Analysis of Microalgal Density Estimation by Using LASSO and Image Texture Features
Abstract
1. Introduction
2. Data Collection
2.1. Monitoring System
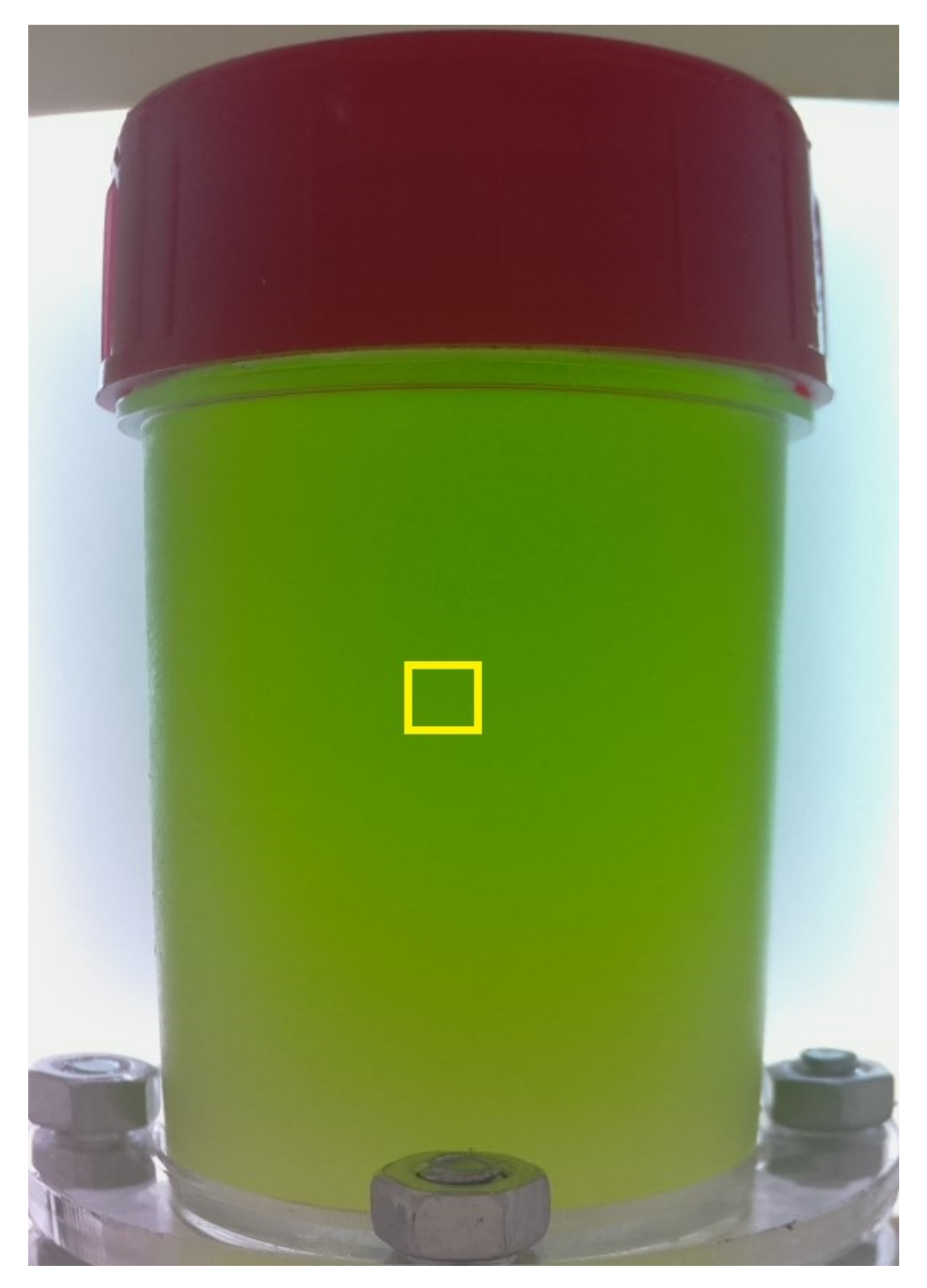
2.2. Preprocessing Data
3. Feature Design
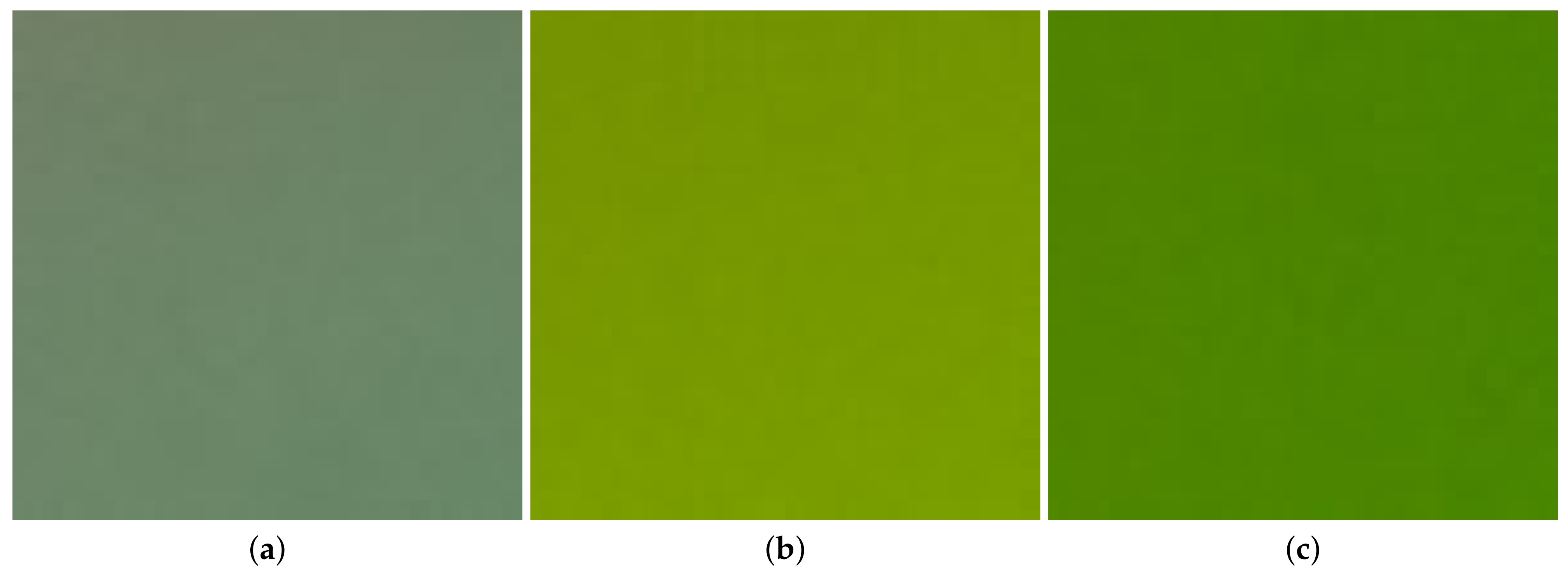
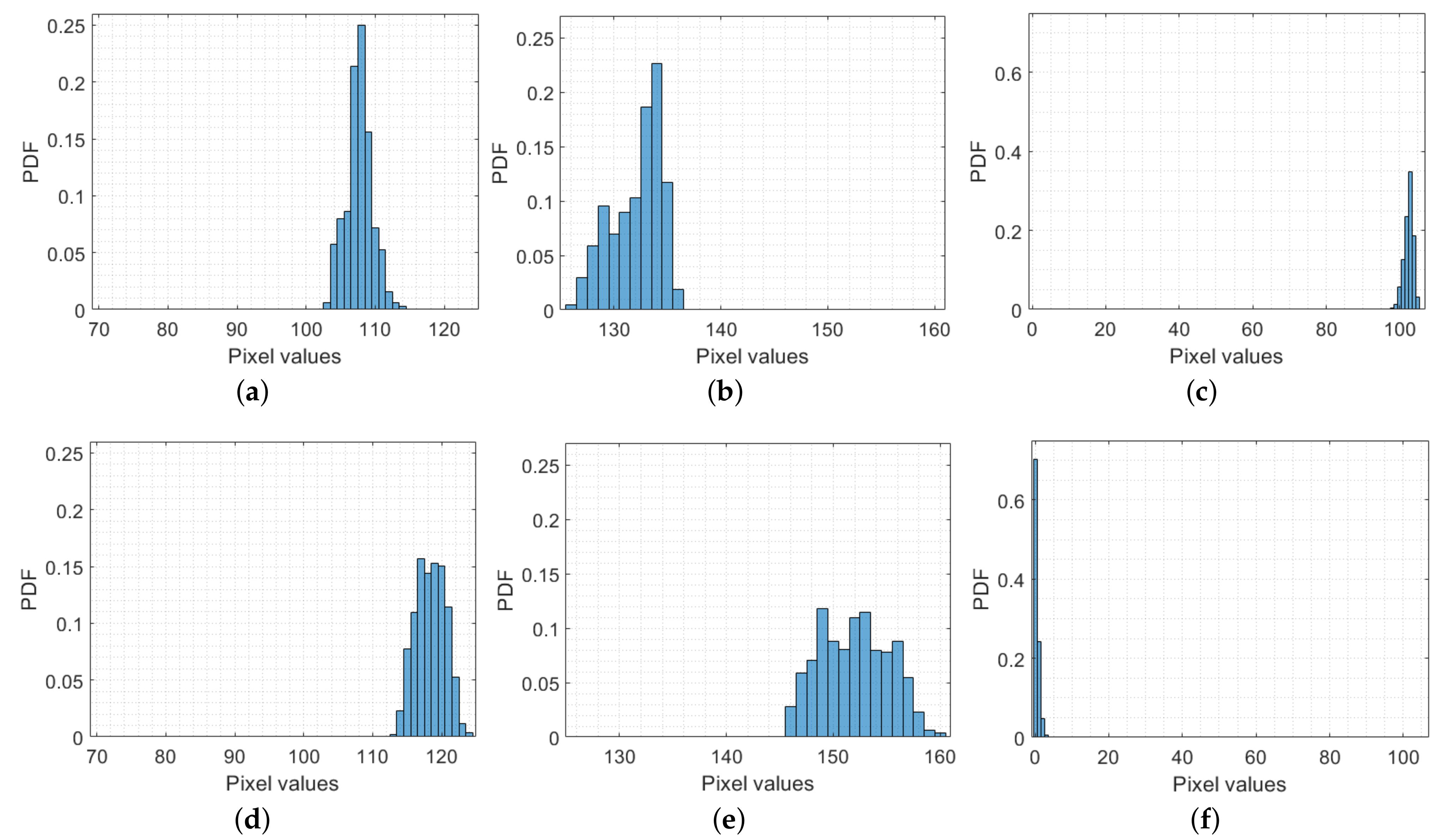
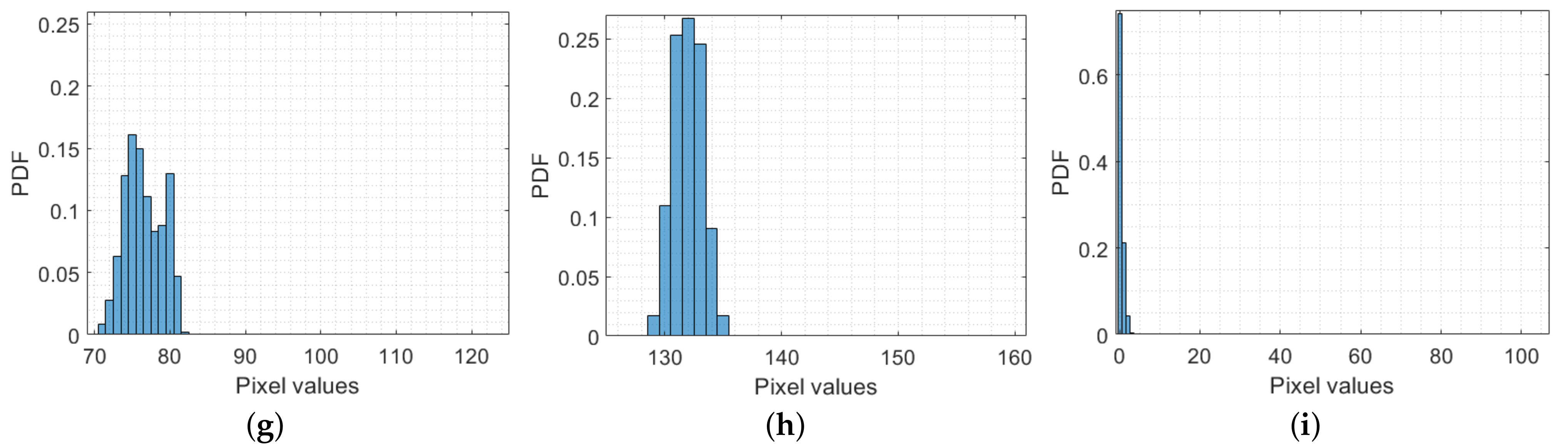
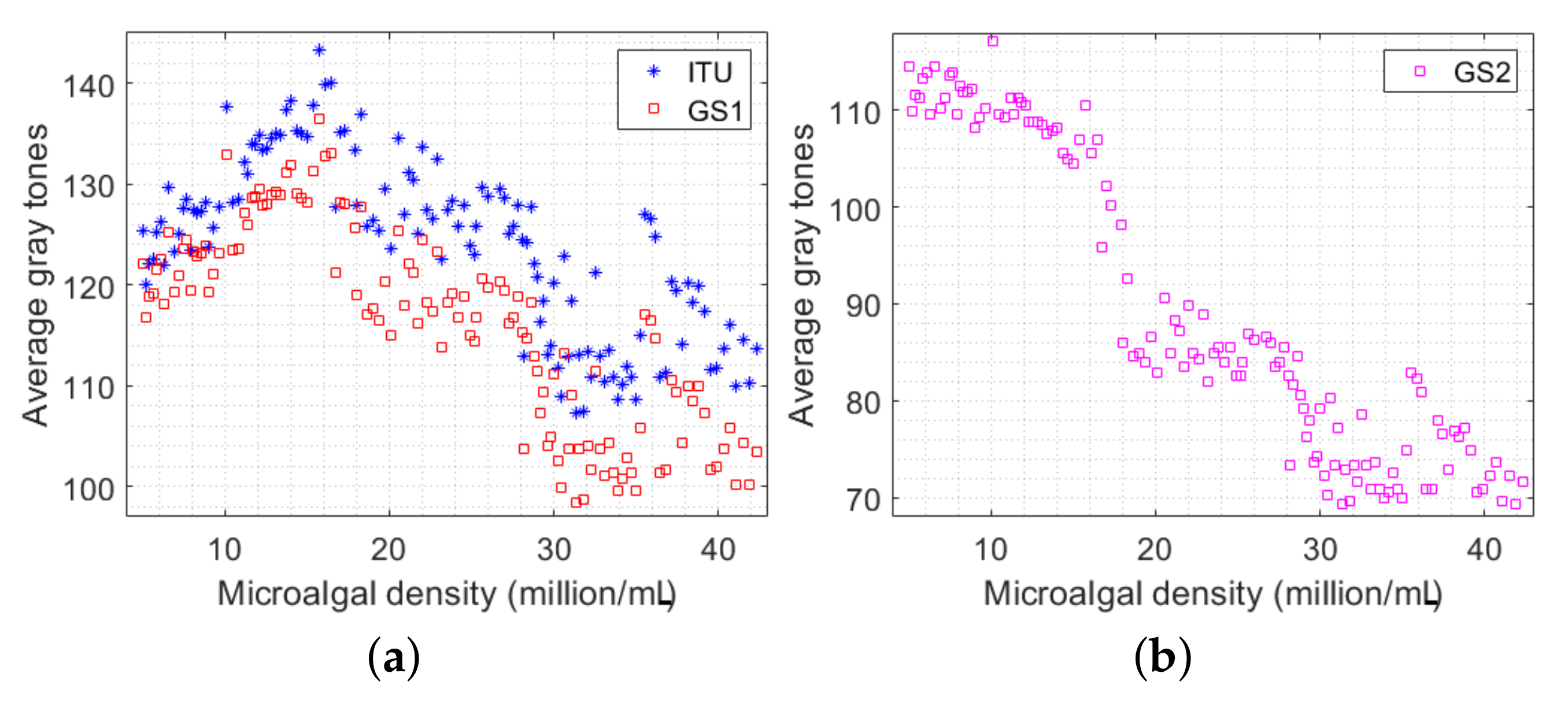
3.1. Averages of Color Channels
3.2. Mean Intervals of Color Channels
3.3. Spatial Frequency
3.4. Entropy
4. L1-Regularization Based Estimation Method
4.1. LASSO
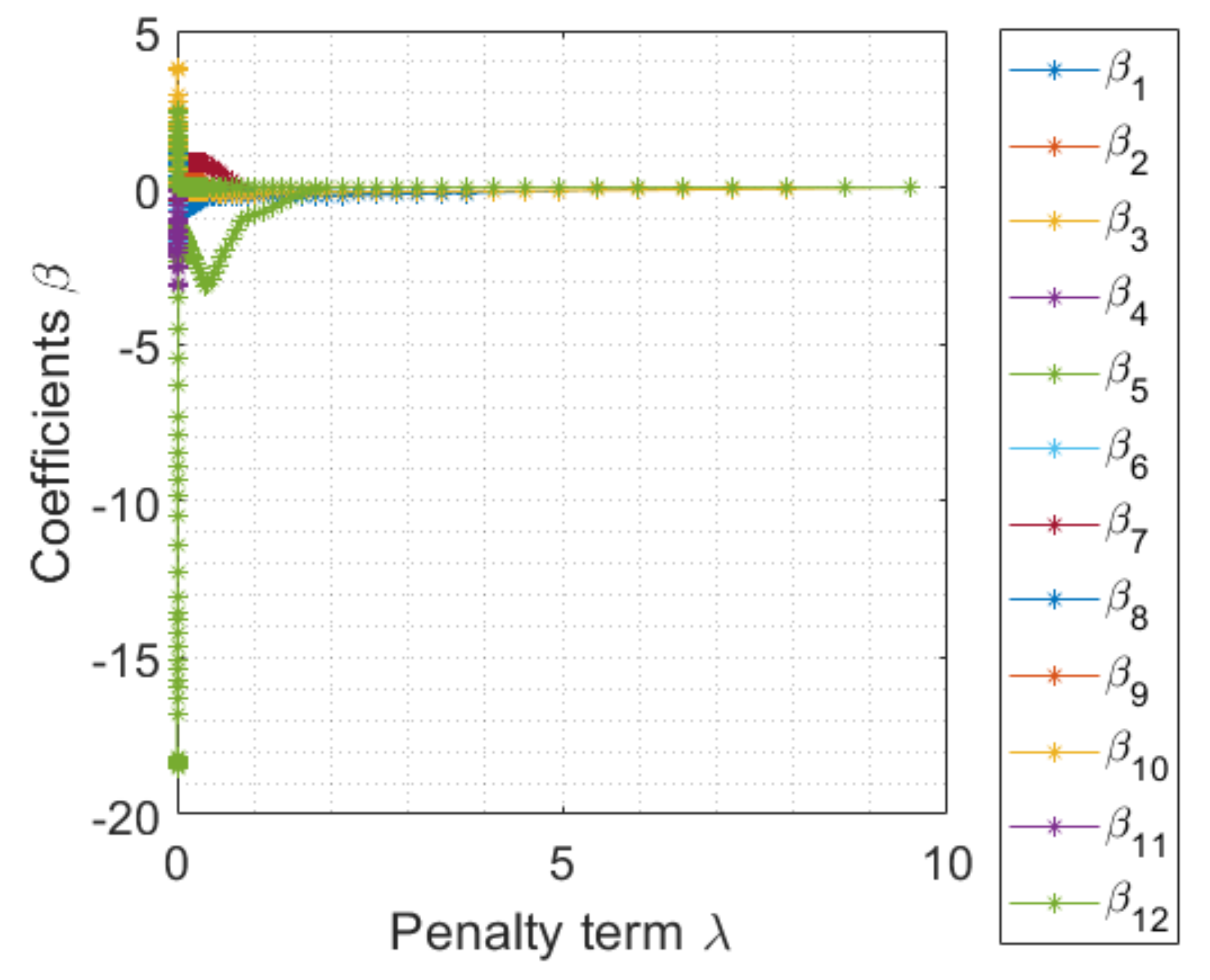
4.2. Penalty Term Learning
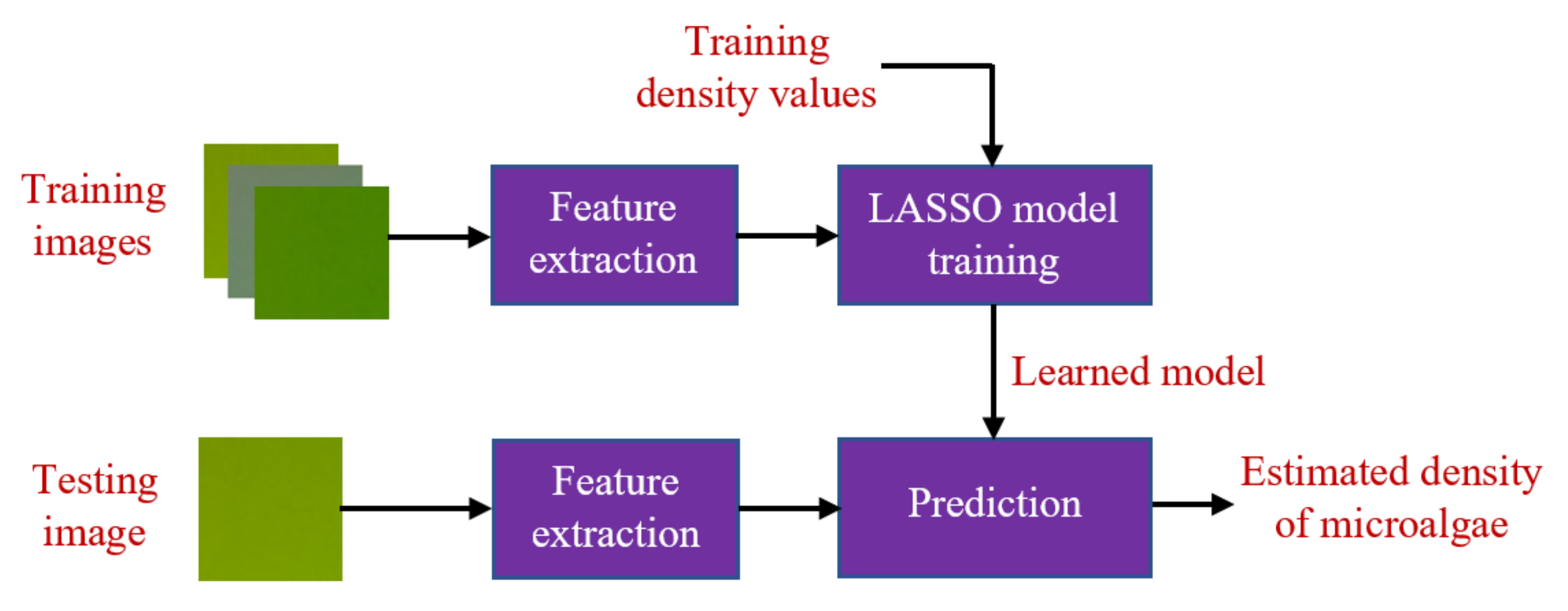
4.3. Microalgal Density Estimation
5. Experimental Result Analysis
5.1. Single Feature Combinations
- S1: Features of average values of all three color channels: R, G and B.
- S2: Six interval features: , , , , , and .
- S3: Three spatial frequency features: , , and .
- S4: Three entropy features: , , and .
5.2. Two-Feature Combinations
- S5: Features of three color channel averages and three entropies: R, G, B, , , and .
- S6: Features of six interval bounds and three entropies: , , , , , , , , and .
- S7: Features of three color channel averages and three spatial frequency powers: R, G, B, , , and .
- S8: Features of three color channel averages and six interval bounds: R, G, B, , , , , , and .
- S9: Features of three spatial frequency powers and three entropies: , , , , , and .
- S10: Features of six interval bounds and three spatial frequency powers: , , , , , , , , and .
5.3. Three and Four-Feature Combinations
- S11: Features of three color channel averages, six interval bounds, and three entropies: R, G, B, , , , , , , , , and .
- S12: Features of three color channel averages, six interval bounds, and three spatial frequency powers: R, G, B, , , , , , , , , and .
- S13: Features of three color channel averages, three entropies, and three spatial frequency powers: R, G, B, , , , , , and .
- S14: Features of six interval bounds, three entropies, and three spatial frequency powers: , , , , , , , , , , , and .
5.4. Higher-Order and Nonlinear Entropy Features
- S16: Features including R, G, B, , , , , , , , , and .
- S17: Features including R, G, B, , , , , , and .
- S18: Features including , , , , , and .
- S19: Features including R, G, B, , , , , , , , , and .
- S20: Features including R, G, B, , , , , , , , , and .
- S21: Features including R, G, B, , , , , , , , , , and .
5.5. Estimation Results
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| LASSO | Least absolute shrinkage and selection operator |
| RGB | Red, green, and blue |
| ITU | International Telecommunication Union |
| DFT | Discrete Fourier transform |
| RMSE | Root mean square error |
| GP | Gaussian process |
| GT | Ground truth |
References
- Christian Barbosa, R.; Soares, J.; Arêdes Martins, M. Low-cost and versatile sensor based on multi-wavelengths for real-time estimation of microalgal biomass concentration in open and closed cultivation systems. Comput. Electron. Agric. 2020, 176, 105641. [Google Scholar] [CrossRef]
- Córdoba-Matson, M.V.; Gutiérrez, J.; Porta-Gándara, M.Á. Evaluation of Isochrysis galbana (clone T-ISO) cell numbers by digital image analysis of color intensity. J. Appl. Phycol. 2010, 22, 427–434. [Google Scholar] [CrossRef]
- Wells, M.; Potin, P.; Craigie, J.; Raven, J.; Merchant, S.; Helliwell, K.; Smith, A.; Camire, M.; Brawley, S. Algae as nutritional and functional food sources: Revisiting our understanding. J. Appl. Phycol. 2017, 29, 949–982. [Google Scholar] [CrossRef] [PubMed]
- Spolaore, P.; Joannis-Cassan, C.; Duran, E.; Isambert, A. Commercial applications of microalgae. J. Biosci. Bioeng. 2006, 101, 87–96. [Google Scholar] [CrossRef]
- Wishkerman, A.; Wishkerman, E. Application note: A novel low-cost open-source LED system for microalgae cultivation. Comput. Electron. Agric. 2017, 132, 56–62. [Google Scholar] [CrossRef]
- Jung, S.K.; Lee, S.B. In Situ Monitoring of Cell Concentration in a Photobioreactor Using Image Analysis: Comparison of Uniform Light Distribution Model and Artificial Neural Networks. Biotechnol. Prog. 2006, 22, 1443–1450. [Google Scholar] [CrossRef]
- Zhou, W.; Wang, Z.; Xu, J.; Ma, L. Cultivation of microalgae Chlorella zofingiensis on municipal wastewater and biogas slurry towards bioenergy. J. Biosci. Bioeng. 2018, 126, 644–648. [Google Scholar] [CrossRef]
- Bitog, J.; Lee, I.B.; Lee, C.G.; Kim, K.S.; Hwang, H.S.; Hong, S.W.; Seo, I.H.; Kwon, K.S.; Mostafa, E. Application of computational fluid dynamics for modeling and designing photobioreactors for microalgae production: A review. Comput. Electron. Agric. 2011, 76, 131–147. [Google Scholar] [CrossRef]
- Laing, I. Cultivation of Marine Unicellular Algae; Ministry of Agriculture, Fisheries and Food Conwy: Conwy, UK, 1991.
- Sananurak, C.; Lirdwitayaprasit, T.; Menasveta, P. Development of a closed-recirculating, continuous culture system for microalga (Tetraselmis sueeica) and rotifer (Brachionus plicatilis) production. ScienceAsia 2009, 35, 118–124. [Google Scholar] [CrossRef]
- Naumann, T.; Çebi, Z.; Podola, B.; Melkonian, M. Growing microalgae as aquaculture feeds on twin-layers: A novel solid-state photobioreactor. J. Appl. Phycol. 2012, 25, 1413–1420. [Google Scholar] [CrossRef]
- Metsoviti, M.; Papapolymerou, G.; Karapanagiotidis, I.; Katsoulas, N. Comparison of Growth Rate and Nutrient Content of Five Microalgae Species Cultivated in Greenhouses. Plants 2019, 8, 279. [Google Scholar] [CrossRef]
- Tham, P.E.; Ng, Y.J.; Vadivelu, N.; Lim, H.R.; Khoo, K.S.; Chew, K.W.; Show, P.L. Sustainable smart photobioreactor for continuous cultivation of microalgae embedded with Internet of Things. Bioresour. Technol. 2022, 346, 126558. [Google Scholar] [CrossRef]
- Lim, H.R.; Khoo, K.S.; Chia, W.Y.; Chew, K.W.; Ho, S.H.; Show, P.L. Smart microalgae farming with internet-of-things for sustainable agriculture. Biotechnol. Adv. 2022, 57, 107931. [Google Scholar] [CrossRef]
- Nguyen, L.; Nguyen, D.K.; Nghiem, T.X.; Nguyen, T. Least square and Gaussian process for image based microalgal density estimation. Comput. Electron. Agric. 2022, 193, 106678. [Google Scholar] [CrossRef]
- Nguyen, D.K.; Nguyen, L.; Le, D.V. A Low-Cost Efficient System for Monitoring Microalgae Density using Gaussian Process. IEEE Trans. Instrum. Meas. 2021, 70, 7504308. [Google Scholar] [CrossRef]
- Havlik, I.; Scheper, T.; Reardon, K.F. Monitoring of Microalgal Processes. In Microalgae Biotechnology; Posten, C., Feng Chen, S., Eds.; Springer International Publishing: Cham, Switzerland, 2016; pp. 89–142. [Google Scholar]
- Cogne, G.; Lasseur, C.; Cornet, J.; Dussap, C.G.; Gros, J. Growth monitoring of a photosynthetic micro-organism (Spirulina platensis) by pressure measurement. Biotechnol. Lett. 2001, 23, 1309–1314. [Google Scholar] [CrossRef]
- Bulgarea, G.; Boukadoum, M. A high-performance instrumentation system to measure the fluorescence kinetics of plants for in vivo photosynthesis research. IEEE Trans. Instrum. Meas. 2001, 50, 679–689. [Google Scholar] [CrossRef]
- Meireles, L.A.; Azevedo, J.L.; Cunha, J.P.; Malcata, F.X. On-line determination of biomass in a microalga bioreactor using a novel computerized flow injection analysis system. Biotechnol. Prog. 2002, 18, 1387–1391. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Goh, Q.Y.; Tan, W.; Hossain, I.; Chen, W.N.; Lau, R. Lumostatic strategy for microalgae cultivation utilizing image analysis and chlorophyll a content as design parameters. Bioresour. Technol. 2011, 102, 6005–6012. [Google Scholar] [CrossRef] [PubMed]
- Uyar, B. A novel non-invasive digital imaging method for continuous biomass monitoring and cell distribution mapping in photobioreactors. J. Chem. Technol. Biotechnol. 2012, 88, 1144–1149. [Google Scholar] [CrossRef]
- Sarrafzadeh, M.H.; La, H.J.; Lee, J.Y.; Cho, D.H.; Shin, S.Y.; Kim, W.J.; Oh, H.M. Microalgae biomass quantification by digital image processing and RGB color analysis. J. Appl. Phycol. 2015, 27, 205–209. [Google Scholar] [CrossRef]
- Winata, H.N.; Nasution, M.A.; Ahamed, T.; Noguchi, R. Prediction of concentration for microalgae using image analysis. Multimed. Tools Appl. 2021, 80, 8541–8561. [Google Scholar] [CrossRef]
- Drabycz, S.; Stockwell, R.G.; Mitchell, J.R. Image Texture Characterization Using the Discrete Orthonormal S-Transform. J. Digit. Imaging 2008, 22, 696–708. [Google Scholar] [CrossRef] [PubMed]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction; Springer: New York, NY, USA, 2009. [Google Scholar]
- Mehr, A.D.; Gandomi, A.H. MSGP-LASSO: An improved multi-stage genetic programming model for streamflow prediction. Inf. Sci. 2021, 561, 181–195. [Google Scholar] [CrossRef]
- Oberdorf, F.; Stein, N.; Flath, C.M. Analytics-enabled escalation management: System development and business value assessment. Comput. Ind. 2021, 131, 103481. [Google Scholar] [CrossRef]
- Nguyen, D.K.; Nguyen, H.Q.; Dang, H.T.T.; Nguyen, V.Q.; Nguyen, L. A low-cost system for monitoring pH, dissolved oxygen and algal density in continuous culture of microalgae. HardwareX 2022, 12, e00353. [Google Scholar] [CrossRef]
- Masojídek, J.; Kopecký, J.; Giannelli, L.; Torzillo, G. Productivity correlated to photobiochemical performance of Chlorella mass cultures grown outdoors in thin-layer cascades. J. Ind. Microbiol. Biotechnol. 2010, 38, 307–317. [Google Scholar] [CrossRef]
- Dekking, F.M.; Kraaikamp, C.; Lopuhaa, H.P.; Meester, L.E. A Modern Introduction to Probability and Statistics: Understanding Why and How; Springer: London, UK, 2005. [Google Scholar]
- Park, C.S. 2D Discrete Fourier Transform on Sliding Windows. IEEE Trans. Image Process. 2015, 24, 901–907. [Google Scholar] [CrossRef]
- Susan, S.; Hanmandlu, M. A non-extensive entropy feature and its application to texture classification. Neurocomputing 2013, 120, 214–225. [Google Scholar] [CrossRef]
- Haralick, R.M.; Shanmugam, K.; Dinstein, I. Textural Features for Image Classification. IEEE Trans. Syst. Man Cybern. 1973; SMC-3, 610–621. [Google Scholar] [CrossRef]
- Silva, L.E.; Duque, J.J.; Felipe, J.C.; Murta, L.O., Jr.; Humeau-Heurtier, A. Two-dimensional multiscale entropy analysis: Applications to image texture evaluation. Signal Process. 2018, 147, 224–232. [Google Scholar] [CrossRef]
- Onchis, H.D.M. A deep learning approach to condition monitoring of cantilever beams via time-frequency extended signatures. Comput. Ind. 2019, 105, 177–181. [Google Scholar] [CrossRef]
- Pan, X.; Xu, Y. A safe reinforced feature screening strategy for lasso based on feasible solutions. Inf. Sci. 2019, 477, 132–147. [Google Scholar] [CrossRef]




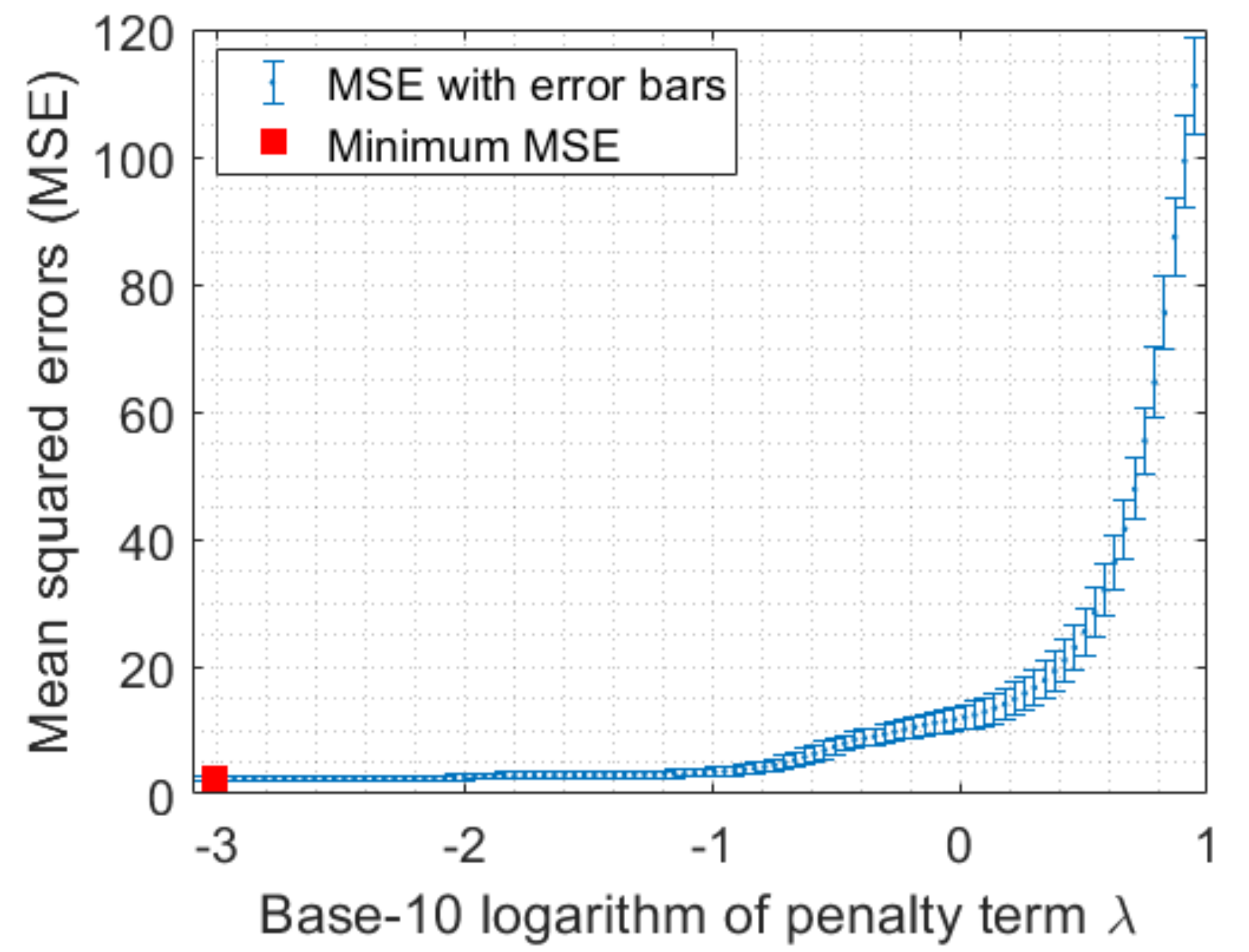

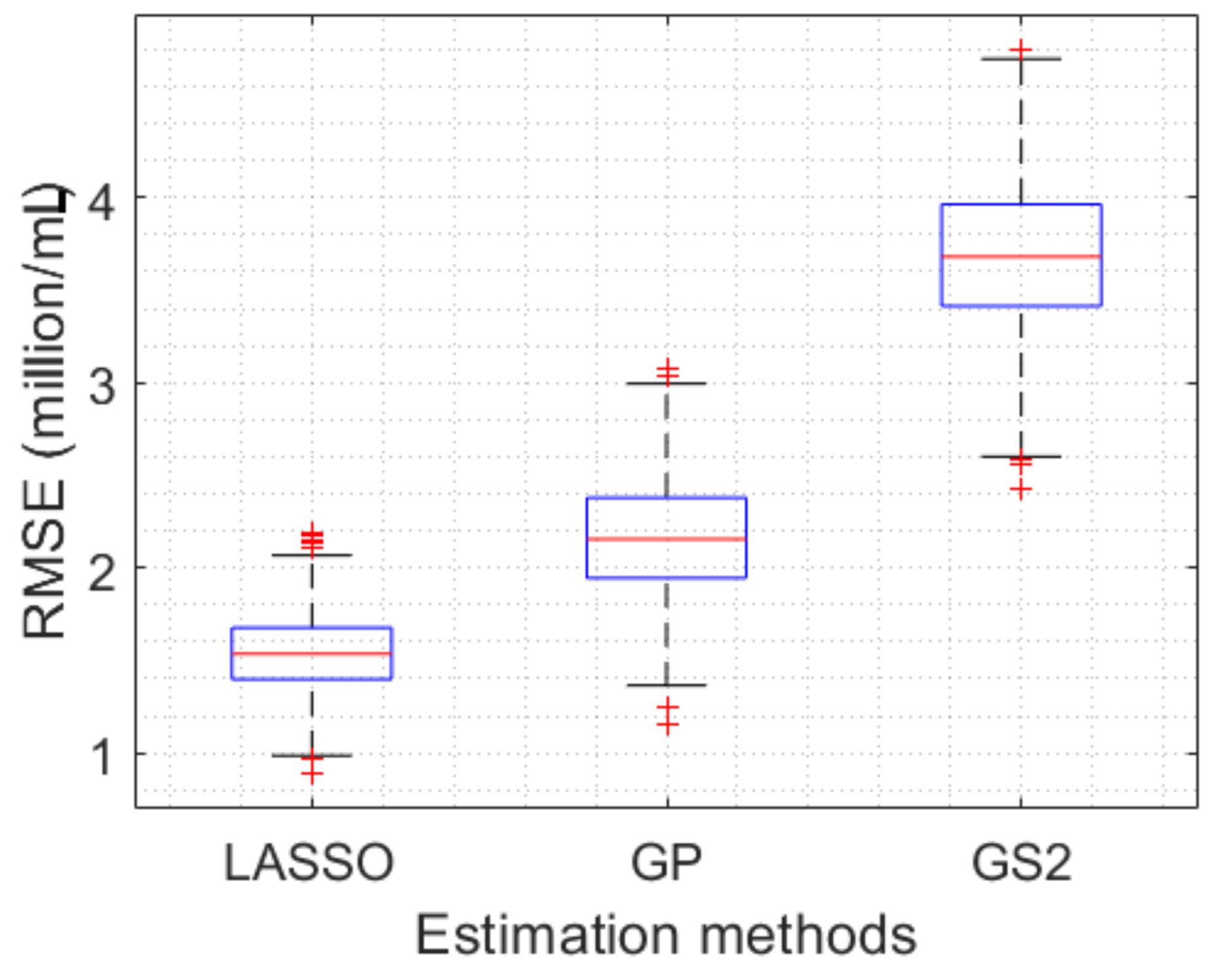
| Minimum MSE | 2.313 | 2.215 | 2.278 |
| 0.001 | 0.001 | 0.001 |
| S1 | S2 | S3 | S4 | |
|---|---|---|---|---|
| mean | 2.40 | 2.12 | 2.36 | 4.90 |
| std | 0.25 | 0.26 | 0.28 | 0.67 |
| S5 | S6 | S7 | S8 | S9 | S10 | |
|---|---|---|---|---|---|---|
| mean | 1.75 | 1.78 | 1.89 | 2.09 | 1.79 | 2.09 |
| std | 0.23 | 0.23 | 0.25 | 0.26 | 0.24 | 0.24 |
| S11 | S12 | S13 | S14 | S15 | |
|---|---|---|---|---|---|
| mean | 1.76 | 2.11 | 1.76 | 1.77 | 1.79 |
| std | 0.24 | 0.23 | 0.24 | 0.23 | 0.24 |
| S16 | S17 | S18 | S19 | S20 | S21 | |
|---|---|---|---|---|---|---|
| mean | 1.63 | 1.64 | 3.76 | 1.67 | 1.54 | 1.56 |
| std | 0.21 | 0.21 | 0.44 | 0.23 | 0.20 | 0.24 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nguyen, L.; Nguyen, D.K.; Nguyen, T.; Nguyen, B.; Nghiem, T.X. Analysis of Microalgal Density Estimation by Using LASSO and Image Texture Features. Sensors 2023, 23, 2543. https://doi.org/10.3390/s23052543
Nguyen L, Nguyen DK, Nguyen T, Nguyen B, Nghiem TX. Analysis of Microalgal Density Estimation by Using LASSO and Image Texture Features. Sensors. 2023; 23(5):2543. https://doi.org/10.3390/s23052543
Chicago/Turabian StyleNguyen, Linh, Dung K. Nguyen, Thang Nguyen, Binh Nguyen, and Truong X. Nghiem. 2023. "Analysis of Microalgal Density Estimation by Using LASSO and Image Texture Features" Sensors 23, no. 5: 2543. https://doi.org/10.3390/s23052543
APA StyleNguyen, L., Nguyen, D. K., Nguyen, T., Nguyen, B., & Nghiem, T. X. (2023). Analysis of Microalgal Density Estimation by Using LASSO and Image Texture Features. Sensors, 23(5), 2543. https://doi.org/10.3390/s23052543










