Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network
Abstract
1. Introduction
2. Materials and Methods
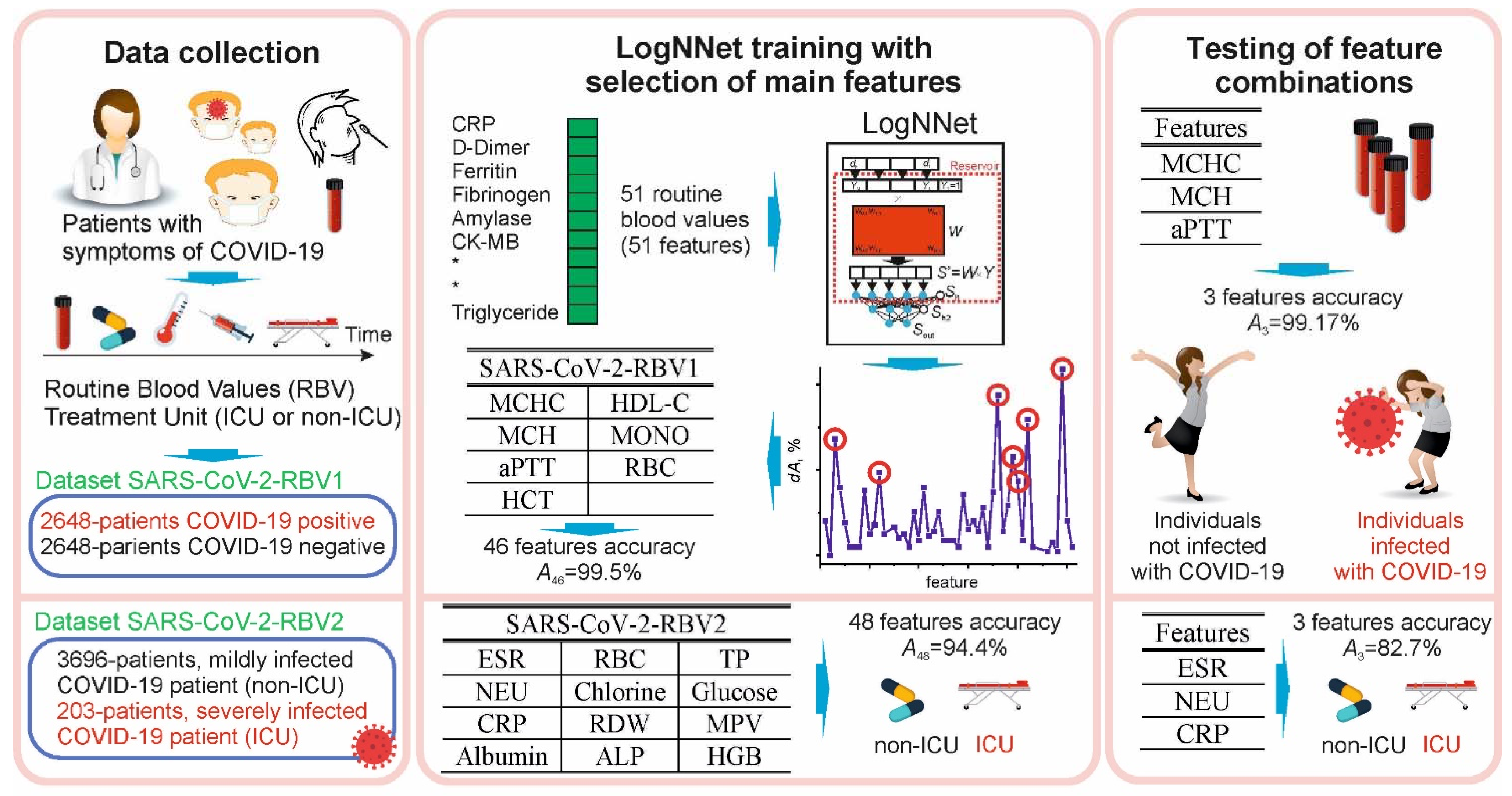
2.1. Characteristic of Participants, Workflow and Define Datasets
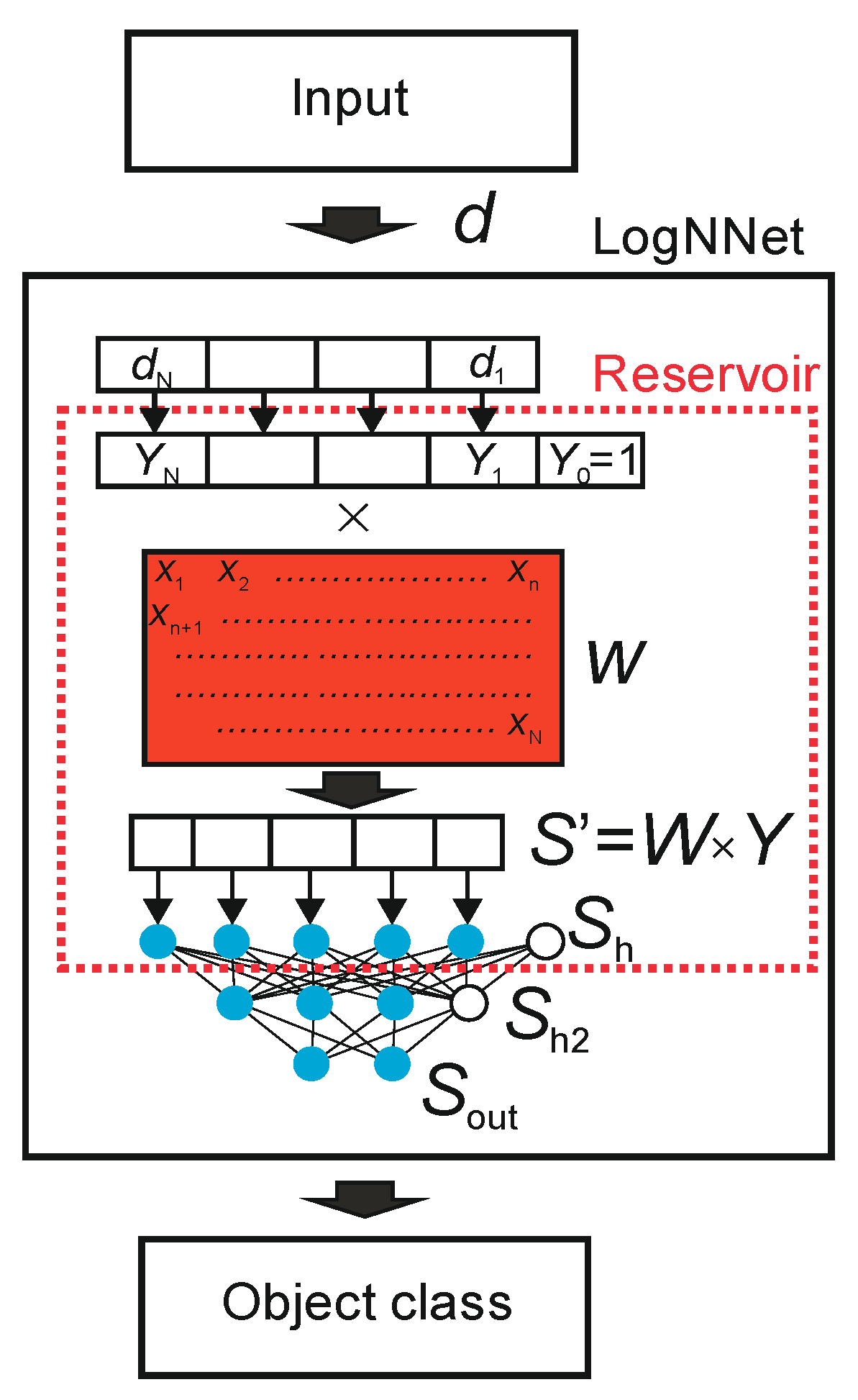
2.2. LogNNet Architecture
| Algorithm 1. Algorithm of matrix W filling. |
| xn: = C; |
| for j: = 1 to P do |
| for i: = 0 to N do |
| begin |
| xn: = (D−K * xn) mod L; // Congruential generator formula |
| W [i,j]: = xn/L; |
| end; |
2.3. Optimization of Reservoir Parameters
2.4. Classification Accuracy, K-Fold Cross-Validation and Balancing Techniques
2.5. Threshold Approach
2.6. Feature Selection Method
3. Results
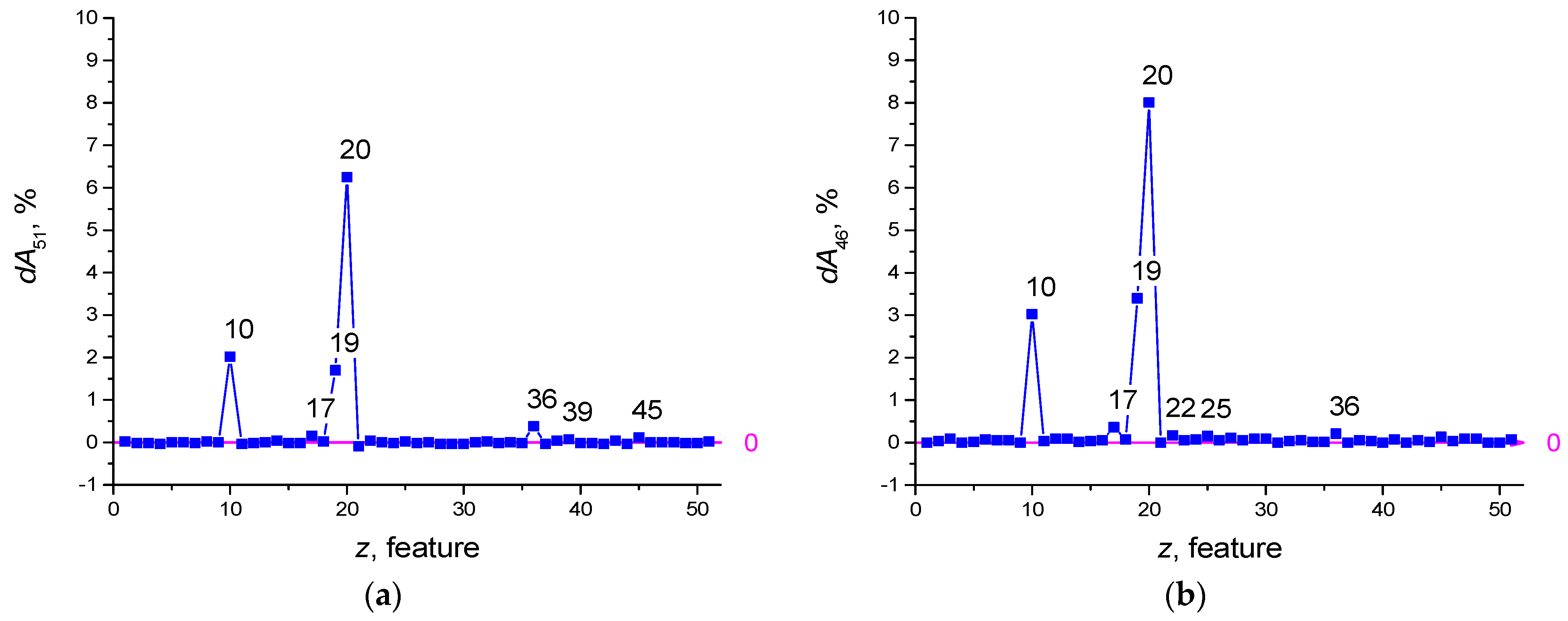
3.1. Dataset SARS-CoV-2-RBV1
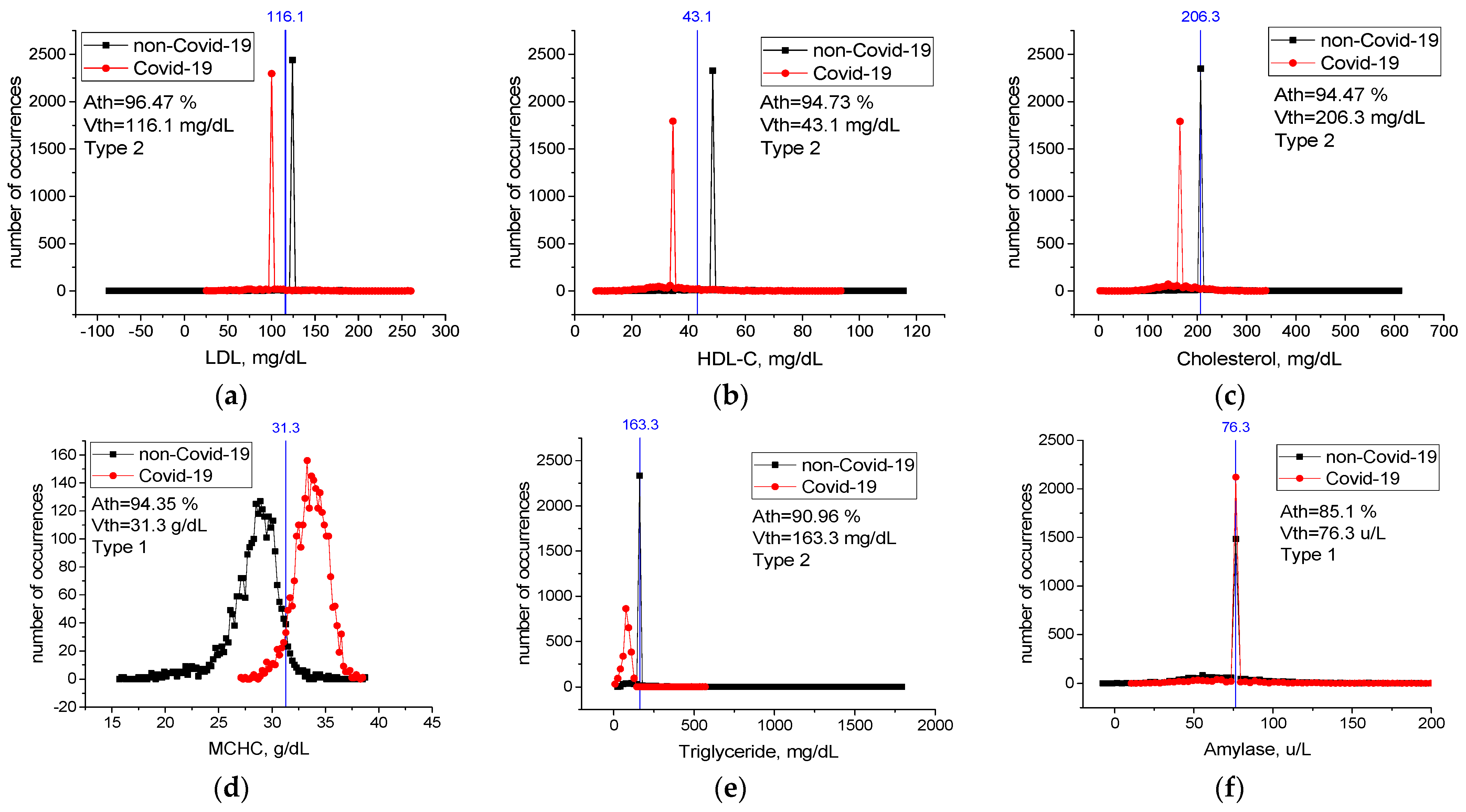
Threshold Accuracy on One Feature
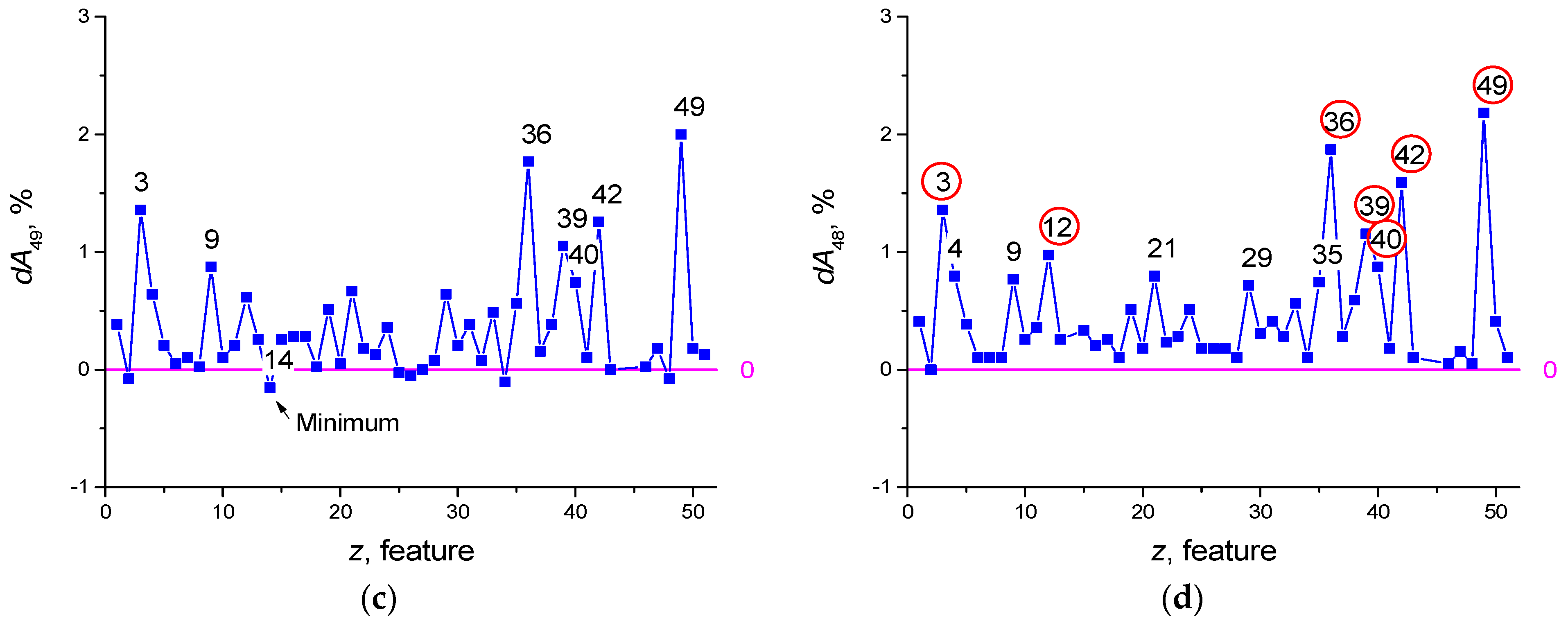
3.2. Dataset SARS-CoV-2-RBV2
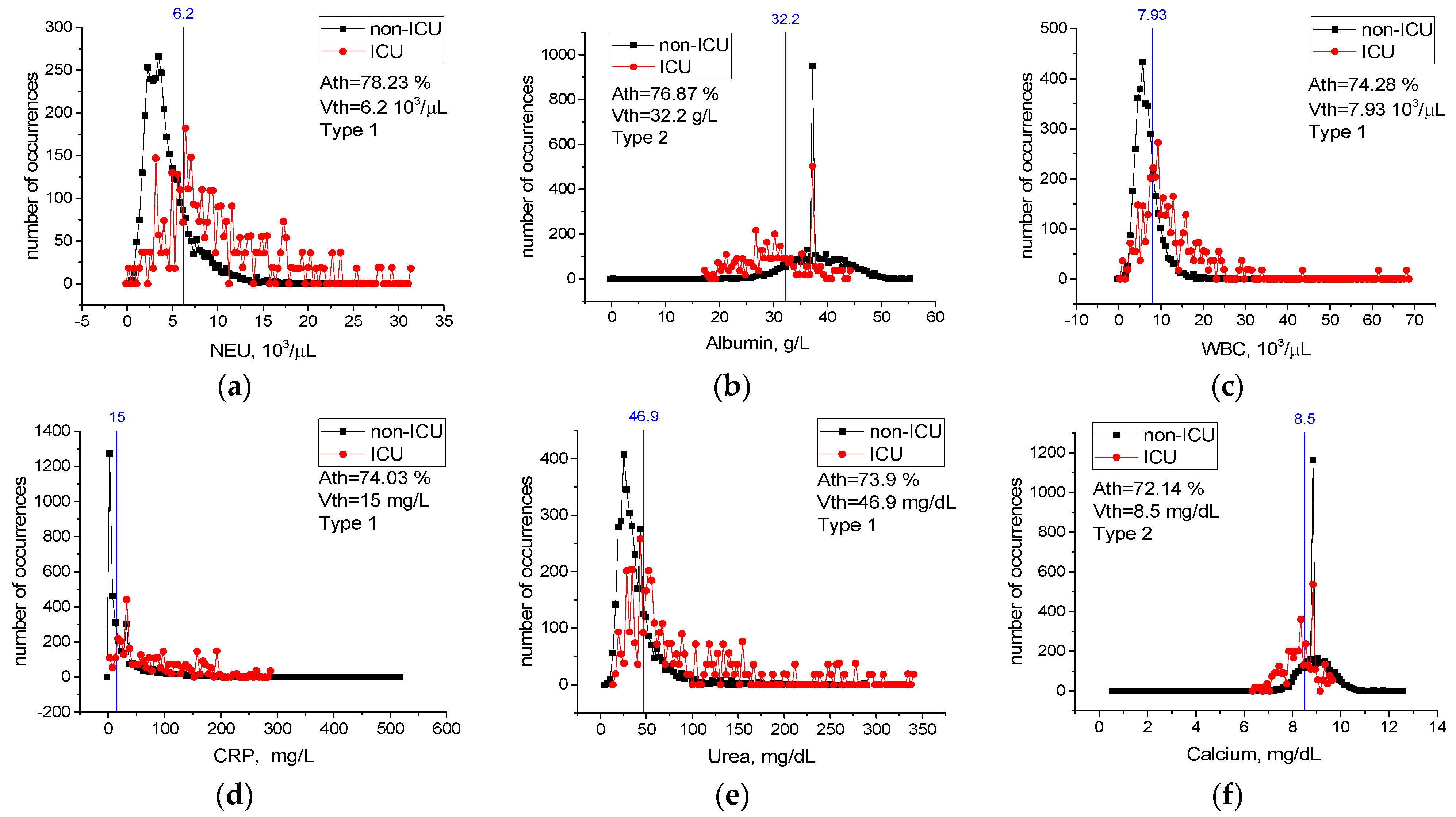
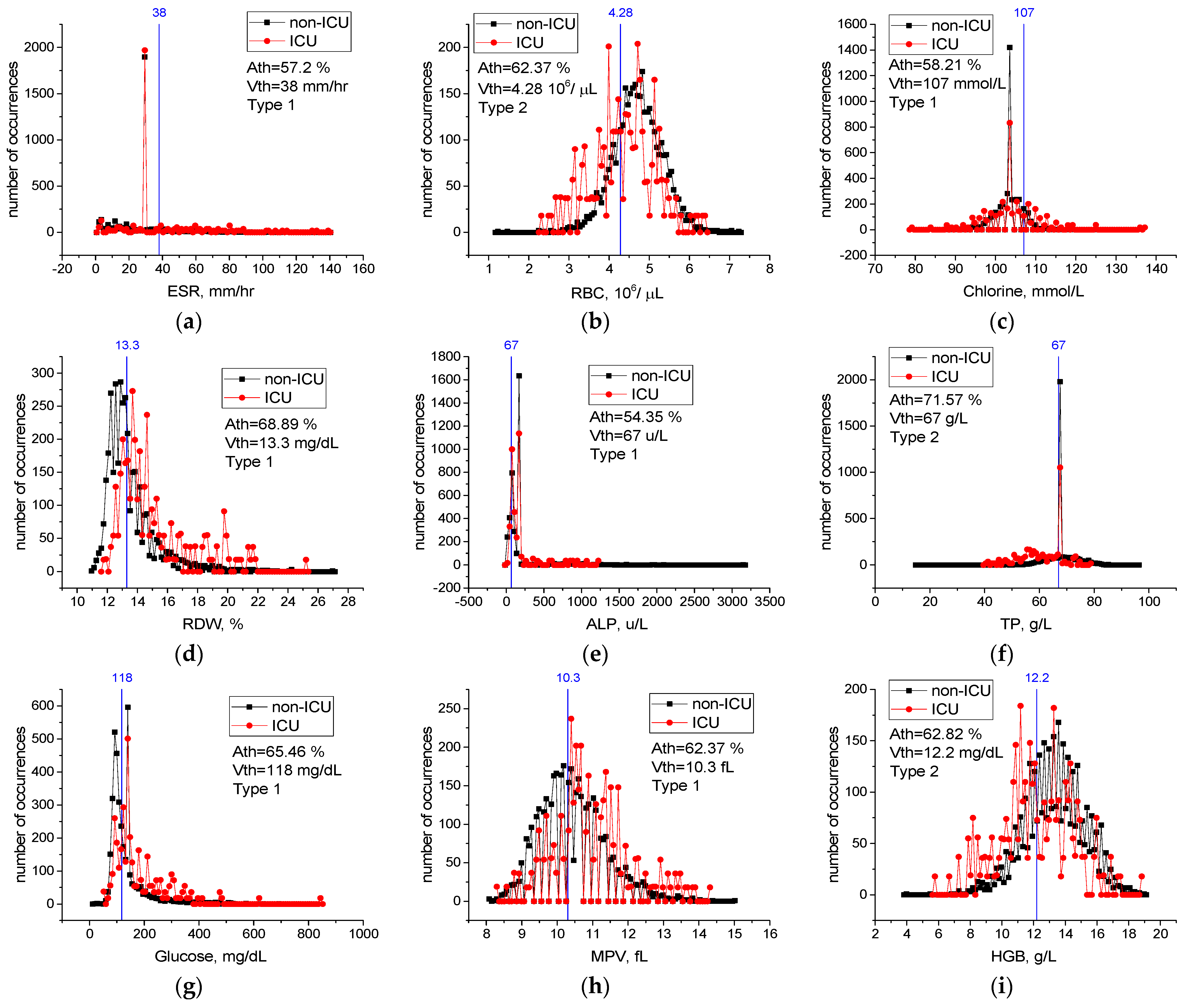
Threshold Accuracy on One Feature
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
Appendix A
| № | Feature | Ath, % | Vth | Units | Type | Min | Max | Bin Size |
|---|---|---|---|---|---|---|---|---|
| 43 | LDL | 96.47 | 116.14 | mg/dL | 2 | −83 | 258 | 3.4 |
| 36 | HDL-C | 94.73 | 43.09 | mg/dL | 2 | 8 | 115 | 1 |
| 39 | Cholesterol | 94.47 | 206.33 | mg/dL | 2 | 5 | 606 | 6 |
| 20 | MCHC | 94.35 | 31.31 | g/dL | 1 | 15.9 | 38.6 | 0.2 |
| 48 | Triglyceride | 90.96 | 163.35 | mg/dL | 2 | 34 | 1782 | 17 |
| 31 | Amylase | 85.1 | 76.35 | u/L | 1 | 0 | 1193 | 3 |
| 51 | UA | 81.12 | 5.39 | mg/dL | 1 | 0 | 14.3 | |
| 47 | TP | 79.68 | 68.05 | g/L | 2 | 15 | 96 | |
| 32 | CK-MB | 78.91 | 19.87 | u/L | 2 | 0 | 685.5 | |
| 42 | LDH | 74.98 | 258.40 | u/L | 1 | 0 | 2749 | |
| 29 | Albumin | 74.91 | 39.61 | g/L | 2 | 0 | 55.87 | |
| 37 | Calcium | 74.21 | 9.01 | mg/dL | 2 | 0 | 12.55 | |
| 30 | ALP | 74.13 | 154.35 | u/L | 1 | 0 | 3150 | |
| 38 | Chlorine | 72.62 | 103.47 | mmol/L | 2 | 79 | 345 | |
| 34 | GGT | 71.6 | 35.51 | u/L | 1 | 0 | 2732 | |
| 1 | CRP | 70.54 | 4.29 | mg/L | 1 | 1 | 1650 | |
| 41 | CK | 70.47 | 111.96 | u/L | 2 | 0 | 4665 | |
| 45 | Sodium | 69.24 | 139.02 | mmol/L | 1 | 108 | 175 | |
| 3 | Ferritin | 68.75 | 49.69 | μg/L | 1 | 0.2 | 1650 | |
| 46 | T-Bil | 68.52 | 0.58 | mg/dL | 2 | −0.35 | 20.95 | |
| 33 | D-Bil | 66.09 | 0.16 | mg/dL | 2 | −0.06 | 20 | |
| 11 | LYM | 66.01 | 1.50 | 103/μL | 2 | 0.08 | 715 | |
| 40 | Creatinine | 64.03 | 1.01 | mg/dL | 1 | 0 | 202 | |
| 7 | PCT | 63.22 | 0.12 | ng/mL | 1 | 0.12 | 1500 | |
| 4 | Fibrinogen | 63.18 | 307.94 | mg/dL | 2 | 10.9 | 668.07 | |
| 35 | Glucose | 62.42 | 122.05 | mg/dL | 1 | 11 | 846 | |
| 49 | eGFR | 61.48 | 87.22 | no unıt | 2 | 3.483 | 561.746 | |
| 27 | ALT | 61.35 | 29.54 | u/L | 1 | 0 | 2110 | |
| 28 | AST | 60.65 | 32.19 | u/L | 1 | 0 | 2927 | |
| 2 | D-Dimer | 60.37 | 385.41 | μg/L | 2 | 1.06 | 9610 | |
| 50 | Urea | 58.19 | 40.99 | mg/dL | 1 | 0 | 427 | |
| 14 | WBC | 58.08 | 5.71 | 103/μL | 2 | 0.4 | 127 | |
| 13 | PLT | 57.46 | 200.26 | 103/μL | 2 | 9 | 768 | |
| 8 | ESR | 57.38 | 14.07 | mm/hr | 1 | 2 | 124 | |
| 16 | EOS | 56.4 | 0 | 103/μL | 1 | 0 | 4.41 | |
| 21 | MCV | 56.25 | 84.03 | fL | 1 | 56.7 | 122.1 | |
| 22 | MONO | 56.25 | 0.54 | 103/μL | 2 | 0.03 | 6.4 | 0.06 |
| 44 | Potassium | 55.63 | 4.36 | mmol/L | 1 | 0 | 59 | |
| 26 | RDW | 55.49 | 13.21 | % | 2 | 0 | 30.8 | |
| 15 | BASO | 55.04 | 0.029 | 103/μL | 2 | 0 | 0.38 | |
| 17 | HCT | 55 | 38.33 | % | 1 | 11.4 | 60.1 | 60 |
| 10 | aPTT | 56.51 | 31.06 | Sec | 1 | 12 | 23,843.7 | 238 |
| 12 | NEU | 54.8 | 2.60 | 103/μL | 2 | 0.49 | 66.43 | |
| 18 | HGB | 54.12 | 12.31 | g/L | 1 | 3.7 | 19 | |
| 5 | INR | 53.15 | 0.735 | no unit | 2 | 0.12 | 88 | |
| 25 | RBC | 53 | 4.29 | 106/μL | 1 | 1.24 | 7.48 | 0.06 |
| 19 | MCH | 52.66 | 28.51 | pg | 1 | 15.9 | 41.9 | 0.2 |
| 24 | PDW | 51.93 | 11.89 | fL | 1 | 0 | 25.3 | |
| 23 | MPV | 51.79 | 9.81 | fL | 1 | 0 | 15 | |
| 6 | PT | 51.79 | 13.09 | Sec | 1 | 2 | 181 | |
| 9 | Troponin | 50.19 | 25 | ng/L | 1 | 0.01 | 25,000 |
| № | Feature | Ath, % | Vth | Units | Type | Min | Max | Bin Size |
|---|---|---|---|---|---|---|---|---|
| 36 | NEU | 78.23 | 6.20 | 103/μL | 1 | 0.1 | 31.26 | 0.3 |
| 3 | Albumin | 76.87 | 32.20 | g/L | 2 | 0.08 | 55 | 0.5 |
| 41 | WBC | 74.28 | 7.93 | 103/μL | 1 | 0.4 | 68.3 | 0.6 |
| 42 | CRP | 74.03 | 15.051 | mg/L | 1 | 0.15 | 514 | 5 |
| 24 | Urea | 73.92 | 46.95 | mg/dL | 1 | 6 | 339 | 3 |
| 11 | Calcium | 72.14 | 8.50 | mg/dL | 2 | 0.6 | 12.43 | 0.1 |
| 21 | TP | 71.57 | 67.00 | g/L | 2 | 15 | 96 | 0.8 |
| 30 | LYM | 71.48 | 1.02 | 103/μL | 2 | 0.08 | 58.87 | |
| 40 | RDW | 68.89 | 13.30 | % | 1 | 11 | 27 | 0.16 |
| 48 | PCT | 67.85 | 0.151 | ng/mL | 1 | 0.052 | 100 | |
| 2 | AST | 66.39 | 44.92 | u/L | 1 | 4 | 2927 | |
| 16 | LDH | 66.11 | 267.37 | u/L | 1 | 20 | 1547 | |
| 9 | Glucose | 65.46 | 118.13 | mg/dL | 1 | 17 | 846 | 8 |
| 7 | D-Bil | 65.04 | 0.209 | mg/dL | 1 | 0.01 | 20 | |
| 44 | Ferritin | 64.17 | 238.116 | μg/L | 1 | 2.4 | 2000 | |
| 15 | CK | 63.66 | 99.92 | u/L | 1 | 2 | 4665 | |
| 43 | D-Dimer | 63.61 | 1074 | μg/L | 1 | 1.06 | 37,000 | |
| 29 | HGB | 62.82 | 12.20 | g/L | 2 | 4 | 19 | 0.15 |
| 47 | PT | 62.78 | 14.30 | Sec | 1 | 9.4 | 129 | |
| 23 | eGFR | 62.55 | 80.47 | no unıt | 2 | 4.724 | 561.746 | |
| 35 | MPV | 62.37 | 10.30 | fL | 1 | 8.1 | 15 | 0.07 |
| 39 | RBC | 62.37 | 4.28 | 106/μL | 2 | 1.24 | 7.22 | 0.06 |
| 50 | Troponin | 61.86 | 10.19 | ng/L | 1 | 1 | 4600 | |
| 20 | T-Bil | 61.81 | 0.58 | mg/dL | 1 | 0.01 | 29 | |
| 8 | GGT | 61.41 | 57.36 | u/L | 1 | 1 | 1085 | |
| 19 | Sodium | 61.01 | 145 | mmol/L | 1 | 112 | 175 | |
| 37 | PDW | 60.86 | 11.51 | fL | 1 | 7.6 | 25.3 | |
| 32 | MCHC | 60.72 | 32.11 | g/dL | 2 | 3.6 | 39.2 | |
| 28 | HCT | 59.71 | 36.63 | % | 2 | 12 | 56.3 | |
| 1 | ALT | 59.02 | 39.80 | u/L | 1 | 0.7 | 1349 | |
| 33 | MCV | 58.79 | 85.93 | fL | 1 | 55.8 | 117.8 | |
| 6 | CK-MB | 58.72 | 19.38 | u/L | 1 | 1 | 575.4 | |
| 14 | Creatinine | 58.39 | 1.26 | mg/dL | 1 | 0.46 | 202 | |
| 12 | Chlorine | 58.21 | 107 | mmol/L | 1 | 79 | 137 | 0.58 |
| 45 | Fibrinogen | 57.22 | 334 | mg/dL | 1 | 70.56 | 681.88 | |
| 49 | ESR | 57.2 | 38.03 | mm/hr | 1 | 2 | 139 | 1.37 |
| 5 | Amylase | 56.46 | 75.7 | 103/μL | 2 | 11 | 874 | |
| 46 | INR | 56.38 | 1.42 | no unit | 1 | 0.77 | 110 | |
| 51 | aPTT | 56.33 | 36.12 | Sec | 2 | 12 | 414 | |
| 25 | UA | 55.92 | 5.412 | mg/dL | 1 | 0.9 | 15 | |
| 38 | PLT | 55.61 | 160 | % | 2 | 5 | 1199 | |
| 34 | MONO | 55.22 | 0.474 | sec | 2 | 0.03 | 6.29 | |
| 18 | Potassium | 54.99 | 3.815 | mmol/L | 2 | 2.4 | 59 | |
| 27 | EOS | 54.72 | 0.111 | 103/Μl | 2 | 0.01 | 4.41 | |
| 4 | ALP | 54.35 | 63.98 | u/L | 1 | 1 | 3150 | 31 |
| 22 | Triglyceride | 53.27 | 141.6 | 106/μL | 1 | 32 | 1402 | |
| 31 | MCH | 53.11 | 28.22 | pg | 2 | 15.6 | 41.9 | |
| 13 | Cholesterol | 53.11 | 170 | mg/dL | 2 | 5 | 354 | |
| 10 | HDL-C | 53.02 | 34.69 | mg/dL | 2 | 8 | 93 | |
| 26 | BASO | 52.75 | 0.01 | 103/μL | 1 | 0.01 | 0.38 | |
| 17 | LDL | 51.26 | 115.1 | mg/dL | 1 | 15 | 258 |
References
- Mertoglu, C.; Huyut, M.; Olmez, H.; Tosun, M.; Kantarci, M.; Coban, T. COVID-19 is more dangerous for older people and its severity is increasing: A case-control study. Med. Gas Res. 2022, 12, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Mertoglu, C.; Huyut, M.T.; Arslan, Y.; Ceylan, Y.; Coban, T.A. How do routine laboratory tests change in coronavirus disease 2019? Scand. J. Clin. Lab. Investig. 2021, 81, 24–33. [Google Scholar] [CrossRef]
- Huyut, M.T.; İlkbahar, F. The effectiveness of blood routine parameters and some biomarkers as a potential diagnostic tool in the diagnosis and prognosis of Covid-19 disease. Int. Immunopharmacol. 2021, 98, 107838. [Google Scholar] [CrossRef] [PubMed]
- Huyut, M.T.; Huyut, Z. Forecasting of Oxidant/Antioxidant levels of COVID-19 patients by using Expert models with biomarkers used in the Diagnosis/Prognosis of COVID-19. Int. Immunopharmacol. 2021, 100, 108127. [Google Scholar] [CrossRef]
- Huyut, M.; Üstündaǧ, H. Prediction of diagnosis and prognosis of COVID-19 disease by blood gas parameters using decision trees machine learning model: A retrospective observational study. Med. Gas Res. 2022, 12, 60–66. [Google Scholar] [CrossRef]
- Tahir Huyut, M.; Huyut, Z.; İlkbahar, F.; Mertoğlu, C. What is the impact and efficacy of routine immunological, biochemical and hematological biomarkers as predictors of COVID-19 mortality? Int. Immunopharmacol. 2022, 105, 108542. [Google Scholar] [CrossRef]
- Guan, W.; Ni, Z.; Hu, Y.; Liang, W.; Ou, C.; He, J.; Liu, L.; Shan, H.; Lei, C.; Hui, D.S.C.; et al. Clinical Characteristics of Coronavirus Disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef]
- Banerjee, A.; Ray, S.; Vorselaars, B.; Kitson, J.; Mamalakis, M.; Weeks, S.; Baker, M.; Mackenzie, L.S. Use of Machine Learning and Artificial Intelligence to predict SARS-CoV-2 infection from Full Blood Counts in a population. Int. Immunopharmacol. 2020, 86, 106705. [Google Scholar] [CrossRef]
- Huyut, M.T.; Soygüder, S. The Multi-Relationship Structure between Some Symptoms and Features Seen during the New Coronavirus 19 Infection and the Levels of Anxiety and Depression post-Covid. East. J. Med. 2022, 27, 1–10. [Google Scholar] [CrossRef]
- Amgalan, A.; Othman, M. Hemostatic laboratory derangements in COVID-19 with a focus on platelet count. Platelets 2020, 31, 740–745. [Google Scholar] [CrossRef]
- Li, X.; Wang, L.; Yan, S.; Yang, F.; Xiang, L.; Zhu, J.; Shen, B.; Gong, Z. Clinical characteristics of 25 death cases with COVID-19: A retrospective review of medical records in a single medical center, Wuhan, China. Int. J. Infect. Dis. 2020, 94, 128–132. [Google Scholar] [CrossRef] [PubMed]
- Kukar, M.; Gunčar, G.; Vovko, T.; Podnar, S.; Černelč, P.; Brvar, M.; Zalaznik, M.; Notar, M.; Moškon, S.; Notar, M. COVID-19 diagnosis by routine blood tests using machine learning. Sci. Rep. 2021, 11, 10738. [Google Scholar] [CrossRef]
- Jiang, S.Q.; Huang, Q.F.; Xie, W.M.; Lv, C.; Quan, X.Q. The association between severe COVID-19 and low platelet count: Evidence from 31 observational studies involving 7613 participants. Br. J. Haematol. 2020, 190, e29–e33. [Google Scholar] [CrossRef]
- Zheng, Y.; Zhang, Y.; Chi, H.; Chen, S.; Peng, M.; Luo, L.; Chen, L.; Li, J.; Shen, B.; Wang, D. The hemocyte counts as a potential biomarker for predicting disease progression in COVID-19: A retrospective study. Clin. Chem. Lab. Med. 2020, 58, 1106–1115. [Google Scholar] [CrossRef] [PubMed]
- Lippi, G.; Plebani, M.; Henry, B.M. Thrombocytopenia is associated with severe coronavirus disease 2019 (COVID-19) infections: A meta-analysis. Clin. Chim. Acta 2020, 506, 145–148. [Google Scholar] [CrossRef]
- Mousavi, S.A.; Rad, S.; Rostami, T.; Rostami, M.; Mousavi, S.A.; Mirhoseini, S.A.; Kiumarsi, A. Hematologic predictors of mortality in hospitalized patients with COVID-19: A comparative study. Hematology 2020, 25, 383–388. [Google Scholar] [CrossRef]
- Beck, B.R.; Shin, B.; Choi, Y.; Park, S.; Kang, K. Predicting commercially available antiviral drugs that may act on the novel coronavirus (SARS-CoV-2) through a drug-target interaction deep learning model. Comput. Struct. Biotechnol. J. 2020, 18, 784–790. [Google Scholar] [CrossRef]
- Xu, X.; Jiang, X.; Ma, C.; Du, P.; Li, X.; Lv, S.; Yu, L.; Ni, Q.; Chen, Y.; Su, J.; et al. A Deep Learning System to Screen Novel Coronavirus Disease 2019 Pneumonia. Engineering 2020, 6, 1122–1129. [Google Scholar] [CrossRef]
- Brinati, D.; Campagner, A.; Ferrari, D.; Locatelli, M.; Banfi, G.; Cabitza, F. Detection of COVID-19 Infection from Routine Blood Exams with Machine Learning: A Feasibility Study. J. Med. Syst. 2020, 44, 135. [Google Scholar] [CrossRef]
- Cabitza, F.; Campagner, A.; Ferrari, D.; Di Resta, C.; Ceriotti, D.; Sabetta, E.; Colombini, A.; De Vecchi, E.; Banfi, G.; Locatelli, M.; et al. Development, evaluation, and validation of machine learning models for COVID-19 detection based on routine blood tests. Clin. Chem. Lab. Med. 2021, 59, 421–431. [Google Scholar] [CrossRef]
- Yang, H.S.; Hou, Y.; Vasovic, L.V.; Steel, P.A.D.; Chadburn, A.; Racine-Brzostek, S.E.; Velu, P.; Cushing, M.M.; Loda, M.; Kaushal, R.; et al. Routine Laboratory Blood Tests Predict SARS-CoV-2 Infection Using Machine Learning. Clin. Chem. 2020, 66, 1396–1404. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.P.; Pejaver, V.; Hammarlund, N.E.; Sung, H.; Kyu, S.; Lee, H.; Scott, G.; Gombar, S.; Shah, N.; Shen, S.; et al. Short communication A predictive tool for identi fi cation of SARS-CoV-2 PCR-negative emergency department patients using routine test results. J. Clin. Virol. 2020, 129, 104502. [Google Scholar] [CrossRef] [PubMed]
- Mei, X.; Lee, H.C.; Diao, K.Y.; Huang, M.; Lin, B.; Liu, C.; Xie, Z.; Ma, Y.; Robson, P.M.; Chung, M.; et al. Artificial intelligence–enabled rapid diagnosis of patients with COVID-19. Nat. Med. 2020, 26, 1224–1228. [Google Scholar] [CrossRef]
- Soares, F. A novel specific artificial intelligence-based method to identify COVID-19 cases using simple blood exams. medRxiv 2020. [Google Scholar] [CrossRef]
- Soltan, A.A.; Kouchaki, S.; Zhu, T.; Kiyasseh, D.; Taylor, T.; Hussain, Z.B.; Peto, T.; Brent, A.J.; Eyre, D.W.; Clifton, D. Artificial intelligence driven assessment of routinely collected healthcare data is an effective screening test for COVID-19 in patients presenting to hospital. medRxiv 2020. [Google Scholar] [CrossRef]
- Remeseiro, B.; Bolon-Canedo, V. A review of feature selection methods in medical applications. Comput. Biol. Med. 2019, 112, 103375. [Google Scholar] [CrossRef]
- Bikku, T. Multi-layered deep learning perceptron approach for health risk prediction. J. Big Data 2020, 7, 50. [Google Scholar] [CrossRef]
- Battineni, G.; Chintalapudi, N.; Amenta, F. Machine learning in medicine: Performance calculation of dementia prediction by support vector machines (SVM). Inform. Med. Unlocked 2019, 16, 100200. [Google Scholar] [CrossRef]
- Xing, W.; Bei, Y. Medical Health Big Data Classification Based on KNN Classification Algorithm. IEEE Access 2020, 8, 28808–28819. [Google Scholar] [CrossRef]
- Hoodbhoy, Z.; Noman, M.; Shafique, A.; Nasim, A.; Chowdhury, D.; Hasan, B. Use of machine learning algorithms for prediction of fetal risk using cardiotocographic data. Int. J. Appl. Basic Med. Res. 2019, 9, 226. [Google Scholar] [CrossRef]
- Alam, M.Z.; Rahman, M.S.; Rahman, M.S. A Random Forest based predictor for medical data classification using feature ranking. Inform. Med. Unlocked 2019, 15, 100180. [Google Scholar] [CrossRef]
- Schober, P.; Vetter, T.R. Logistic Regression in Medical Research. Anesth. Analg. 2021, 132, 365–366. [Google Scholar] [CrossRef] [PubMed]
- Podgorelec, V.; Kokol, P.; Stiglic, B.; Rozman, I. Decision trees: An overview and their use in medicine. J. Med. Syst. 2002, 26, 445–463. [Google Scholar] [CrossRef]
- Guyon, I.; Gunn, S.; Nikravesh, M.; Zadeh, L.A. Feature Extraction: Foundations and Applications; Studies in Fuzziness and Soft Computing; Springer: Berlin/Heidelberg, Germany, 2008; ISBN 9783540354888. [Google Scholar]
- Hall, M.A. Correlation-based Feature Selection for Machine Learning. Ph.D. Thesis, Department of Computer Science, The University of Waikato, Hamilton, NewZealand, April 1999; pp. 51–69. [Google Scholar]
- Dash, M.; Liu, H. Consistency-based search in feature selection. Artif. Intell. 2003, 151, 155–176. [Google Scholar] [CrossRef]
- Zhao, Z.; Liu, H. Searching for interacting features. In Proceedings of the 20th International Joint Conference on Artificial Intelligence, Hyderabad, India, 6–12 January 2007; pp. 1156–1161. [Google Scholar]
- Hall, M.A.; Smith, L.A. Practical feature subset selection for machine learning. In Proceedings of the Computer Science ’98, 21st Australasian Computer Science Conference ACSC’98, Perth, Australia, 4–6 February 1998; Volume 20, pp. 181–191. [Google Scholar]
- Kononenko, I. Estimating attributes: Analysis and extensions of RELIEF. In European Conference on Machine Learning; Springer: Berlin/Heidelberg, Germany, 1994; Volume 784, pp. 171–182. [Google Scholar] [CrossRef]
- Le Thi, H.A.; Nguyen, V.V.; Ouchani, S. Gene selection for cancer classification using DCA. In International Conference on Advanced Data Mining and Applications; Springer: Berlin/Heidelberg, Germany, 2008; Volume 5139, pp. 62–72. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression Shrinkage and Selection via the Lasso. J. R. Stat. Soc. Ser. B 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Velichko, A. Neural network for low-memory IoT devices and MNIST image recognition using kernels based on logistic map. Electronics 2020, 9, 1432. [Google Scholar] [CrossRef]
- Velichko, A. A method for medical data analysis using the lognnet for clinical decision support systems and edge computing in healthcare. Sensors 2021, 21, 6209. [Google Scholar] [CrossRef]
- Velichko, A.; Heidari, H. A Method for Estimating the Entropy of Time Series Using Artificial Neural Networks. Entropy 2021, 23, 1432. [Google Scholar] [CrossRef]
- Izotov, Y.A.; Velichko, A.A.; Boriskov, P.P. Method for fast classification of MNIST digits on Arduino UNO board using LogNNet and linear congruential generator. J. Phys. Conf. Ser. 2021, 2094, 32055. [Google Scholar] [CrossRef]
- Heidari, H.; Velichko, A. An improved LogNNet classifier for IoT application. J. Phys. Conf. Ser. 2021, 2094, 032015. [Google Scholar] [CrossRef]
- Mattiuzzi, C.; Lippi, G. Which lessons shall we learn from the 2019 novel coronavirus outbreak? Ann. Transl. Med. 2020, 8, 48. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Kim, D.-M.; Lee, B. Insufficient Sensitivity of RNA Dependent RNA Polymerase Gene of SARS-CoV-2 Viral Genome as Confirmatory Test using Korean COVID-19 Cases. Preprints 2020, 1–4. [Google Scholar] [CrossRef]
- Zhang, J.J.; Cao, Y.Y.; Tan, G.; Dong, X.; Wang, B.C.; Lin, J.; Yan, Y.Q.; Liu, G.H.; Akdis, M.; Akdis, C.A.; et al. Clinical, radiological, and laboratory characteristics and risk factors for severity and mortality of 289 hospitalized COVID-19 patients. Allergy Eur. J. Allergy Clin. Immunol. 2021, 76, 533–550. [Google Scholar] [CrossRef] [PubMed]
- Teymouri, M.; Mollazadeh, S.; Mortazavi, H.; Naderi Ghale-noie, Z.; Keyvani, V.; Aghababaei, F.; Hamblin, M.R.; Abbaszadeh-Goudarzi, G.; Pourghadamyari, H.; Hashemian, S.M.R.; et al. Recent advances and challenges of RT-PCR tests for the diagnosis of COVID-19. Pathol. Res. Pract. 2021, 221, 153443. [Google Scholar] [CrossRef]
- D’Cruz, R.J.; Currier, A.W.; Sampson, V.B. Laboratory Testing Methods for Novel Severe Acute Respiratory Syndrome-Coronavirus-2 (SARS-CoV-2). Front. Cell Dev. Biol. 2020, 8, 468. [Google Scholar] [CrossRef]
- Yang, A.P.; Liu, J.P.; Tao, W.Q.; Li, H.M. The diagnostic and predictive role of NLR, d-NLR and PLR in COVID-19 patients. Int. Immunopharmacol. 2020, 84, 106504. [Google Scholar] [CrossRef]
- Huyut, M.T. Automatic Detection of Severely and Mildly Infected COVID-19 Patients with Supervised Machine Learning Models. IRBM 2022, 1, 1–12. [Google Scholar] [CrossRef]
- Zhang, C.; Shi, L.; Wang, F.S. Liver injury in COVID-19: Management and challenges. Lancet Gastroenterol. Hepatol. 2020, 5, 428–430. [Google Scholar] [CrossRef]
- Zheng, M.; Gao, Y.; Wang, G.; Song, G.; Liu, S.; Sun, D.; Xu, Y.; Tian, Z. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell. Mol. Immunol. 2020, 17, 533–535. [Google Scholar] [CrossRef]
- Luo, J.; Zhou, L.; Feng, Y.; Li, B.; Guo, S. The selection of indicators from initial blood routine test results to improve the accuracy of early prediction of COVID-19 severity. PLoS ONE 2021, 16, e0253329. [Google Scholar] [CrossRef]
- Ma, Y.; Hou, L.; Yang, X.; Huang, Z.; Yang, X.; Zhao, N.; He, M.; Shi, Y.; Kang, Y.; Yue, J.; et al. The association between frailty and severe disease among COVID-19 patients aged over 60 years in China: A prospective cohort study. BMC Med. 2020, 18, 274. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.C.; Shih, T.P.; Ko, W.C.; Tang, H.J.; Hsueh, P.R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int. J. Antimicrob. Agents 2020, 55, 105924. [Google Scholar] [CrossRef] [PubMed]
- Feld, J.; Tremblay, D.; Thibaud, S.; Kessler, A.; Naymagon, L. Ferritin levels in patients with COVID-19: A poor predictor of mortality and hemophagocytic lymphohistiocytosis. Int. J. Lab. Hematol. 2020, 42, 773–779. [Google Scholar] [CrossRef]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Chen, G.; Wu, D.; Guo, W.; Cao, Y.; Huang, D.; Wang, H.; Wang, T.; Zhang, X.; Chen, H.; Yu, H.; et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J. Clin. Investig. 2020, 130, 2620–2629. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Li, H.; Li, L.; Liu, C.; Yan, S.; Chen, H.; Li, Y. Ferritin in the coronavirus disease 2019 (COVID-19): A systematic review and meta-analysis. J. Clin. Lab. Anal. 2020, 34, 1–18. [Google Scholar] [CrossRef]
- Cavalcante-Silva, L.H.A.; Carvalho, D.C.M.; Lima, É.D.A.; Galvão, J.G.; da Silva, J.S.d.F.; de Sales-Neto, J.M.; Rodrigues-Mascarenhas, S. Neutrophils and COVID-19: The road so far. Int. Immunopharmacol. 2021, 90, 107233. [Google Scholar] [CrossRef]
- Pan, F.; Ye, T.; Sun, P.; Gui, S.; Liang, B.; Li, L.; Zheng, D.; Wang, J.; Hesketh, R.L.; Yang, L.; et al. Time Course of Lung Changes on Chest CT During Recovery From 2019 Novel Coronavirus (COVID-19) Pneumonia. Radiology 2020, 295, 200370. [Google Scholar] [CrossRef]
- Zhao, D.; Yao, F.; Wang, L.; Zheng, L.; Gao, Y.; Ye, J.; Guo, F.; Zhao, H.; Gao, R. A Comparative Study on the Clinical Features of Coronavirus 2019 (COVID-19) Pneumonia with Other Pneumonias. Clin. Infect. Dis. 2020, 71, 756–761. [Google Scholar] [CrossRef]
- Rabanser, S.; Günnemann, S.; Lipton, Z.C. Failing loudly: An empirical study of methods for detecting dataset shift. Adv. Neural Inf. Process. Syst. 2019, 32. [Google Scholar] [CrossRef]
- Al-Aubidy, K.M.; Derbas, A.M.; Al-Mutairi, A.W. Real-time patient health monitoring and alarming using wireless-sensor-network. In Proceedings of the 2016 13th International Multi-Conference on Systems, Signals & Devices (SSD), Leipzig, Germany, 21–24 March 2016; pp. 416–423. [Google Scholar] [CrossRef]
- Taiwo, O.; Ezugwu, A.E. Smart healthcare support for remote patient monitoring during Covid-19 quarantine. Inform. Med. Unlocked 2020, 20, 100428. [Google Scholar] [CrossRef] [PubMed]
- Lamonaca, F.; Balestrieri, E.; Tudosa, I.; Picariello, F.; Carnì, D.L.; Scuro, C.; Bonavolontà, F.; Spagnuolo, V.; Grimaldi, G.; Colaprico, A. An Overview on Internet of Medical Things in Blood Pressure Monitoring. In Proceedings of the 2019 IEEE International Symposium on Medical Measurements and Applications (MeMeA), Istanbul, Turkey, 26–28 June 2019; pp. 1–6. [Google Scholar]
- Hu, F.; Xiao, Y.; Hao, Q. Congestion-aware, loss-resilient bio-monitoring sensor networking for mobile health applications. IEEE J. Sel. Areas Commun. 2009, 27, 450–465. [Google Scholar] [CrossRef]
- Vizbaras, A.; Simonyte, I.; Droz, S.; Torcheboeuf, N.; Miasojedovas, A.; Trinkunas, A.; Buciunas, T.; Dambrauskas, Z.; Gulbinas, A.; Boiko, D.L.; et al. GaSb Swept-Wavelength Lasers for Biomedical Sensing Applications. IEEE J. Sel. Top. Quantum Electron. 2019, 25, 1–12. [Google Scholar] [CrossRef]
- Clifton, L.; Clifton, D.A.; Pimentel, M.A.F.; Watkinson, P.J.; Tarassenko, L. Predictive monitoring of mobile patients by combining clinical observations with data from wearable sensors. IEEE J. Biomed. Health Inform. 2014, 18, 722–730. [Google Scholar] [CrossRef]
- Pfeil, J.; Nechyporenko, A.; Frohme, M.; Hufert, F.T.; Schulze, K. Examination of blood samples using deep learning and mobile microscopy. BMC Bioinform. 2022, 23, 1–14. [Google Scholar] [CrossRef]
- Chan, J.; Michaelsen, K.; Estergreen, J.K.; Sabath, D.E.; Gollakota, S. Micro-mechanical blood clot testing using smartphones. Nat. Commun. 2022, 13, 1–12. [Google Scholar] [CrossRef]
- Farooqi, M.H.; Abdelmannan, D.K.; Mubarak, M.; Abdalla, M.; Hamed, A.; Xavier, M.; Joyce, T.; Cadiz, S.; Nawaz, F.A. The Impact of Telemonitoring on Improving Glycemic and Metabolic Control in Previously Lost-to-Follow-Up Patients with Type 2 Diabetes Mellitus: A Single-Center Interventional Study in the United Arab Emirates. Int. J. Clin. Pract. 2022, 2022, 6286574. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Y.; Li, H.; Cao, Y.; Han, S.; Zhang, K.; He, W. Covalent Biosensing Polymer Chain Reaction Enabling Periphery Blood Testing to Predict Tumor Invasiveness with a Platelet Procancerous Protein. Anal. Chem. 2022, 94, 1983–1989. [Google Scholar] [CrossRef]



| № | Feature | № | Feature | № | Feature | № | Feature | № | Feature |
|---|---|---|---|---|---|---|---|---|---|
| 1 | CRP | 12 | NEU | 23 | MPV | 34 | GGT | 45 | Sodium |
| 2 | D-Dimer | 13 | PLT | 24 | PDW | 35 | Glucose | 46 | T-Bil |
| 3 | Ferritin | 14 | WBC | 25 | RBC | 36 | HDL-C | 47 | TP |
| 4 | Fibrinogen | 15 | BASO | 26 | RDW | 37 | Calcium | 48 | Triglyceride |
| 5 | INR | 16 | EOS | 27 | ALT | 38 | Chlorine | 49 | eGFR |
| 6 | PT | 17 | HCT | 28 | AST | 39 | Cholesterol | 50 | Urea |
| 7 | PCT | 18 | HGB | 29 | Albumin | 40 | Creatinine | 51 | UA |
| 8 | ESR | 19 | MCH | 30 | ALP | 41 | CK | ||
| 9 | Troponin | 20 | MCHC | 31 | Amylase | 42 | LDH | ||
| 10 | aPTT | 21 | MCV | 32 | CK-MB | 43 | LDL | ||
| 11 | LYM | 22 | MONO | 33 | D-Bil | 44 | Potassium |
| № | Feature | № | Feature | № | Feature | № | Feature | № | Feature |
|---|---|---|---|---|---|---|---|---|---|
| 1 | ALT | 12 | Chlorine | 23 | eGFR | 34 | MONO | 45 | Fibrinogen |
| 2 | AST | 13 | Cholesterol | 24 | Urea | 35 | MPV | 46 | INR |
| 3 | Albumin | 14 | Creatinine | 25 | UA | 36 | NEU | 47 | PT |
| 4 | ALP | 15 | CK | 26 | BASO | 37 | PDW | 48 | PCT |
| 5 | Amylase | 16 | LDH | 27 | EOS | 38 | PLT | 49 | ESR |
| 6 | CK-MB | 17 | LDL | 28 | HCT | 39 | RBC | 50 | Troponin |
| 7 | D-Bil | 18 | Potassium | 29 | HGB | 40 | RDW | 51 | aPTT |
| 8 | GGT | 19 | Sodium | 30 | LYM | 41 | WBC | ||
| 9 | Glucose | 20 | T-Bil | 31 | MCH | 42 | CRP | ||
| 10 | HDL-C | 21 | TP | 32 | MCHC | 43 | D-Dimer | ||
| 11 | Calcium | 22 | Triglyceride | 33 | MCV | 44 | Ferritin |
| Chaotic Map | List of Optimized Parameters (Limits) | Equation | |
|---|---|---|---|
| Congruent generator | K (−100 to 100) D (−100 to 100) L (2 to 10,000) C (−100 to 100) | (1) |
| Dataset SARS-CoV-2-RBV1 | Dataset SARS-CoV-2-RBV2 | ||||||
|---|---|---|---|---|---|---|---|
| K | D | L | C | K | D | L | C |
| 93 | 68 | 9276 | 73 | 47 | 99 | 8941 | 56 |
| Ep | A46(FR [21,37,40,42,49]) | Precision “Non-COVID-19” | Precision “COVID-19” | Recall “Non-COVID-19” | Recall “COVID-19” | F1 “Non-COVID-19” | F1 “COVID-19” |
|---|---|---|---|---|---|---|---|
| 10 | 98.376 | 0.978 | 0.99 | 0.991 | 0.977 | 0.984 | 0.984 |
| 30 | 99.339 | 0.992 | 0.995 | 0.995 | 0.992 | 0.993 | 0.993 |
| 100 | 99.509 | 0.994 | 0.996 | 0.996 | 0.994 | 0.995 | 0.995 |
| 150 | 99.49 | 0.994 | 0.996 | 0.996 | 0.994 | 0.995 | 0.995 |
| 200 | 99.471 | 0.994 | 0.995 | 0.995 | 0.994 | 0.995 | 0.995 |
| Number | dA46 | Features |
|---|---|---|
| 20 | 8.007 | MCHC |
| 19 | 3.399 | MCH |
| 10 | 3.022 | aPTT |
| 17 | 0.359 | HCT |
| 36 | 0.208 | HDL-C |
| 22 | 0.17 | MONO |
| 25 | 0.151 | RBC |
| Combinations of Features | A | Precision “Non-COVID-19” | Precision “COVID-19” | Recall “Non-COVID-19” | Recall “COVID-19” | F1 “Non-COVID-19” | F1 “COVID-19” |
|---|---|---|---|---|---|---|---|
| A46(FR [21,37,40,42,49]) | 99.509 | 0.994 | 0.996 | 0.996 | 0.994 | 0.995 | 0.995 |
| A7(FS [10,17,19,20,22,25,36]) | 99.358 | 0.991 | 0.996 | 0.996 | 0.991 | 0.994 | 0.994 |
| A1(FS [>20]) | 94.279 | 0.930 | 0.958 | 0.959 | 0.926 | 0.944 | 0.942 |
| A1(FS [>19]) | 52.418 | 0.526 | 0.524 | 0.500 | 0.548 | 0.509 | 0.532 |
| A1(FS [10]) | 52.398 | 0.516 | 0.947 | 0.972 | 0.075 | 0.672 | 0.100 |
| A1(FS [36]) | 94.429 | 0.935 | 0.955 | 0.956 | 0.932 | 0.945 | 0.943 |
| A2(FS [19,20]) | 99.150 | 0.989 | 0.994 | 0.994 | 0.989 | 0.992 | 0.991 |
| A2(FS [20,36]) | 97.583 | 0.973 | 0.979 | 0.979 | 0.972 | 0.976 | 0.976 |
| A2(FS [19,36]) | 94.373 | 0.934 | 0.955 | 0.957 | 0.931 | 0.945 | 0.943 |
| A3(FS [10,19,20]) | 99.169 | 0.989 | 0.995 | 0.995 | 0.989 | 0.992 | 0.992 |
| A5(FS [10,17,19,22,25]) | 51.699 | 0.526 | 0.546 | 0.784 | 0.250 | 0.604 | 0.277 |
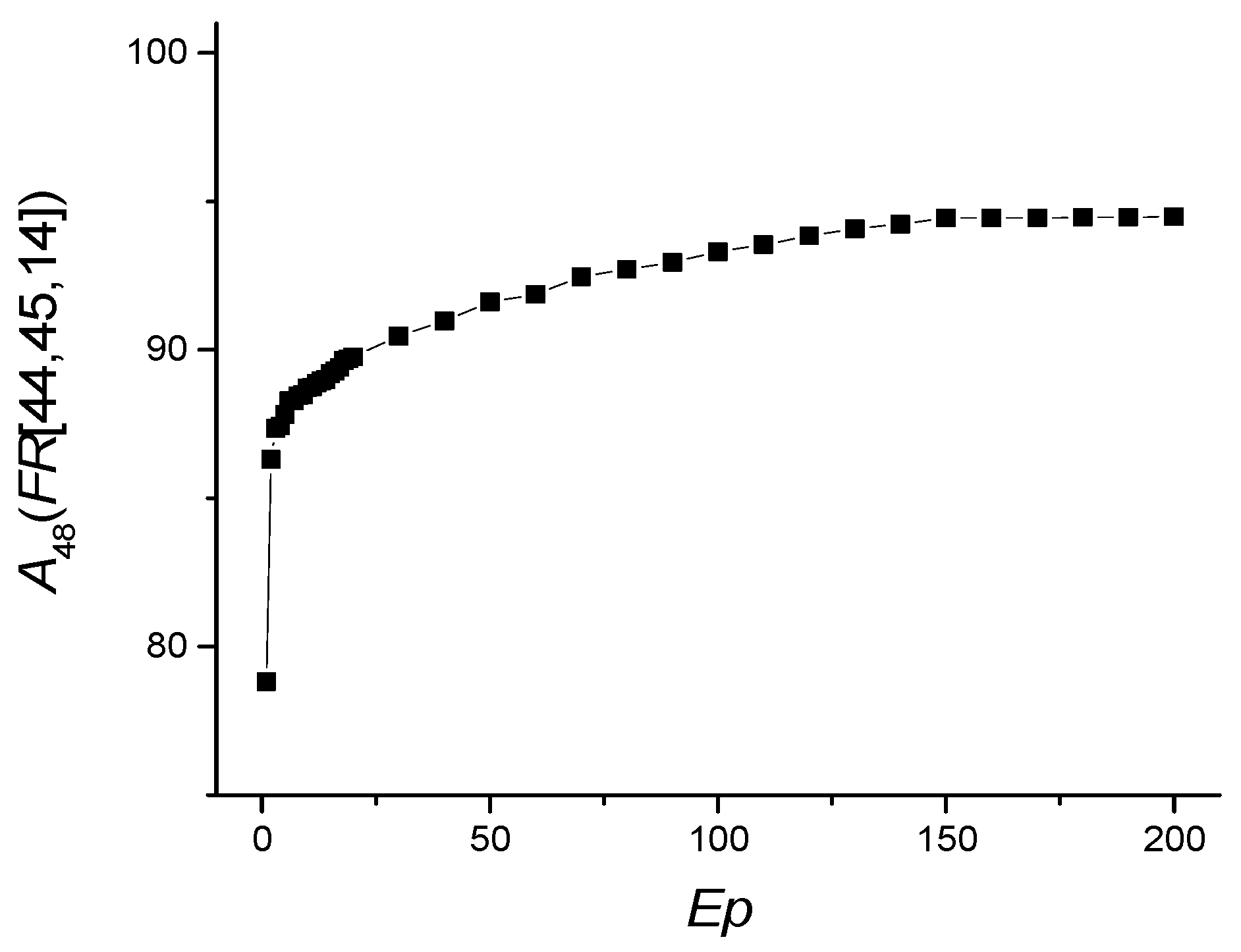
| Ep | A48(FR [14,44,45]) | Precision “Non-ICU” | Precision “ICU” | Recall “Non-ICU” | Recall “ICU” | F1 “Non-ICU” | F2 “ICU” |
|---|---|---|---|---|---|---|---|
| 10 | 88.715 | 0.993 | 0.307 | 0.887 | 0.881 | 0.937 | 0.451 |
| 30 | 90.459 | 0.993 | 0.347 | 0.906 | 0.876 | 0.947 | 0.492 |
| 100 | 93.306 | 0.990 | 0.433 | 0.939 | 0.821 | 0.964 | 0.562 |
| 150 | 94.434 | 0.989 | 0.49 | 0.952 | 0.797 | 0.97 | 0.599 |
| 200 | 94.486 | 0.987 | 0.495 | 0.955 | 0.767 | 0.97 | 0.592 |
| Number | dA48 | Features |
|---|---|---|
| 49 | 2.18 | ESR |
| 36 | 1.872 | NEU |
| 42 | 1.59 | CRP |
| 3 | 1.359 | Albumin |
| 39 | 1.154 | RBC |
| 12 | 0.974 | Chlorine |
| 40 | 0.872 | RDW |
| 4 | 0.795 | ALP |
| 21 | 0.795 | TP |
| 9 | 0.769 | Glucose |
| 35 | 0.744 | MPV |
| 29 | 0.718 | HGB |
| Combinations of Features | A | Precision “Non-ICU” | Precision “ICU” | Recall “Non-ICU” | Recall “ICU” | F1 “Non-ICU” | F1 “ICU” |
|---|---|---|---|---|---|---|---|
| A48(FR [14,44,45]) | 94.434 | 0.989 | 0.49 | 0.952 | 0.797 | 0.97 | 0.599 |
| A12(FS [3,4,9,12,21,29,35,36,39,40,42,49]) | 90.946 | 0.990 | 0.364 | 0.914 | 0.831 | 0.950 | 0.499 |
| A1(FS [49]) | 59.598 | 0.950 | 0.059 | 0.605 | 0.418 | 0.694 | 0.097 |
| A1(FS [49]) | 75.040 | 0.955 | 0.085 | 0.773 | 0.341 | 0.851 | 0.133 |
| A3(FS [36,42,49]) | 82.712 | 0.989 | 0.210 | 0.827 | 0.826 | 0.900 | 0.334 |
| A7(FS [3,12,36,39,40,42,49]) | 89.355 | 0.991 | 0.341 | 0.896 | 0.846 | 0.940 | 0.469 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huyut, M.T.; Velichko, A. Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network. Sensors 2022, 22, 4820. https://doi.org/10.3390/s22134820
Huyut MT, Velichko A. Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network. Sensors. 2022; 22(13):4820. https://doi.org/10.3390/s22134820
Chicago/Turabian StyleHuyut, Mehmet Tahir, and Andrei Velichko. 2022. "Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network" Sensors 22, no. 13: 4820. https://doi.org/10.3390/s22134820
APA StyleHuyut, M. T., & Velichko, A. (2022). Diagnosis and Prognosis of COVID-19 Disease Using Routine Blood Values and LogNNet Neural Network. Sensors, 22(13), 4820. https://doi.org/10.3390/s22134820






