Machine Learning Sensors for Diagnosis of COVID-19 Disease Using Routine Blood Values for Internet of Things Application
Abstract
1. Introduction
2. Related Studies
3. Data and Methods
3.1. Characteristic of Participants, Workflow and Datasets
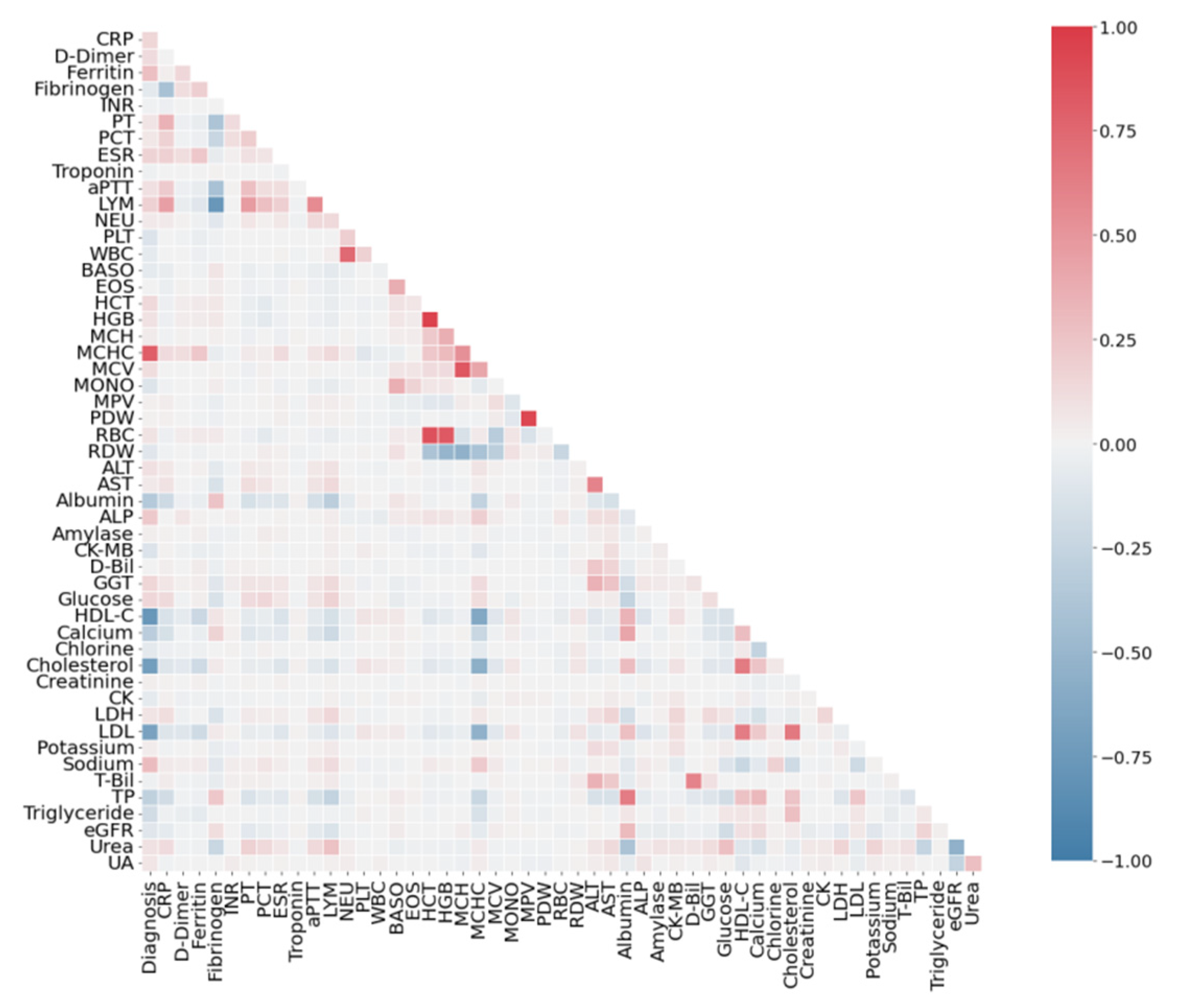
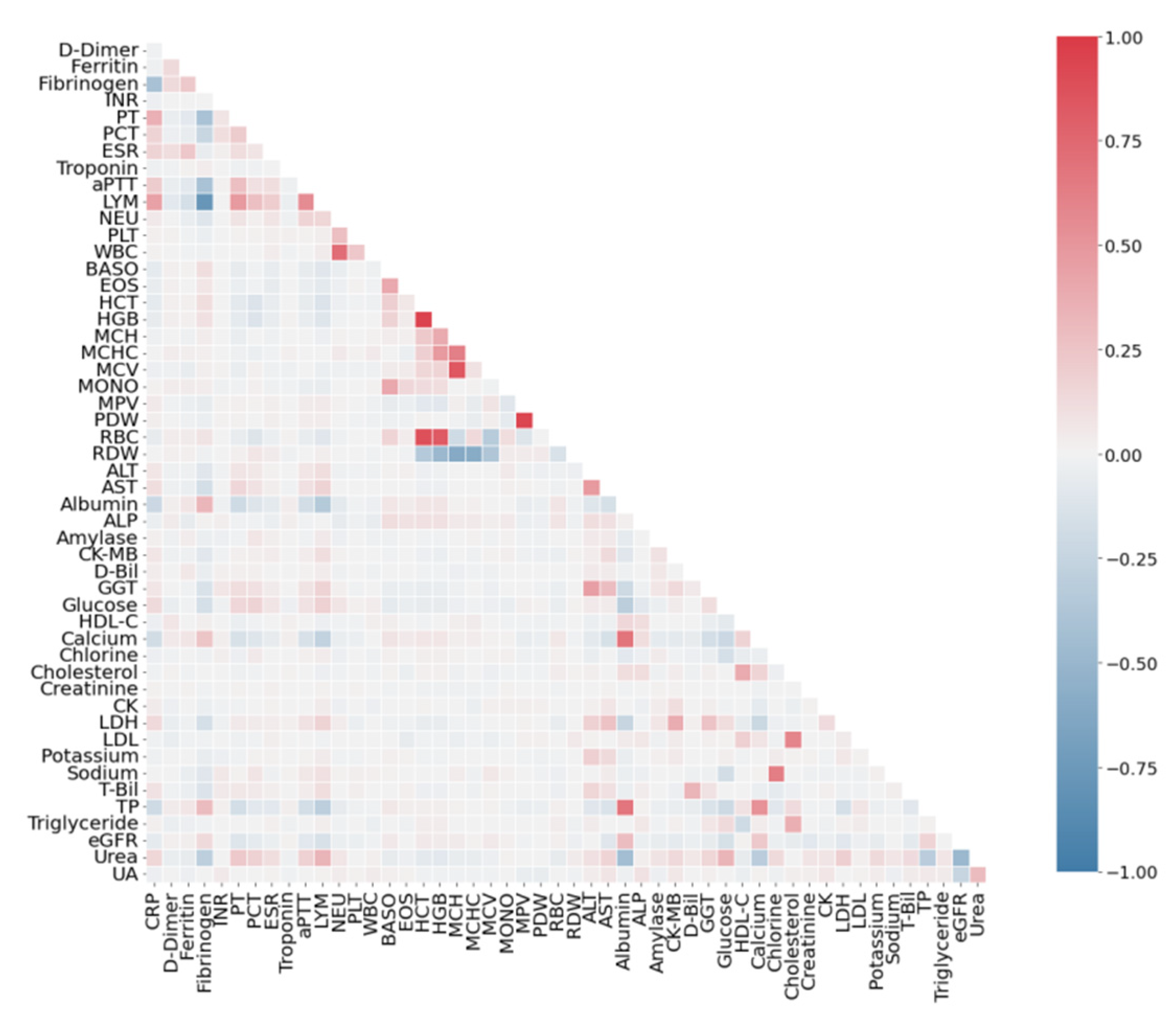
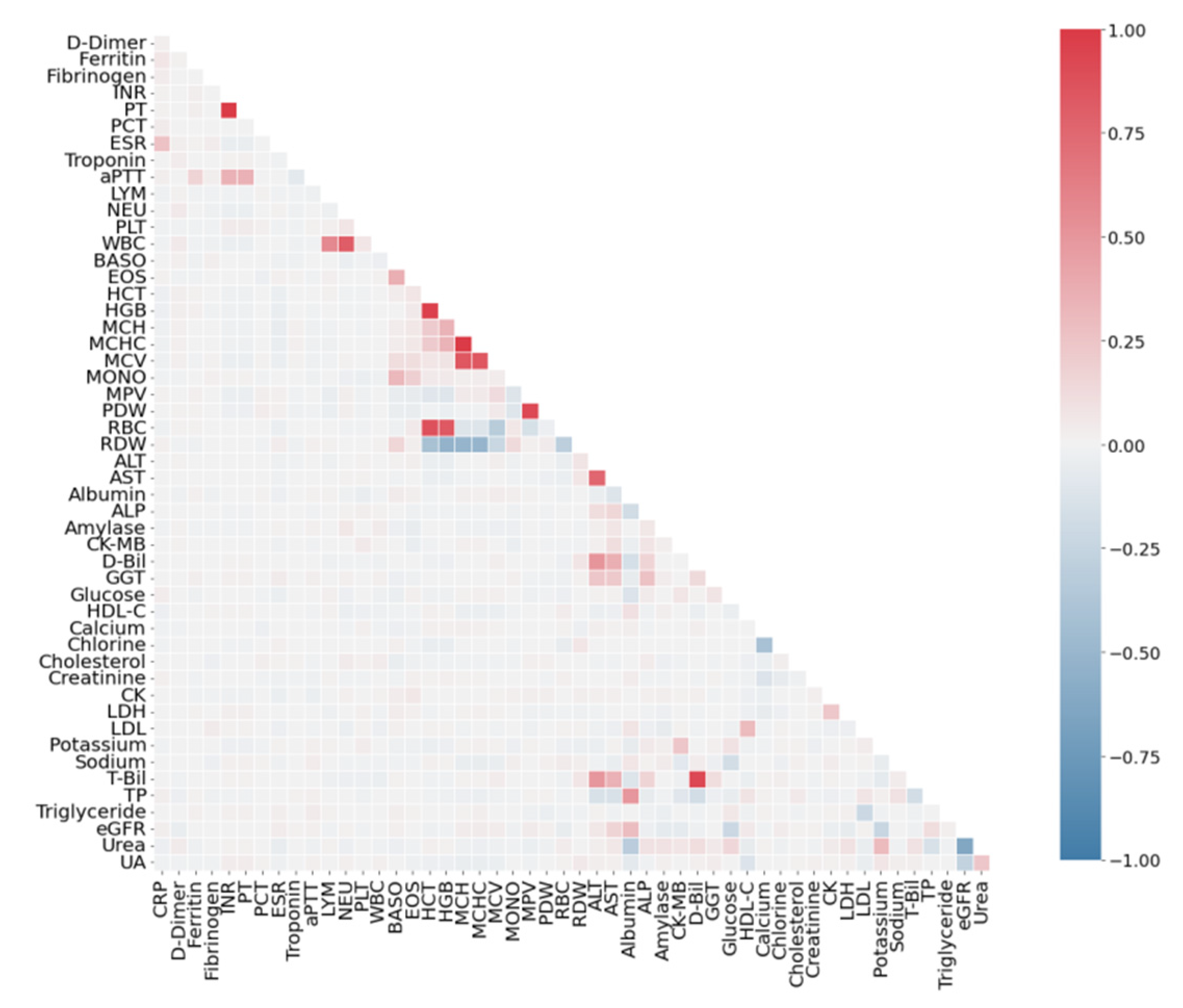
3.2. Correlation Analysis of Features
3.3. Machine Learning Methods, Hyperparameters, Accuracy Estimation
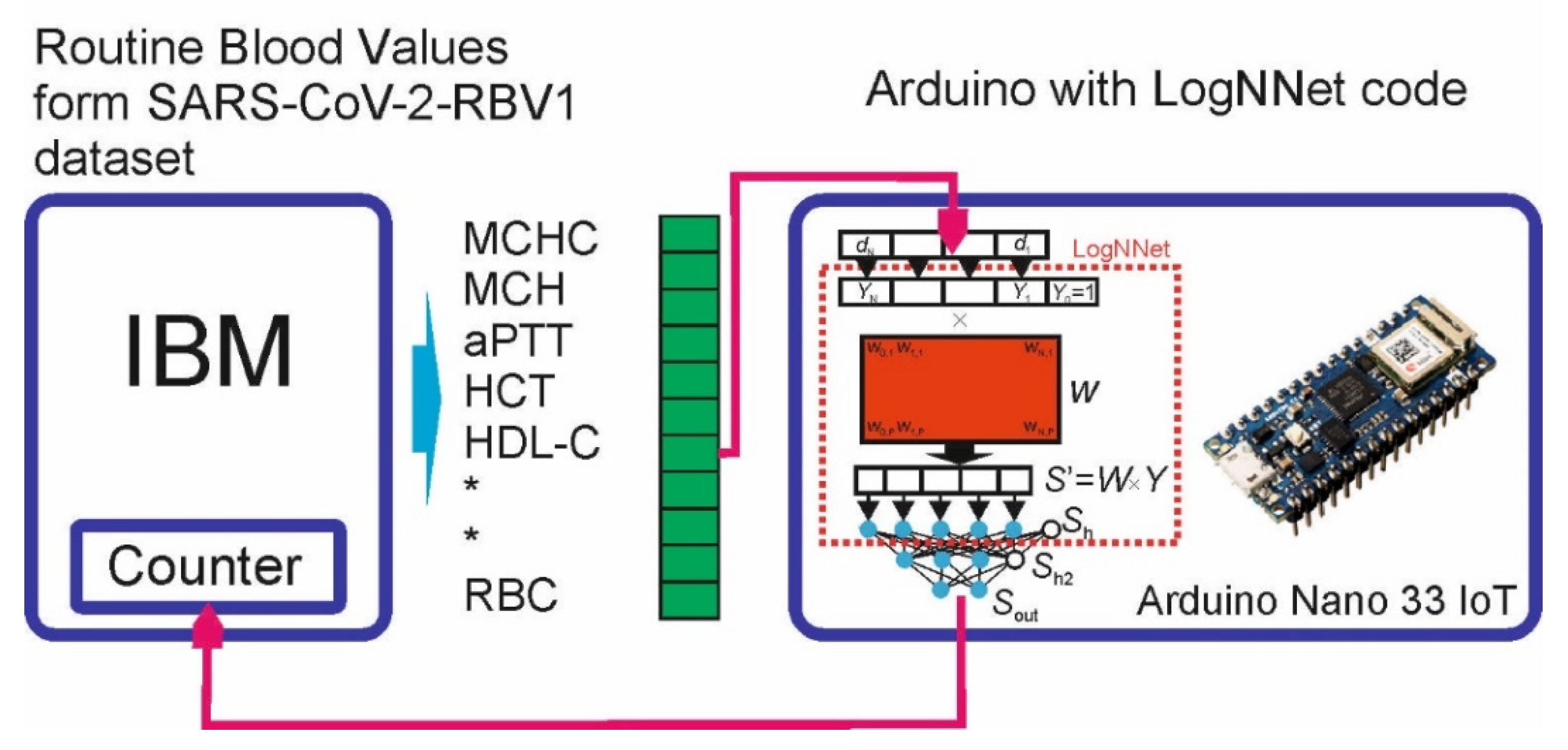
3.4. Implementing LogNNet on an Arduino Board
3.4.1. LogNNet Program for Arduino Board
- Function “Fun_activ”—activation function, lines 10–12;
- Procedure “Reservoir”—calculation of coefficients of reservoir matrix W by congruent generator formula, multiplication of arrays and calculation of neurons in layer Sh, lines 14–28;
- Procedure “Hidden_Layer”—calculation of neurons in the hidden layer Sh2, lines 30–39;
- Function “Output_Layer_Layer”—calculation of the output layer Sout, lines 41–54;
- The “void loop” block is an executable loop, lines 61–77;
- “void setup” block—initialization block, lines 61–77.
3.4.2. Test Scheme
4. Results
4.1. Correlation Analysis of Dataset SARS-CoV-2-RBV1
4.2. Classification Results for Dataset SARS-CoV-2-RBV1
4.2.1. Investigation of the Effectiveness of the HGB Model Operating on One Feature
4.2.2. Investigation of the Effectiveness of the HGB Model Operating on Two Features
4.2.3. The Study of the Most Significant Combination of Three Features of the HGB Model
4.2.4. The Study of the Most Significant Combination of 11 Features of the HGB Model
4.3. LogNNet Implementation on Arduino for Edge Computing
4.4. Machine Learning COVID-19 Sensor for IoT
5. Discussion
5.1. Analysis of Results from a Medical Perspective
5.2. Analysis of Results from IoT Perspective
6. Limitations of the Study
7. Conclusions and Future Studies
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
Appendix A
| Algorithm A1. LogNNet neural network executable code on Arduino Nano IoT | |||
| 1 | #include “LogNNet.h” | 40 | |
| 2 | 41 | byte Output_Layer() { | |
| 3 | float Y[S+1]; | 42 | float Sout[N+1]; byte digit = 0; |
| 4 | float Sh[P+1]; | 43 | for (int j = 0; j <= N; j++) { |
| 5 | float Sh2[M+1]; | 44 | Sout[j] = 0; |
| 6 | 45 | for (int i = 0; i <= M; i++) | |
| 7 | int i = 0; | 46 | Sout[j] = Sout[j] + Sh2[i] * |
| 8 | String data; | 47 | ((float)W2[i][j]/scale_factor); |
| 9 | 48 | Sout[j] = Fun_activ(Sout[j]); | |
| 10 | float Fun_activ(float x) { | 49 | } |
| 11 | return 1 / (1 + exp(-1*x)); | 50 | for (int j = 0; j <= N; j++) { |
| 12 | } | 51 | if (Sout[j] > Sout[digit]) |
| 13 | 52 | digit = j; | |
| 14 | void Reservoir(float *Y) { | 53 | } |
| 15 | long W = C; | 54 | return digit; |
| 16 | Sh[0] = 1; | 55 | } |
| 17 | for (int j = 1; j <= P; j++) { | 56 | |
| 18 | Sh[j] = 0; | 57 | void setup() { |
| 19 | for (int i = 0; i <= S; i++) { | 58 | Serial.begin(115200); |
| 20 | W = (D - K * W) % L; | 59 | } |
| 21 | Sh[j] = Sh[j] + ((float)W/L) * Y[i]; | 60 | |
| 22 | } | 61 | void loop() { |
| 23 | Sh[j] = ((Sh[j] - (float)minS[j-1]/ | 62 | if (Serial.available() > 0) { |
| 24 | scale_factor) / ((float)(maxS[j-1] | 63 | data = Serial.readStringUntil(‘T’); |
| 25 | - minS[j-1])/scale_factor)) - 0.5 | 64 | |
| 26 | - (float)meanS[j-1]/(scale_factor*10); | 65 | if (data != “FN”) { |
| 27 | } | 66 | Y[i] = data.toFloat(); |
| 28 | } | 67 | i++; |
| 29 | 68 | } | |
| 30 | void Hidden_Layer() { | 69 | else { |
| 31 | Sh2[0] = 1; | 70 | i = 0; |
| 32 | for (int j = 1; j <= M; j++) { | 71 | Reservoir(Y); |
| 33 | Sh2[j] = 0; | 72 | Hidden_Layer(); |
| 34 | for (int i = 0; i <= P; i++) | 73 | byte Digit = Output_Layer(); |
| 35 | Sh2[j] = Sh2[j] + Sh[i] * | 74 | Serial.print(String(Digit)); |
| 36 | ((float)W1[i][j]/scale_factor); | 75 | } |
| 37 | Sh2[j] = Fun_activ(Sh2[j]); | 76 | } |
| 38 | } | 77 | } |
| 39 | } | 78 | |
References
- Mertoglu, C.; Huyut, M.; Olmez, H.; Tosun, M.; Kantarci, M.; Coban, T. COVID-19 Is More Dangerous for Older People and Its Severity Is Increasing: A Case-Control Study. Med. Gas Res. 2022, 12, 51–54. [Google Scholar] [CrossRef]
- Mertoglu, C.; Huyut, M.T.; Arslan, Y.; Ceylan, Y.; Coban, T.A. How Do Routine Laboratory Tests Change in Coronavirus Disease 2019? Scand. J. Clin. Lab. Investig. 2021, 81, 24–33. [Google Scholar] [CrossRef]
- Zhou, C.; Chen, Y.; Ji, Y.; He, X.; Xue, D. Increased Serum Levels of Hepcidin and Ferritin Are Associated with Severity of COVID-19. Med. Sci. Monit. 2020, 26, e926178. [Google Scholar] [CrossRef]
- Huyut, M.T.; Huyut, Z. Forecasting of Oxidant/Antioxidant Levels of COVID-19 Patients by Using Expert Models with Biomarkers Used in the Diagnosis/Prognosis of COVID-19. Int. Immunopharmacol. 2021, 100, 108127. [Google Scholar] [CrossRef]
- Huyut, M.T.; İlkbahar, F. The Effectiveness of Blood Routine Parameters and Some Biomarkers as a Potential Diagnostic Tool in the Diagnosis and Prognosis of Covid-19 Disease. Int. Immunopharmacol. 2021, 98, 107838. [Google Scholar] [CrossRef]
- Huyut, M.; Üstündaǧ, H. Prediction of Diagnosis and Prognosis of COVID-19 Disease by Blood Gas Parameters Using Decision Trees Machine Learning Model: A Retrospective Observational Study. Med. Gas Res. 2022, 12, 60–66. [Google Scholar] [CrossRef] [PubMed]
- Cabitza, F.; Campagner, A.; Ferrari, D.; Di Resta, C.; Ceriotti, D.; Sabetta, E.; Colombini, A.; De Vecchi, E.; Banfi, G.; Locatelli, M.; et al. Development, Evaluation, and Validation of Machine Learning Models for COVID-19 Detection Based on Routine Blood Tests. Clin. Chem. Lab. Med. 2021, 59, 421–431. [Google Scholar] [CrossRef] [PubMed]
- Vogels, C.B.F.; Brito, A.F.; Wyllie, A.L.; Fauver, J.R.; Ott, I.M.; Kalinich, C.C.; Petrone, M.E.; Casanovas-Massana, A.; Catherine Muenker, M.; Moore, A.J.; et al. Analytical Sensitivity and Efficiency Comparisons of SARS-CoV-2 RT–QPCR Primer–Probe Sets. Nat. Microbiol. 2020, 5, 1299–1305. [Google Scholar] [CrossRef]
- Booth, A.L.; Abels, E.; McCaffrey, P. Development of a Prognostic Model for Mortality in COVID-19 Infection Using Machine Learning. Mod. Pathol. 2021, 34, 522–531. [Google Scholar] [CrossRef]
- Ko, J.; Baldassano, S.N.; Loh, P.-L.; Kording, K.; Litt, B.; Issadore, D. Machine Learning To Detect Signatures of Disease in Liquid Biopsies-A User’s Guide Graphical Abstract HHS Public Access. Lab Chip 2018, 18, 395–405. [Google Scholar] [CrossRef]
- Huyut, M.T. Automatic Detection of Severely and Mildly Infected COVID-19 Patients with Supervised Machine Learning Models. IRBM 2022, 1, 35673548. [Google Scholar] [CrossRef]
- Huyut, M.T.; Velichko, A. Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network. Sensors 2022, 22, 4820. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and Clinical Characteristics of 99 Cases of 2019 Novel Coronavirus Pneumonia in Wuhan, China: A Descriptive Study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Tahir Huyut, M.; Huyut, Z.; İlkbahar, F.; Mertoğlu, C. What Is the Impact and Efficacy of Routine Immunological, Biochemical and Hematological Biomarkers as Predictors of COVID-19 Mortality? Int. Immunopharmacol. 2022, 105, 108542. [Google Scholar] [CrossRef] [PubMed]
- Huyut, M.T.; Kocaturk, İ. The Effect of Some Symptoms and Features During the İnfection Period on the Level of Anxiety and Depression of Adults After Recovery From COVID-19. Curr. Psychiatry Res. Rev. 2022, 18, 151–163. [Google Scholar] [CrossRef]
- Guan, W.; Ni, Z.; Hu, Y.; Liang, W.; Ou, C.; He, J.; Liu, L.; Shan, H.; Lei, C.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef]
- Banerjee, A.; Ray, S.; Vorselaars, B.; Kitson, J.; Mamalakis, M.; Weeks, S.; Baker, M.; Mackenzie, L.S. Use of Machine Learning and Artificial Intelligence to Predict SARS-CoV-2 Infection from Full Blood Counts in a Population. Int. Immunopharmacol. 2020, 86, 106705. [Google Scholar] [CrossRef] [PubMed]
- Huyut, M.T.; Soygüder, S. The Multi-Relationship Structure between Some Symptoms and Features Seen during the New Coronavirus 19 Infection and the Levels of Anxiety and Depression Post-Covid. East. J. Med. 2022, 27, 1–10. [Google Scholar] [CrossRef]
- Perricone, C.; Bartoloni, E.; Bursi, R.; Cafaro, G.; Guidelli, G.M.; Shoenfeld, Y.; Gerli, R. COVID-19 as Part of the Hyperferritinemic Syndromes: The Role of Iron Depletion Therapy. Immunol. Res. 2020, 68, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J. COVID-19: Consider Cytokine Storm Syndromes and Immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Karakike, E.; Giamarellos-Bourboulis, E.J. Macrophage Activation-like Syndrome: A Distinct Entity Leading to Early Death in Sepsis. Front. Immunol. 2019, 10, 55. [Google Scholar] [CrossRef]
- Tural Onur, S.; Altın, S.; Sokucu, S.N.; Fikri, B.İ.; Barça, T.; Bolat, E.; Toptaş, M. Could Ferritin Level Be an Indicator of COVID-19 Disease Mortality? J. Med. Virol. 2021, 93, 1672–1677. [Google Scholar] [CrossRef] [PubMed]
- Rosário, C.; Zandman-Goddard, G.; Meyron-Holtz, E.G.; D’Cruz, D.P.; Shoenfeld, Y. The Hyperferritinemic Syndrome: Macrophage Activation Syndrome, Still’s Disease, Septic Shock and Catastrophic Antiphospholipid Syndrome. BMC Med. 2013, 11, 185. [Google Scholar] [CrossRef]
- Lippi, G.; Plebani, M.; Henry, B.M. Thrombocytopenia Is Associated with Severe Coronavirus Disease 2019 (COVID-19) Infections: A Meta-Analysis. Clin. Chim. Acta 2020, 506, 145–148. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Li, H.; Li, L.; Liu, C.; Yan, S.; Chen, H.; Li, Y. Ferritin in the Coronavirus Disease 2019 (COVID-19): A Systematic Review and Meta-Analysis. J. Clin. Lab. Anal. 2020, 34, e23618. [Google Scholar] [CrossRef] [PubMed]
- Feld, J.; Tremblay, D.; Thibaud, S.; Kessler, A.; Naymagon, L. Ferritin Levels in Patients with COVID-19: A Poor Predictor of Mortality and Hemophagocytic Lymphohistiocytosis. Int. J. Lab. Hematol. 2020, 42, 773–779. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical Course and Risk Factors for Mortality of Adult Inpatients with COVID-19 in Wuhan, China: A Retrospective Cohort Study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Winata, S.; Kurniawan, A. Coagulopathy in COVID-19: A Systematic Review. Medicinus 2021, 8, 72. [Google Scholar] [CrossRef]
- Hariyanto, T.I.; Japar, K.V.; Kwenandar, F.; Damay, V.; Siregar, J.I.; Lugito, N.P.H.; Tjiang, M.M.; Kurniawan, A. Inflammatory and hematologic markers as predictors of severe outcomes in COVID-19 infection: A systematic review and meta-analysis. Am. J. Emerg. Med. 2021, 41, 110–119. [Google Scholar] [CrossRef]
- Remeseiro, B.; Bolon-Canedo, V. A Review of Feature Selection Methods in Medical Applications. Comput. Biol. Med. 2019, 112, 103375. [Google Scholar] [CrossRef]
- Guyon, I.; Gunn, S.; Nikravesh, M.; Zadeh, L.A. Feature Extraction: Foundations and Applications; Studies in Fuzziness and Soft Computing; Springer: Berlin/Heidelberg, Germany, 2008; ISBN 9783540354888. [Google Scholar]
- Jiang, S.Q.; Huang, Q.F.; Xie, W.M.; Lv, C.; Quan, X.Q. The Association between Severe COVID-19 and Low Platelet Count: Evidence from 31 Observational Studies Involving 7613 Participants. Br. J. Haematol. 2020, 190, e29–e33. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Zhang, Y.; Chi, H.; Chen, S.; Peng, M.; Luo, L.; Chen, L.; Li, J.; Shen, B.; Wang, D. The Hemocyte Counts as a Potential Biomarker for Predicting Disease Progression in COVID-19: A Retrospective Study. Clin. Chem. Lab. Med. 2020, 58, 1106–1115. [Google Scholar] [CrossRef] [PubMed]
- Hall, M.A. Correlation-Based Feature Selection for Machine Learning. Ph.D. Thesis, The University of Waikato, Hamilton, New Zealand, 1999. [Google Scholar]
- Dash, M.; Liu, H. Consistency-Based Search in Feature Selection. Artif. Intell. 2003, 151, 155–176. [Google Scholar] [CrossRef]
- Zhao, Z.; Liu, H. Searching for Interacting Features. In Proceedings of the 20th International Joint Conference on Artificial Intelligence, Hyderabad, India, 6–12 January 2007; pp. 1156–1161. [Google Scholar]
- Hall, M.A.; Smith, L.A. Practical Feature Subset Selection for Machine Learning. In Proceedings of the the 21st Australasian Computer Science Conference ACSC’98, Perth, Australia, 4–6 February 1998; Volume 20, pp. 181–191. [Google Scholar]
- Kononenko, I. Estimating Attributes: Analysis and Extensions of RELIEF. Machine Learning: ECML-94; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 1994; Volume 784, pp. 171–182. [Google Scholar] [CrossRef]
- Le Thi, H.A.; Nguyen, V.V.; Ouchani, S. Gene Selection for Cancer Classification Using DCA. Advanced Data Mining and Applications; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2008; Volume 5139, pp. 62–72. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression Shrinkage and Selection via the Lasso. J. R. Stat. Society. Ser. B Methodol. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Zhu, J.S.; Ge, P.; Jiang, C.; Zhang, Y.; Li, X.; Zhao, Z.; Zhang, L.; Duong, T.Q. Deep-Learning Artificial Intelligence Analysis of Clinical Variables Predicts Mortality in COVID-19 Patients. J. Am. Coll. Emerg. Physicians Open 2020, 1, 1364–1373. [Google Scholar] [CrossRef] [PubMed]
- Meigal, A.Y.; Korzun, D.G.; Moschevikin, A.P.; Reginya, S.; Gerasimova-Meigal, L.I. Ambient Assisted Living At-Home Laboratory for Motor Status Diagnostics in Parkinson’s Disease Patients and Aged People. In Research Anthology on Supporting Healthy Aging in a Digital Society; IGI Global: Hershey, PA, USA, 2022; pp. 836–862. Available online: https://services.igi-global.com/resolvedoi/resolve.aspx?doi=10.4018/978-1-6684-5295-0.ch047 (accessed on 1 October 2022). [CrossRef]
- Fernandes, J.M.; Silva, J.S.; Rodrigues, A.; Boavida, F. A Survey of Approaches to Unobtrusive Sensing of Humans. ACM Comput. Surv. 2022, 55, 1–28. [Google Scholar] [CrossRef]
- Kostakos, V.; Rogstadius, J.; Ferreira, D.; Hosio, S.; Goncalves, J. Participatory Sensing, Opinions and Collective Awareness, 1st ed.; Loreto, V., Haklay, M., Hotho, A., Servedio, V.D.P., Stumme, G., Theunis, J., Tria, F., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 69–92. ISBN 978-3-319-25658-0. [Google Scholar]
- Korzun, D.G. Internet of Things Meets Mobile Health Systems in Smart Spaces: An Overview. In Internet of Things and Big Data Technologies for Next Generation Healthcare; Bhatt, C., Dey, N., Ashour, A.S., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 111–129. ISBN 978-3-319-49736-5. [Google Scholar]
- Korzun, D.; Balandina, E.; Kashevnik, A.; Balandin, S.; Viola, F. Ambient Intelligence Services in IoT Environments: Emerging Research and Opportunities; IGI Global: Hershey, PA, USA, 2019; ISBN 9781522589730. [Google Scholar]
- Warden, P.; Stewart, M.; Plancher, B.; Banbury, C.; Prakash, S.; Chen, E.; Asgar, Z.; Katti, S.; Reddi, V.J. Machine Learning Sensors. arXiv 2022. [Google Scholar] [CrossRef]
- Chinchole, S.; Patel, S. Artificial Intelligence and Sensors Based Assistive System for the Visually Impaired People. In Proceedings of the 2017 International Conference on Intelligent Sustainable Systems (ICISS), Palladam, India, 7–8 December 2017; pp. 16–19. [Google Scholar]
- Gulzar Ahmad, S.; Iqbal, T.; Javaid, A.; Ullah Munir, E.; Kirn, N.; Ullah Jan, S.; Ramzan, N. Sensing and Artificial Intelligent Maternal-Infant Health Care Systems: A Review. Sensors 2022, 22, 4362. [Google Scholar] [CrossRef]
- Machine Learning Sensors: Truly Data-Centric AI | Towards Data Science. Available online: https://towardsdatascience.com/machine-learning-sensors-truly-data-centric-ai-8f6b9904633a (accessed on 23 August 2022).
- Ma, Y.; Hou, L.; Yang, X.; Huang, Z.; Yang, X.; Zhao, N.; He, M.; Shi, Y.; Kang, Y.; Yue, J.; et al. The Association between Frailty and Severe Disease among COVID-19 Patients Aged over 60 Years in China: A Prospective Cohort Study. BMC Med. 2020, 18, 274. [Google Scholar] [CrossRef]
- Fan, B.E. Hematologic Parameters in Patients with COVID-19 Infection: A Reply. Am. J. Hematol. 2020, 95, E215. [Google Scholar] [CrossRef]
- Ferrari, D.; Motta, A.; Strollo, M.; Banfi, G.; Locatelli, M. Routine Blood Tests as a Potential Diagnostic Tool for COVID-19. Clin. Chem. Lab. Med. 2020, 58, 1095–1099. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhang, P.; Zhang, L.; Meng, W.; Li, J.; Tong, C.; Li, Y.; Cai, J.; Yang, Z.; Zhu, J.; et al. Rapid and Accurate Identification of COVID-19 Infection through Machine Learning Based on Clinical Available Blood Test Results. medRxiv 2020. [Google Scholar] [CrossRef]
- Shakeel, P.M.; Burhanuddin, M.A.; Desa, M.I. Lung Cancer Detection from CT Image Using Improved Profuse Clustering and Deep Learning Instantaneously Trained Neural Networks. Meas. J. Int. Meas. Confed. 2019, 145, 702–712. [Google Scholar] [CrossRef]
- Cui, H.; Wang, X.; Feng, D. Automated Localization and Segmentation of Lung Tumor from PET-CT Thorax Volumes Based on Image Feature Analysis. In Proceedings of the 2012 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, San Diego, CA, USA, 28 August–1 September 2012; pp. 5384–5387. [Google Scholar] [CrossRef]
- Tomita, K.; Nagao, R.; Touge, H.; Ikeuchi, T.; Sano, H.; Yamasaki, A.; Tohda, Y. Deep Learning Facilitates the Diagnosis of Adult Asthma. Allergol. Int. 2019, 68, 456–461. [Google Scholar] [CrossRef]
- Ryu, K.S.; Lee, S.W.; Batbaatar, E.; Lee, J.W.; Choi, K.S.; Cha, H.S. A Deep Learning Model for Estimation of Patients with Undiagnosed Diabetes. Appl. Sci. 2020, 10, 421. [Google Scholar] [CrossRef]
- Kolachalama, V.B.; Singh, P.; Lin, C.Q.; Mun, D.; Belghasem, M.E.; Henderson, J.M.; Francis, J.M.; Salant, D.J.; Chitalia, V.C. Association of Pathological Fibrosis With Renal Survival Using Deep Neural Networks. Kidney Int. Rep. 2018, 3, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.C.; Yeh, W.C.; Hsu, W.D.; Islam, M.M.; Nguyen, P.A.A.; Poly, T.N.; Wang, Y.C.; Yang, H.C.; Li, Y.C.J. Prediction of Fatty Liver Disease Using Machine Learning Algorithms. Comput. Methods Programs Biomed. 2019, 170, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Oguntimilehin, A.; Adetunmbi, A.O.; Abiola, O.B. A Machine Learning Approach to Clinical Diagnosis of Typhoid Fever. Int. J. Comput. Inf. Technol. 2013, 4, 671–676. [Google Scholar]
- Kouchaki, S.; Yang, Y.Y.; Walker, T.M.; Walker, A.S.; Wilson, D.J.; Peto, T.E.A.; Crook, D.W.; Clifton, D.A.; Hoosdally, S.J.; Gibertoni Cruz, A.L.; et al. Application of Machine Learning Techniques to Tuberculosis Drug Resistance Analysis. Bioinformatics 2019, 35, 2276–2282. [Google Scholar] [CrossRef]
- Taylor, R.A.; Moore, C.L.; Cheung, K.H.; Brandt, C. Predicting Urinary Tract Infections in the Emergency Department with Machine Learning. PLoS ONE 2018, 13, e0194085. [Google Scholar] [CrossRef]
- Yang, H.S.; Hou, Y.; Vasovic, L.V.; Steel, P.A.D.; Chadburn, A.; Racine-Brzostek, S.E.; Velu, P.; Cushing, M.M.; Loda, M.; Kaushal, R.; et al. Routine Laboratory Blood Tests Predict SARS-CoV-2 Infection Using Machine Learning. Clin. Chem. 2020, 66, 1396–1404. [Google Scholar] [CrossRef] [PubMed]
- Brinati, D.; Campagner, A.; Ferrari, D.; Locatelli, M.; Banfi, G.; Cabitza, F. Detection of COVID-19 Infection from Routine Blood Exams with Machine Learning: A Feasibility Study. J. Med. Syst. 2020, 44, 135. [Google Scholar] [CrossRef]
- Zhang, S.; Huang, S.; Liu, J.; Dong, X.; Meng, M.; Chen, L.; Wen, Z.; Zhang, L.; Chen, Y.; Du, H.; et al. Identification and Validation of Prognostic Factors in Patients with COVID-19: A Retrospective Study Based on Artificial Intelligence Algorithms. J. Intensive Med. 2021, 1, 103–109. [Google Scholar] [CrossRef]
- Alle, S.; Karthikeyan, A.; Kanakan, A.; Siddiqui, S.; Garg, A.; Mehta, P.; Mishra, N.; Chattopadhyay, P.; Devi, P.; Waghdhare, S.; et al. COVID-19 Risk Stratification and Mortality Prediction in Hospitalized Indian Patients: Harnessing Clinical Data for Public Health Benefits. PLoS ONE 2022, 17, e0264785. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Chen, L.; Chi, J.; Zeng, S.; Feng, X.; Li, H.; Liu, D.; Feng, X.; Wang, S.; Wang, Y.; et al. Development and Validation of an Online Model to Predict Critical COVID-19 with Immune-Inflammatory Parameters. J. Intensive Care 2021, 9, 19. [Google Scholar] [CrossRef]
- Vaishnav, P.K.; Sharma, S.; Sharma, P. Analytical Review Analysis for Screening COVID-19 Disease. Int. J. Mod. Res. 2021, 1, 22–29. [Google Scholar]
- Kukar, M.; Gunčar, G.; Vovko, T.; Podnar, S.; Černelč, P.; Brvar, M.; Zalaznik, M.; Notar, M.; Moškon, S.; Notar, M. COVID-19 Diagnosis by Routine Blood Tests Using Machine Learning. Sci. Rep. 2021, 11, 10738. [Google Scholar] [CrossRef] [PubMed]
- Mei, X.; Lee, H.C.; Diao, K.Y.; Huang, M.; Lin, B.; Liu, C.; Xie, Z.; Ma, Y.; Robson, P.M.; Chung, M.; et al. Artificial Intelligence–Enabled Rapid Diagnosis of Patients with COVID-19. Nat. Med. 2020, 26, 1224–1228. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Pastora, J.; Weigand, M.; Kim, J.; Wu, X.; Strayer, J.; Palmer, A.F.; Zborowski, M.; Yazer, M.; Chalmers, J.J. Hyperferritinemia in Critically Ill COVID-19 Patients—Is Ferritin the Product of Inflammation or a Pathogenic Mediator? Clin. Chim. Acta 2020, 509, 249–251. [Google Scholar] [CrossRef]
- Boslaugh, S. Statistics in a Nutshell, 2nd ed.; O’Reilly Media, Incorporated: Newton, MA, USA, 2012; ISBN 9781449361129. [Google Scholar]
- Weiss, G.; Ganz, T.; Goodnough, L.T. Anemia of Inflammation. Blood 2019, 133, 40–50. [Google Scholar] [CrossRef]
- Harrington, P. Machine Learning in Action; Simon and Schuster: New York, NY, USA, 2012; ISBN 978-1617290183. [Google Scholar]
- Feurer, M.; Eggensperger, K.; Falkner, S.; Lindauer, M.; Hutter, F. Auto-Sklearn 2.0: Hands-Free AutoML via Meta-Learning. arXiv 2020, arXiv:2007.04074. [Google Scholar]
- Kim, S.; Kim, D.-M.; Lee, B. Insufficient Sensitivity of RNA Dependent RNA Polymerase Gene of SARS-CoV-2 Viral Genome as Confirmatory Test Using Korean COVID-19 Cases. Preprints 2020, 1–14. [Google Scholar] [CrossRef]
- Teymouri, M.; Mollazadeh, S.; Mortazavi, H.; Naderi Ghale-noie, Z.; Keyvani, V.; Aghababaei, F.; Hamblin, M.R.; Abbaszadeh-Goudarzi, G.; Pourghadamyari, H.; Hashemian, S.M.R.; et al. Recent Advances and Challenges of RT-PCR Tests for the Diagnosis of COVID-19. Pathol. Res. Pract. 2021, 221, 153443. [Google Scholar] [CrossRef]
- Djakpo, D.K.; Wang, Z.; Zhang, R.; Chen, X.; Chen, P.; Ketisha Antoine, M.M.L. Blood Routine Test in Mild and Common 2019 Coronavirus (COVID-19) Patients. Biosci. Rep. 2020, 40, BSR20200817. [Google Scholar] [CrossRef]
- Fang, B.; Meng, Q.H. The Laboratory’s Role in Combating COVID-19. Crit. Rev. Clin. Lab. Sci. 2020, 57, 400–414. [Google Scholar] [CrossRef]
- Bertolini, A.; van de Peppel, I.P.; Bodewes, F.A.J.A.; Moshage, H.; Fantin, A.; Farinati, F.; Fiorotto, R.; Jonker, J.W.; Strazzabosco, M.; Verkade, H.J.; et al. Abnormal Liver Function Tests in Patients With COVID-19: Relevance and Potential Pathogenesis. Hepatology 2020, 72, 1864–1872. [Google Scholar] [CrossRef]
- Terpos, E.; Ntanasis-Stathopoulos, I.; Elalamy, I.; Kastritis, E.; Sergentanis, T.N.; Politou, M.; Psaltopoulou, T.; Gerotziafas, G.; Dimopoulos, M.A. Hematological Findings and Complications of COVID-19. Am. J. Hematol. 2020, 95, 834–847. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.; Dai, R.; Du, R.C.; Zhu, Y.; Shui, L.P.; Luo, X.H. Hematologic Changes Predict Clinical Outcome in Recovered Patients with COVID-19. Ann. Hematol. 2021, 100, 675–689. [Google Scholar] [CrossRef]
- Kočar, E.; Režen, T.; Rozman, D. Cholesterol, Lipoproteins, and COVID-19: Basic Concepts and Clinical Applications. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2021, 1866, 209–214. [Google Scholar] [CrossRef]
- Zinellu, A.; Paliogiannis, P.; Fois, A.G.; Solidoro, P.; Carru, C.; Mangoni, A.A. Cholesterol and Triglyceride Concentrations, COVID-19 Severity, and Mortality: A Systematic Review and Meta-Analysis With Meta-Regression. Front. Public Health 2021, 9, 705916. [Google Scholar] [CrossRef]
- Wei, X.; Zeng, W.; Su, J.; Wan, H.; Yu, X.; Cao, X.; Tan, W.; Wang, H. Hypolipidemia Is Associated with the Severity of COVID-19. J. Clin. Lipidol. 2020, 14, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Stephens, J.R.; Wong, J.L.C.; Broomhead, R.; Mpfle, R.S.; Waheed, U.; Patel, P.; Brett, S.J.; Soni, S. Raised Serum Amylase in Patients with COVID-19 May Not Be Associated with Pancreatitis. Br. J. Surg. 2021, 108, e152–e153. [Google Scholar] [CrossRef]
- Wu, M.Y.; Yao, L.; Wang, Y.; Zhu, X.Y.; Wang, X.F.; Tang, P.J.; Chen, C. Clinical Evaluation of Potential Usefulness of Serum Lactate Dehydrogenase (LDH) in 2019 Novel Coronavirus (COVID-19) Pneumonia. Respir. Res. 2020, 21, 171. [Google Scholar] [CrossRef] [PubMed]
- Hu, F.; Guo, Y.; Lin, J.; Zeng, Y.; Wang, J.; Li, M.; Cong, L. Association of Serum Uric Acid Levels with COVID-19 Severity. BMC Endocr. Disord. 2021, 21, 97. [Google Scholar] [CrossRef] [PubMed]
- Zinellu, A.; Sotgia, S.; Fois, A.G.; Mangoni, A.A. Serum CK-MB, COVID-19 Severity and Mortality: An Updated Systematic Review and Meta-Analysis with Meta-Regression. Adv. Med. Sci. 2021, 66, 304–314. [Google Scholar] [CrossRef]
- Afra, H.S.; Amiri-Dashatan, N.; Ghorbani, F.; Maleki, I. Positive Association between Severity of COVID-19 Infection and Liver Damage: A Systematic Review and Meta-Analysis. Gastroenterol. Hepatol. Bed Bench 2020, 13, 292–304. [Google Scholar] [CrossRef]
- Huang, Y.; Yang, R.; Xu, Y.; Gong, P. Clinical Characteristics of 36 Non-Survivors with COVID-19 in Wuhan, China. medRxiv 2020. [Google Scholar] [CrossRef]
- González-Cruz, C.; Jofre, M.A.; Podlipnik, S.; Combalia, M.; Gareau, D.; Gamboa, M.; Vallone, M.G.; Faride Barragán-Estudillo, Z.; Tamez-Peña, A.L.; Montoya, J.; et al. Machine Learning in Melanoma Diagnosis. Limitations About to Be Overcome. Actas Dermo-Sifiliográficas (Engl. Ed.) 2020, 111, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.; Reddy, V.; Myers, B.; Thibodeaux, Q.; Brownstone, N.; Liao, W. Machine Learning in Dermatology: Current Applications, Opportunities, and Limitations. Dermatol. Ther. 2020, 10, 365–386. [Google Scholar] [CrossRef] [PubMed]
- Sáez, C.; Romero, N.; Conejero, J.A.; García-Gómez, J.M. Potential Limitations in COVID-19 Machine Learning Due to Data Source Variability: A Case Study in the NCov2019 Dataset. J. Am. Med. Inform. Assoc. 2021, 28, 360–364. [Google Scholar] [CrossRef]
- Amin, S.U.; Hossain, M.S. Edge Intelligence and Internet of Things in Healthcare: A Survey. IEEE Access 2021, 9, 45–59. [Google Scholar] [CrossRef]
- Sruthi, P.L.; Raju, K.B. Prediction of the COVID-19 Pandemic with Machine Learning Models. In Proceedings of the 2021 Fifth International Conference on I-SMAC (IoT in Social, Mobile, Analytics and Cloud) (I-SMAC), Palladam, India, 11–13 November 2021; pp. 474–481. [Google Scholar]
- Anjum, N.; Alibakhshikenari, M.; Rashid, J.; Jabeen, F.; Asif, A.; Mohamed, E.M.; Falcone, F. IoT-Based COVID-19 Diagnosing and Monitoring Systems: A Survey. IEEE Access 2022, 10, 87168–87181. [Google Scholar] [CrossRef]
- Mukati, N.; Namdev, N.; Dilip, R.; Hemalatha, N.; Dhiman, V.; Sahu, B. Healthcare Assistance to COVID-19 Patient Using Internet of Things (IoT) Enabled Technologies. Mater. Today Proc. 2021. [Google Scholar] [CrossRef]
- Meigal, A.Y.; Korzun, D.G.; Gerasimova-Meigal, L.I.; Borodin, A.V.; Zavyalova, Y.V. Ambient Intelligence At-Home Laboratory for Human Everyday Life. Int. J. Embed. Real-Time Commun. Syst. 2019, 10, 117–134. Available online: https://services.igi-global.com/resolvedoi/resolve.aspx?doi=10.4018/IJERTCS.2019040108 (accessed on 14 September 2022). [CrossRef]
- Vetter, B.; Sampath, R.; Carmona, S. Landscape of Point-of-Care Devices for Testing of Cardiometabolic Diseases; FIND: Geneva, Switzerland, 2020. [Google Scholar]
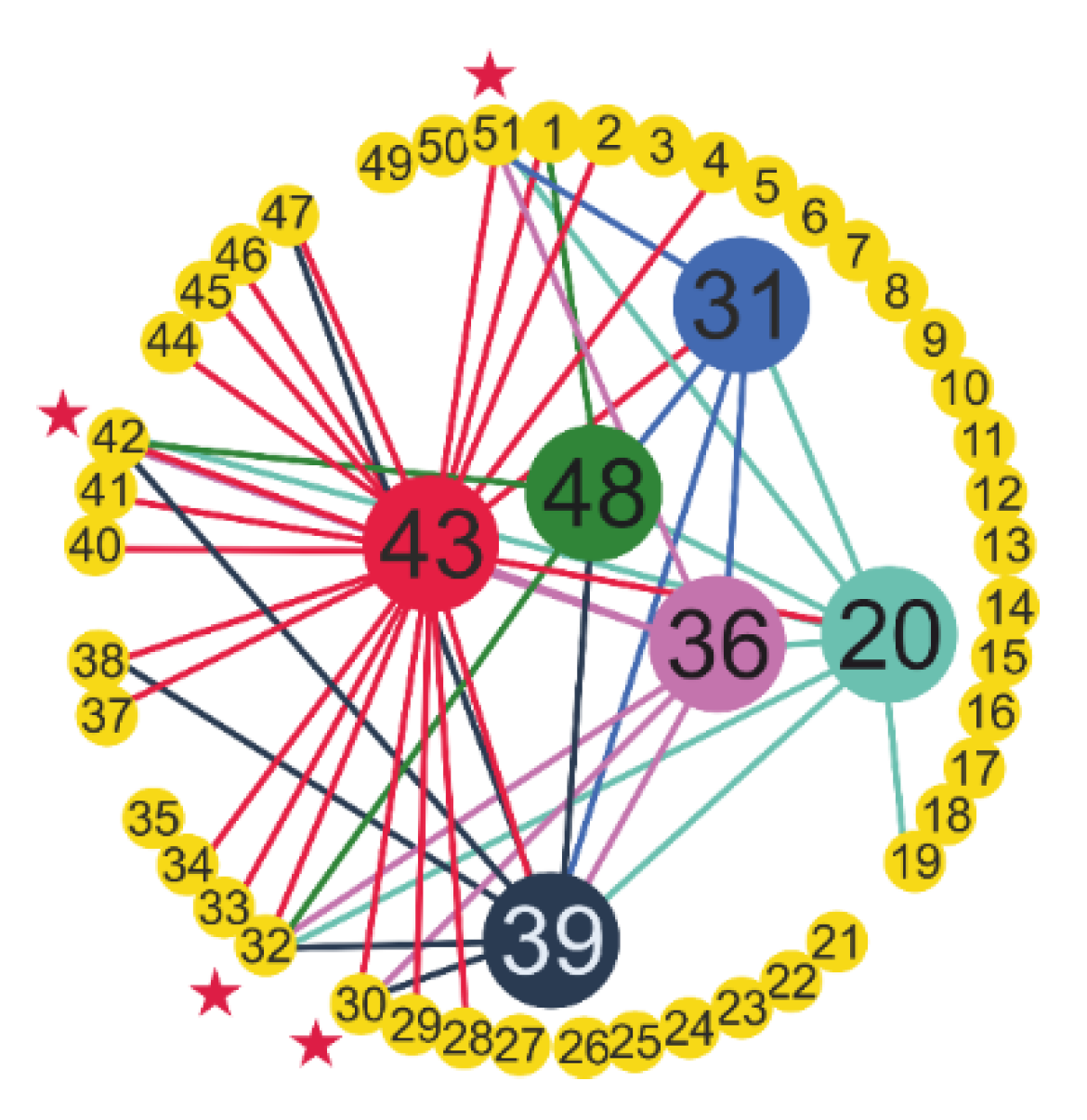
| № | Feature | № | Feature | № | Feature | № | Feature | № | Feature |
|---|---|---|---|---|---|---|---|---|---|
| 1 | CRP | 12 | NEU | 23 | MPV | 34 | GGT | 45 | Sodium |
| 2 | D-Dimer | 13 | PLT | 24 | PDW | 35 | Glucose | 46 | T-Bil |
| 3 | Ferritin | 14 | WBC | 25 | RBC | 36 | HDL-C | 47 | TP |
| 4 | Fibrinogen | 15 | BASO | 26 | RDW | 37 | Calcium | 48 | Triglyceride |
| 5 | INR | 16 | EOS | 27 | ALT | 38 | Chlorine | 49 | eGFR |
| 6 | PT | 17 | HCT | 28 | AST | 39 | Cholesterol | 50 | Urea |
| 7 | PCT | 18 | HGB | 29 | Albumin | 40 | Creatinine | 51 | UA |
| 8 | ESR | 19 | MCH | 30 | ALP | 41 | CK | ||
| 9 | Troponin | 20 | MCHC | 31 | Amylase | 42 | LDH | ||
| 10 | aPTT | 21 | MCV | 32 | CK-MB | 43 | LDL | ||
| 11 | LYM | 22 | MONO | 33 | D-Bil | 44 | Potassium |
| Chaotic Map | List of Parameters | Equation | |
|---|---|---|---|
| Congruent generator | K = 93 D = 68 L = 9276 C = 73 | (1) |
| Feature (Point-Biserial Correlation Coefficient (rpb)) | Feature (Threshold Accuracy of Classification Ath from [12]) |
|---|---|
| MCHC (0.8) | MCHC (94.35%) |
| HDL-C (−0.77) | HDL-C (94.73%) |
| Cholesterol (−0.71) | Cholesterol (94.47%) |
| LDL (−0.68) | LDL (96.47%) |
| Triglyceride (90.96%) | |
| Amylase (85.1%) | |
| UA (81.12%) | |
| TP (79.68%) | |
| CK-MB (78.91%) | |
| LDH (74.98%) | |
| Albumin (72.91%) |
| Pair Feature–Feature | Pearson’s Coefficient for COVID-19 Diagnosis | Pearson’s Coefficient for Positive COVID-19 | Pearson’s Coefficient for Negative COVID-19 |
|---|---|---|---|
| Type High–High | |||
| HCT–HGB | 0.96 | 0.95 (High) | 0.97(High) |
| MPV–PDW | 0.93 | 0.94 | 0.92 |
| HCT–RBC | 0.87 | 0.88 | 0.87 |
| MCH–MCV | 0.84 | 0.84 | 0.84 |
| HGB–RBC | 0.83 | 0.83 | 0.83 |
| NEU–WBC | 0.74 | 0.71 | 0.81 |
| Albumin–TP | 0.64 | 0.67 | 0.5 |
| MCH–MCHC | 0.53 | 0.62 | 0.99 |
| MCH–RDW | −0.55 | −0.61 | −0.51 |
| Type High–Low | |||
| Fibrinogen–LYM | −0.77 | −0.78 (High) | −0.01 (Low) |
| Cholesterol–LDL | 0.65 | 0.59 | 0.012 |
| Cholesterol–HDL-C | 0.64 | 0.39 | −0.024 |
| Chlorine–Sodium | 0.18 | 0.63 | −0.025 |
| Type Low–High | |||
| MCHC–MCV | 0.41 | 0.09 1(Low) | 0.84 (High) |
| ALT–AST | 0.6 | 0.48 | 0.76 |
| eGFR–Urea | −0.55 | −0.49 | −0.63 |
| INR–PT | 0.12 | 0.075 | 1 |
| D-Bil–T-Bil | 0.6 | 0.33 | 0.91 |
| HDL-C–LDL | 0.63 | 0.19 | 0.3 |
| Classification Algorithm | Average Model Accuracy A51,% | Average Learning Time, s | Average Inference Time, μs | Normalization Method | Methods for Generating Additional Features |
|---|---|---|---|---|---|
| Histogram-based Gradient Boosting | 100 | 6.39 | 11.6 | - | - |
| Random Forest | 99.943 | 13.15 | 21.9 | QT | - |
| K-nearest neighbors | 99.924 | 3.17 | 22.1 | QT | ETP |
| Extra Trees classifier | 99.905 | 18.73 | 24.5 | RS | - |
| Multilayer Perceptron | 99.886 | 3.99 | 2.2 | RS | LSVMP |
| Multinomial Naive Bayes | 99.792 | 2.48 | 11.7 | QT | RTE |
| Linear Discriminant Analysis | 99.773 | 9.15 | 7.5 | QT | PN |
| Support Vector Machine with non-linear kernel | 99.754 | 222.41 | 43.2 | QT | NS |
| Decision Tree | 99.660 | 1.46 | 1.2 | RS | LSVMP |
| Passive-Aggressive | 99.641 | 2.91 | 13.1 | QT | RTE |
| Bernoulli Naive Bayes | 99.622 | 2.59 | 11.3 | QT | RTE |
| Support Vector Machine with linear kernel | 99.584 | 5.21 | 1242 | MM | PN |
| Gaussian Naive Bayes | 98.565 | 1.68 | 3.5 | QT | ICA |
| LogNNet [12] | 99.509 | 100 | 3 | - | - |
| № | Feature | A1,% | № | Feature | A1,% | № | Feature | A1,% | № | Feature | A1,% |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 43 | LDL | 96.84 | 4 | Fibrinogen | 76.03 | 50 | Urea | 68.10 | 21 | MCV | 56.43 |
| 39 | Cholesterol | 95.07 | 29 | Albumin | 75.3 | 7 | PCT | 63.25 | 22 | MONO | 56.26 |
| 36 | HDL-C | 94.99 | 44 | Potassium | 75.22 | 27 | ALT | 62.33 | 5 | INR | 56.19 |
| 20 | MCHC | 94.35 | 3 | Ferritin | 74.45 | 35 | Glucose | 62.17 | 6 | PT | 56.04 |
| 48 | Triglyceride | 93.76 | 38 | Chlorine | 73.18 | 49 | eGFR | 62.04 | 17 | HCT | 55.75 |
| 31 | Amylase | 90.01 | 46 | T-Bil | 72.77 | 14 | WBC | 61.91 | 26 | RDW | 55.62 |
| 51 | UA | 87.91 | 34 | GGT | 72.62 | 16 | EOS | 61.40 | 9 | Troponin | 54.07 |
| 42 | LDH | 85.76 | 41 | CK | 70.97 | 13 | PLT | 61.25 | 18 | HGB | 53.94 |
| 47 | TP | 80.41 | 2 | D-Dimer | 70.46 | 28 | AST | 60.55 | 25 | RBC | 53.43 |
| 37 | Calcium | 80.40 | 33 | D-Bil | 70.37 | 8 | ESR | 59.12 | 23 | MPV | 53.13 |
| 32 | CK-MB | 79.73 | 11 | LYM | 69.90 | 15 | BASO | 58.72 | 24 | PDW | 53.09 |
| 1 | CRP | 77.81 | 45 | Sodium | 69.35 | 12 | NEU | 57.51 | 19 | MCH | 52.13 |
| 30 | ALP | 77,71 | 40 | Creatinine | 69,24 | 10 | aPTT | 56.53 |
| № | First Feature | Second Feature | Average Accuracy A2,% |
|---|---|---|---|
| 20-19 | MCHC | MCH | 99.81 |
| 43-32 | LDL | CK-MB | 99.62 |
| 36-32 | HDL-C | CK-MB | 99.49 |
| 48-32 | Triglyceride | CK-MB | 99.45 |
| 43-39 | LDL | Cholesterol | 99.43 |
| 43-20 | LDL | MCHC | 99.22 |
| 39-36 | Cholesterol | HDL-C | 99.18 |
| 39-48 | Cholesterol | Triglyceride | 99.11 |
| 43-42 | LDL | LDH | 99.05 |
| 43-31 | LDL | Amylase | 99.03 |
| 36-20 | HDL-C | MCHC | 98.98 |
| 43-51 | LDL | UA | 98.86 |
| 36-31 | HDL-C | Amylase | 98.81 |
| 39-20 | Cholesterol | MCHC | 98.73 |
| 20-48 | MCHC | Triglyceride | 98.65 |
| 39-38 | Cholesterol | Chlorine | 98.62 |
| 43-38 | LDL | Chlorine | 98.43 |
| 20-31 | MCHC | Amylase | 98.28 |
| 36-42 | HDL-C | LDH | 98.16 |
| 48-42 | Triglyceride | LDH | 98.14 |
| № | First Feature | Second Feature | Third Feature | Average Accuracy A3,% |
|---|---|---|---|---|
| 39-48-32 | Cholesterol | Triglyceride | CK-MB | 99.91 |
| 39-36-32 | Cholesterol | HDL-C | CK-MB | 99.91 |
| 43-20-19 | LDL | MCHC | MCH | 99.91 |
| 20-31-19 | MCHC | Amylase | MCH | 99.85 |
| 43-51-32 | LDL | UA | CK-MB | 99.85 |
| 39-20-19 | Cholesterol | MCHC | MCH | 99.83 |
| 48-42-32 | Triglyceride | LDH | CK-MB | 99.83 |
| 36-20-19 | HDL-C | MCHC | MCH | 99.79 |
| 36-42-32 | HDL-C | LDH | CK-MB | 99.79 |
| 43-38-51 | LDL | Cholesterol | UA | 99.79 |
| 20-48-19 | MCHC | Triglyceride | MCH | 99.77 |
| 39-48-31 | Cholesterol | Triglyceride | Amylase | 99.77 |
| 39-36-38 | Cholesterol | HDL-C | Chlorine | 99.75 |
| 36-31-51 | HDL-C | Amylase | UA | 99.75 |
| 39-36-42 | Cholesterol | HDL-C | LDH | 99.75 |
| 20-51-19 | MCHC | UA | MCH | 99.74 |
| 39-48-38 | Cholesterol | Triglyceride | Chlorine | 99.72 |
| 39-31-51 | Cholesterol | Amylase | UA | 99.70 |
| 39-48-42 | Cholesterol | Triglyceride | LDH | 99.66 |
| 48-31-42 | Triglyceride | Amylase | LDH | 99.51 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Velichko, A.; Huyut, M.T.; Belyaev, M.; Izotov, Y.; Korzun, D. Machine Learning Sensors for Diagnosis of COVID-19 Disease Using Routine Blood Values for Internet of Things Application. Sensors 2022, 22, 7886. https://doi.org/10.3390/s22207886
Velichko A, Huyut MT, Belyaev M, Izotov Y, Korzun D. Machine Learning Sensors for Diagnosis of COVID-19 Disease Using Routine Blood Values for Internet of Things Application. Sensors. 2022; 22(20):7886. https://doi.org/10.3390/s22207886
Chicago/Turabian StyleVelichko, Andrei, Mehmet Tahir Huyut, Maksim Belyaev, Yuriy Izotov, and Dmitry Korzun. 2022. "Machine Learning Sensors for Diagnosis of COVID-19 Disease Using Routine Blood Values for Internet of Things Application" Sensors 22, no. 20: 7886. https://doi.org/10.3390/s22207886
APA StyleVelichko, A., Huyut, M. T., Belyaev, M., Izotov, Y., & Korzun, D. (2022). Machine Learning Sensors for Diagnosis of COVID-19 Disease Using Routine Blood Values for Internet of Things Application. Sensors, 22(20), 7886. https://doi.org/10.3390/s22207886









