Protein Detection with Aptamer Biosensors
Abstract
:1. Introduction
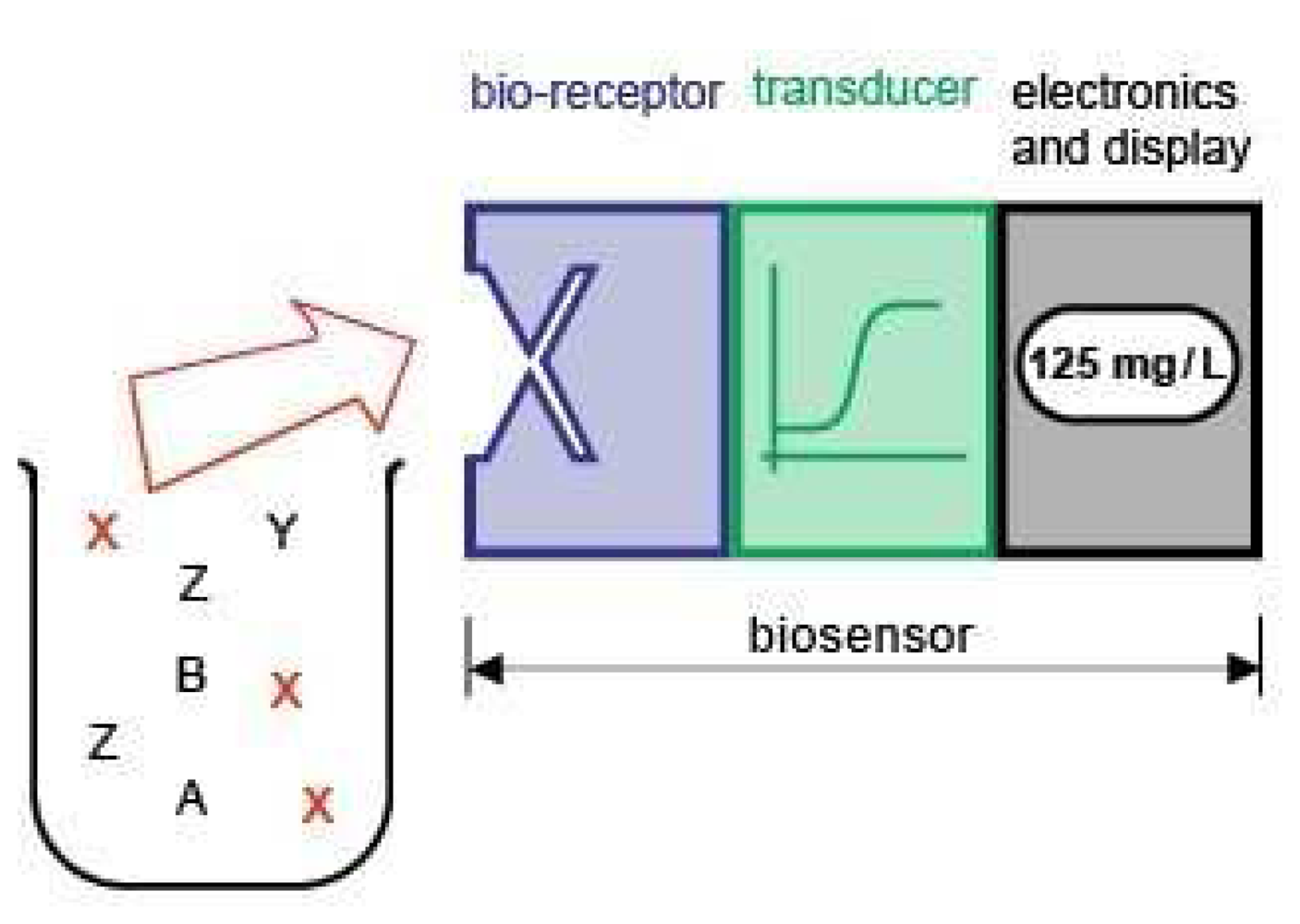
2. Biosensor
3. Protein biosensor detection principles based on aptamers
3.1. Electrochemical aptasensors
3.2. Optical aptasensors
3.3. Mass sensitive aptasensors
3.4. Potentiometric aptasensors
4. Aptamer biosensors for protein detection
5. Conclusions
Acknowledgments
References and Notes
- O'Sullivan, C.K. Aptasensors - the future of biosensing. Anal. Bioanal. Chem. 2002, 372, 44–48. [Google Scholar]
- Stadtherr, K.; Wolf, H.; Lindner, P. An aptamer-based protein biochip. Anal. Chem. 2005, 77, 3437–3443. [Google Scholar]
- Jenison, R.D.; Gill, S.C.; Pardi, A.; Polisky, B. High-resolution molecular discrimination by RNA. Science. 1994, 263, 1425–1429. [Google Scholar]
- Michaud, M.; Jourdan, E.; Villet, A.; Ravel, A.; Grosset, C.; Peyrin, E. A DNA aptamer as a new target-specific chiral selector for HPLC. J. Am. Chem. Soc. 2003, 125, 8672–8679. [Google Scholar]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990, 346, 818–822. [Google Scholar]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990, 249, 505–510. [Google Scholar]
- Stoltenburg, R.; Reinemann, C.; Strehlitz, B. SELEX-A (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomol. Eng. 2007, 24, 381–403. [Google Scholar]
- Mukhopadhyay, R. Aptamers are ready for the spotlight. Anal. Chem. 2005, 77, 114A–118A. [Google Scholar]
- Jayasena, S.D. Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar]
- Willner, I.; Zayats, M. Electronic Aptamer-Based Sensors. Angew. Chem. Int. Ed Engl. 2007, 46, 6408–6418. [Google Scholar]
- Thevenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical Biosensors: Recommended Definitions and Classification. Pure Appl. Chem. 1999, 71, 2333–2348. [Google Scholar]
- McCauley, T.G.; Hamaguchi, N.; Stanton, M. Aptamer-based biosensor arrays for detection and quantification of biological macromolecules. Anal. Biochem. 2003, 319, 244–250. [Google Scholar]
- Kubik, M.F.; Bell, C.; Fitzwater, T.; Watson, S.R.; Tasset, D.M. Isolation and characterization of 2′-fluoro-, 2′-amino-, and 2′-fluoro/amino-modified RNA ligands to human IFN-gamma that inhibit receptor binding. J. Immunol. 1997, 159, 259–267. [Google Scholar]
- Kujau, M.J.; Wolfl, S. Intramolecular derivatization of 2′-amino-pyrimidine modified RNA with functional groups that is compatible with re-amplification. Nucleic Acids Res. 1998, 26, 1851–1853. [Google Scholar]
- Minunni, M.; Tombelli, S.; Gullotto, A.; Luzi, E.; Mascini, M. Development of biosensors with aptamers as bio-recognition element: the case of HIV-1 Tat protein. Biosens. Bioelectron. 2004, 20, 1149–1156. [Google Scholar]
- Xu, D.K.; Xu, D.W.; Yu, X.B.; Liu, Z.H.; He, W.; Ma, Z.Q. Label-free electrochemical detection for aptamer-based array electrodes. Anal. Chem. 2005, 77, 5107–5113. [Google Scholar]
- Schlecht, U.; Malave, A.; Gronewold, T.; Tewes, M.; Lohndorf, M. Comparison of antibody and aptamer receptors for the specific detection of thrombin with a nanometer gap-sized impedance biosensor. Anal. Chim. Acta. 2006, 573, 65–68. [Google Scholar]
- Rodriguez, M.C.; Kawde, A.N.; Wang, J. Aptamer biosensor for label-free impedance spectroscopy detection of proteins based on recognition-induced switching of the surface charge. Chem. Commun. 2005, 4267–4269. [Google Scholar]
- Cheng, A.K.; Ge, B.; Yu, H.Z. Aptamer-based biosensors for label-free voltammetric detection of lysozyme. Anal. Chem. 2007, 79, 5158–5164. [Google Scholar]
- Radi, A.E.; Sanchez, J.L.A.; Baldrich, E.; O'Sullivan, C.K. Reusable impedimetric aptasensor. Anal. Chem. 2005, 77, 6320–6323. [Google Scholar]
- Bang, G.S.; Cho, S.; Kim, B.G. A novel electrochemical detection method for aptamer biosensors. Biosens. Bioelectron. 2005, 21, 863–870. [Google Scholar]
- Mir, M.; Katakis, I. Aptamers as elements of bioelectronic devices. Mol. Biosyst. 2007, 3, 620–622. [Google Scholar]
- Xiao, Y.; Lubin, A.A.; Heeger, A.J.; Plaxco, K.W. Label-free electronic detection of thrombin in blood serum by using an aptamer-based sensor. Angew. Chem. Int. Ed Engl. 2005, 44, 5456–5459. [Google Scholar]
- Ikebukuro, K.; Kiyohara, C.; Sode, K. Novel electrochemical sensor system for protein using the aptamers in sandwich manner. Biosens. Bioelectron. 2005, 20, 2168–2172. [Google Scholar]
- Baldrich, E.; Restrepo, A.; O'Sullivan, C.K. Aptasensor development: Elucidation of critical parameters for optimal aptamer performance. Anal. Chem. 2004, 76, 7053–7063. [Google Scholar]
- Wang, Z.; Wilkop, T.; Xu, D.; Dong, Y.; Ma, G.; Cheng, Q. Surface plasmon resonance imaging for affinity analysis of aptamer-protein interactions with PDMS microfluidic chips. Anal. Bioanal. Chem. 2007, 389, 819–825. [Google Scholar]
- Tombelli, S.; Minunni, A.; Luzi, E.; Mascini, M. Aptamer-based biosensors for the detection of HIV-1 Tat protein. Bioelectrochemistry. 2005, 67, 135–141. [Google Scholar]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature. 1992, 355, 564–566. [Google Scholar]
- Potyrailo, R.A.; Conrad, R.C.; Ellington, A.D.; Hieftje, G.M. Adapting selected nucleic acid ligands (aptamers) to biosensors. Anal. Chem. 1998, 70, 3419–3425. [Google Scholar]
- Tang, J.; Yu, T.; Guo, L.; Xie, J.; Shao, N.; He, Z. In vitro selection of DNA aptamer against abrin toxin and aptamer-based abrin direct detection. Biosens. Bioelectron. 2007, 22, 2456–2463. [Google Scholar]
- Pavlov, V.; Xiao, Y.; Shlyahovsky, B.; Willner, I. Aptamer-functionalized Au nanoparticles for the amplified optical detection of thrombin. J. Am. Chem. Soc. 2004, 126, 11768–11769. [Google Scholar]
- Maehashi, K.; Katsura, T.; Kerman, K.; Takamura, Y.; Matsumoto, K.; Tamiya, E. Label-free protein biosensor based on aptamer-modified carbon nanotube field-effect transistors. Anal. Chem. 2007, 79, 782–787. [Google Scholar]
- Mir, M.; Vreeke, M.; Katakis, L. Different strategies to develop an electrochemical thrombin aptasensor. Electrochem. Commun. 2006, 8, 505–511. [Google Scholar]
- Hansen, J.A.; Wang, J.; Kawde, A.N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar]
- Liss, M.; Petersen, B.; Wolf, H.; Prohaska, E. An aptamer-based quartz crystal protein biosensor. Anal. Chem. 2002, 74, 4488–4495. [Google Scholar]
| Abbreviations | |
|---|---|
| bFGF | Basic fibroblast growth factor |
| CNT-FET | Carbon nanotube field-effect transistor |
| CMOS | Complementary metal–oxide–semiconductor |
| DNA, ssDNA | Desoxyribonucleic acid, single stranded desoxyribonucleic acid |
| ELISA | Enzyme linked immunosorbent assay |
| FET | Field effect transistor |
| FIS | Faradaic Impedance Spectroscopy |
| HSA | Human serum albumin |
| IMPDH | Inosine monophosphate dehydrogenase |
| IUPAC | International Union of Pure and Applied Chemistry |
| Kd | Dissociation constant |
| mAb | Monoclonal Antibody |
| MB | Methylene Blue |
| QCM | Quartz crystal microbalance |
| RNA | Ribonucleic acid |
| RNAse | Ribonuclease |
| SELEX | Systematic evolution of ligands by exponential enrichment |
| SPR | Surface plasmon resonance |
| VEGF | Vascular endothelial growth factor |

| Target Protein | Aptamer | Type of Sensor, Reporter Unit | Detect. Limit, Linear Range | Ref |
|---|---|---|---|---|
| Thrombin | DNA beacon | ec, differential pulsevoltammetry, methyleneblue intercalator | 11 nM 0 … 50.8 nM | [21] |
| Thrombin | DNA | ec, impedancespectroscopy, [Fe(CN)6]3-/4- | 2 nM 5 … 35 nM | [20] |
| Thrombin | DNA thiolated/biotinylated | ec, differential pulsepolarography, p-nitroaniline/peroxidase/HRP | 80 nM/ 3.5 nM n.s. | [33] |
| Thrombin | DNA labeled with methylene blue | ec, alternating currentvoltammetry, methylene blue | n.s. n.s. (logarithmic dependence) | [23] |
| Thrombin | DNA labeled with pyrroquinoline quinone glucose dehydrogenase (PQQ)GDH, sandwich assay | ec, amperometry, glucose;single shot sensor | 10 nM 40 … 100 nM | [24] |
| Thrombin | DNA ferrocene labeled | optical combined with ec(cyclic voltammetry), eSPR/ ec, amperometry with co-immobilized microperoxidase | n.s. n.s. | [22] |
| Thrombin | DNA thiolated/ biotinylated | optical, SPR (Biacore™) | n.s. n.s. | [25] |
| Thrombin/ Lysozyme | n.s., thiolated | ec, square wave stripping voltammetry | 0.5 pM (20 … 500 ng/L)1 | [34] |
| Lysozyme | DNA | ec impedance spectroscopy, [Fe(CN)6]3-/4- | [18] | |
| Lysozyme | DNA | ec, [Ru(NH3)6]3+ cv peak decrease with target binding | 0.5 μg/ml 0.5 … 50 μg/ml | [19] |
| IgE | DNA thiolated | optical, SPR | 2 nM 8.4 … 84 nM | [26] |
| IgE | DNA biotinylated | mass sensitive, QCM | 100 μg/L 0 … 10 mg/L | [35] |
| IgE | DNA | carbon nanotube FET | 250 pM 250 pM … 20 nM | [32] |
| IgE | DNA | ec impedance spectroscopy, array | 0.1 nM 2.5 … 100 nM | [16] |
| HIV-Tat protein | RNA biotinylated | optical, SPR/ mass sensitive, QCM | n.s./ 0.25 ppm 0… 2.5 ppm/ 0 … 1.25 ppm | [27] |
| HIV-Tat 1 protein | RNA biotinylated | mass sensitive, QCM | 0.65 ppm 0… 2.5 ppm | [15] |
| Abrin toxin | DNA | optical, luminescence, molecular light switching intercalator | 1 nM 1 … 400 nM | [30] |
| Thrombin, bFGF, IMPDH, VEGF | RNA, DNA, fluorescently labeled | optical array, fluorescence polarization anisotropy | n.s. | [12] |
© 2008 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Strehlitz, B.; Nikolaus, N.; Stoltenburg, R. Protein Detection with Aptamer Biosensors. Sensors 2008, 8, 4296-4307. https://doi.org/10.3390/s8074296
Strehlitz B, Nikolaus N, Stoltenburg R. Protein Detection with Aptamer Biosensors. Sensors. 2008; 8(7):4296-4307. https://doi.org/10.3390/s8074296
Chicago/Turabian StyleStrehlitz, Beate, Nadia Nikolaus, and Regina Stoltenburg. 2008. "Protein Detection with Aptamer Biosensors" Sensors 8, no. 7: 4296-4307. https://doi.org/10.3390/s8074296
APA StyleStrehlitz, B., Nikolaus, N., & Stoltenburg, R. (2008). Protein Detection with Aptamer Biosensors. Sensors, 8(7), 4296-4307. https://doi.org/10.3390/s8074296







