Ecology of Anti-Biofilm Agents I: Antibiotics versus Bacteriophages
Abstract
:1. Introduction
“Chronic infections… are very difficult, if not impossible, to cure with antibiotics.”—T. Bjarnsholt [1]
Antibiotics and Biofilm Disruption
- (1)
- Biofilms at least in part tend to be inherently resistant to antibiotics, that is, selectively toxic chemical agents that are not applied at extremely high concentrations (though antibiotics such as colistin do exist which are effective at targeting less metabolically active bacteria, though in this case there is also noticeable toxicity to human tissue as well);
- (2)
- Highly efficacious, broadly acting anti-biofilm compounds may be difficult for organisms to produce or deploy without harming themselves (see, however, the newly discovered, broadly acting anti-biofilm protein, BL-DZ1 [38]);
- (3)
- Highly efficacious but narrowly acting anti-biofilm agents may not possess sufficient ranges of activity to justify the costs to organisms of producing them, or for us to develop them as pharmaceuticals (though narrowly acting anti-biofilm agents nonetheless do exist, such as bacteriocins); or
- (4)
- The utility of breaking up existing, intact biofilms through the use of antibiotics alone might not be sufficiently compelling to antibiotic-producing organisms to result in the evolution of antibiotics with highly effective anti-biofilm activities (though, in fact, there are numerous suggestions that the competitiveness of biofilm-producing bacteria may be enhanced through the production of antibacterial substances).
2. Biofilm Disruption by Microorganisms
2.1. Differentiating among Potential Utilities of Antibacterial Action
| State of Targeted Biofilm (below): | Use in Defense (δ) (resource protection) | Use as Offense (ω) (resource acquisition) |
|---|---|---|
| “Before” (B) biofilms have formed as the target state | δB: Protection of antibiotic-producing organisms from death or displacement that may be mediated by target, disseminating bacteria (ωD-1 or ωD-3 represent what potentially is being protected against) | ωB: Destruction of target, disseminating bacteria in order to obtain nutrients that are directly associated with those bacteria (δD-1 could serve as a potential counter measure mediated by these target bacteria) |
| “During” (D) biofilm sessile existence as the target state | δD-1: Protection of antibiotic-producing disseminating organism from target, biofilm bacteria (ωB represents what potentially is being protected against); δD-2: Protection of antibiotic-producing organisms as found within biofilms from encroachment or consumption by adjacent, target, biofilm bacteria (ωD-2 or ωD-3 represent what potentially is being protected against) | ωD-1: Displacement of target, biofilm bacteria by disseminating, antibiotic-producing bacteria (in order to obtain “Space”); ωD-2: Encroachment by antibiotic-producing, biofilm bacteria on adjacent, target, biofilm bacteria (in order to obtain “Space”); ωD-3: Destruction of target, biofilm bacteria by antibiotic-producing organism in order to obtain nutrients from those target bacteria |
| “After” (A) biofilms have been disrupted as the target state | δA: Destruction of target bacteria that have been displaced from biofilms, in order to prevent competition for nutrients | ωA: Destruction of target bacteria that have been displaced from biofilms, in order to obtain nutrients from those bacteria |

2.2. Limitations on Antibiotic Anti-Biofilm Activity, a Genetics Perspective
3. Scenarios of Antibiotic Anti-Biofilm Ecology
| One Target Cell | Many Target Cells | |
|---|---|---|
| One Producing Cell | Not involving biofilm: Scenario 3 (“Before”, δB or ωB) | Effecting biofilm invasion: Scenario 1 (“During”, ωD, but also δD) |
| Many Producing Cells | Effecting biofilm protection: Scenario 2 (“Before”, δB, but also ωB) | Within-biofilm competition: Scenario 4 (“During”, δD or ωD) |
3.1. Scenario 1, Disseminating Antibiotic-Producing Cell, Biofilm Bacteria as Targets
3.1.1. Ineffectiveness of Small Invading Forces
3.1.2. Insufficiency of Soluble Antibacterials as Facilitators of Biofilm Invasion
3.1.3. Requirements for Effective Biofilm Invasion and Displacement
3.2. Scenario 2, Antibiotic-Producing Biofilm, Disseminating Target Bacterium
3.3. Scenario 3, Disseminating Antibiotic-Producing Cell, Disseminating Target Bacterium
3.4. Scenario 4, Antibiotic-Producing Biofilm, Biofilm Bacteria as Targets
3.4.1. Evidence of Within-Biofilm Antibacterial Effectiveness
3.4.2. Does Killing or Removal of Biofilm Bacteria Occur via Antibacterial Action Alone?
3.4.3. The Issue of Public Goods
3.4.4. What Works and What Doesn’t
3.5. Scenario 5, Antibacterial Action Following Biofilm Physical Disruption
4. Bacteriophage Anti-Biofilm Activity
4.1. Bacteriophages as Anti-Biofilm Agents
4.2. Biofilms as Targets of Phage Action
4.3. Concentrating Phage Antibacterial Activity
4.3.1. Trojan Horse Strategy Number 1
4.3.2. Trojan Horse Strategy Number 2
4.3.3. Phages as Anti-Biofilm Agents, a Summary
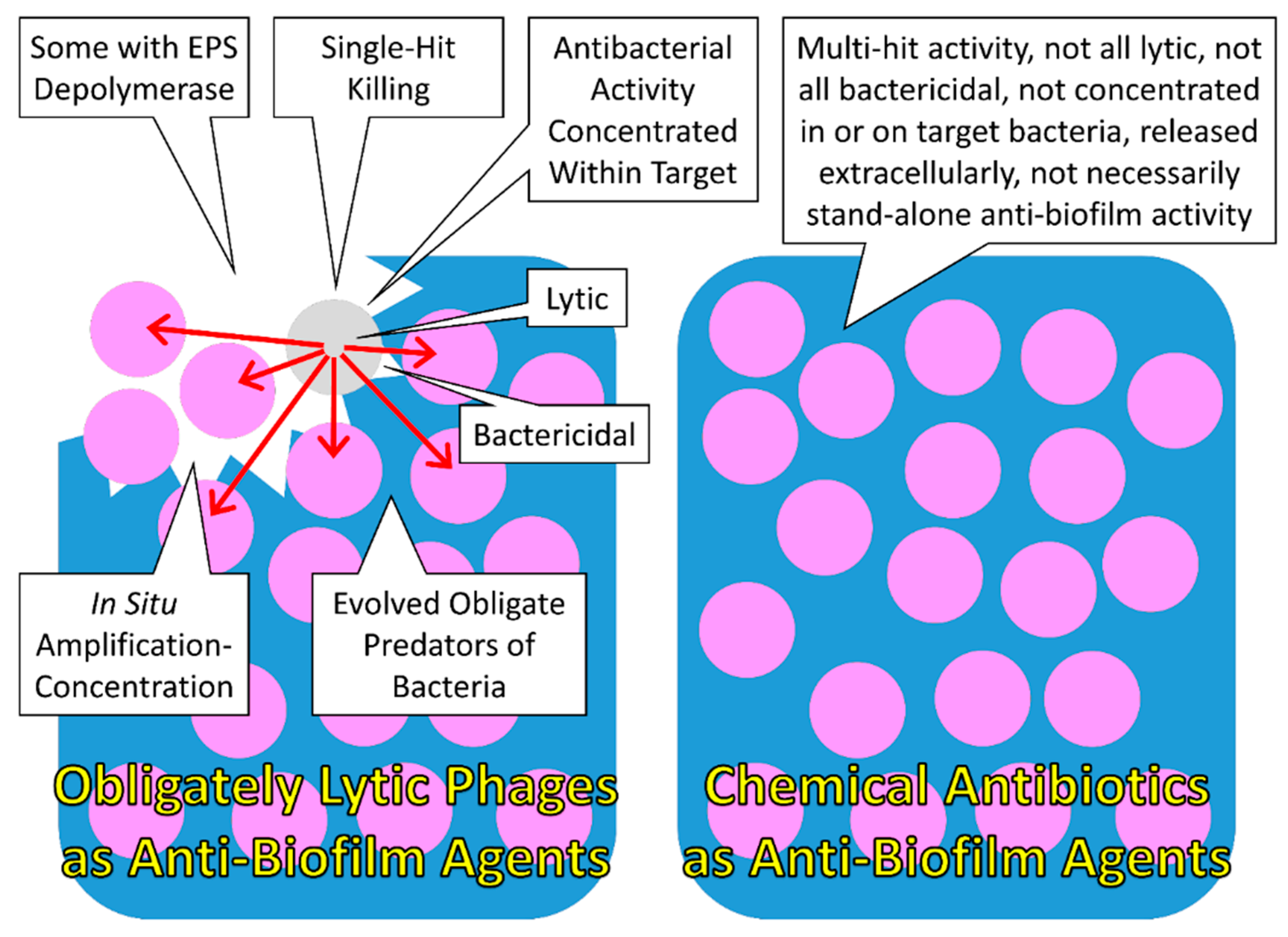
| Property of Anti-biofilm Agent | As Considered in Terms of Bacteriophages | As Considered in Terms of Antibiotic Producers |
|---|---|---|
| Inherent predators of bacteria | Particularly for obligately lytic phages, their ability to replicate is closely associated with their ability to kill target bacteria, resulting in an antibacterial activity which is under strong selection, as evidenced by all lytic phages obligately killing target bacteria to produce new phage virions | Particularly for organisms that are not obligate predators of bacteria, their reproduction likely is not explicitly dependent on an ability to kill bacteria, suggesting that antibiotic production is not under as strong selection in non-predatory organisms as it is for predatory ones |
| Obligate predators of bacteria | The concept that losing a meal is less costly than becoming a meal, to explain differential selective pressures acting on predators versus prey [129], is less applicable to organisms that tend to die if they fail to succeed in exploiting a given meal, once obtained, and this tends to be the case for parasites and, by extension, for phages, i.e., as host-killing parasites | For antibiotic-producing organisms, the cost associated with an antibiotic being less efficacious likely is lower than the equivalent costs to phages for less than optimal antibacterial activity because ongoing replication of antibiotic-producing organisms mostly is not absolutely dependent on inhibition of target bacteria metabolism |
| Concentration of antibacterial activity within the vicinity of individual target bacteria | Antibacterial action tends to be concentration dependent, as too can be antibacterial toxicity, and phages are able to concentrate their antibacterial activity not just in the vicinity of target bacteria, but within target bacteria | Concentration of antibiotics on specific targets can be more difficult to achieve for organisms that release antibiotics randomly in all directions and/or for which antibiotic release is not triggered by contact with target organisms |
| Concentration of antibacterial activity within spatially associated groups of target bacteria | An ability to replicate in the course of effecting antibacterial activity can allow phages to concentrate their activity spatially within phage-sensitive microcolonies or phage-sensitive cellular arrangements | Antibiotic-producing organisms also are capable of replication, including in the vicinity of target organisms, though replication by binary fission can be slower than that achievable by phages in the presence of high target-bacteria densities |
| Bactericidal activity | For lytic phages the death of target bacteria tends to be highly associated with antibacterial activity | Even among effective antibiotics, not all result directly in the death of target bacteria, i.e., bacteriostatic agents |
| Lytic activity | For lytic phages the lysis of target bacteria is highly associated with antibacterial activity and can lead to sequential removal of biofilm material (e.g., leading to “Active penetration” [120]) | Not all antibiotics give rise directly to the lysis of target bacteria so therefore do not necessarily directly give rise to destruction of biofilm physical structure |
| EPS depolymerases | Certain phages deploy enzymes that are capable of breaking down biofilm extracellular matrix | Antibiotics in and of themselves will not likely possess EPS depolymerase functions |
| Single-hit killing kinetics | Generally the death of sensitive bacteria follows the adsorption of only a single phage | Generally the death of sensitive bacteria requires exposure to large numbers of molecules of individual antibiotic types |
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Bjarnsholt, T. The role of bacterial biofilms in chronic infections. APMIS Suppl. 2013, 121, 1–51. [Google Scholar] [CrossRef]
- Hokkanen, H.M.T.; Sailer, R.I. Success in classical biological control. Crit. Rev. Plant Sci. 1985, 1, 35–72. [Google Scholar] [CrossRef]
- Harper, D.R. Biological control by microorganisms. In The Encyclopedia of Life Sciences; John Wiley & Sons: England, UK, 2006; pp. 1–10. [Google Scholar]
- Jordan, K.; Dalmasso, M.; Zentek, J.; Mader, A.; Bruggeman, G.; Wallace, J.; De, M.D.; Fiore, A.; Prukner-Radovcic, E.; Lukac, M.; et al. Microbes versus microbes: Control of pathogens in the food chain. J. Sci. Food Agric. 2014, 94, 3079–3089. [Google Scholar] [CrossRef]
- Gottlieb, D. The production and role of antibiotics in soil. J. Antibiot. 1976, 29, 987–1000. [Google Scholar] [CrossRef]
- Audrain, B.; Farag, M.A.; Ryu, C.M.; Ghigo, J.M. Role of bacterial volatile compounds in bacterial biology. FEMS Microbiol. Rev. 2015, 39, 222–233. [Google Scholar] [CrossRef]
- Schmidt, R.; Cordovez, V.; De Boer, W.; Raaijmakers, J.; Garbeva, P. Volatile affairs in microbial interactions. ISME J. 2015. [Google Scholar] [CrossRef]
- Cotter, P.D.; Ross, R.P.; Hill, C. Bacteriocins—a viable alternative to antibiotics? Nat. Rev. Microbiol. 2013, 11, 95–105. [Google Scholar] [CrossRef]
- Gill, J.J.; Young, R. Therapeutic applications of phage biology: History, practice and recommendations. In Emerging Trends in Antibacterial Discovery: Answering the Call to Arms; Miller, A.A., Miller, P.F., Eds.; Caister Academic Press: Norfolk, UK, 2011; pp. 367–410. [Google Scholar]
- Leiman, P.G.; Shneider, M.M. Contractile tail machines of bacteriophages. Adv. Exp. Med. Biol. 2012, 726, 93–114. [Google Scholar]
- Calendar, R.; Abedon, S.T. The Bacteriophages, 2nd ed.; Oxford University Press: Oxford, UK, 2006. [Google Scholar]
- Pirnay, J.P.; De, V.D.; Verbeken, G.; Merabishvili, M.; Chanishvili, N.; Vaneechoutte, M.; Zizi, M.; Laire, G.; Lavigne, R.; Huys, I.; et al. The phage therapy paradigm: Prêt-à-porter or sur-mesure? Pharm. Res. 2011, 28, 934–937. [Google Scholar] [CrossRef]
- Chan, B.K.; Abedon, S.T. Phage therapy pharmacology: Phage cocktails. Adv. Appl. Microbiol. 2012, 78, 1–23. [Google Scholar]
- Chan, B.K.; Abedon, S.T.; Loc-Carrillo, C. Phage cocktails and the future of phage therapy. Future Microbiol. 2013, 8, 769–783. [Google Scholar] [CrossRef]
- Schmerer, M.; Molineux, I.J.; Bull, J.J. Synergy as a rationale for phage therapy using phage cocktails. PeerJ 2014, 2, e590. [Google Scholar] [CrossRef]
- Ceri, H.; Olson, M.E.; Stremick, C.; Read, R.R.; Morck, D.; Buret, A. The calgary biofilm device: New technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J. Clin. Microbiol. 1999, 37, 1771–1776. [Google Scholar]
- Macia, M.D.; Rojo-Molinero, E.; Oliver, A. Antimicrobial susceptibility testing in biofilm-growing bacteria. Clin. Microbiol. Infect. 2014, 20, 981–990. [Google Scholar] [CrossRef]
- Sengupta, S.; Chattopadhyay, M.K.; Grossart, H.P. The multifaceted roles of antibiotics and antibiotic resistance in nature. Front Microbiol. 2013, 4, 47. [Google Scholar] [CrossRef]
- Abedon, S.T. Ecology of anti-biofilm agents II. Bacteriophage exploitation and biocontrol of biofilm bacteria. Pharmaceuticals 2015, 8, 559–589. [Google Scholar]
- Jacqueline, C.; Caillon, J. Impact of bacterial biofilm on the treatment of prosthetic joint infections. J. Antimicrob. Chemother. 2014, 69, i37–i40. [Google Scholar] [CrossRef]
- Pulcini, C.; Mainardi, J.L. Antimicrobial stewardship: An international emergency. Clin. Microbiol. Infect. 2014, 20, 947–948. [Google Scholar] [CrossRef] [Green Version]
- Smith, R.A.; M'ikanatha, N.M.; Read, A.F. Antibiotic resistance: a primer and call to action. Health Commun. 2015, 30, 309–314. [Google Scholar] [CrossRef]
- Varughese, C.A.; Vakil, N.H.; Phillips, K.M. Antibiotic-associated diarrhea: A refresher on causes and possible prevention with probiotics—continuing education article. J. Pharm. Pract. 2013, 26, 476–482. [Google Scholar] [CrossRef]
- Vangay, P.; Ward, T.; Gerber, J.S.; Knights, D. Antibiotics, pediatric dysbiosis, and disease. Cell Host Microbe 2015, 17, 553–564. [Google Scholar] [CrossRef]
- Gullberg, E.; Cao, S.; Berg, O.G.; Ilback, C.; Sandegren, L.; Hughes, D.; Andersson, D.I. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 2011, 7, e1002158. [Google Scholar] [CrossRef]
- Tello, A.; Austin, B.; Telfer, T.C. Selective pressure of antibiotic pollution on bacteria of importance to public health. Environ. Health Perspect. 2012, 120, 1100–1106. [Google Scholar] [CrossRef]
- Huang, R.; Ding, P.; Huang, D.; Yang, F. Antibiotic pollution threatens public health in China. Lancet 2015, 385, 773–774. [Google Scholar] [CrossRef]
- Allen, H.K.; Trachsel, J.; Looft, T.; Casey, T.A. Finding alternatives to antibiotics. Ann. N.Y. Acad. Sci. 2014, 1323, 91–100. [Google Scholar] [CrossRef]
- Penesyan, A.; Gillings, M.; Paulsen, I.T. Antibiotic discovery: Combatting bacterial resistance in cells and in biofilm communities. Molecules 2015, 20, 5286–5298. [Google Scholar] [CrossRef]
- Jolivet-Gougeon, A.; Bonnaure-Mallet, M. Biofilms as a mechanism of bacterial resistance. Drug Discov. Today Technol. 2014, 11, 49–56. [Google Scholar] [CrossRef]
- Olsen, I. Biofilm-specific antibiotic tolerance and resistance. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 877–886. [Google Scholar] [CrossRef]
- Davies, J. Are antibiotics naturally antibiotics? J. Ind. Microbiol. Biotechnol. 2006, 33, 496–499. [Google Scholar] [CrossRef]
- Hibbing, M.E.; Fuqua, C.; Parsek, M.R.; Peterson, S.B. Bacterial competition: Surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 2010, 8, 15–25. [Google Scholar] [CrossRef]
- Curtright, A.J.; Abedon, S.T. Phage therapy: Emergent property pharmacology. J. Bioanal. Biomed. 2011, S6. [Google Scholar] [CrossRef]
- Beloin, C.; Renard, S.; Ghigo, J.M.; Lebeaux, D. Novel approaches to combat bacterial biofilms. Curr. Opin. Pharmacol. 2014, 18, 61–68. [Google Scholar] [CrossRef]
- Prax, M.; Bertram, R. Metabolic aspects of bacterial persisters. Front Cell Infect. Microbiol. 2014, 4, 148. [Google Scholar] [CrossRef]
- Conlon, B.P.; Rowe, S.E.; Lewis, K. Persister cells in biofilm associated infections. Adv. Exp. Med. Biol. 2015, 831, 1–9. [Google Scholar]
- Dusane, D.H.; Damare, S.R.; Nancharaiah, Y.V.; Ramaiah, N.; Venugopalan, V.P.; Kumar, A.R.; Zinjarde, S.S. Disruption of microbial biofilms by an extracellular protein isolated from epibiotic tropical marine strain of Bacillus licheniformis. PLoS ONE 2013, 8, e64501. [Google Scholar] [CrossRef]
- Stallings, J.H. Soil produced antibiotics—plant disease and insect control. Bacteriol. Rev. 1954, 18, 131–146. [Google Scholar]
- Williams, D.H.; Stone, M.J.; Hauck, P.R.; Rahman, S.K. Why are secondary metabolites (natural products) biosynthesized? J. Nat. Prod. 1989, 52, 1189–1208. [Google Scholar] [CrossRef]
- Aminov, R.I. The role of antibiotics and antibiotic resistance in nature. Environ. Microbiol. 2009, 11, 2970–2988. [Google Scholar] [CrossRef]
- Aminov, R.I. Biotic acts of antibiotics. Front Microbiol. 2013, 4, 241. [Google Scholar] [CrossRef]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious disease. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Kaplan, J.B. Biofilm dispersal: Mechanisms, clinical implications, and potential therapeutic uses. J. Dent. Res. 2010, 89, 205–218. [Google Scholar] [CrossRef]
- Williams, G.C. Pleiotropy, natural selection, and the evolution of senescence. Evolution 1957, 11, 398–411. [Google Scholar] [CrossRef]
- Presloid, J.B.; Ebendick-Corp, B.E.; Zarate, S.; Novella, I.S. Antagonistic pleiotropy involving promoter sequences in a virus. J. Mol. Biol. 2008, 382, 342–352. [Google Scholar] [CrossRef]
- Rodriguez-Verdugo, A.; Carrillo-Cisneros, D.; Gonzalez-Gonzalez, A.; Gaut, B.S.; Bennett, A.F. Different tradeoffs result from alternate genetic adaptations to a common environment. Proc. Natl. Acad. Sci. USA 2014, 111, 12121–12126. [Google Scholar] [CrossRef]
- Elena, S.F.; Sanjuán, R. Climb every mountain? Science 2003, 302, 2074–2075. [Google Scholar] [CrossRef]
- Bleich, R.; Watrous, J.D.; Dorrestein, P.C.; Bowers, A.A.; Shank, E.A. Thiopeptide antibiotics stimulate biofilm formation in Bacillus subtilis. Proc. Natl. Acad. Sci. USA 2015, 112, 3086–3091. [Google Scholar] [CrossRef]
- Narisawa, N.; Haruta, S.; Arai, H.; Ishii, M.; Igarashi, Y. Coexistence of antibiotic-producing and antibiotic-sensitive bacteria in biofilms is mediated by resistant bacteria. Appl. Environ. Microbiol. 2008, 74, 3887–3894. [Google Scholar] [CrossRef]
- Gilbert, P.; Allison, D.G.; Jacob, A.; Korber, D.; Wolfaart, G.; Foley, I. Immigration of planktonic Enterococcus faecalis cells into mature E. facealis biofilms. In Biofilms: Community Interactions and Control; Wimpenny, J.W.T., Handley, P., Gilbert, P., Lappin-Scott, H.M., Eds.; Bioline: Cardiff, UK, 1997; pp. 133–141. [Google Scholar]
- Lanchester, F.W. Aircraft in Warfare, The Dawn of the Fourth Arm; Constable and Company Limited: London, UK, 1916. [Google Scholar]
- Tzu, S.; Conners, S.; Giles, L. The Art of War by Sun Tzu, classic ed.; El Paso Norte Press: El Paso, TX, USA, 2009. [Google Scholar]
- Borges, R.M. Warring ants: Lessons from Lanchester's laws of combat? J. Biosci. 2002, 27, 75–78. [Google Scholar] [CrossRef]
- Oliveira, N.M.; Martinez-Garcia, E.; Xavier, J.; Durham, W.M.; Kolter, R.; Kim, W.; Foster, K.R. Biofilm formation as a response to ecological competition. PLoS. Biol. 2015, 13, e1002191. [Google Scholar] [CrossRef]
- Price-Whelan, A.; Dietrich, L.E.; Newman, D.K. Rethinking “secondary” metabolism: Physiological roles for phenazine antibiotics. Nat. Chem. Biol. 2006, 2, 71–78. [Google Scholar] [CrossRef]
- Tait, K.; Sutherland, I.W. Antagonistic interactions amongst bacteriocin-producing enteric bacteria in dual species biofilms. J. Appl. Microbiol. 2002, 93, 345–352. [Google Scholar] [CrossRef]
- Houry, A.; Gohar, M.; Deschamps, J.; Tischenko, E.; Aymerich, S.; Gruss, A.; Briandet, R. Bacterial swimmers that infiltrate and take over the biofilm matrix. Proc. Natl. Acad. Sci. USA 2012, 109, 13088–13093. [Google Scholar] [CrossRef]
- Brown, M.R.W.; Allison, D.G.; Gilbert, G. Resistance of bacterial biofilms to antibiotics: A growth-rate related effect? J. Antimicrob. Chemother. 1988, 22, 777–780. [Google Scholar] [CrossRef]
- Rafii, F.; Hart, M.E. Antimicrobial resistance in clinically important biofilms. World J. Pharmacol. 2015, 4, 31–46. [Google Scholar] [CrossRef]
- Burmolle, M.; Webb, J.S.; Rao, D.; Hansen, L.H.; Sorensen, S.J.; Kjelleberg, S. Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Appl. Environ. Microbiol. 2006, 72, 3916–3923. [Google Scholar] [CrossRef]
- Elias, S.; Banin, E. Multi-species biofilms: Living with friendly neighbors. FEMS Microbiol. Rev. 2012, 36, 990–1004. [Google Scholar] [CrossRef]
- Connell, J.L.; Ritschdorff, E.T.; Whiteley, M.; Shear, J.B. 3D printing of microscopic bacterial communities. Proc. Natl. Acad. Sci. USA 2013, 110, 18380–18385. [Google Scholar] [CrossRef]
- Burmolle, M.; Ren, D.; Bjarnsholt, T.; Sorensen, S.J. Interactions in multispecies biofilms: Do they actually matter? Trends Microbiol. 2014, 22, 84–91. [Google Scholar] [CrossRef]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef]
- Matz, C. Biofilms as refuge against predation. In The Biofilm Mode of Life: Mechanisms and Adaptations; Kjelleberg, S., Givskov, M., Eds.; Horizon Bioscience: Norfolk, UK, 2007; pp. 195–213. [Google Scholar]
- Rao, D.; Webb, J.S.; Kjelleberg, S. Competitive interactions in mixed-species biofilms containing the marine bacterium Pseudoalteromonas tunicata. Appl. Environ. Microbiol. 2005, 71, 1729–1736. [Google Scholar] [CrossRef]
- Xiao, Y.; Wei, X.; Ebright, R.; Wall, D. Antibiotic production by myxobacteria plays a role in predation. J. Bacteriol. 2011, 193, 4626–4633. [Google Scholar] [CrossRef]
- McBride, M.J.; Zusman, D.R. Behavioral analysis of single cells of Myxococcus xanthus in response to prey cells of Escherichia coli. FEMS Microbiol. Lett. 1996, 137, 227–231. [Google Scholar] [CrossRef]
- Kadouri, D.; O'Toole, G.A. Susceptibility of biofilms to Bdellovibrio bacteriovorus attack. Appl. Environ. Microbiol. 2005, 71, 4044–4051. [Google Scholar] [CrossRef]
- Berleman, J.E.; Allen, S.; Danielewicz, M.A.; Remis, J.P.; Gorur, A.; Cunha, J.; Hadi, M.Z.; Zusman, D.R.; Northen, T.R.; Witkowska, H.E.; et al. The lethal cargo of Myxococcus xanthus outer membrane vesicles. Front Microbiol. 2014, 5, 474. [Google Scholar] [CrossRef]
- Iebba, V.; Totino, V.; Santangelo, F.; Gagliardi, A.; Ciotoli, L.; Virga, A.; Ambrosi, C.; Pompili, M.; De Biase, R.V.; Selan, L.; et al. Bdellovibrio bacteriovorus directly attacks Pseudomonas aeruginosa and Staphylococcus aureus cystic fibrosis isolates. Front Microbiol. 2014, 5, 280. [Google Scholar] [CrossRef]
- Al-Bakri, A.G.; Gilbert, P.; Allison, D.G. Immigration and emigration of Burkholderia cepacia and Pseudomonas aeruginosa between and within mixed biofilm communities. J. Appl. Microbiol. 2004, 96, 455–463. [Google Scholar] [CrossRef]
- Banks, M.K.; Bryers, J.D. Bacterial species dominance within a binary culture biofilm. Appl. Environ. Microbiol. 1991, 57, 1974–1979. [Google Scholar]
- Bull, J.J.; Regoes, R.R. Pharmacodynamics of non-replicating viruses, bacteriocins and lysins. Proc. R. Soc. Lond. B Biol. Sci. 2006, 273, 2703–2712. [Google Scholar] [CrossRef]
- Abedon, S. Phage therapy pharmacology: Calculating phage dosing. Adv. Appl. Microbiol. 2011, 77, 1–40. [Google Scholar]
- Stewart, F.M.; Levin, B.R. The population biology of bacterial viruses: Why be temperate. Theor. Pop. Biol. 1984, 26, 93–117. [Google Scholar] [CrossRef]
- Paul, J.H. Prophages in marine bacteria: Dangerous molecular time bombs or the key to survival in the seas? ISME J. 2008, 2, 579–589. [Google Scholar] [CrossRef]
- Gardner, A.; West, S.A.; Buckling, A. Bacteriocins, spite and virulence. Proc. Roy. Soc. Lond. B 2004, 271, 1529–1535. [Google Scholar] [CrossRef]
- Nedelcu, A.M.; Driscoll, W.W.; Durand, P.M.; Herron, M.D.; Rashidi, A. On the paradigm of altruistic suicide in the unicellular world. Evolution 2011, 65, 3–20. [Google Scholar] [CrossRef]
- Solheim, H.T.; Sekse, C.; Urdahl, A.M.; Wasteson, Y.; Nesse, L.L. Biofilm as an environment for dissemination of stx genes by transduction. Appl. Environ. Microbiol. 2013, 79, 896–900. [Google Scholar] [CrossRef]
- Yan, L.; Boyd, K.G.; Adams, D.R.; Burgess, J.G. Biofilm-specific cross-species induction of antimicrobial compounds in bacilli. Appl. Environ. Microbiol. 2003, 69, 3719–3727. [Google Scholar] [CrossRef]
- Sarkar, S.; Roy, D.; Mukherjee, J. Production of a potentially novel antimicrobial compound by a biofilm-forming marine Streptomyces sp. in a niche-mimic rotating disk bioreactor. Bioprocess. Biosyst. Eng. 2010, 33, 207–217. [Google Scholar] [CrossRef]
- Kreth, J.; Merritt, J.; Shi, W.; Qi, F. Competition and coexistence between Streptococcus mutans and Streptococcus sanguinis in the dental biofilm. J. Bacteriol. 2005, 187, 7193–7203. [Google Scholar] [CrossRef]
- Farmer, J.T.; Shimkevitch, A.V.; Reilly, P.S.; Mlynek, K.D.; Jensen, K.S.; Callahan, M.T.; Bushaw-Newton, K.L.; Kaplan, J.B. Environmental bacteria produce abundant and diverse antibiofilm compounds. J. Appl. Microbiol. 2014, 117, 1663–1673. [Google Scholar] [CrossRef]
- Nadell, C.D.; Drescher, K.; Wingreen, N.S.; Bassler, B.L. Extracellular matrix structure governs invasion resistance in bacterial biofilms. ISME J. 2015, 9, 1700–1709. [Google Scholar] [CrossRef]
- Kroiss, J.; Kaltenpoth, M.; Schneider, B.; Schwinger, M.G.; Hertweck, C.; Maddula, R.K.; Strohm, E.; Svatos, A. Symbiotic streptomycetes provide antibiotic combination prophylaxis for wasp offspring. Nat. Chem. Biol. 2010, 6, 261–263. [Google Scholar] [CrossRef]
- Moons, P.; Michiels, C.W.; Aertsen, A. Bacterial interactions in biofilms. Crit. Rev. Microbiol. 2009, 35, 157–168. [Google Scholar] [CrossRef]
- Prol Garcia, M.J.; D'Alvise, P.W.; Rygaard, A.M.; Gram, L. Biofilm formation is not a prerequisite for production of the antibacterial compound tropodithietic acid in Phaeobacter inhibens DSM17395. J. Appl. Microbiol. 2014, 117, 1592–1600. [Google Scholar] [CrossRef]
- Rendueles, O.; Beloin, C.; Latour-Lambert, P.; Ghigo, J.M. A new biofilm-associated colicin with increased efficiency against biofilm bacteria. ISME J. 2014, 8, 1275–1288. [Google Scholar] [CrossRef]
- Moons, P.; Van, H.R.; Aertsen, A.; Vanoirbeek, K.; Engelborghs, Y.; Michiels, C.W. Role of quorum sensing and antimicrobial component production by Serratia plymuthica in formation of biofilms, including mixed biofilms with Escherichia coli. Appl. Environ. Microbiol. 2006, 72, 7294–7300. [Google Scholar] [CrossRef]
- Schluter, J.; Nadell, C.D.; Bassler, B.L.; Foster, K.R. Adhesion as a weapon in microbial competition. ISME J. 2015, 9, 139–149. [Google Scholar] [CrossRef]
- Nadell, C.D.; Bassler, B.L. A fitness trade-off between local competition and dispersal in Vibrio cholerae biofilms. Proc. Natl. Acad. Sci. USA 2011, 108, 14181–14185. [Google Scholar] [CrossRef]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef]
- Scali, C.; Kunimoto, B. An update on chronic wounds and the role of biofilms. J. Cutan. Med. Surg. 2013, 17, 371–376. [Google Scholar]
- Cooper, R.A.; Bjarnsholt, T.; Alhede, M. Biofilms in wounds: A review of present knowledge. J. Wound. Care 2014, 23, 570–580. [Google Scholar] [CrossRef]
- Levin, S.A. Public goods in relation to competition, cooperation, and spite. Proc. Natl. Acad. Sci. USA 2014, 111, 10838–10845. [Google Scholar] [CrossRef]
- Mitri, S.; Foster, K.R. The genotypic view of social interactions in microbial communities. Ann. Rev. Genet. 2013, 47, 247–273. [Google Scholar] [CrossRef]
- Drescher, K.; Nadell, C.D.; Stone, H.A.; Wingreen, N.S.; Bassler, B.L. Solutions to the public goods dilemma in bacterial biofilms. Curr. Biol. 2014, 24, 50–55. [Google Scholar] [CrossRef]
- Cordero, O.X.; Wildschutte, H.; Kirkup, B.; Proehl, S.; Ngo, L.; Hussain, F.; Le, R.F.; Mincer, T.; Polz, M.F. Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 2012, 337, 1228–1231. [Google Scholar] [CrossRef]
- Hwang, I.Y.; Tan, M.H.; Koh, E.; Ho, C.L.; Poh, C.L.; Chang, M.W. Reprogramming microbes to be pathogen-seeking killers. ACS Synth. Biol. 2014, 3, 228–237. [Google Scholar] [CrossRef]
- Baelo, A.; Levato, R.; Julian, E.; Crespo, A.; Astola, J.; Gavalda, J.; Engel, E.; Mateos-Timoneda, M.A.; Torrents, E. Disassembling bacterial extracellular matrix with DNase-coated nanoparticles to enhance antibiotic delivery in biofilm infections. J. Control Release 2015, 209, 150–158. [Google Scholar] [CrossRef]
- Martin, C.; Low, W.L.; Gupta, A.; Amin, M.C.; Radecka, I.; Britland, S.T.; Raj, P.; Kenward, K.M. Strategies for antimicrobial drug delivery to biofilm. Curr. Pharm. Des. 2015, 21, 43–66. [Google Scholar] [CrossRef]
- Alhede, M.; Kragh, K.N.; Qvortrup, K.; Allesen-Holm, M.; van Gennip, M.; Christensen, L.D.; Jensen, P.O.; Nielsen, A.K.; Parsek, M.; Wozniak, D.; et al. Phenotypes of non-attached Pseudomonas aeruginosa aggregates resemble surface attached biofilm. PLoS ONE 2011, 6, e27943. [Google Scholar] [CrossRef]
- Derlon, N.; Koch, N.; Eugster, B.; Posch, T.; Pernthaler, J.; Pronk, W.; Morgenroth, E. Activity of metazoa governs biofilm structure formation and enhances permeate flux during gravity-driven membrane (GDM) filtration. Water Res. 2013, 47, 2085–2095. [Google Scholar] [CrossRef]
- Erken, M.; Farrenschon, N.; Speckmann, S.; Arndt, H.; Weitere, M. Quantification of individual flagellate-bacteria interactions within semi-natural biofilms. Protist 2012, 163, 632–642. [Google Scholar] [CrossRef]
- Abedon, S.T. Bacteriophages and Biofilms: Ecology, Phage Therapy, Plaques; Nova Science Publishers: New York, NY, USA, 2011. [Google Scholar]
- Brüssow, H. Bacteriophage-host interaction: From splendid isolation into a messy reality. Curr. Opin. Microbiol. 2013, 16, 500–506. [Google Scholar] [CrossRef]
- Fan, X.; Li, W.; Zheng, F.; Xie, J. Bacteriophage inspired antibiotics discovery against infection involved biofilm. Crit. Rev. Eukaryot. Gene Expr. 2013, 23, 317–326. [Google Scholar] [CrossRef]
- Harper, D.R.; Parracho, H.M.R.; Walker, J.; Sharp, R.; Hughes, G.; Werthrén, M.; Lehman, S.; Morales, S. Bacteriophages and biofilms. Antibiotics 2014, 3, 270–284. [Google Scholar] [CrossRef]
- Parasion, S.; Kwiatek, M.; Gryko, R.; Mizak, L.; Malm, A. Bacteriophages as an alternative strategy for fighting biofilm development. Pol. J. Microbiol. 2014, 63, 137–145. [Google Scholar]
- Sillankorva, S.; Azeredo, J. Bacteriophage attack as an anti-biofilm strategy. Methods Mol. Biol. 2014, 1147, 277–285. [Google Scholar]
- Sillankorva, S.; Azeredo, J. The use of bacteriophages and bacteriophage-derived enzymes for clinically relevant biofilm control. In Phage Therapy: Current Research and Applications; Borysowski, J., Miedzybrodzki, R., Górski, A., Eds.; Caister Academic Press: Norfolk, UK, 2014. [Google Scholar]
- Chan, B.K.; Abedon, S.T. Bacteriophages and their enzymes in biofilm control. Curr. Pharm. Des. 2015, 21, 85–99. [Google Scholar] [CrossRef]
- Abedon, S.T. Kinetics of phage-mediated biocontrol of bacteria. Foodborne Pathog. Dis. 2009, 6, 807–815. [Google Scholar] [CrossRef]
- Kutter, E.; De Vos, D.; Gvasalia, G.; Alavidze, Z.; Gogokhia, L.; Kuhl, S.; Abedon, S.T. Phage therapy in clinical practice: treatment of human infections. Curr. Pharm. Biotechnol. 2010, 11, 69–86. [Google Scholar] [CrossRef]
- Abedon, S.T.; Kuhl, S.J.; Blasdel, B.G.; Kutter, E.M. Phage treatment of human infections. Bacteriophage 2011, 1, 66–85. [Google Scholar] [CrossRef]
- Abedon, S.T. Phage therapy of pulmonary infections. Bacteriophage 2015. [Google Scholar] [CrossRef]
- Olszowska-Zaremba, N.; Borysowski, J.; Dabrowska, J.; Górski, A. Phage translocation, safety, and immunomodulation. In Bacteriophages in Health and Disease; Hyman, P., Abedon, S.T., Eds.; CABI Press: Wallingford, UK, 2012; pp. 168–184. [Google Scholar]
- Abedon, S.T.; Thomas-Abedon, C. Phage therapy pharmacology. Curr. Pharm. Biotechnol. 2010, 11, 28–47. [Google Scholar] [CrossRef]
- Abedon, S.T. Phage therapy: Eco-physiological pharmacology. Scientifica 2014, 2014, 581639. [Google Scholar] [CrossRef]
- Abedon, S.T. Spatial vulnerability: Bacterial arrangements, microcolonies, and biofilms as responses to low rather than high phage densities. Viruses 2012, 4, 663–687. [Google Scholar] [CrossRef]
- Shen, Y.; Mitchell, M.S.; Donovan, D.M.; Nelson, D.C. Phage-based enzybiotics. In Bacteriophages in Health and Disease; Hyman, P., Abedon, S.T., Eds.; CABI Press: Wallingford, UK, 2012; pp. 217–239. [Google Scholar]
- Hyman, P.; Abedon, S.T. Bacteriophage host range and bacterial resistance. Adv. Appl. Microbiol. 2010, 70, 217–248. [Google Scholar]
- Abedon, S.T. Envisaging bacteria as phage targets. Bacteriophage 2011, 1, 228–230. [Google Scholar] [CrossRef]
- Abedon, S.T. Bacterial “immunity” against bacteriophages. Bacteriophage 2012, 2, 50–54. [Google Scholar] [CrossRef]
- Johnston, N. Viral Trojan horse for combating tuberculosis. Drug Discov. Today 2002, 7, 333–335. [Google Scholar] [CrossRef]
- Broxmeyer, L.; Sosnowskai, D.; Miltner, E.; Chacón, O.; Wagner, D.; McGarvey, J.; Barletta, R.G.; Bermudez, L.E. Killing of Mycobacterium avium and Mycobacterium tuberculosis by a mycobacteriophage delivered by a nonvirulent mycobacterium: A model for phage therapy of intracellular bacterial pathogens. J. Infect. Dis. 2002, 186, 1155–1160. [Google Scholar] [CrossRef]
- Dawkins, R.; Krebs, J.R. Arms races between and within species. Proc. Soc. Exp. Biol. Med. 1979, 205, 489–511. [Google Scholar] [CrossRef]
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abedon, S.T. Ecology of Anti-Biofilm Agents I: Antibiotics versus Bacteriophages. Pharmaceuticals 2015, 8, 525-558. https://doi.org/10.3390/ph8030525
Abedon ST. Ecology of Anti-Biofilm Agents I: Antibiotics versus Bacteriophages. Pharmaceuticals. 2015; 8(3):525-558. https://doi.org/10.3390/ph8030525
Chicago/Turabian StyleAbedon, Stephen T. 2015. "Ecology of Anti-Biofilm Agents I: Antibiotics versus Bacteriophages" Pharmaceuticals 8, no. 3: 525-558. https://doi.org/10.3390/ph8030525
APA StyleAbedon, S. T. (2015). Ecology of Anti-Biofilm Agents I: Antibiotics versus Bacteriophages. Pharmaceuticals, 8(3), 525-558. https://doi.org/10.3390/ph8030525