Best Fit DNA-Based Cryptographic Keys: The Genetic Algorithm Approach
Abstract
1. Introduction
- Inculcate the benefits of Genetic Algorithms in DNA cryptography instead of Traditional Cryptography.
- Categorize the initial population of keys as strong or weak. The strong keys are used as it is for encryption. The weak keys instead of getting dropped are strengthened by the Genetic Algorithm. This step reduces the key generation time by only applying the scheme to weak keys. It also reduces key wastage.
- Propose suitable fitness functions by checking the frequency and gap of occurrence of the four nitrogenous bases to convert the weak keys into their fitter counterparts. It also reduces key wastage and enhances their efficiency for effective DNA-based cryptographic schemes.
2. Related Work
- The majority of the existing schemes are based on traditional binary keys and much less emphasis has been made on DNA-based keys.
- Most existing algorithms discussed are applying their proposed methodology to the initial key population which makes the key generation process lengthy and difficult.
- Based on the suitability of their proposal, each algorithm has defined its fitness test and selection, crossover, and mutation are the predominant genetic operators used.
- To choose the appropriate fitness test to be used as four different nitrogenous bases are involved in DNA cryptosystems.
- To decide whether the methodology is to be applied to the initial key population or not. For this, the fitness test is applied, and keys are categorized as strong or weak. If found strong, they are directly used for encryption. Only the weak keys are acted upon and thus the number of keys to be acted upon is reduced and the time complexity will reduce.
- To reduce key wastage by strengthening the weak keys and removing visible patterns instead of completely discarding them.
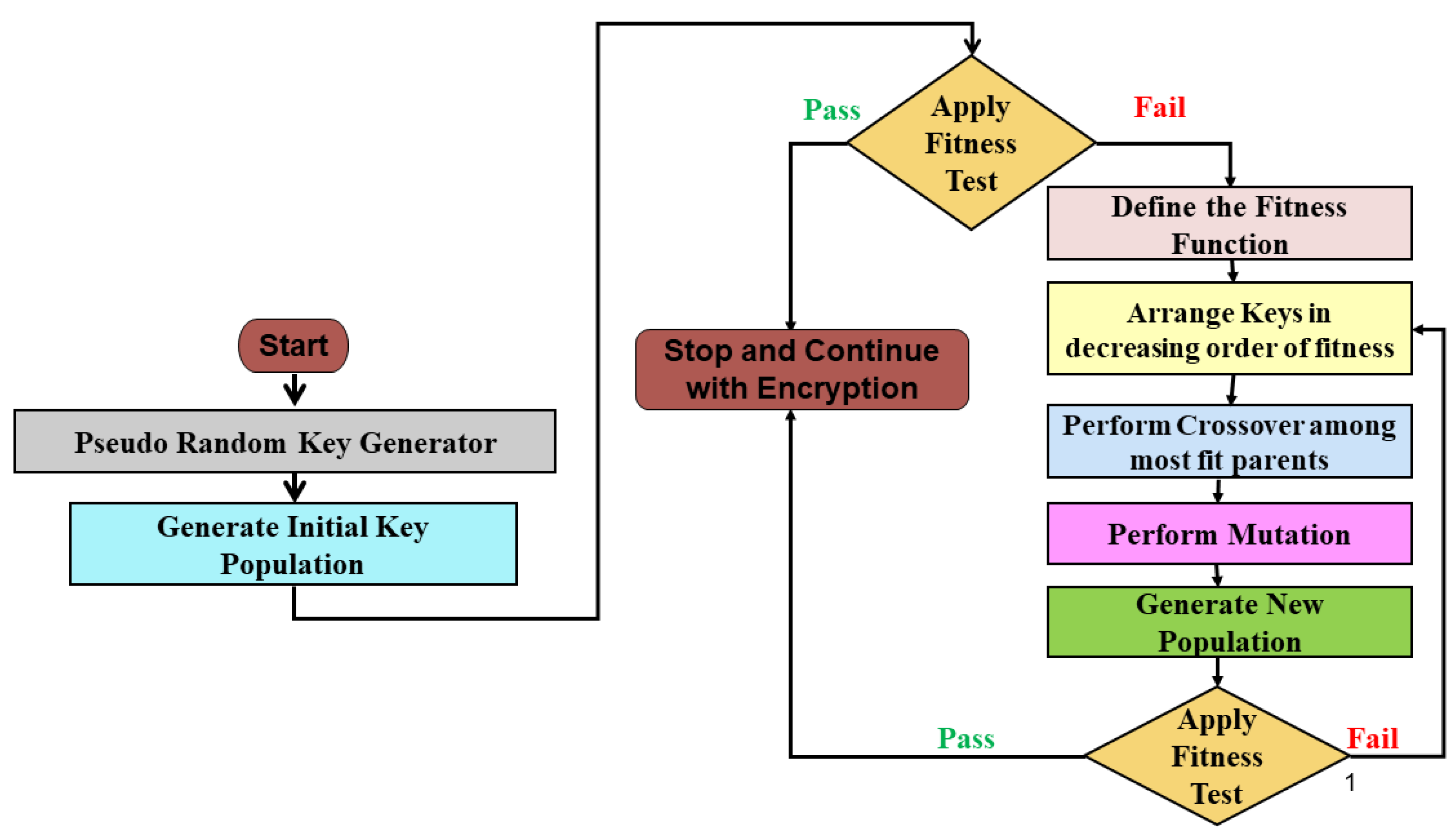
3. Proposed Methodology
3.1. Generating the Initial Population
3.2. Applying Fitness Tests
3.3. Defining Fitness Functions for Weak Keys
3.4. Arranging in Decreasing Order of Fitness Function
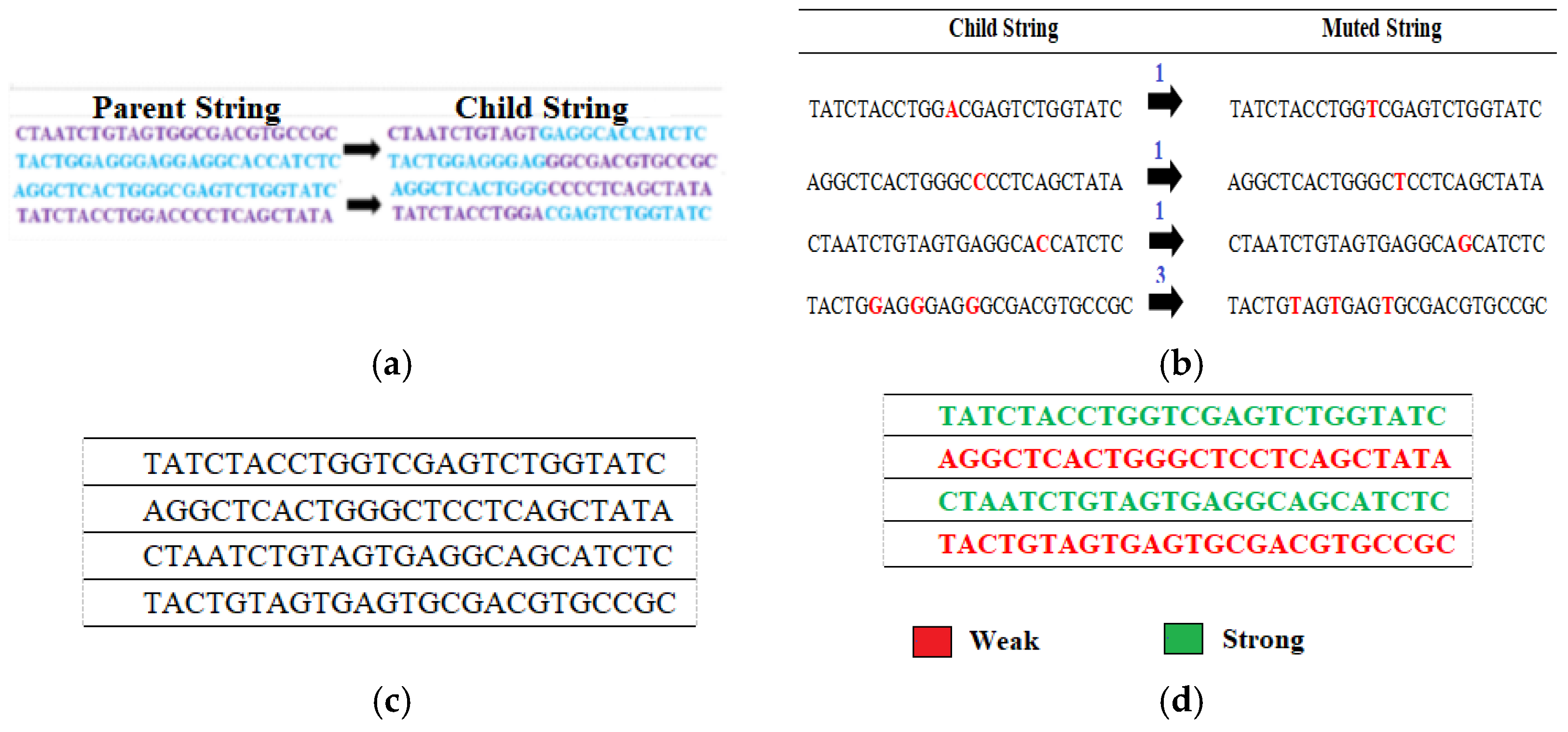
3.5. Perform Crossover Operation
3.6. Perform Mutation Operation
3.7. Generate the New Population
3.8. Reapply Fitness Test and Repeat the Entire Process
4. Results and Calculations
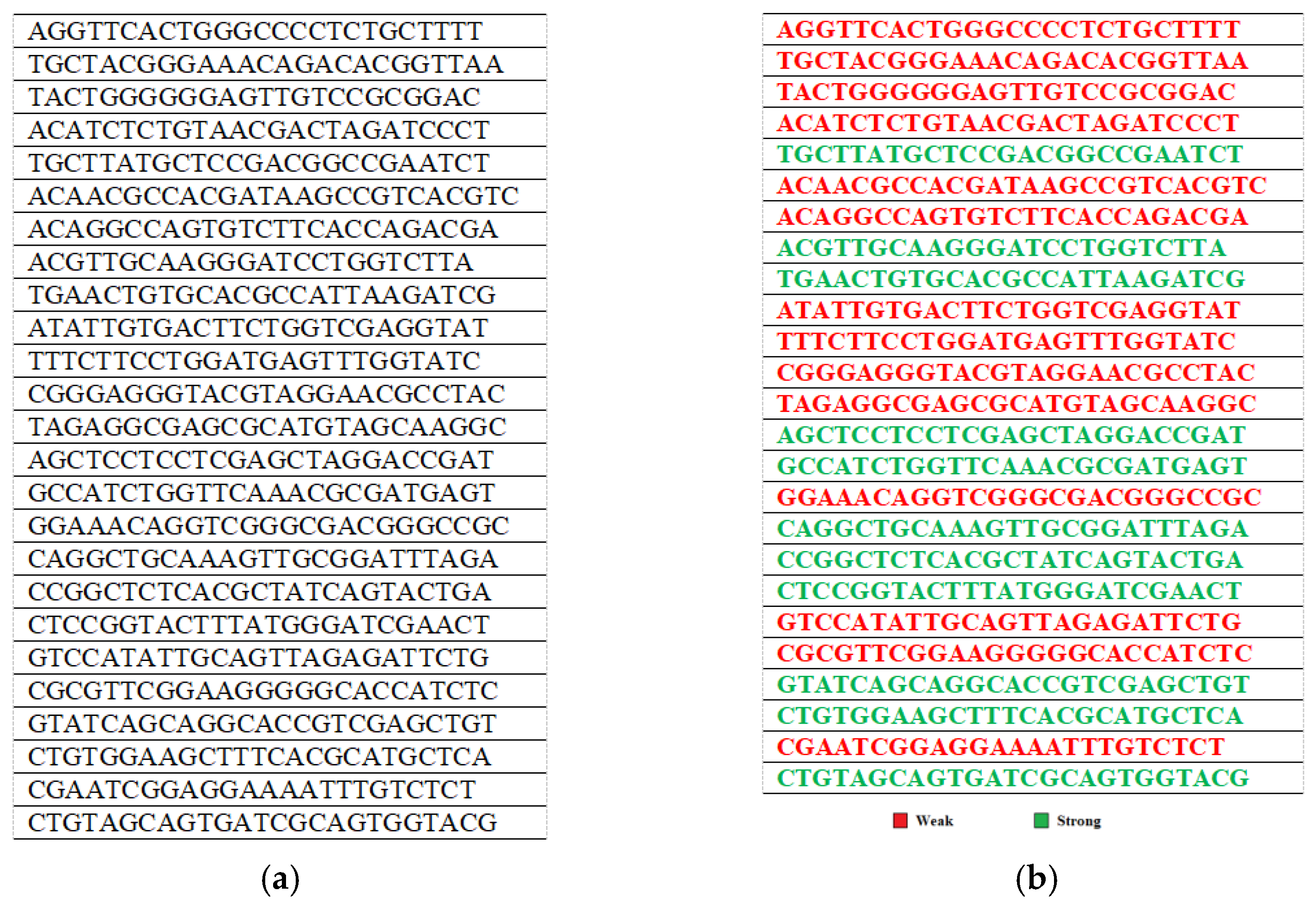
4.1. Generating the Initial Population
4.2. Applying Fitness Tests
4.3. Defining Fitness Functions for Weak Keys
4.4. Arranging in Decreasing Order of Fitness Function
4.5. Perform Crossover Operation
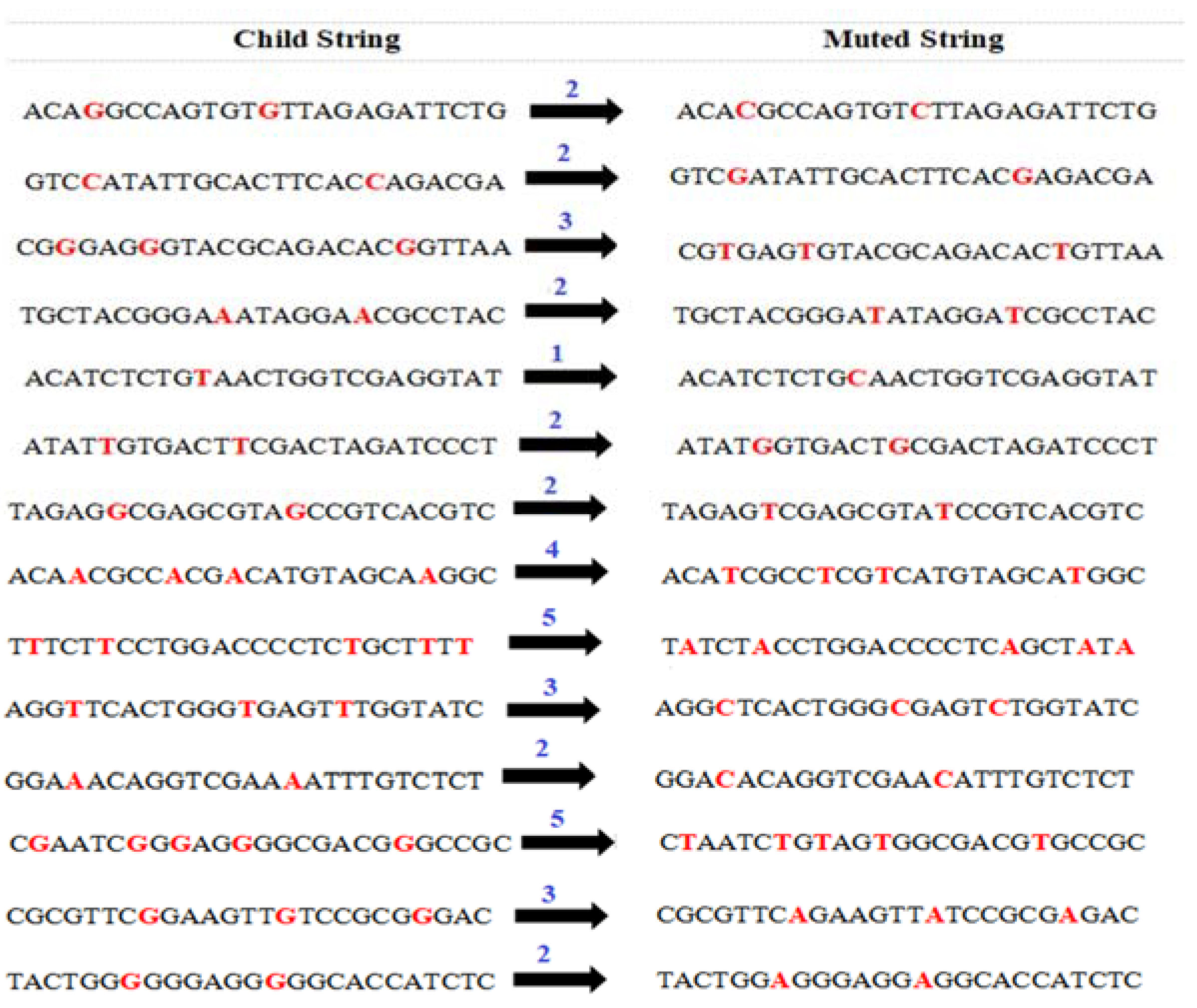
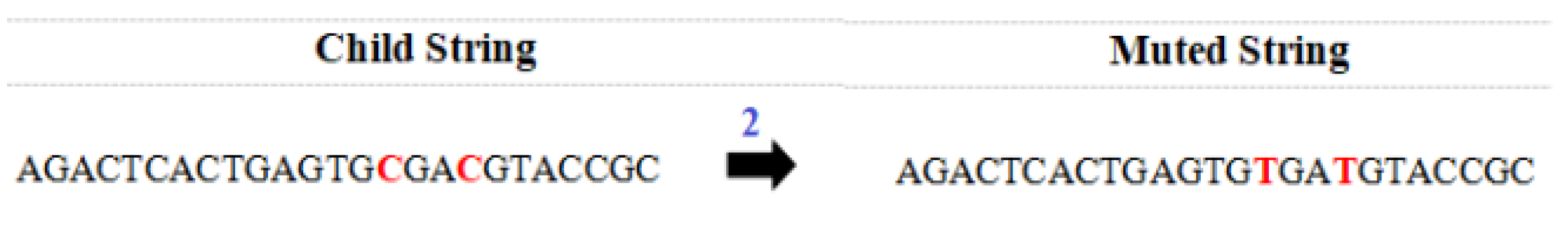
4.6. Perform Mutation Operation
4.7. Generate the New Population
4.8. Reapply Fitness Test and Repeat the Entire Process
5. Analysis of Proposed Methodology
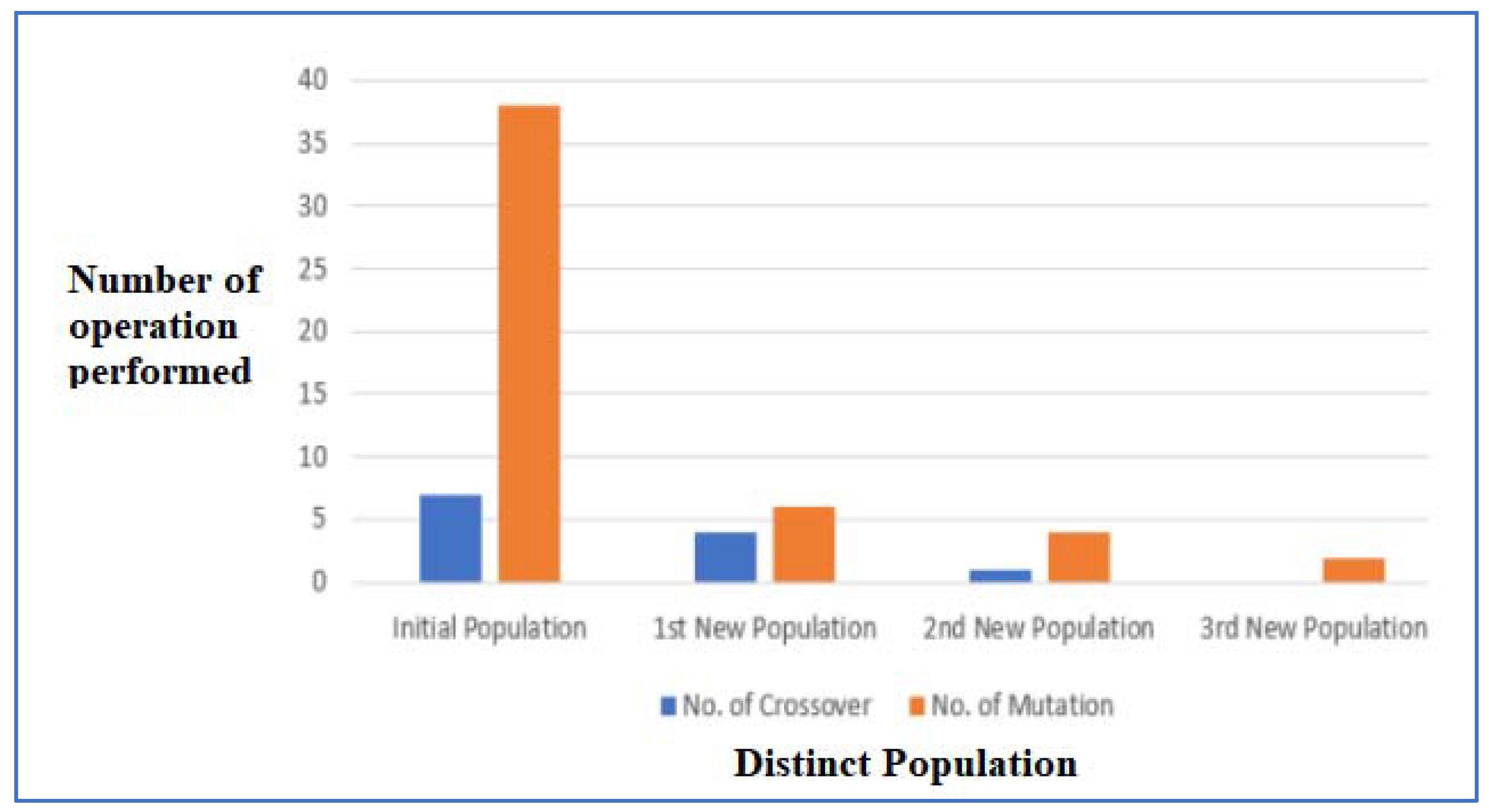
5.1. Number of Crossover and Mutation
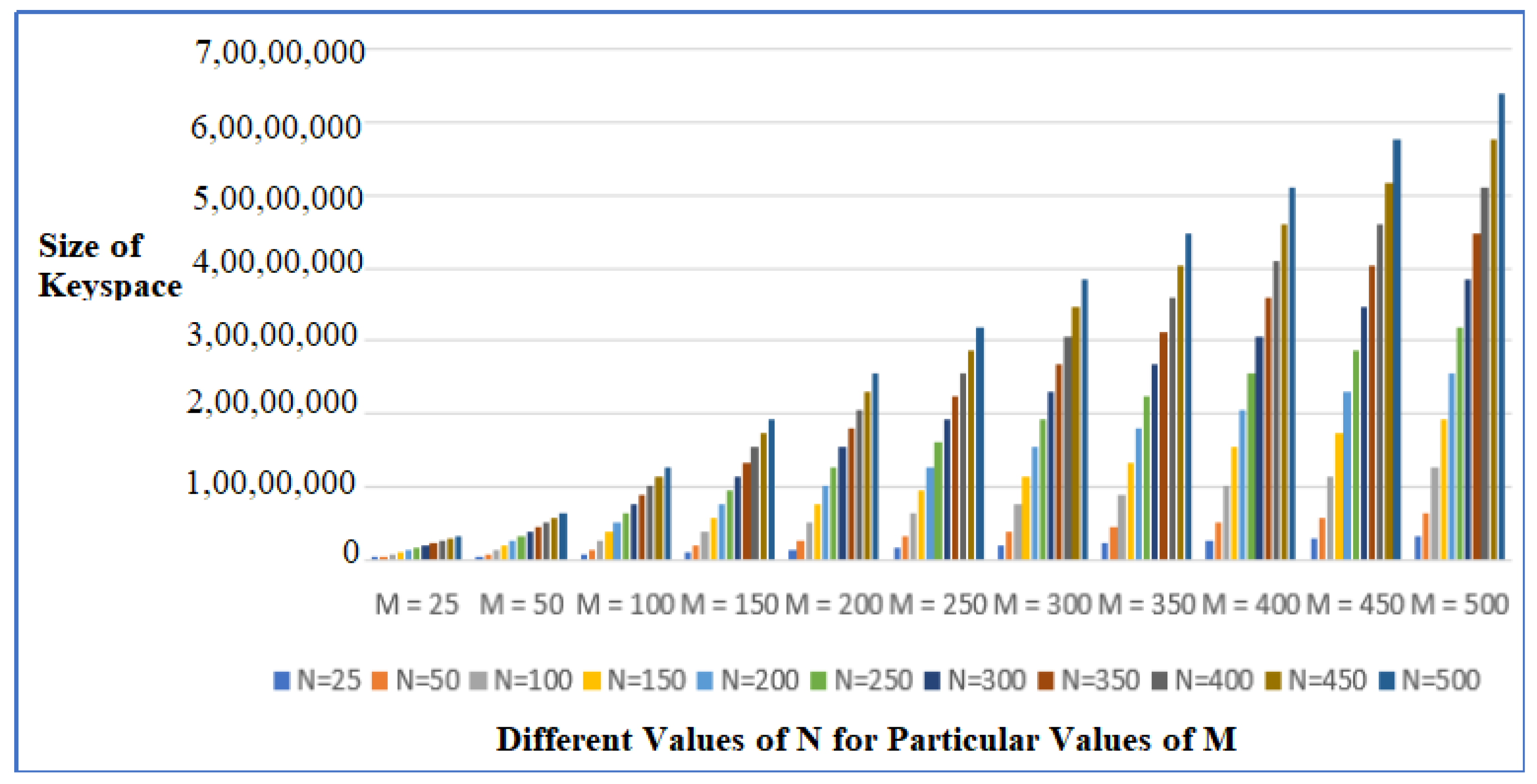
5.2. Effect of Different Values of N and M on the Number of Weak Keys Achieved
5.3. The Comparison of Number of Populations Generated to Strengthen Weak Keys Using Proposed Algorithm
5.4. Immunity to Security Attacks
5.5. Complexity Analysis
5.6. Practical Application of Proposed Scheme
6. Conclusions and Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gehani, A.; LaBean, T.; Reif, J. DNA-Based Cryptography. In Aspects of Molecular Computing; Springer: Berlin/Heidelberg, Germany, 2003; pp. 167–188. [Google Scholar]
- Xiao, G.; Lu, M.; Qin, L.; Lai, X. New field of cryptography: DNA cryptography. Chin. Sci. Bull. 2006, 51, 1413–1420. [Google Scholar] [CrossRef]
- Borda, M.; Tornea, O. DNA secret writing techniques. In Proceedings of the 2010 8th International Conference on Communications, Bucharest, Romania, 10–12 June 2010; IEEE: Piscataway, NJ, USA, 2010; pp. 451–456. [Google Scholar]
- Nandy, N.; Banerjee, D.; Pradhan, C. Color image encryption using DNA based cryptography. Int. J. Inf. Technol. 2018, 13, 533–540. [Google Scholar] [CrossRef]
- Cherian, A.; Raj, S.R.; Abraham, A. A survey on different DNA cryptographic methods. Int. J. Sci. Res. 2013, 2, 167–169. [Google Scholar]
- Pramanik, S.; Setua, S.K. DNA cryptography. In Proceedings of the 2012 7th International Conference on Electrical and Computer Engineering, Dhaka, Bangladesh, 20–22 December 2012; IEEE: Piscataway, NJ, USA, 2012; pp. 551–554. [Google Scholar]
- Katoch, S.; Chauhan, S.S.; Kumar, V. A review on genetic algorithm: Past, present, and future. Multimed. Tools Appl. 2021, 80, 8091–8126. [Google Scholar] [CrossRef]
- Mirjalili, S. Genetic algorithm. In Evolutionary Algorithms and Neural Networks; Springer: Cham, Switzerland, 2019; pp. 43–55. [Google Scholar]
- Bottaci, L. A genetic algorithm fitness function for mutation testing. In Proceedings of the SEMINALL-Workshop at the 23rd International Conference on Software Engineering, Toronto, ON, Canada, 12–19 May 2001. [Google Scholar]
- Poon, P.W.; Carter, J.N. Genetic algorithm crossover operators for ordering applications. Comput. Oper. Res. 1995, 22, 135–147. [Google Scholar] [CrossRef]
- Razali, N.M.; Geraghty, J. Genetic algorithm performance with different selection strategies in solving TSP. In Proceedings of the World Congress on Engineering, London, UK, 6–8 July 2011; International Association of Engineers: Hong Kong, China, 2011; Volume 2, pp. 1–6. [Google Scholar]
- Syswerda, G. Simulated crossover in genetic algorithms. In Foundations of Genetic Algorithms; Elsevier: Amsterdam, The Netherlands, 1993; Volume 2, pp. 239–255. [Google Scholar]
- De Falco, I.; Della Cioppa, A.; Tarantino, E. Mutation-based genetic algorithm: Performance evaluation. Appl. Soft Comput. 2002, 1, 285–299. [Google Scholar] [CrossRef]
- Soni, A.; Agrawal, S. Using genetic algorithm for symmetric key generation in image encryption. Int. J. Adv. Res. Comput. Eng. Technol. IJARCET 2012, 1, 137–140. [Google Scholar]
- Singh, D.; Rani, P.; Kumar, R. To design a genetic algorithm for cryptography to enhance the security. Int. J. Innov. Eng. Technol. 2013, 2, 380–385. [Google Scholar]
- Mishra, S.; Bali, S. Public key cryptography using genetic algorithm. Int. J. RecentTechnol. Eng. 2013, 2, 150–154. [Google Scholar]
- Jhingran, R.; Thada, V.; Dhaka, S. A study on cryptography using genetic algorithm. Int. J. Comput. Appl. 2015, 118, 10–14. [Google Scholar] [CrossRef]
- Malhotra, N.; Nagpal, G. Genetic Symmetric Key Generation for IDEA. JIPS 2015, 11, 239–247. [Google Scholar]
- Jain, A.; Chaudhari, N.S. An improved genetic algorithm for developing deterministic OTP key generator. Complexity 2017, 2017, 1–17. [Google Scholar] [CrossRef]
- Chunka, C.; Goswami, R.S.; Banerjee, S. An efficient mechanism to generate dynamic keys based on genetic algorithm. Secur. Priv. 2018, 4, e37. [Google Scholar] [CrossRef]
- Nazeer, M.I.; Mallah, G.A.; Shaikh, N.A.; Bhatra, R.; Memon, R.A.; Mangrio, M.I. Implication of genetic algorithm in cryptography to enhance security. Int. J. Adv. Comput. Sci. Appl. 2018, 9, 375–379. [Google Scholar] [CrossRef]
- Kalsi, S.; Kaur, H.; Chang, V. DNA cryptography and deep learning using genetic algorithm with NW algorithm for key generation. J. Med. Syst. 2018, 42, 17. [Google Scholar] [CrossRef]
- Turčaník, M.; Javurek, M. Cryptographic Key Generation by Genetic Algorithms. Inf. Secur. 2019, 43, 54–61. [Google Scholar] [CrossRef]
- Vidhya, E.; Rathipriya, R. Key Generation for DNA Cryptography Using Genetic Operators and Diffie-Hellman Key Exchange Algorithm. Comput. Sci. 2020, 15, 1109–1115. [Google Scholar]
- Tahir, M.; Sardaraz, M.; Mehmood, Z.; Muhammad, S. CryptoGA: A cryptosystem based on genetic algorithm for cloud data security. Clust. Comput. 2021, 24, 739–752. [Google Scholar] [CrossRef]
- Abduljabbar, R.B.; Hamid, O.K.; Alhyani, N.J. Features of genetic algorithm for plain text encryption. Int. J. Electr. Comput. Eng. 2021, 11, 434. [Google Scholar] [CrossRef]
- Alhassan, S. Audio Cryptography via Enhanced Genetic Algorithm. Int. J. Multimed. Appl. IJMA 2021, 13, 37–45. [Google Scholar] [CrossRef]
- Garg, D.; Bhatia, K.K.; Gupta, S. A novel Genetic Algorithm based Encryption Technique for Securing Data on Fog Network Using DNA Cryptography. In Proceedings of the 2022 2nd International Conference on Innovative Practices in Technology and Management (ICIPTM), Pradesh, India, 23–25 February 2022; IEEE: Piscataway, NJ, USA, 2022; Volume 2, pp. 362–368. [Google Scholar]
- Hussein, A.A.; Ayoob, N.K. Key Generation for Vigenere Ciphering Based on Genetic Algorithm. J. Univ. Babylon Pure Appl. Sci. 2022, 30, 200–208. [Google Scholar] [CrossRef]
- Shivani, S.; Patel, S.C.; Arora, V.; Sharma, B.; Jolfaei, A.; Srivastava, G. Real-time cheating immune secret sharing for remote sensing images. J. Real-Time Image Process. 2021, 18, 1493–1508. [Google Scholar] [CrossRef]
- Garg, H.; Sharma, B.; Shekhar, S.; Agarwal, R. Spoofing detection system for e-health digital twin using EfficientNet Convolution Neural Network. Multimed. Tools Appl. 2022, 81, 26873–26888. [Google Scholar] [CrossRef]
- Shekhar, S.; Garg, H.; Agrawal, R.; Shivani, S.; Sharma, B. Hatred and trolling detection transliteration framework using hierarchical LSTM in code-mixed social media text. Complex Intell. Syst. 2021, 1–14. [Google Scholar] [CrossRef]
- Gupta, M.; Patel, R.B.; Jain, S.; Garg, H.; Sharma, B. Lightweight branched blockchain security framework for Internet of Vehicles. Trans. Emerg. Telecommun. Technol. 2022, e4520. [Google Scholar] [CrossRef]
- Gupta, M.; Kumar, R.; Shekhar, S.; Sharma, B.; Patel, R.B.; Jain, S.; Dhaou, I.B.; Iwendi, C. Game Theory-Based Authentication Framework to Secure Internet of Vehicles with Blockchain. Sensors 2022, 22, 5119. [Google Scholar] [CrossRef]
- Agarwal, R.; Jalal, A.S.; Arya, K.V. Enhanced Binary Hexagonal Extrema Pattern (EBHXEP) Descriptor for Iris Liveness Detection. Wirel. Pers. Commun. 2020, 115, 2627–2643. [Google Scholar] [CrossRef]
- Agarwal, R.; Jalal, A.S.; Arya, K.V. Local binary hexagonal extrema pattern (LBHXEP): A new feature descriptor for fake iris detection. Vis. Comput. 2021, 37, 1357–1368. [Google Scholar] [CrossRef]
- Agarwal, R.; Jalal, A.S.; Arya, K.V. A review on presentation attack detection system for fake fingerprint. Mod. Phys. Lett. B 2020, 34, 2030001. [Google Scholar] [CrossRef]
- Pavithran, P.; Mathew, S.; Namasudra, S.; Srivastava, G. A novel cryptosystem based on DNA cryptography, hyperchaotic systems and a randomly generated Moore machine for cyber physical systems. Comput. Commun. 2022, 188, 1–12. [Google Scholar] [CrossRef]
- Rupa, C.; Harshita, M.; Srivastava, G.; Gadekallu, T.R.; Maddikunta, P.K. Securing Multimedia using a Deep Learning based Chaotic Logistic Map. IEEEJ. Biomed. HealthInform. 2022. [Google Scholar] [CrossRef] [PubMed]




| Author Name | Type of Cryptosystem | Genetic Operators Used | Fitness Test Applied | Whether GA-Applied on Complete Initial Key Population |
|---|---|---|---|---|
| Soni et al. (2012) | Traditional | Selection Crossover Mutation | Nil | Yes |
| Singh et al. (2013) | Traditional | Crossover | Nil | Yes |
| Mishra et al. (2013) | Traditional | Selection Crossover Mutation | Pearson’s Coefficient of auto-correlation | Yes |
| Jhingran et al. (2015) | Traditional | Selection Crossover Mutation | Nil | Yes |
| Malhotra et al. (2015) | Traditional | Selection Crossover Mutation | Comparing with parents | No |
| Jain et al. (2017) | Traditional | Selection Crossover Mutation | Frequency Test. Serial Test, Autocorrelation Test, Poker Test | Yes |
| Chunka et al. (2018) | Traditional | Selection Crossover Mutation | Frequency test, Block frequency, Runs test, Cumulative sums forward, Cumulative sums backward | Yes |
| Nazeer et al. (2018) | Traditional | Selection Crossover Mutation | Shannon Key Entropy | Yes |
| Kalsi et al. (2018) | DNA | Selection Crossover Mutation | Run Test and Needleman- Wunsch Algorithm | Yes |
| Turčaník et al. (2019) | Traditional | Selection Crossover Mutation | Frequency Test | Yes |
| Vidhya et al. (2020) | DNA | Selection Crossover Mutation | Shanon Key Entropy | Yes |
| Tahir et al. (2021) | Traditional | Selection Crossover Mutation | Shanon key Entropy | Yes |
| Abduljabbar et al. (2021) | Traditional | Selection Crossover Mutation | Nil | Yes |
| Salamudeen et al. (2021) | Audio | Bits fission Switching Mutation Fusion Deconditioning | Fission-Fusion Scheme | Yes |
| Garg et al. (2022) | DNA | Crossover Mutation | NA | Yes |
| Hussein et al. (2022) | Traditional | Crossover Mutation | Entropy Test | Yes |
| Weak Key | a | t | c | g | σA | σT | σC | σG | λ1 |
|---|---|---|---|---|---|---|---|---|---|
| AGGTTCACTGGGCCCCTCTGCTTTT | 2 | 9 | 8 | 6 | 1.069 | 0.802 | 0.535 | 0 | 0.6015 |
| TGCTACGGGAAACAGACACGGTTAA | 9 | 4 | 5 | 7 | 0.802 | 0.535 | 0.266 | 0.266 | 0.4673 |
| TACTGGGGGGAGTTGTCCGCGGGAC | 3 | 5 | 5 | 12 | 0.802 | 0.266 | 0.266 | 1.603 | 0.7342 |
| ACATCTCTGTAACGACTAGATCCCT | 7 | 7 | 8 | 3 | 0.266 | 0.266 | 0.535 | 0.802 | 0.4673 |
| ACAACGCCACGATAGCCGTCACGTC | 7 | 3 | 10 | 5 | 0.266 | 0.802 | 1.069 | 0.266 | 0.6008 |
| ACAGGCCAGTGTCTTCACCAGACGA | 7 | 4 | 8 | 6 | 0.266 | 0.535 | 0.535 | 0 | 0.3340 |
| ATATTGTGACTTCTGGTCGAGGTAT | 5 | 10 | 3 | 6 | 0.266 | 1.069 | 0.802 | 0 | 0.5343 |
| TTTCTTCCTGGATGAGTTTGGTATC | 3 | 12 | 4 | 6 | 0.802 | 1.603 | 0.535 | 0 | 0.7350 |
| CGGGAGGGTACGTAGGAACGCCTAC | 6 | 3 | 6 | 9 | 0 | 0.802 | 0 | 0.802 | 0.4010 |
| TAGAGGCGAGCGCATGTAGCAAGGC | 7 | 3 | 5 | 9 | 0.266 | 0.802 | 0.266 | 0.802 | 0.5340 |
| GGAAACAGGTCGGGCGACGGGCCGC | 5 | 1 | 7 | 12 | 0.266 | 1.336 | 0.266 | 1.603 | 0.8677 |
| GTCCATATTGCAGTTAGAGATTCTG | 6 | 9 | 4 | 6 | 0 | 0.802 | 0.535 | 0 | 0.3343 |
| CGCGTTCGGAAGGGGGCACCATCTC | 4 | 4 | 8 | 9 | 0.535 | 0.535 | 0.535 | 0.802 | 0.6018 |
| CGAATCGGGAGGAAAATTTGTCTCT | 7 | 7 | 4 | 7 | 0.266 | 0.266 | 0.535 | 0.266 | 0.3332 |
| Weak Key | λ2 |
|---|---|
| AGGTTCACTGGGCCCCTCTGCTTTT | 1 |
| TGCTACGGGAAACAGACACGGTTAA | 0 |
| TACTGGGGGGAGTTGTCCGCGGGAC | 1 |
| ACATCTCTGTAACGACTAGATCCCT | 0 |
| ACAACGCCACGATAGCCGTCACGTC | 0 |
| ACAGGCCAGTGTCTTCACCAGACGA | 0 |
| ATATTGTGACTTCTGGTCGAGGTAT | 0 |
| TTTCTTCCTGGATGAGTTTGGTATC | 0 |
| CGGGAGGGTACGTAGGAACGCCTAC | 0 |
| TAGAGGCGAGCGCATGTAGCAAGGC | 0 |
| GGAAACAGGTCGGGCGACGGGCCGC | 0 |
| GTCCATATTGCAGTTAGAGATTCTG | 0 |
| CGCGTTCGGAAGGGGGCACCATCTC | 1 |
| CGAATCGGGAGGAAAATTTGTCTCT | 1 |
| Weak Key | λ1 | λ2 | λ | F |
|---|---|---|---|---|
| AGGTTCACTGGGCCCCTCTGCTTTT | 0.6015 | 1 | 1.6015 | 0.1868 |
| TGCTACGGGAAACAGACACGGTTAA | 0.4673 | 0 | 0.4673 | 0.3852 |
| TACTGGGGGGAGTTGTCCGCGGGAC | 0.7342 | 1 | 1.7342 | 0.1500 |
| ACATCTCTGTAACGACTAGATCCCT | 0.4673 | 0 | 0.4673 | 0.3852 |
| ACAACGCCACGATAGCCGTCACGTC | 0.6008 | 0 | 0.6008 | 0.3541 |
| ACAGGCCAGTGTCTTCACCAGACGA | 0.3340 | 0 | 0.3340 | 0.4181 |
| ATATTGTGACTTCTGGTCGAGGTAT | 0.5343 | 0 | 0.5343 | 0.3695 |
| TTTCTTCCTGGATGAGTTTGGTATC | 0.7350 | 0 | 0.7350 | 0.3241 |
| CGGGAGGGTACGTAGGAACGCCTAC | 0.4010 | 0 | 0.4010 | 0.4011 |
| TAGAGGCGAGCGCATGTAGCAAGGC | 0.5340 | 0 | 0.5340 | 0.3695 |
| GGAAACAGGTCGGGCGACGGGCCGC | 0.8677 | 0 | 0.8677 | 0.2958 |
| GTCCATATTGCAGTTAGAGATTCTG | 0.3343 | 0 | 0.3343 | 0.4171 |
| CGCGTTCGGAAGGGGGCACCATCTC | 0.6018 | 1 | 1.6018 | 0.1677 |
| CGAATCGGGAGGAAAATTTGTCTCT | 0.3332 | 1 | 1.3332 | 0.2113 |
| Weak Key | F |
|---|---|
| ACAGGCCAGTGTCTTCACCAGACGA | 0.4181 |
| GTCCATATTGCAGTTAGAGATTCTG | 0.4171 |
| CGGGAGGGTACGTAGGAACGCCTAC | 0.4011 |
| TGCTACGGGAAACAGACACGGTTAA | 0.3852 |
| ACATCTCTGTAACGACTAGATCCCT | 0.3852 |
| ATATTGTGACTTCTGGTCGAGGTAT | 0.3695 |
| TAGAGGCGAGCGCATGTAGCAAGGC | 0.3695 |
| ACAACGCCACGATAGCCGTCACGTC | 0.3541 |
| TTTCTTCCTGGATGAGTTTGGTATC | 0.3241 |
| AGGTTCACTGGGCCCCTCTGCTTTT | 0.1868 |
| GGAAACAGGTCGGGCGACGGGCCGC | 0.2958 |
| CGAATCGGGAGGAAAATTTGTCTCT | 0.2113 |
| CGCGTTCGGAAGGGGGCACCATCTC | 0.1677 |
| TACTGGGGGGAGTTGTCCGCGGGAC | 0.1500 |
| Child String | a | t | c | g | i | m |
|---|---|---|---|---|---|---|
| ACAGGCCAGTGTGTTAGAGATTCTG | 6 | 7 | 4 | 8 | 6 | 2 |
| GTCCATATTGCACTTCACCAGACGA | 7 | 6 | 8 | 4 | 6 | 2 |
| CGGGAGGGTACGCAGACACGGTTAA | 7 | 3 | 5 | 10 | 6 | 3 |
| TGCTACGGGAAATAGGAACGCCTAC | 8 | 4 | 6 | 7 | 6 | 2 |
| ACATCTCTGTAACTGGTCGAGGTAT | 6 | 8 | 5 | 6 | 6 | 1 |
| ATATTGTGACTTCGACTAGATCCCT | 6 | 9 | 6 | 4 | 6 | 2 |
| TAGAGGCGAGCGTAGCCGTCACGTC | 5 | 4 | 7 | 9 | 6 | 2 |
| ACAACGCCACGACATGTAGCAAGGC | 9 | 2 | 8 | 6 | 6 | 4 |
| TTTCTTCCTGGACCCCTCTGCTTTT | 1 | 12 | 9 | 3 | 6 | 5 |
| AGGTTCACTGGGTGAGTTTGGTATC | 4 | 9 | 3 | 9 | 6 | 3 |
| GGAAACAGGTCGAAAATTTGTCTCT | 8 | 7 | 4 | 6 | 6 | 2 |
| CGAATCGGGAGGGGCGACGGGCCGC | 4 | 1 | 7 | 13 | 6 | 5 |
| CGCGTTCGGAAGTTGTCCGCGGGAC | 3 | 5 | 7 | 10 | 6 | 3 |
| TACTGGGGGGAGGGGGCACCATCTC | 4 | 4 | 6 | 11 | 6 | 2 |
| Weak Key | a | t | c | g | σA | σT | σC | σG | λ1 | λ2 | λ | F |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TATCTACCTGGACCCCTCAGCTATA | 6 | 7 | 9 | 3 | 0 | 0.266 | 0.802 | 0.802 | 0.4675 | 1 | 1.4675 | 0.1873 |
| AGGCTCACTGGGCGAGTCTGGTATC | 4 | 6 | 6 | 9 | 0.535 | 0 | 0 | 0.802 | 0.3342 | 0 | 0.3342 | 0.4172 |
| CTAATCTGTAGTGGCGACGTGCCGC | 4 | 6 | 7 | 8 | 0.535 | 0 | 0.266 | 0.535 | 0.3340 | 0 | 0.3340 | 0.4172 |
| TACTGGAGGGAGGAGGCACCATCTC | 6 | 4 | 6 | 9 | 0 | 0.535 | 0 | 0.802 | 0.3342 | 0 | 0.3342 | 0.4172 |
| Child String | a | t | c | g | i | m |
|---|---|---|---|---|---|---|
| TATCTACCTGGACGAGTCTGGTATC | 5 | 8 | 6 | 6 | 6 | 1 |
| AGGCTCACTGGGCCCCTCAGCTATA | 5 | 5 | 9 | 6 | 6 | 1 |
| CTAATCTGTAGTGAGGCACCATCTC | 6 | 7 | 7 | 5 | 6 | 1 |
| TACTGGAGGGAGGGCGACGTGCCGC | 4 | 3 | 6 | 12 | 6 | 3 |
| Child String | a | t | c | g | i | m |
|---|---|---|---|---|---|---|
| AGGCTCACTGGGTGCGACGTGCCGC | 3 | 4 | 8 | 10 | 6 | 3 |
| TACTGTAGTGAGCTCCTCAGCTATA | 6 | 8 | 6 | 5 | 6 | 1 |
| Child String | a | t | c | g | i | m |
|---|---|---|---|---|---|---|
| AGACTCACTGAGTGCGACGTACCGC | 6 | 4 | 8 | 7 | 6 | 2 |
| M = 25 | M = 50 | M = 100 | M = 150 | M = 200 | M = 250 | M = 300 | M = 350 | M = 400 | M = 450 | M = 500 | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| N= 25 | 14 | 22 | 24 | 24 | 24 | 25 | 25 | 25 | 25 | 25 | 25 |
| N= 50 | 22 | 38 | 44 | 49 | 49 | 49 | 50 | 50 | 50 | 50 | 50 |
| N= 100 | 34 | 87 | 94 | 97 | 98 | 98 | 99 | 99 | 100 | 100 | 100 |
| N= 150 | 60 | 89 | 95 | 112 | 139 | 146 | 147 | 148 | 149 | 150 | 150 |
| N= 200 | 79 | 98 | 126 | 157 | 164 | 179 | 198 | 198 | 199 | 200 | 200 |
| N= 250 | 89 | 99 | 135 | 168 | 173 | 191 | 240 | 248 | 249 | 250 | 250 |
| N= 300 | 107 | 115 | 142 | 196 | 248 | 289 | 291 | 298 | 299 | 299 | 300 |
| N= 350 | 137 | 141 | 156 | 198 | 249 | 324 | 335 | 340 | 350 | 350 | 350 |
| N= 400 | 148 | 159 | 175 | 180 | 237 | 329 | 367 | 384 | 400 | 400 | 400 |
| N= 450 | 158 | 173 | 192 | 226 | 290 | 316 | 384 | 437 | 450 | 450 | 450 |
| N= 500 | 173 | 213 | 246 | 287 | 314 | 384 | 453 | 488 | 497 | 500 | 500 |
| M = 25 | M = 50 | M = 100 | M = 150 | M = 200 | M = 250 | M = 300 | M = 350 | M = 400 | M = 450 | M = 500 | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| N= 25 | 4 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 |
| N= 50 | 5 | 5 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 |
| N= 100 | 5 | 7 | 7 | 7 | 7 | 7 | 7 | 7 | 7 | 7 | 7 |
| N= 150 | 6 | 6 | 7 | 7 | 7 | 7 | 7 | 7 | 7 | 7 | 7 |
| N= 200 | 6 | 7 | 7 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 250 | 7 | 7 | 7 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 300 | 7 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 350 | 7 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 400 | 7 | 7 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 450 | 7 | 7 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
| N= 500 | 7 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 | 8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mukherjee, P.; Garg, H.; Pradhan, C.; Ghosh, S.; Chowdhury, S.; Srivastava, G. Best Fit DNA-Based Cryptographic Keys: The Genetic Algorithm Approach. Sensors 2022, 22, 7332. https://doi.org/10.3390/s22197332
Mukherjee P, Garg H, Pradhan C, Ghosh S, Chowdhury S, Srivastava G. Best Fit DNA-Based Cryptographic Keys: The Genetic Algorithm Approach. Sensors. 2022; 22(19):7332. https://doi.org/10.3390/s22197332
Chicago/Turabian StyleMukherjee, Pratyusa, Hitendra Garg, Chittaranjan Pradhan, Soumik Ghosh, Subrata Chowdhury, and Gautam Srivastava. 2022. "Best Fit DNA-Based Cryptographic Keys: The Genetic Algorithm Approach" Sensors 22, no. 19: 7332. https://doi.org/10.3390/s22197332
APA StyleMukherjee, P., Garg, H., Pradhan, C., Ghosh, S., Chowdhury, S., & Srivastava, G. (2022). Best Fit DNA-Based Cryptographic Keys: The Genetic Algorithm Approach. Sensors, 22(19), 7332. https://doi.org/10.3390/s22197332







