Matrix Metalloproteinases in Human Decidualized Endometrial Stromal Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. ESCs Collection
2.3. Human ESC Culture and Treatment with Steroid Hormones
2.4. siRNA-Meditated HAND2 Silencing
2.5. Quantitative Reverse Transcription PCR
2.6. Immunoblot Analysis
2.7. Statistical Analyses
3. Results
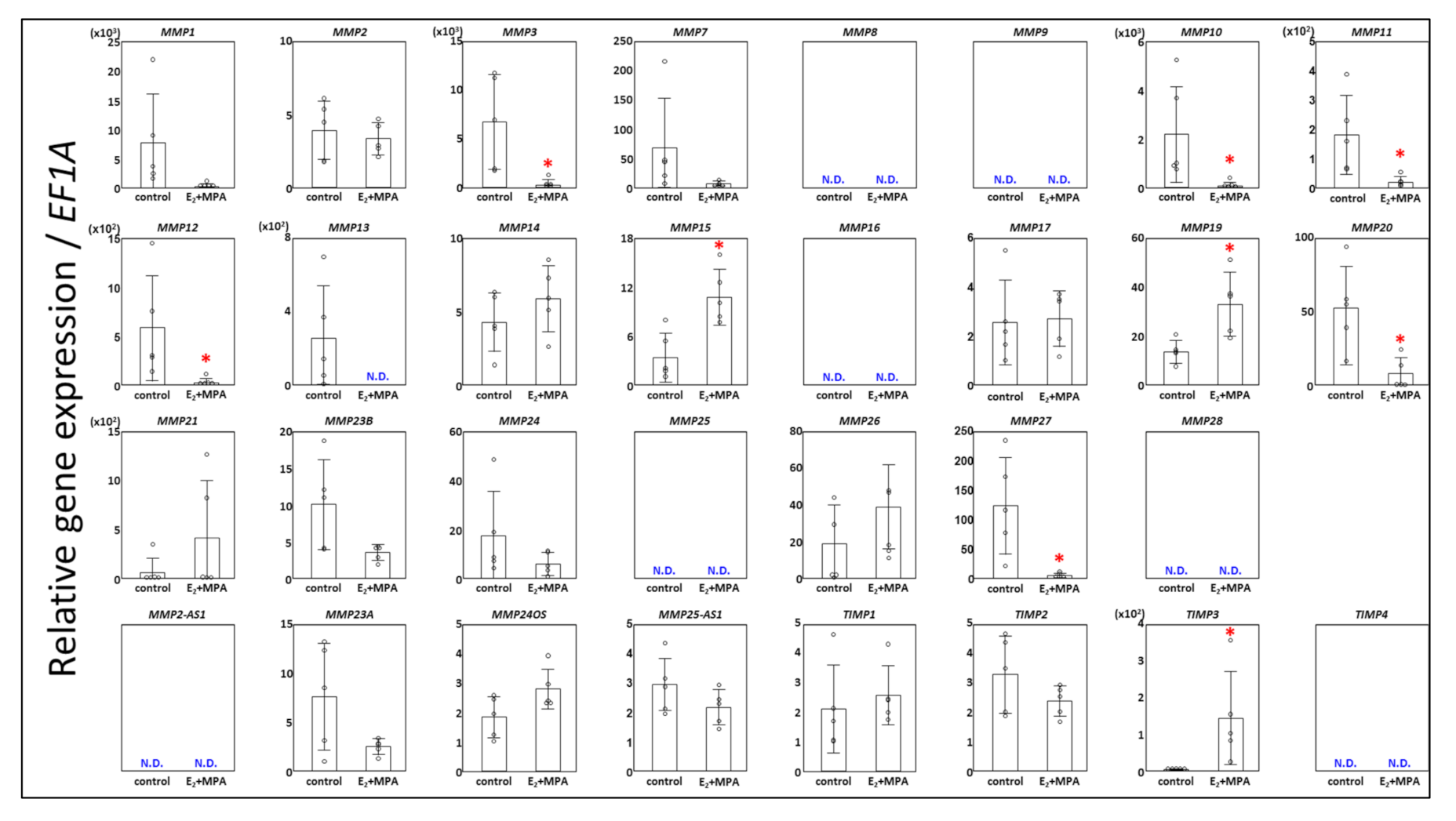
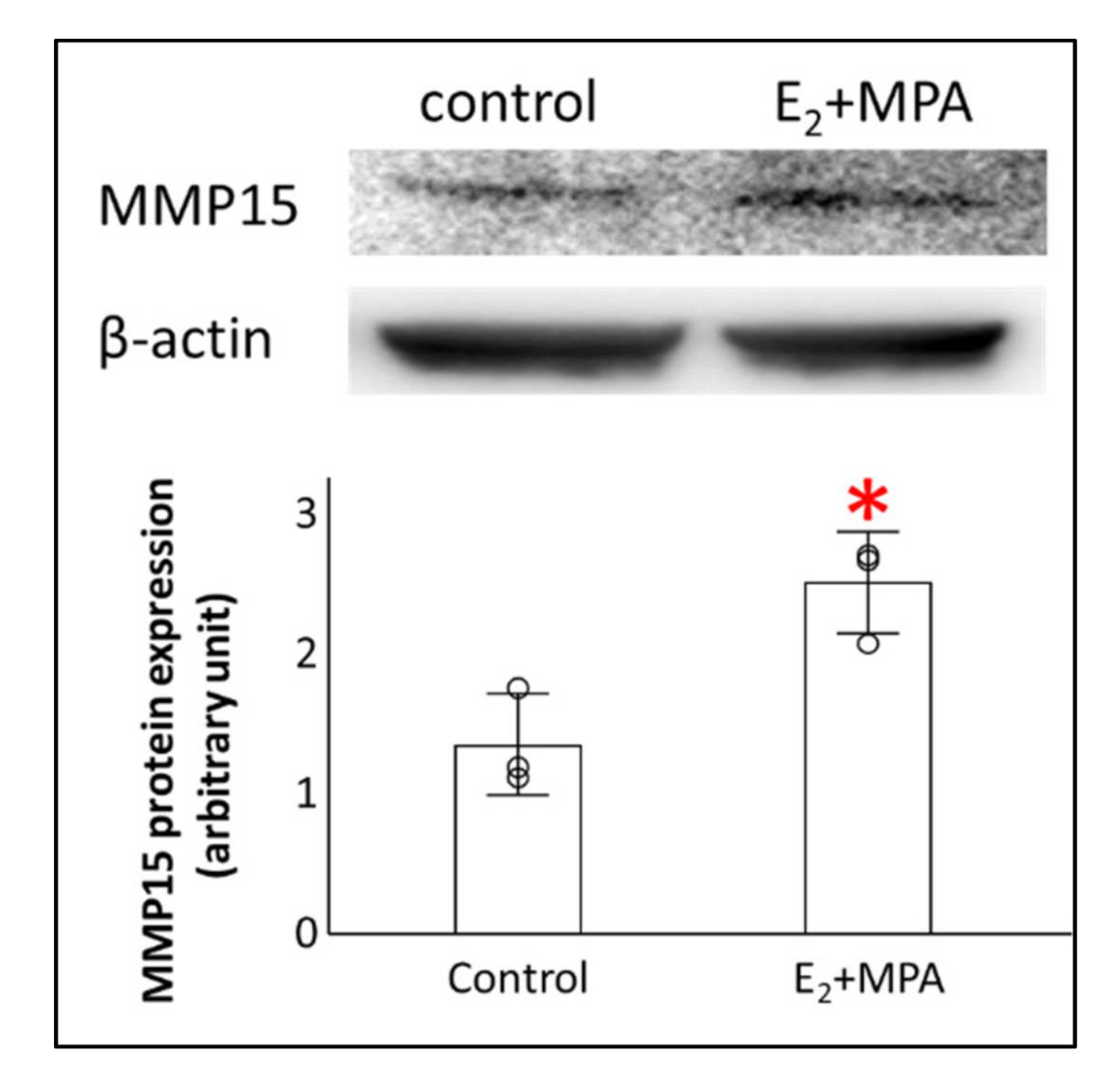
3.1. Expression Changes in MMPs
3.2. Expression Chages in TIMPs
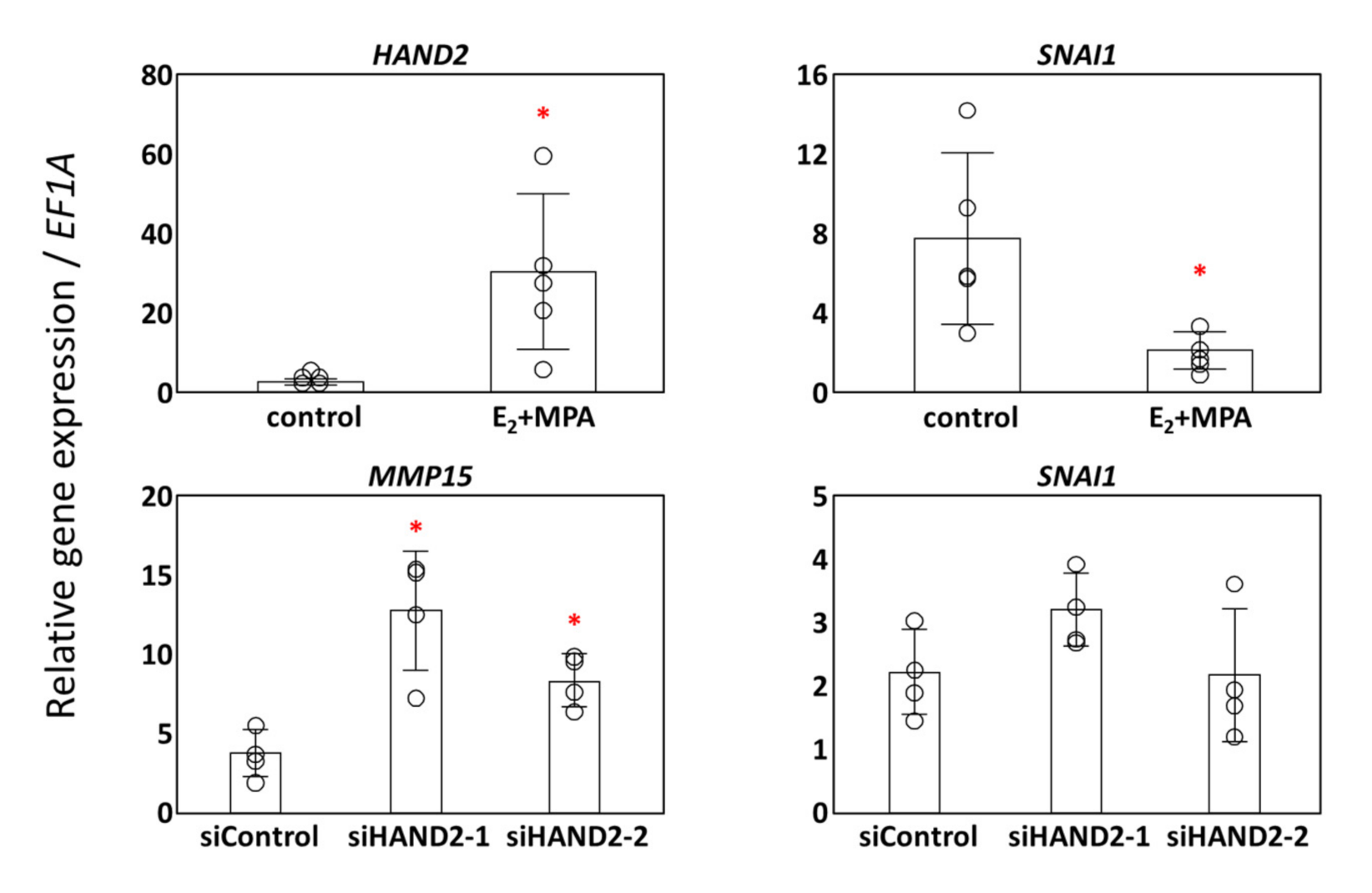
3.3. Potential Transcription Factors for MMP15
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cha, J.; Sun, X.; Dey, S.K. Mechanisms of implantation: Strategies for successful pregnancy. Nat. Med. 2012, 18, 1754–1767. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.; Salamonsen, L.A.; Winship, A.; Menkhorst, E.; Nie, G.; Gargett, C.E.; Dimitriadis, E. Fertile ground: Human endometrial programming and lessons in health and disease. Nat. Rev. Endocrinol. 2016, 12, 654–667. [Google Scholar] [CrossRef] [PubMed]
- Fox, C.; Morin, S.; Jeong, J.W.; Scott, R.T., Jr.; Lessey, B.A. Local and systemic factors and implantation: What is the evidence? Fertil. Steril. 2016, 105, 873–884. [Google Scholar] [CrossRef]
- Patel, B.; Elguero, S.; Thakore, S.; Dahoud, W.; Bedaiwy, M.; Mesiano, S. Role of nuclear progesterone receptor isoforms in uterine pathophysiology. Hum. Reprod. Update 2015, 21, 155–173. [Google Scholar] [CrossRef] [PubMed]
- Vinketova, K.; Mourdjeva, M.; Oreshkova, T. Human Decidual Stromal Cells as a Component of the Implantation Niche and a Modulator of Maternal Immunity. J. Pregnancy 2016, 2016, 8689436. [Google Scholar] [CrossRef]
- Mori, M.; Bogdan, A.; Balassa, T.; Csabai, T.; Szekeres-Bartho, J. The decidua-the maternal bed embracing the embryo-maintains the pregnancy. Semin. Immunopathol. 2016, 38, 635–649. [Google Scholar] [CrossRef] [PubMed]
- Burton, G.J.; Woods, A.W.; Jauniaux, E.; Kingdom, J.C. Rheological and physiological consequences of conversion of the maternal spiral arteries for uteroplacental blood flow during human pregnancy. Placenta 2009, 30, 473–482. [Google Scholar] [CrossRef]
- Fitzgerald, J.S.; Germeyer, A.; Huppertz, B.; Jeschke, U.; Knofler, M.; Moser, G.; Scholz, C.; Sonderegger, S.; Toth, B.; Markert, U.R. Governing the invasive trophoblast: Current aspects on intra- and extracellular regulation. Am. J. Reprod. Immunol. 2010, 63, 492–505. [Google Scholar] [CrossRef] [PubMed]
- Anacker, J.; Segerer, S.E.; Hagemann, C.; Feix, S.; Kapp, M.; Bausch, R.; Kammerer, U. Human decidua and invasive trophoblasts are rich sources of nearly all human matrix metalloproteinases. Mol. Hum. Reprod. 2011, 17, 637–652. [Google Scholar] [CrossRef]
- Dong, J.C.; Dong, H.; Campana, A.; Bischof, P. Matrix metalloproteinases and their specific tissue inhibitors in menstruation. Reproduction 2002, 123, 621–631. [Google Scholar] [CrossRef]
- Anacker, J.; Feix, S.; Kapp, M.; Bausch, R.; Kammerer, U. Expression pattern of matrix metalloproteinases (MMPs) in human decidua during pregnancy. J. Reprod. Immunol. 2010, 86, 79. [Google Scholar] [CrossRef]
- Murphy, G.; Nagase, H. Progress in matrix metalloproteinase research. Mol. Asp. Med. 2008, 29, 290–308. [Google Scholar] [CrossRef] [PubMed]
- Kessenbrock, K.; Plaks, V.; Werb, Z. Matrix metalloproteinases: Regulators of the tumor microenvironment. Cell 2010, 141, 52–67. [Google Scholar] [CrossRef]
- Murata, H.; Tsuzuki, T.; Kido, T.; Kakita-Kobayashi, M.; Kida, N.; Hisamatsu, Y.; Okada, H. Progestin-induced heart and neural crest derivatives-expressed transcript 2 inhibits angiopoietin 2 via fibroblast growth factor 9 in human endometrial stromal cells. Reprod. Biol. 2019, 19, 14–21. [Google Scholar] [CrossRef]
- Okada, H.; Nakajima, T.; Yoshimura, T.; Yasuda, K.; Kanzaki, H. The inhibitory effect of dienogest, a synthetic steroid, on the growth of human endometrial stromal cells in vitro. Mol. Hum. Reprod. 2001, 7, 341–347. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.H.; Huang, J.R.; Mazella, J.; Rosenberg, M.; Tseng, L. Differential effects of progestin and relaxin on the synthesis and secretion of immunoreactive prolactin in long term culture of human endometrial stromal cells. J. Clin. Endocrinol. Metab. 1990, 71, 889–899. [Google Scholar] [CrossRef] [PubMed]
- Shindoh, H.; Okada, H.; Tsuzuki, T.; Nishigaki, A.; Kanzaki, H. Requirement of heart and neural crest derivatives-expressed transcript 2 during decidualization of human endometrial stromal cells in vitro. Fertil. Steril. 2014, 101, 1781–1790. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Cho, H.; Okada, H.; Tsuzuki, T.; Nishigaki, A.; Yasuda, K.; Kanzaki, H. Progestin-induced heart and neural crest derivatives expressed transcript 2 is associated with fibulin-1 expression in human endometrial stromal cells. Fertil. Steril. 2013, 99, 248–255. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Wahlestedt, C. Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat. Rev. Drug Discov. 2013, 12, 433–446. [Google Scholar] [CrossRef]
- Mazur, E.C.; Vasquez, Y.M.; Li, X.; Kommagani, R.; Jiang, L.; Chen, R.; Lanz, R.B.; Kovanci, E.; Gibbons, W.E.; DeMayo, F.J. Progesterone receptor transcriptome and cistrome in decidualized human endometrial stromal cells. Endocrinology 2015, 156, 2239–2253. [Google Scholar] [CrossRef]
- Oki, S.; Ohta, T.; Shioi, G.; Hatanaka, H.; Ogasawara, O.; Okuda, Y.; Kawaji, H.; Nakaki, R.; Sese, J.; Meno, C. ChIP-Atlas: A data-mining suite powered by full integration of public ChIP-seq data. EMBO Rep. 2018, 19, e46255. [Google Scholar] [CrossRef]
- Okada, H.; Tsuzuki, T.; Murata, H. Decidualization of the human endometrium. Reprod. Med. Biol. 2018, 17, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Murata, H.; Tanaka, S.; Tsuzuki-Nakao, T.; Kido, T.; Kakita-Kobayashi, M.; Kida, N.; Hisamatsu, Y.; Tsubokura, H.; Hashimoto, Y.; Kitada, M.; et al. The transcription factor HAND2 up-regulates transcription of the IL15 gene in human endometrial stromal cells. J. Biol. Chem. 2020, 295, 9596–9605. [Google Scholar] [CrossRef]
- Laurent, F.; Girdziusaite, A.; Gamart, J.; Barozzi, I.; Osterwalder, M.; Akiyama, J.A.; Lincoln, J.; Lopez-Rios, J.; Visel, A.; Zuniga, A.; et al. HAND2 Target Gene Regulatory Networks Control Atrioventricular Canal and Cardiac Valve Development. Cell Rep. 2017, 19, 1602–1613. [Google Scholar] [CrossRef] [PubMed]
- Tao, G.; Levay, A.K.; Gridley, T.; Lincoln, J. Mmp15 is a direct target of Snai1 during endothelial to mesenchymal transformation and endocardial cushion development. Dev. Biol. 2011, 359, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Ding, L.; Wang, L.; Cao, Y.; Zhu, H.; Lu, J.; Li, X.; Song, T.; Hu, Y.; Dai, J. Umbilical cord-derived mesenchymal stem cells on scaffolds facilitate collagen degradation via upregulation of MMP-9 in rat uterine scars. Stem Cell Res. Ther. 2017, 8, 84. [Google Scholar] [CrossRef]
- Spitzer, T.L.; Rojas, A.; Zelenko, Z.; Aghajanova, L.; Erikson, D.W.; Barragan, F.; Meyer, M.; Tamaresis, J.S.; Hamilton, A.E.; Irwin, J.C.; et al. Perivascular human endometrial mesenchymal stem cells express pathways relevant to self-renewal, lineage specification, and functional phenotype. Biol. Reprod. 2012, 86, 58. [Google Scholar] [CrossRef]
- Majali-Martinez, A.; Hiden, U.; Ghaffari-Tabrizi-Wizsy, N.; Lang, U.; Desoye, G.; Dieber-Rotheneder, M. Placental membrane-type metalloproteinases (MT-MMPs): Key players in pregnancy. Cell Adhes. Migr. 2016, 10, 136–146. [Google Scholar] [CrossRef]
- Pollheimer, J.; Fock, V.; Knofler, M. Review: The ADAM metalloproteinases—Novel regulators of trophoblast invasion? Placenta 2014, 35, S57–S63. [Google Scholar] [CrossRef] [PubMed]
- Takino, T.; Sato, H.; Shinagawa, A.; Seiki, M. Identification of the second membrane-type matrix metalloproteinase (MT-MMP-2) gene from a human placenta cDNA library. MT-MMPs form a unique membrane-type subclass in the MMP family. J. Biol. Chem. 1995, 270, 23013–23020. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Barrantes, S.; Bernardo, M.; Toth, M.; Fridman, R. Regulation of membrane type-matrix metalloproteinases. Semin. Cancer Biol. 2002, 12, 131–138. [Google Scholar] [CrossRef] [PubMed]
- Majali-Martinez, A.; Hoch, D.; Tam-Amersdorfer, C.; Pollheimer, J.; Glasner, A.; Ghaffari-Tabrizi-Wizsy, N.; Beristain, A.G.; Hiden, U.; Dieber-Rotheneder, M.; Desoye, G. Matrix metalloproteinase 15 plays a pivotal role in human first trimester cytotrophoblast invasion and is not altered by maternal obesity. FASEB J. 2020, 34, 10720–10730. [Google Scholar] [CrossRef]
- Smith, S.D.; Choudhury, R.H.; Matos, P.; Horn, J.A.; Lye, S.J.; Dunk, C.E.; Aplin, J.D.; Jones, R.L.; Harris, L.K. Changes in vascular extracellular matrix composition during decidual spiral arteriole remodeling in early human pregnancy. Histol. Histopathol. 2016, 31, 557–571. [Google Scholar] [CrossRef]
- Huppertz, B.; Kertschanska, S.; Demir, A.Y.; Frank, H.G.; Kaufmann, P. Immunohistochemistry of matrix metalloproteinases (MMP), their substrates, and their inhibitors (TIMP) during trophoblast invasion in the human placenta. Cell Tissue Res. 1998, 291, 133–148. [Google Scholar] [CrossRef] [PubMed]
- Halari, C.D.; Nandi, P.; Jeyarajah, M.J.; Renaud, S.J.; Lala, P.K. Decorin production by the human decidua: Role in decidual cell maturation. Mol. Hum. Reprod. 2020, 26, 784–796. [Google Scholar] [CrossRef] [PubMed]
- Huyen, D.V.; Bany, B.M. Evidence for a conserved function of heart and neural crest derivatives expressed transcript 2 in mouse and human decidualization. Reproduction 2011, 142, 353–368. [Google Scholar] [CrossRef]
- Murata, H.; Tanaka, S.; Hisamatsu, Y.; Tsubokura, H.; Hashimoto, Y.; Kitada, M.; Okada, H. Transcriptional regulation of LGALS9 by HAND2 and FOXO1 in human endometrial stromal cells in women with regular cycles. Mol. Hum. Reprod. 2021, 27, gaab063. [Google Scholar] [CrossRef]
- Stormo, G.D. Modeling the specificity of protein-DNA interactions. Quant. Biol. 2013, 1, 115–130. [Google Scholar] [CrossRef]
- Morrison, C.J.; Butler, G.S.; Bigg, H.F.; Roberts, C.R.; Soloway, P.D.; Overall, C.M. Cellular activation of MMP-2 (gelatinase A) by MT2-MMP occurs via a TIMP-2-independent pathway. J. Biol. Chem. 2001, 276, 47402–47410. [Google Scholar] [CrossRef]
- Morrison, C.J.; Overall, C.M. TIMP independence of matrix metalloproteinase (MMP)-2 activation by membrane type 2 (MT2)-MMP is determined by contributions of both the MT2-MMP catalytic and hemopexin C domains. J. Biol. Chem. 2006, 281, 26528–26539. [Google Scholar] [CrossRef] [PubMed]
- Ogiwara, K.; Takahashi, T. Involvement of the nuclear progestin receptor in LH-induced expression of membrane type 2-matrix metalloproteinase required for follicle rupture during ovulation in the medaka, Oryzias latipes. Mol. Cell. Endocrinol. 2017, 450, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Taylor, R.N.; Bagchi, I.C.; Bagchi, M.K. Regulation of human endometrial stromal proliferation and differentiation by C/EBPbeta involves cyclin E-cdk2 and STAT3. Mol. Endocrinol. 2012, 26, 2016–2030. [Google Scholar] [CrossRef] [PubMed]
- Tsunoda, T.; Takagi, T. Estimating transcription factor bindability on DNA. Bioinformatics 1999, 15, 622–630. [Google Scholar] [CrossRef] [PubMed]
- Carver, E.A.; Jiang, R.; Lan, Y.; Oram, K.F.; Gridley, T. The mouse snail gene encodes a key regulator of the epithelial-mesenchymal transition. Mol. Cell. Biol. 2001, 21, 8184–8188. [Google Scholar] [CrossRef] [PubMed]
- Schroer, U.; Volk, G.F.; Liedtke, T.; Thanos, S. Translin-associated factor-X (Trax) is a molecular switch of growth-associated protein (GAP)-43 that controls axonal regeneration. Eur. J. Neurosci. 2007, 26, 2169–2178. [Google Scholar] [CrossRef]
- Liu, Y.; Sun, X.; Feng, J.; Deng, L.L.; Li, B.; Zhu, M.; Lu, C.; Zhou, L. MT2-MMP induces proteolysis and leads to EMT in carcinomas. Oncotarget 2016, 7, 48193–48205. [Google Scholar] [CrossRef] [PubMed]
- Kaitu’u-Lino, T.J.; Palmer, K.; Tuohey, L.; Ye, L.; Tong, S. MMP-15 is upregulated in preeclampsia, but does not cleave endoglin to produce soluble endoglin. PLoS ONE 2012, 7, e39864. [Google Scholar] [CrossRef]
- Kolb, C.; Mauch, S.; Krawinkel, U.; Sedlacek, R. Matrix metalloproteinase-19 in capillary endothelial cells: Expression in acutely, but not in chronically, inflamed synovium. Exp. Cell Res. 1999, 250, 122–130. [Google Scholar] [CrossRef]
- Brauer, R.; Beck, I.M.; Roderfeld, M.; Roeb, E.; Sedlacek, R. Matrix metalloproteinase-19 inhibits growth of endothelial cells by generating angiostatin-like fragments from plasminogen. BMC Biochem. 2011, 12, 38. [Google Scholar] [CrossRef]
- Mattos, R.; Staples, C.R.; Thatcher, W.W. Effects of dietary fatty acids on reproduction in ruminants. Rev. Reprod. 2000, 5, 38–45. [Google Scholar] [CrossRef]
- Killeen, A.P.; Morris, D.G.; Kenny, D.A.; Mullen, M.P.; Diskin, M.G.; Waters, S.M. Global gene expression in endometrium of high and low fertility heifers during the mid-luteal phase of the estrous cycle. BMC Genom. 2014, 15, 234. [Google Scholar] [CrossRef]
- Quintero-Ronderos, P.; Mercier, E.; Fukuda, M.; Gonzalez, R.; Suarez, C.F.; Patarroyo, M.A.; Vaiman, D.; Gris, J.C.; Laissue, P. Novel genes and mutations in patients affected by recurrent pregnancy loss. PLoS ONE 2017, 12, e0186149. [Google Scholar] [CrossRef]
- Staskus, P.W.; Masiarz, F.R.; Pallanck, L.J.; Hawkes, S.P. The 21-kDa protein is a transformation-sensitive metalloproteinase inhibitor of chicken fibroblasts. J. Biol. Chem. 1991, 266, 449–454. [Google Scholar] [CrossRef]
- Pavloff, N.; Staskus, P.W.; Kishnani, N.S.; Hawkes, S.P. A new inhibitor of metalloproteinases from chicken: ChIMP-3. A third member of the TIMP family. J. Biol. Chem. 1992, 267, 17321–17326. [Google Scholar] [CrossRef]
- Leco, K.J.; Khokha, R.; Pavloff, N.; Hawkes, S.P.; Edwards, D.R. Tissue inhibitor of metalloproteinases-3 (TIMP-3) is an extracellular matrix-associated protein with a distinctive pattern of expression in mouse cells and tissues. J. Biol. Chem. 1994, 269, 9352–9360. [Google Scholar] [CrossRef]
- Apte, S.S.; Olsen, B.R.; Murphy, G. The gene structure of tissue inhibitor of metalloproteinases (TIMP)-3 and its inhibitory activities define the distinct TIMP gene family. J. Biol. Chem. 1995, 270, 14313–14318. [Google Scholar] [CrossRef] [PubMed]
- Brew, K.; Nagase, H. The tissue inhibitors of metalloproteinases (TIMPs): An ancient family with structural and functional diversity. Biochim. Biophys. Acta 2010, 1803, 55–71. [Google Scholar] [CrossRef] [PubMed]
- Moore, L.; Fan, D.; Basu, R.; Kandalam, V.; Kassiri, Z. Tissue inhibitor of metalloproteinases (TIMPs) in heart failure. Heart Fail. Rev. 2012, 17, 693–706. [Google Scholar] [CrossRef]
- Jackson, H.W.; Defamie, V.; Waterhouse, P.; Khokha, R. TIMPs: Versatile extracellular regulators in cancer. Nat. Rev. Cancer 2017, 17, 38–53. [Google Scholar] [CrossRef] [PubMed]
- Qi, J.H.; Ebrahem, Q.; Moore, N.; Murphy, G.; Claesson-Welsh, L.; Bond, M.; Baker, A.; Anand-Apte, B. A novel function for tissue inhibitor of metalloproteinases-3 (TIMP3): Inhibition of angiogenesis by blockage of VEGF binding to VEGF receptor-2. Nat. Med. 2003, 9, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Klenotic, P.A.; Munier, F.L.; Marmorstein, L.Y.; Anand-Apte, B. Tissue inhibitor of metalloproteinases-3 (TIMP-3) is a binding partner of epithelial growth factor-containing fibulin-like extracellular matrix protein 1 (EFEMP1). Implications for macular degenerations. J. Biol. Chem. 2004, 279, 30469–30473. [Google Scholar] [CrossRef] [PubMed]
- Nour, N.; Mayer, G.; Mort, J.S.; Salvas, A.; Mbikay, M.; Morrison, C.J.; Overall, C.M.; Seidah, N.G. The cysteine-rich domain of the secreted proprotein convertases PC5A and PACE4 functions as a cell surface anchor and interacts with tissue inhibitors of metalloproteinases. Mol. Biol. Cell 2005, 16, 5215–5226. [Google Scholar] [CrossRef][Green Version]
- Kang, K.H.; Park, S.Y.; Rho, S.B.; Lee, J.H. Tissue inhibitor of metalloproteinases-3 interacts with angiotensin II type 2 receptor and additively inhibits angiogenesis. Cardiovasc. Res. 2008, 79, 150–160. [Google Scholar] [CrossRef] [PubMed]
- Scilabra, S.D.; Troeberg, L.; Yamamoto, K.; Emonard, H.; Thogersen, I.; Enghild, J.J.; Strickland, D.K.; Nagase, H. Differential regulation of extracellular tissue inhibitor of metalloproteinases-3 levels by cell membrane-bound and shed low density lipoprotein receptor-related protein 1. J. Biol. Chem. 2013, 288, 332–342. [Google Scholar] [CrossRef]
- Huttlin, E.L.; Ting, L.; Bruckner, R.J.; Gebreab, F.; Gygi, M.P.; Szpyt, J.; Tam, S.; Zarraga, G.; Colby, G.; Baltier, K.; et al. The BioPlex Network: A Systematic Exploration of the Human Interactome. Cell 2015, 162, 425–440. [Google Scholar] [CrossRef] [PubMed]
- Huttlin, E.L.; Bruckner, R.J.; Paulo, J.A.; Cannon, J.R.; Ting, L.; Baltier, K.; Colby, G.; Gebreab, F.; Gygi, M.P.; Parzen, H.; et al. Architecture of the human interactome defines protein communities and disease networks. Nature 2017, 545, 505–509. [Google Scholar] [CrossRef]
- Kwasna, D.; Abdul Rehman, S.A.; Natarajan, J.; Matthews, S.; Madden, R.; De Cesare, V.; Weidlich, S.; Virdee, S.; Ahel, I.; Gibbs-Seymour, I.; et al. Discovery and Characterization of ZUFSP/ZUP1, a Distinct Deubiquitinase Class Important for Genome Stability. Mol. Cell 2018, 70, 150–164, e156. [Google Scholar] [CrossRef]
| Sample No. | Materials | Methods | Age | Menstrual Cycle Phase at the Time of Collection | |
|---|---|---|---|---|---|
| 1 | Primary culture ESCs | Treated with E2 + MPA for 12 days | RT-qPCR | 50 | Proliferative |
| 2 | Primary culture ESCs | Treated with E2 + MPA for 12 days | RT-qPCR | 45 | Mid-secretory |
| 3 | Primary culture ESCs | Treated with E2 + MPA for 12 days | RT-qPCR | 48 | Late-secretory |
| 4 | Primary culture ESCs | Treated with E2 + MPA for 12 days | RT-qPCR | 50 | Mid-secretory |
| 5 | Primary culture ESCs | Treated with E2 + MPA for 12 days | RT-qPCR | 44 | Late-secretory |
| 6 | Primary culture ESCs | Transfection of HAND2-siRNA | RT-qPCR | 49 | Mid-secretory |
| 7 | Primary culture ESCs | Transfection of HAND2-siRNA | RT-qPCR | 46 | Late-secretory |
| 8 | Primary culture ESCs | Transfection of HAND2-siRNA | RT-qPCR | 42 | Proliferative |
| 9 | Primary culture ESCs | Transfection of HAND2-siRNA | RT-qPCR | 47 | Late-secretory |
| 10 | Primary culture ESCs | Treated with E2 + MPA for 12 days | Immunoblot analysis | 45 | Mid-Secretory |
| 11 | Primary culture ESCs | Treated with E2 + MPA for 12 days | Immunoblot analysis | 42 | Proliferative |
| 12 | Primary culture ESCs | Treated with E2 + MPA for 12 days | Immunoblot analysis | 49 | Proliferative |
| MMP1 | MMP1_884F | 5′-ACAAACCCCAAAAGCGTGTG-3′ |
| MMP1_993R | 5′-AGAAGGGATTTGTGCGCATG-3′ | |
| MMP2 | MMP2_697F | 5′-ACAGTGGATGATGCCTTTGC-3 |
| MMP2_808R | 5′-AGCGGCCAAAGTTGATCATG-3′ | |
| MMP3 | MMP3_1578F | 5′-TTCGTTTTCTCCTGCCTGTG-3′ |
| MMP3_1693R | 5′-AGCAGCAGCCCATTTGAATG-3′ | |
| MMP7 | MMP7_362F | 5′-TTCCAAAGTGGTCACCTACAGG-3′ |
| MMP7_459R | 5′-TGCCCCACATGTTTAAAGCC-3′ | |
| MMP8 | MMP8_355F | 5′-ATGAAAAAGCCTCGCTGTGG-3′ |
| MMP8_436R | 5′-AGTTAGTGCGTTCCCACTTG-3′ | |
| MMP9 | MMP9_1560F | 5′-ATGCCTGCAACGTGAACATC-3′ |
| MMP9_1646R | 5′-AGAATCGCCAGTACTTCCCATC-3′ | |
| MMP10 | MMP10_373F | 5′-CCTTACATACAGGATTGTGAATTATACACC-3′ |
| MMP10_519R | 5′-GAGATCATTATATCAGCCTCTCCTTCATAC-3′ | |
| MMP11 | MMP11_558F | 5′-ATGCCTTCTTCCCCAAGACTC-3′ |
| MMP11_684R | 5′-AGCACGTGGCCAAATTCATG-3′ | |
| MMP12 | MMP12_440F | 5′-ACGCAATCCGGAAAGCTTTC-3′ |
| MMP12_527R | 5′-ACCAAAATGTCAGCCATGCC-3′ | |
| MMP13 | MMP13_864F | 5′-AACGCCAGACAAATGTGACC-3′ |
| MMP13_991R | 5′-AAAACAGCTCCGCATCAACC-3′ | |
| MMP14 | MMP14_461F | 5′-AACAGGCAAAGCTGATGCAG-3′ |
| MMP14_567R | 5′-AGCGCTTCCTTCGAACATTG-3′ | |
| MMP15 | MMP15_1187F | 5′-TCATGGTACTCTTTGCCTCTGG-3′ |
| MMP15_1310R | 5′-TCTGCGTCAAAATGGGTGTC-3′ | |
| MMP16 | MMP16_654F | 5′-AATGGCAGCACAAGCACATC-3′ |
| MMP16_748R | 5′-ATCAAAGGCACGGCGAATAG-3′ | |
| MMP17 | MMP17_1095F | 5′-ATGCAGCACTCACTTTGACG-3′ |
| MMP17_1169R | 5′-CGCCAGAAGTACTTGCCTTTG-3′ | |
| MMP19 | MMP19_1606F | 5′-AGCCACAGAAACCACGTTTG-3′ |
| MMP19_1698R | 5′-AAATGAAAGGGTGGGTGGTG-3′ | |
| MMP20 | MMP20_1053F | 5′-TGGATGCAGCTTACGAAGTG-3′ |
| MMP20_1195R | 5′-TATTTGCTGCACGTGCCTTG-3′ | |
| MMP21 | MMP21_889F | 5′-ACGGGATCCATAATGCAACC-3′ |
| MMP21_1025R | 5′-TTGCGAATCCAGTCAAACGC-3′ | |
| MMP23B | MMP23B_1021F | 5′-CCTCCACAAGAAAGGGAAAGTG-3′ |
| MMP23B_1151R | 5′-TGTAGGTGCCCTCATTGACG-3′ | |
| MMP24 | MMP24_847F | 5′-AGGAAATGCCAACCATGACG-3′ |
| MMP24_990R | 5′-AGCTTGAAGTTGTGCGTCTC-3′ | |
| MMP25 | MMP25_3278F | 5′-TGGCTGTTTCGTGGCATTTC-3′ |
| MMP25_3407R | 5’-TGGACAGCAACTTAGGAAGTGG-3′ | |
| MMP26 | MMP26_1127F | 5′-AAAGCACTAGAGCAGCCTTG-3′ |
| MMP26_1215R | 5′-AGCGTTTTGAGTGTCGGTTC-3′ | |
| MMP27 | MMP27_853F | 5′-ACCTGCTAAGCCAAAGGAAC-3′ |
| MMP27_932R | 5′-TGCGGAAAGTTGTGATAGCG-3′ | |
| MMP28 | MMP28_1214F | 5′-AAACGCAGGGCCCTAAATAC-3′ |
| MMP28_1283R | 5′-TACAGTTGCTGTTGCCTGTC-3′ | |
| TIMP1 | TIMP1_490F | 5′-AGGAATGCACAGTGTTTCCC-3′ |
| TIMP1_592R | 5′-AAGCCCTTTTCAGAGCCTTG-3′ | |
| TIMP2 | TIMP2_2430F | 5-ACACGCAATGAAACCGAAGC-3′ |
| TIMP2_2503R | 5′-AACAGGCTAAGGTGGCTTTG-3′ | |
| TIMP3 | TIMP3_1700F | 5′-TGCCTGCCTTGTACAAAAGC-3′ |
| TIMP3_1805R | 5′-TGGCCAAATCTACCAAAGCG-3′ | |
| TIMP4 | TIMP4_611F | 5′-ACTGGCTGTTGGAACGAAAG-3′ |
| TIMP4_681R | 5′-GCCGTCAACATGCTTCATACAG-3′ | |
| MMP2-AS1 | MMP2-AS1_541F | 5′-ATGTTGTGAGCAGCCCAATG-3′ |
| MMP2-AS1_682R | 5′-TTGCCACTCAGCATCATCAC-3′ | |
| MMP23A | MMP23A_682F | 5′-CCTCCACAAGAAAGGGAAAGTG-3′ |
| MMP23A_812R | 5′-TGTAGGTGCCCTCATTGACG-3′ | |
| MMP24OS | MMP24OS_1029F | 5′-ATCCCAGGGAAATGACACACTC-3′ |
| MMP24OS_1144R | 5′-GGGACATCACAGCATTTCAGTG-3′ | |
| MMP25-AS1 | MMP25-AS1_2017F | 5′-AGTTCCGGAATGCAAAACCC-3′ |
| MMP25-AS1_2086R | 5′-AGGACCTTGAAAGCATGTGG-3′ | |
| HAND2 | hHAND2-Forward | 5′-AGAGGAAGAAGGAGCTGAACGA-3′ |
| hHAND2-Reverse | 5′-CGTCCGGCCTTTGGTTTT-3′ | |
| SNAI1 | SNAI1_1405F | 5′-TTTCAGCCTCCTGTTTGGTG-3′ |
| SNAI1_1489R | 5′-TGACAGCCATTACTCACAGTCC-3′ | |
| EF1A | hEF1A_F | 5′-TCTGGTTGGAATGGTGACAACATGC-3′ |
| hEF1A_R | 5′-AGAGCTTCACTCAAAGCTTCATGG-3′ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hisamatsu, Y.; Murata, H.; Tsubokura, H.; Hashimoto, Y.; Kitada, M.; Tanaka, S.; Okada, H. Matrix Metalloproteinases in Human Decidualized Endometrial Stromal Cells. Curr. Issues Mol. Biol. 2021, 43, 2111-2123. https://doi.org/10.3390/cimb43030146
Hisamatsu Y, Murata H, Tsubokura H, Hashimoto Y, Kitada M, Tanaka S, Okada H. Matrix Metalloproteinases in Human Decidualized Endometrial Stromal Cells. Current Issues in Molecular Biology. 2021; 43(3):2111-2123. https://doi.org/10.3390/cimb43030146
Chicago/Turabian StyleHisamatsu, Yoji, Hiromi Murata, Hiroaki Tsubokura, Yoshiko Hashimoto, Masaaki Kitada, Susumu Tanaka, and Hidetaka Okada. 2021. "Matrix Metalloproteinases in Human Decidualized Endometrial Stromal Cells" Current Issues in Molecular Biology 43, no. 3: 2111-2123. https://doi.org/10.3390/cimb43030146
APA StyleHisamatsu, Y., Murata, H., Tsubokura, H., Hashimoto, Y., Kitada, M., Tanaka, S., & Okada, H. (2021). Matrix Metalloproteinases in Human Decidualized Endometrial Stromal Cells. Current Issues in Molecular Biology, 43(3), 2111-2123. https://doi.org/10.3390/cimb43030146






