death 4 Inhibits Breast Cancer Cell Invasion by Increasing Tissue Inhibitor of Metalloproteinases-2 Expression. Breast Cancer Res. Treat. 2009, 114, 203–209. [Google Scholar] [CrossRef]
- Jansen, A.P.; Camalier, C.E.; Colburn, N.H. Epidermal Expression of the Translation Inhibitor Programmed Cell Death 4 Suppresses Tumorigenesis. Cancer Res. 2005, 65, 6034–6041. [Google Scholar] [CrossRef]
- Asangani, I.A.; Rasheed, S.A.; Nikolova, D.A.; Leupold, J.H.; Colburn, N.H.; Post, S. MicroRNA-21 (miR-21) Post-transcriptionally Downregulates Tumor Suppressor PDCD4 and Stimulates Invasion, Intravasation and Metastasis in Colorectal Cancer. Oncogene 2008, 27, 2128–2136. [Google Scholar]
- Chen, Y.; Liu, W.; Chao, T.; Zhang, Y.; Yan, X.; Gong, Y.; Qiang, B.; Yuan, J.; Sun, M.; Peng, X.; et al. MicroRNA-21 Down-regulates the Expression of Tumor Suppressor PDCD4 in Human Glioblastoma Cell T98G. Cancer Lett. 2008, 272, 197–205. [Google Scholar] [CrossRef]
- Wang, X.; Wei, Z.; Gao, F.; Zhang, X.; Zhou, C.; Zhu, F.; Wang, Q.; Gao, Q.; Ma, C.; Sun, W.; et al. Expression and Prognostic Significance of PDCD4 in Human Epithelial Ovarian Carcinoma. Anticancer Res. 2008, 28, 2991–2996. [Google Scholar]
- Zhang, H.; Ozaki, I.; Mizuta, T.; Hamajima, H.; Yasutake, T.; Eguchi, Y.; Ideguchi, H.; Yamamoto, K.; Matsuhashi, S. Involvement of Programmed Cell Death 4 in Transforming Growth Factor-β 1-Induced Apoptosis in Human Hepatocellular Carcinoma. Oncogene 2006, 25, 6101–6112. [Google Scholar] [CrossRef]
- Dorrello, N.V.; Peschiaroli, A.; Guardavaccaro, D.; Colburn, N.H.; Sherman, N.E.; Pagano, M. S6K1- and βTRCP-Mediated Degradation of PDCD4 Promotes Protein Translation and Cell Growth. Science 2006, 314, 467–471. [Google Scholar] [CrossRef]
- Schmid, T.; Jansen, A.P.; Baker, A.R.; Hegamyer, G.; Hagan, J.P.; Colburn, N.H. Translation Inhibitor PDCD4 is Targeted for Degradation during Tumor Promotion. Cancer Res. 2008, 68, 1254–1260. [Google Scholar]
- Schmid, T.; Bajer, M.M.; Blees, J.S.; Eifler, L.K.; Milke, L.; Rübsamen, D.; Schulz, K.; Weigert, A.; Baker, A.R.; Colburn, N.H.; et al. Inflammation-induced Loss of PDCD4 is Mediated by Phosphorylation-Dependent Degradation. Carcinogenesis 2011, 32, 1427–1433. [Google Scholar] [CrossRef]
- Zhao, L.-X.; Huang, S.-X.; Tang, S.-K.; Jiang, C.-L.; Duan, Y.; Beutler, J.A.; Henrich, C.J.; McMahon, J.B.; Schmid, T.; Blees, J.S.; et al. Actinopolysporins A-C and Tubercidin as a PDCD4 Stabilizer from the Halophilic Actinomycete Actinopolyspora erythraea YIM 90600. J. Nat. Prod. 2011, 74, 1990–1995. [Google Scholar] [CrossRef]
- Blees, J.S.; Schmid, T.; Thomas, C.L.; Baker, A.R.; Benson, L.; Evans, J.R.; Goncharova, E.I.; Colburn, N.H.; McMahon, J.B.; Henrich, C.J.; et al. Development of a High-throughput Cell-based Reporter Assay to Identify Stabilizers of Tumor Suppressor PDCD4. J. Biomol. Screen. 2010, 15, 21–29. [Google Scholar]
- Grkovic, T.; Blees, J.S.; Colburn, N.H.; Schmid, T.; Thomas, C.L.; Henrich, C.J.; McMahon, J.B.; Gustafson, K.R. Cryptocaryols A–H, A-Pyrone-Containing 1,3-Polyols from Cryptocarya Sp. Implicated in Stabilizing The Tumor Suppressor PDCD4. J. Nat. Prod. 2011, 74, 1015–1020. [Google Scholar] [CrossRef]
- Blees, J.S.; Bokesch, H.R.; Rübsamen, D.; Schulz, K.; Milke, L.; Bajer, M.M.; Gustafson, K.R.; Henrich, C.J.; McMahon, J.B.; Colburn, N.H.; et al. Erioflorin Stabilizes the Tumor Suppressor PDCD4 by Inhibiting its Interaction with the E3-ligase -TrCP1. PLoS One 2012, 7, e46567. [Google Scholar]
- Bajer, M.M.; Kunze, M.M.; Blees, J.S.; Bokesch, H.R.; Chen, H.; Dong, Z.; Gustafson, K.R.; Biondi, R.M.; Henrich, C.J.; McMahon, J.B.; et al. Characterization of Pomiferin Triacetate as a Novel mTOR and Translation Inhibitor. Biochem. Pharmacol. 2014, 88, 313–321. [Google Scholar] [CrossRef]
- Capon, R.J.; Miller, M.; Rooney, F. Mirabilin G: A New Alkaloid from a Southern Australian Marine Sponge, Clathria species. J. Nat. Prod. 2001, 64, 643–644. [Google Scholar] [CrossRef]
- El-Naggar, M.; Conte, M.; Capon, R.J. Mirabilins Revisited: Polyketide Alkaloids from a Southern Australian Marine Sponge, Clathria sp. Org. Biomol. Chem. 2010, 8, 407–412. [Google Scholar]
- Gros, E.; Al-Mourabit, A.; Martin, M.-T.; Sorres, J.; Vacelet, J.; Frederich, M.; Aknin, M.; Kashman, Y.; Gauvin-Bialecki, A. Netamines H-N, Tricyclic Alkaloids from the Marine Sponge Biemna laboutei and their Antimalarial Activity. J. Nat. Prod. 2014. [Google Scholar] [CrossRef]
- Harbour, G.C.; Tymiak, A.A.; Rinehart, K.L.; Shaw, P.D.; Hughes, R.G.; Mizsak, S.A.; Coats, J.H.; Zurenko, G.E.; Li, L.H.; Kuentzel, S.L.; et al. Ptilocaulin and Isoptilocaulin, Antimicrobial and Cytotoxic Cyclic Guanidines from the Caribbean Sponge Ptilocaulis aff. P. spiculifer (Lamarck, 1814). J. Am. Chem. Soc. 1981, 103, 5604–5606. [Google Scholar] [CrossRef]
- Tavares, R.; Daloze, D.; Braekman, J.C.; Hajdu, E.; van Soest, R.W.M. 8β-Hydroxyptilocaulin, a New Guanidine Alkaloid from the Sponge Monfloata arbuscula. J. Nat. Prod. 1995, 58, 1139–1142. [Google Scholar] [CrossRef]
- Patil, A.D.; Freyer, A.J.; Offen, P.; Bean, M.F.; Johnson, R.K. Three New Tricyclic Guanidine Alkaloids from the Sponge Batzella sp. J. Nat. Prod. 1997, 60, 704–707. [Google Scholar] [CrossRef]
- Sorek, H.; Rudi, A.; Gueta, S.; Reyes, F.; Martin, M.J.; Aknin, M.; Gaydou, E.; Vacelet, J.; Kashman, Y. Netamines A–G: Seven New Tricyclic Guanidine Alkaloids from the Marine Sponge Biemna laboutei. Tetrahedron 2006, 62, 8838–8843. [Google Scholar] [CrossRef]
- Barrow, R.A.; Murray, L.M.; Lim, T.K.; Capon, R.J. Mirabilins (A–F): New Alkaloids from a Southern Australian Marine Sponge, Arenochalina mirabilis. Aust. J. Chem. 1996, 49, 767–773. [Google Scholar]
- McCloud, T.G. High Throughput Extraction of Plant, Marine and Fungal Specimens for Preservation of Biologically Active Molecules. Molecules 2010, 15, 4526–4563. [Google Scholar] [CrossRef]
- Wang, Y.; O’Doherty, G.A. Cryptocaryols A and B: Total Synthesis, Stereochemical Revision, Structural Elucidation, and Structure-Activity Relationship. J. Am. Chem. Soc. 2013, 135, 9334–9337. [Google Scholar] [CrossRef]
- Reddy, D.S.; Mohapatra, D.K. Total Synthesis and Structure Confirmation of Cryptocaryol A. Eur. J. Org. Chem. 2013, 2013, 1051–1057. [Google Scholar] [CrossRef]
- Cuccarese, M.F.; Wanf, Y.; Beuning, P.J.; O’Doherty, G.A. Cryptocaryol Structure-activity Relationship Study of Cancer Cell Cytotoxicity and Ability to Stabilize PDCD4. ACS Med. Chem. Lett. 2014, 5, 522–526. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Grkovic, T.; Blees, J.S.; Bayer, M.M.; Colburn, N.H.; Thomas, C.L.; Henrich, C.J.; Peach, M.L.; McMahon, J.B.; Schmid, T.; Gustafson, K.R. Tricyclic Guanidine Alkaloids from the Marine Sponge Acanthella cavernosa that Stabilize the Tumor Suppressor PDCD4. Mar. Drugs 2014, 12, 4593-4601. https://doi.org/10.3390/md12084593
Grkovic T, Blees JS, Bayer MM, Colburn NH, Thomas CL, Henrich CJ, Peach ML, McMahon JB, Schmid T, Gustafson KR. Tricyclic Guanidine Alkaloids from the Marine Sponge Acanthella cavernosa that Stabilize the Tumor Suppressor PDCD4. Marine Drugs. 2014; 12(8):4593-4601. https://doi.org/10.3390/md12084593
Chicago/Turabian StyleGrkovic, Tanja, Johanna S. Blees, Magdalena M. Bayer, Nancy H. Colburn, Cheryl L. Thomas, Curtis J. Henrich, Megan L. Peach, James B. McMahon, Tobias Schmid, and Kirk R. Gustafson. 2014. "Tricyclic Guanidine Alkaloids from the Marine Sponge Acanthella cavernosa that Stabilize the Tumor Suppressor PDCD4" Marine Drugs 12, no. 8: 4593-4601. https://doi.org/10.3390/md12084593
APA StyleGrkovic, T., Blees, J. S., Bayer, M. M., Colburn, N. H., Thomas, C. L., Henrich, C. J., Peach, M. L., McMahon, J. B., Schmid, T., & Gustafson, K. R. (2014). Tricyclic Guanidine Alkaloids from the Marine Sponge Acanthella cavernosa that Stabilize the Tumor Suppressor PDCD4. Marine Drugs, 12(8), 4593-4601. https://doi.org/10.3390/md12084593