Group Theory of Syntactical Freedom in DNA Transcription and Genome Decoding
Abstract
1. Introduction
2. Materials and Methods
2.1. Finitely Generated Groups
2.2. Free Groups and Their Conjugacy Classes
2.3. Content of the Paper
3. Results
3.1. The TATA Box, the Hecke Groups and More
3.2. Gilbert’s Syndrome
3.3. Single Nucleotide Polymorphism
3.4. A Few DNA/Protein Complexes and Their Transcription Factors
3.4.1. Immediate Early Genes and Their Motifs
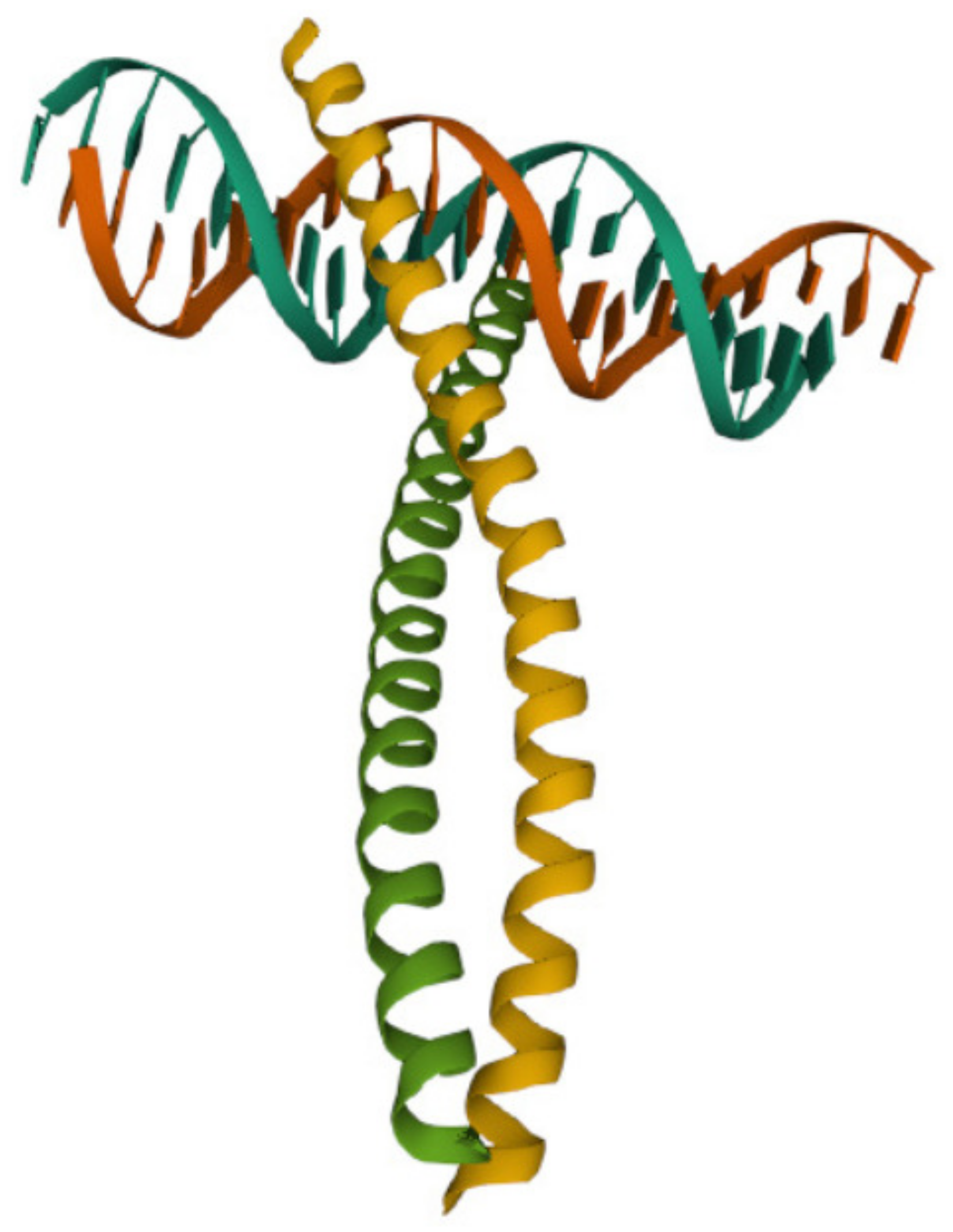
3.4.2. The DNA-Binding Domain Fos
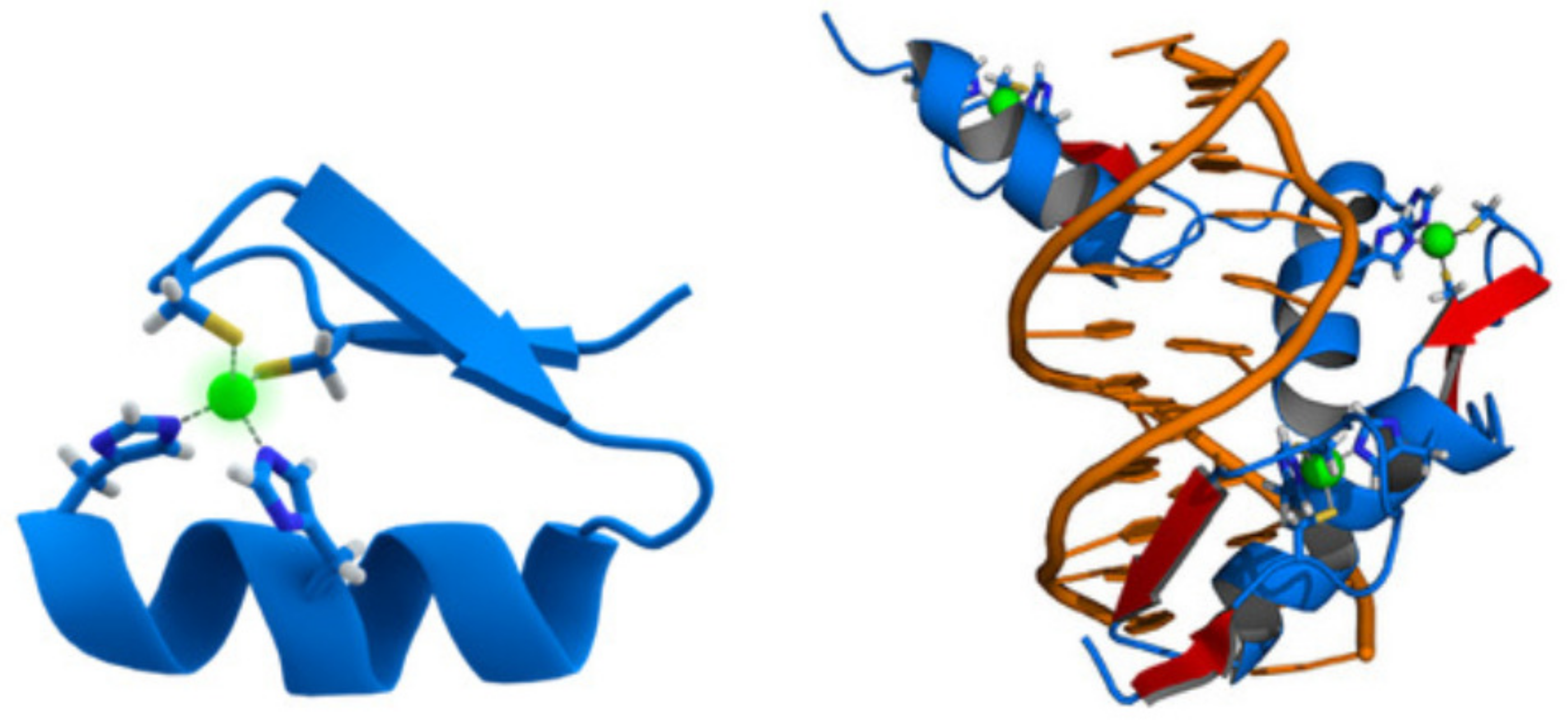
3.4.3. The DNA-Binding Domain EGR1
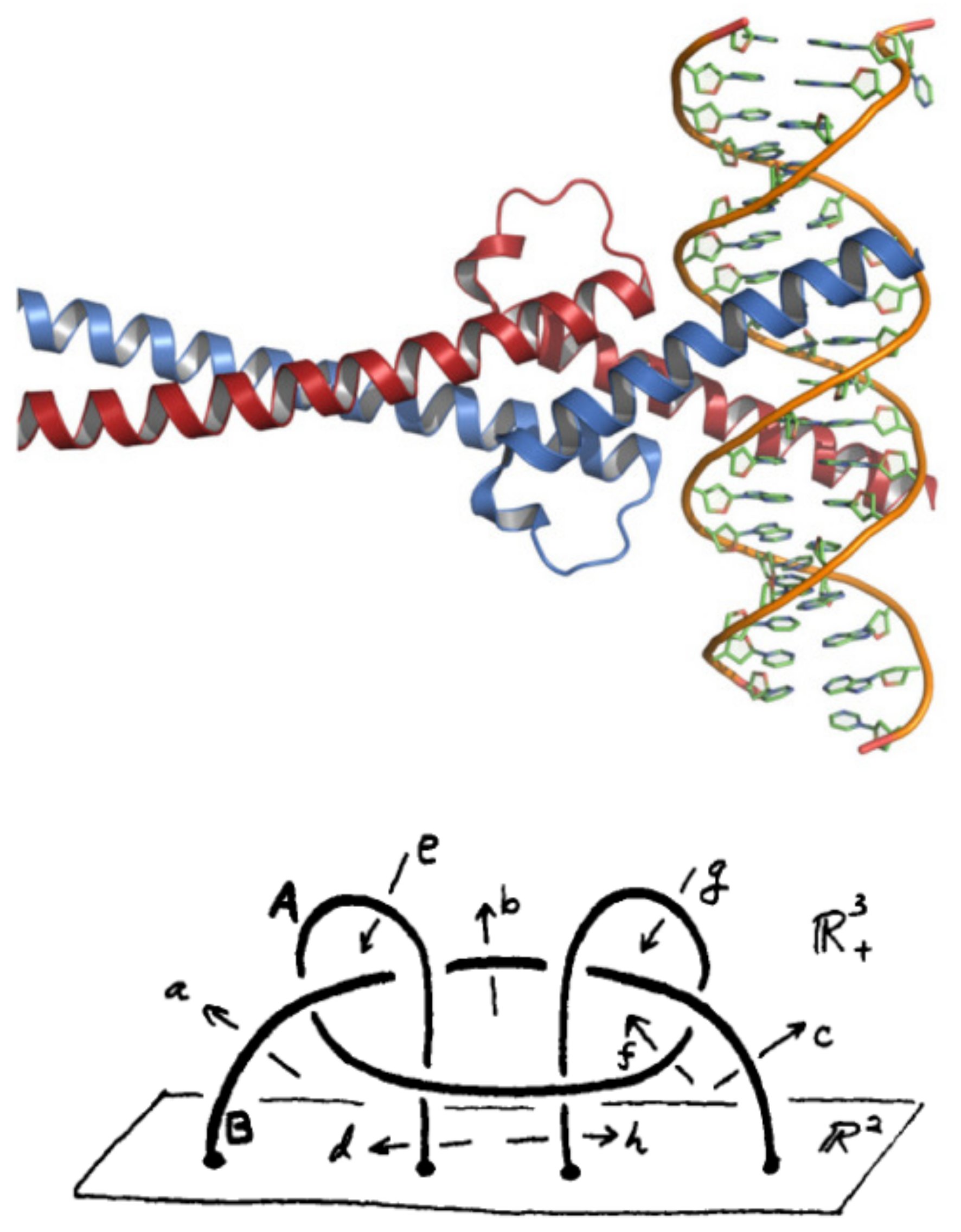
3.4.4. The DNA-Binding Domain Myc
3.5. Genes Whose Transcription Factors Have a Group Structure Away from a Free Group
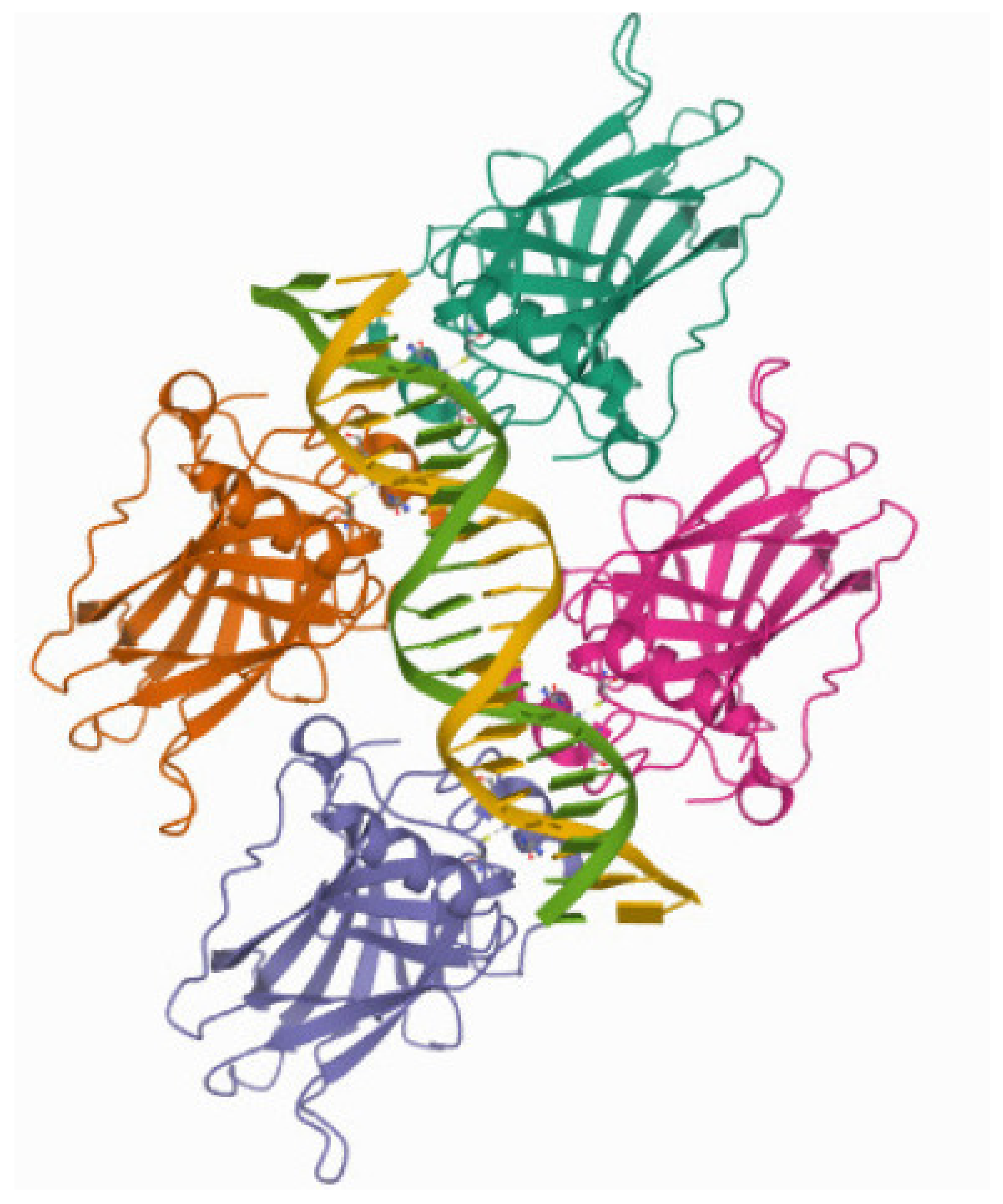
The DNA-Binding Domain of p53
4. Discussion
4.1. Aperiodicity of Substitutions
4.1.1. A Two-Letter Sequence for the Transcription Factor of Gene DBX in Drosophila Melanogaster
4.1.2. A Three-Letter Sequence for the Transcription Factor of Gene EGR1
4.1.3. A Four-Letter Sequence for the Transcription Factor of the Fos Gene
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Irwin, K. The code-theoretic axiom; the third ontology. Rep. Adv. Phys. Sci. 2019, 3, 39. [Google Scholar] [CrossRef]
- Planat, M.; Aschheim, R.; Amaral, M.M.; Fang, F.; Irwin, K. Quantum information in the protein codes, 3-manifolds and the Kummer surface. Symmetry 2020, 13, 1146. [Google Scholar] [CrossRef]
- Planat, M.; Aschheim, R.; Amaral, M.M.; Fang, F.; Irwin, K. Graph coverings for investigating non local structures in protein, music and poems. Science 2021, 3, 39. [Google Scholar] [CrossRef]
- Lambert, S.A.; Jolma, A.; Campitelli, L.F.; Das, P.K.; Yin, Y.; Albu, M.; Chen, X.; Talpale, J.; Hughes, T.R.; Weirauch, M.T. The human transcription factors. Cell 2018, 172, 650–665. Available online: http://www.edgar-wingender.de/huTF_classification.html (accessed on 1 September 2021).
- Wingender, E.; Schoeps, T.; Dönitz, J. TFClass: An expandable hierarchical classification of human transcription factors. Nucleic Acids Res. 2013, T1, D165–D170. [Google Scholar] [CrossRef] [PubMed]
- Sandelin, A.; Alkema, W.; Engström, P.; Wasserman, W.W.; Lenhard, B. JASPAR: An open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 2004, 32, D91–D94. Available online: https://jaspar.genereg.net/ (accessed on 1 September 2021). [CrossRef]
- Planat, M.; Giorgetti, A.; Holweck, F. Saniga, M. Quantum contextual finite geometries from dessins d’enfants. Int. J. Geom. Meth. Mod. Phys. 2015, 12, 1550067. [Google Scholar] [CrossRef]
- Hall, M., Jr. Subgroups of finite index in free groups. Can. J. Math. 1949, 1, 187–190. [Google Scholar] [CrossRef]
- Kwak, J.H.; Nedela, R. Graphs and their coverings. Lect. Notes Ser. 2007, 17, 118. [Google Scholar]
- The Modular Group. Available online: https://en.wikipedia.org/wiki/Modular_group (accessed on 1 October 2021).
- Suzuki, Y.; Tsunoda, T.; Sese, J.; Taira, H.; Mizushima-Sugano, J.; Hata, H.; Ota, T.; Isogai, T.; Tanaka, T.; Nakamura, Y.; et al. Identification and characterization of the potential promoter regions of 1031 kinds of human genes. Genome Res. 2001, 11, 677–684. [Google Scholar] [CrossRef]
- TATA Box. Available online: https://en.wikipedia.org/wiki/TATA_box (accessed on 1 September 2021).
- Wang, Y.; Jensen, R.C.; Stumph, W.E. Role of TATA box sequence and orientation in determining RNA polymerase II/III transcription specificity. Nucleic Acids Res. 1996, 24, 3100–3106. [Google Scholar] [CrossRef][Green Version]
- Li, Y.; Buckley, D.; Wang, S.; Klaassen, C.D.; Zhong, X.B. Phenobarbital-Responsive Enhancer Module of the UGT1A1. Drug Metab. Disp. 2009, 37, 1978–1986. [Google Scholar] [CrossRef] [PubMed]
- Chadaeva, I.V.; Ponomarenko, P.M.; Rasskazov, D.A.; Sharypova, E.B.; Kashina, E.V.; Zhechev, D.A.; Drachkova, I.A.; Arkova, O.V.; Savinkova, L.K.; Ponomarenko, M.P.; et al. Candidate SNP markers of reproductive potential are predicted by a significant change in the affinity of TATA-binding protein for human gene promoters. BMC Genom. 2018, 19, 16–38. [Google Scholar] [CrossRef] [PubMed]
- Zerbino, D.R.; Wilder, S.P.; Johnson, N.; Juettemann, T.; Flicek, P.R. The ensembl Regulatory Build. Genome Biol. 2015, 16, 56. [Google Scholar] [CrossRef]
- Hodgson, C.D.; Weeks, J.R. Symmetries, Isometries and length spectra of closed hyperbolic three-manifolds. Exp. Math. 1994, 3, 261–274. [Google Scholar] [CrossRef]
- Gallo, F.T.; Katche, C.; Morici, J.F.; Medina, J.H.; Weisstaub, N.V. Immediate early genes, memory and psychiatric disorders: Focus on c-Fos, Egr1 and Arc. Front. Behav. Neurosci. 1998, 12, 79. [Google Scholar] [CrossRef] [PubMed]
- Glover, J.N.; Harrison, S.C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos-c-Jun bound to DNA. Nature 1995, 373, 257–261. [Google Scholar] [CrossRef]
- Hashimoto, H.; Olanrewaju, Y.; Zheng, Y.; Wilson, G.G.; Zhang, X.; Cheng, X. Wilms tumor protein recognizes 5-carboxylcytosine within a specific DNA sequence. Genes Dev. 2019, 28, 2304–2313. [Google Scholar] [CrossRef]
- Nair, S.K.; Burley, S.K. X-ray structures of Myc-Max and Mad-Max recognizing DNA: Molecular bases of regulation by proto-oncogenic transcription factors. Cell 2003, 112, 193–205. [Google Scholar] [CrossRef]
- Zeeman, E.C. Linking spheres. Abh. Math. Sem. Univ. Hamburg 1960, 24, 149–153. [Google Scholar] [CrossRef]
- Rolfsen, D. Knots and Links; AMS Chelsea Publishing: Providence, RI, USA, 2000. [Google Scholar]
- Schaeffer, L.N.; Huchet-Dymanus, M.; Changeux, J.P. Implication of a multisubunit Ets-related transcription factor in synaptic expression of the nicotinic acetylcholine receptor. EMBO J. 1998, 17, 3078–3090. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Grimm, S.A.; Bushel, P.R.; Li, J.; Li, Y.; Bennett, B.D.; Lavender, C.A.; Ward, J.M.; Fargo, D.C.; Anderson, C.W.; et al. Revealing a human p53 universe. Nucl. Acids Res. 2018, 46, 8153–8167. [Google Scholar] [CrossRef]
- Nakamivhi, N.; Yoneda, Y. Transcription factors and drugs in the brain. Jpn J. Pharmacol. 2002, 89, 337–348. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, Y.; Zhang, X.; Dantas Machado, A.C.; Ding, Y.; Chen, Z.; Qin, P.Z.; Rohs, R.; Chen, L. Structure of p53 binding to the BAX response element reveals DNA unwinding and compression to accommodate base-pair insertion. Nucleic Acids Res. 2013, 41, 8368–8376. [Google Scholar] [CrossRef] [PubMed]
- Baake, M.; Grimm, U. Aperiodic Order, Volume I: A Mathematical Invitation; Cambridge University Press: Cambridge, UK, 2013. [Google Scholar]
- Planat, M.; Aschheim, R.; Amaral, M.M.; Fang, F.; Irwin, K. Complete quantum information in the DNA genetic code. Symmetry 2020, 12, 1993. [Google Scholar] [CrossRef]
- Planat, M.; Chester, D.; Aschheim, R.; Amaral, M.M.; Fang, F.; Irwin, K. Finite groups for the Kummer surface: The genetic code and quantum gravity. Quantum Rep. 2021, 3, 68–79. [Google Scholar] [CrossRef]
- Grandy, J.K. The three neurogenetic phases of human consciousness. J. Conscious Evol. 2018, 9, 24. [Google Scholar]
- Changeux, J.P. Allosteric receptors: From electric organ to cognition. Annu. Rev. Pharmacol. 2010, 50, 1–38. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, T.E.; Mallatt, J. The evolutionary and genetic origin of consciousness in the Cambrian Period over 500 million years ago. Front. Psychol. 2013, 4, 667. [Google Scholar] [CrossRef]
- Amaral, M.M.; Fang, F.; Hammock, D.; Irwin, K. Geometric state sum models from quasicrystals. Foundations 2021, 1, 155–168. [Google Scholar] [CrossRef]
- Amaral, M.M.; Fang, F.; Aschheim, R.; Irwin, K. On the Emergence of Space Time and Matter from Model Sets. Preprint 2021, 2021110359. [Google Scholar] [CrossRef]
| r | d = 1 | d = 2 | d = 3 | d = 4 | d = 5 | d = 6 | d = 7 |
|---|---|---|---|---|---|---|---|
| 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 2 | 1 | 3 | 7 | 26 | 97 | 624 | 4163 |
| 3 | 1 | 7 | 41 | 604 | 13,753 | 504,243 | 24,824,785 |
| 4 | 1 | 15 | 235 | 14,120 | 1,712,845 | 371,515,454 | 127,635,996,839 |
| 5 | 1 | 31 | 1361 | 334,576 | 207,009,649 | 268,530,771,271 | 644,969,015,852,641 |
| Rel: Cons Seq | Card Struct of cc of Subgroups | Group | Literature |
|---|---|---|---|
| TATAAAA | [6] (MA0108.1) | ||
| TATAAAAA | [13] | ||
| A(TA)TAA | [14] | ||
| A(TA)TAA | . | ||
| A(TA)TAA | . | ||
| A(TA)TAA | . |
| Gene | Rel: Marker | Card Seq of cc of Subgroups | Literature |
|---|---|---|---|
| ESR2 | TTAAAAGGAA | Table 1 in [15], B | |
| HSD17B1 | AGCCCAGAGC | ., A | |
| . | CAAGCCCAGA | ., A | |
| PGR | AAAGGAGCCG | ., A | |
| GSTM3 | GGGTATAAAG | ., E | |
| . | CCCCTCCCGC | ., C | |
| . | CCCTCCCGCT | . | ., C |
| IL1B | AAAACAGCGA | Table 2 in [15], A | |
| CYP2A6 | AAAGGCAAC | ., A | |
| DHFR | GGGACGAGGG | ., A | |
| . | GGACGAGGGG | . | ., A |
| LEP | GGGGCGGGA | Table 3 in [15], C | |
| GCG | TGCGCCTTGG | ., B | |
| GH1 | TATAAAAAGG | ] | ., E |
| . | GTATAAAAAG | . | ., D |
| . | GGTATAAAAA | . | ., E |
| . | AGGGCCCACA | ., A | |
| . | AAAGGGCCCC | ., A | |
| . | AAAGGGCCA | . | ., A |
| NOS2 | TCTTGGCTGC | Table 4 in [15], A | |
| TPI1 | ATATAAGTGG | ., B | |
| GJA5 | TATTAAACAC | ., E | |
| HBD | AAAAGGCAGG | Table 5 in [15], A | |
| F2 | AACCCAGAGG | ., A | |
| F8 | GGAAGAGGGA | * | Table 6 in [15], A |
| F3 | GCGCGGGGCA | ., A | |
| F11 | TTTTTAGTAA | . | ., D |
| . | TTTTTAGTAA | ., A | |
| . | AAGGAAATTT | ., A | |
| AR | GTGGAAGATT | Table 7 in [15], A | |
| . | CCACGACCCG | ., D | |
| MTHFR | TCCCTCCCA | ., A | |
| DMNT1 | TGTGTGGCCCG | . | ., A |
| . | GTGTGTGCCC | . | ., A |
| . | GACGAGCCCA | ., A | |
| NR5A1 | ACAAGAGAAA | ., A | |
| . | GGTGTGAGAG | ., A |
| Gene | Rel: Motif | Card Seq | Literature |
|---|---|---|---|
| Fos | TGAGTCA | [19] | |
| . | TGACTCA | [6], MA MA0099.2 | |
| EGR1 | GCGTGGGCG | [6], MA0162.1 | |
| . | CCGCCCCCG | ., MA0162.2 | |
| . | CCGCCCCCGC | ., . | |
| . | ACGCCCACGCA | ., MA0162.3 | |
| . | GGCCCACGC | . | ., MA0162.4 |
| EGR2 | CCGCCCACGC | . | ., MA0472.1 |
| . | ACGCCCACGCA | . | ., MA0472.2 |
| EGR3,EGR4 | ACGCCCACGCA | . | ., [ MA0732.1, MA0733.1] |
| Myc | CACGTG | [19] | |
| . | CGCACGTGGT | . | [6], MA0147.1 |
| . | CCCACGTGCTT | . | ., MA0147.2 |
| . | CCACGTGC | . | ., MA0147.3 |
| Mycn, Max::Myc, etc | GACCACGTGGT, etc. | . | ., [MA0104.1, etc.] |
| Gene | Rel: Motif | Card Seq | Literature |
|---|---|---|---|
| NKX6-2 | TAATTAA | [6], [MA0675.1, MA0675.2] | |
| HoxA1, HoxA2 | TAATTA | [6], [MA1495.1, MA0900.1] | |
| POU6F1, Vax | . | . | ., [MAO628.1, MA0722.1] |
| RUNX1 | TGTGGT | . | ., MA0511.1 |
| RUNX1 | TGTGGTT | [6], MA0002.2 | |
| EHF | CCTTCCTC | . | ., MA0598.1 |
| POU6F1 | TAATGAG | [6] MA1549.1 | |
| PITX2 | TAATCCC | . | ., [MA1547.1, MA1547.2] |
| ELK4 | CTTCCGG | . | ., MA0076.2 |
| OTX2, Dmbx1 | GGATTA | [6], [MA0712.2, MA0883.1] | |
| PitX1, PitX2, PitX3, OTX1 | TAATCC | . | .,[MA0682.1, MA0711.1] |
| N-box | TTCCGG | . | [24] |
| p53 | CACATGTCCA | [25] | |
| GZF1 | TGCGCGTCTATA | . | [4] |
| NF-kappa-B | GGGAATTTCC | . | [6], [MA0107.1, MA1911.1] |
| STAT1 | TTTCCCGGAA | . | ., MA0137.2 |
| . | TTCCAGGAA | . | ., MA0137.3 |
| STAT4 | TTCCAGGAAA | . | ., MA0518.1 |
| FOSL1::Jun | ATGACGTCAT | [6], MA1129.1 | |
| USF2 | GTCATGTGACC | . | . , MA0626.1 |
| PAX1 | CGTCACGCATGA | . | . , MA0779.1 |
| STAT2 | TTCCAGGAAG | . | . , MA0144.1 |
| FOS | GATGACGTCATCA | [6], MA1951.1 | |
| MAFA, MAFF,MAFK | TGCTGAGTCAGCA | . | ., [MA1521.1, MA0495.2, MA0946.2] |
| CREB | TGACGTCA | [6], [MA0018.2, MA018.3] | |
| USF2 | GGTCACGTGACC | . | ., MA0526.4 |
| SMAD3, SMAD5 | GTCTAGAC | . | ., [MA0795.1, MA1557.1], [26] |
| Gene | Type | Function | Dysfunction |
|---|---|---|---|
| NKX6-2 | homeobox | central nervous system, pancreas | spastic ataxia |
| HoxA1 | homeobox | embryonic devt of face and heart | autism |
| HoxA2 | . | . | cleft palate |
| Pou6F1 | . | neuroendocrine system | clear cell adenocarcinoma |
| Vax | . | forebrain development | craniofacial malform. |
| RunX1 | Runt-related | cell differentiation, pain neurons | myeloid leukemia |
| EHF | homeobox | epithelial expression | carcinogenesis, asthma |
| PitX2 | . | eye, tooth, abdominal organs | Axenfeld–Rieger syndrome |
| ELK4 | Ets-related | serum response for c-Fos | |
| OTX1,OTX2 | homeobox | brain and sensory organ devt | medulloblastomas |
| Dmbx1 | . | . | farsightedness and strabismus |
| PitX1 | . | organ devt, left–right asymmetry | autism, club foot |
| PitX3 | . | lens formation in eye | congenital cataracts |
| N-box | Ets-related | synaptic expression | drug sensitivity |
| p53 | p53 domain | ‘Guardian of the genome’ | cancers |
| GZF1 | Zinc fingers | protein coding | short stature, myopia |
| NF-kappa-B | . | DNA transcription, cytokines | apoptosis |
| STAT1 | Stat family | signal activator of transcription | immunodeficiency 31 |
| STAT4 | Stat family | signal activator of transcription | rheumatoid arthritis |
| FOSL1::Jun | leucine zipper | cellular proliferation | marker of cancer |
| USF2 | helix-loop-helix | transcription activator | |
| PAX1 | paired box | fetal development | Klippel–Feil syndrome |
| FOS | leucine zipper | cellular proliferation | cancers |
| Maf | . | pancreatic development | congenital cerulean cataract |
| CREB | bZIP | neuronal plasticity | Alzheimer’s disease |
| USF2 | helix-loop-helix | transcription activator | |
| SMAD | homeo domain | cell development and growth | Alzheimer’s disease |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Planat, M.; Amaral, M.M.; Fang, F.; Chester, D.; Aschheim, R.; Irwin, K. Group Theory of Syntactical Freedom in DNA Transcription and Genome Decoding. Curr. Issues Mol. Biol. 2022, 44, 1417-1433. https://doi.org/10.3390/cimb44040095
Planat M, Amaral MM, Fang F, Chester D, Aschheim R, Irwin K. Group Theory of Syntactical Freedom in DNA Transcription and Genome Decoding. Current Issues in Molecular Biology. 2022; 44(4):1417-1433. https://doi.org/10.3390/cimb44040095
Chicago/Turabian StylePlanat, Michel, Marcelo M. Amaral, Fang Fang, David Chester, Raymond Aschheim, and Klee Irwin. 2022. "Group Theory of Syntactical Freedom in DNA Transcription and Genome Decoding" Current Issues in Molecular Biology 44, no. 4: 1417-1433. https://doi.org/10.3390/cimb44040095
APA StylePlanat, M., Amaral, M. M., Fang, F., Chester, D., Aschheim, R., & Irwin, K. (2022). Group Theory of Syntactical Freedom in DNA Transcription and Genome Decoding. Current Issues in Molecular Biology, 44(4), 1417-1433. https://doi.org/10.3390/cimb44040095







