Phybrata Sensors and Machine Learning for Enhanced Neurophysiological Diagnosis and Treatment
Abstract
:1. Introduction
1.1. Challenges in Concussion Diagnosis and Management
1.2. Machine Learning and Wearable Motion Sensors in Concussion Management
1.3. Phybrata Sensing
1.4. Distinguishing Neurological vs. Vestibular Impairments
1.5. Present Study
2. Materials and Methods
2.1. Study Population, Data Collection, Derivation of Phybrata Biomarkers
2.2. Data Preprocessing
- Time-Series Averaging (TSA): For each Eo and Ec patient test phase, the three phybrata time-series signals (x, y, z) and the phybrata power (calculated using the vector sum of the three acceleration components [58,69]) were averaged over one-second time-steps (100 samples per step), reducing the dimensionality of each time series from 2000 samples to 20 samples. Once averaged, the data were either used in their existing form for CNNs or converted such that each time-step represents a column instead of a row for classical ML models. There are two reasons for using this averaging approach as an alternative to using the raw signal. First, the raw data contains 6000 measurements per patient test (100 Hz sampling over 20 s for each of the x, y, and z axes), which presents challenges for training classical ML models, since the number of data features greatly exceeds the number of patients. This excessive number of features can lead to models that overfit and generalize poorly to data from new patients. Second, the computational advantages in using an averaged time-series instead of a full time-series signal recording can enable much faster and lower computational complexity training and classification, allowing the use of remote sensor devices that do not require cloud connectivity for computational support. No frequency features were extracted from the TSA preprocessed data. Further details of the phybrata power calculations and data processing are included in the Appendix A.
- Non-Time-Series (NTS) Feature Extraction: Standard statistical measures (variance, mean, standard deviation, min, max and median) were calculated for each of the three phybrata time-series signals (x, y, and z accelerations) and several additional power and frequency features extracted for both Eo and Ec test phases, including phybrata powers within the physiological-system-specific frequency bands discussed above. To extract the power features, the phybrata power was first calculated at each value in the accelerometer time-series data. The power values were then summed for each respective test phase (e.g., Eo Power and Ec Power) and the powers for the two phases were averaged (e.g., (Ec + Eo)/2). Phybrata signal PSD curves were also calculated using Welch’s method [78], and these PSD curves were then used to calculate phybrata powers within specific frequency bands. PSD variations within specific spectral bands, as well as correlated PSD variations across multiple spectral bands, were shown to help quantify the sensory reweighting that often accompanies many neurophysiological impairments [58] and may thus also serve as useful ML classification features. A more detailed description of Welch’s method is included in the Appendix A.
2.3. Modeling
- The performance of four different ML models (SVM, RF, XGB, CNN) was assessed using standard open-source implementations [79,80,81]. Model training, testing, and validation were carried using a standard leave-one-out K-fold cross-validation procedure [36,82,83,84,85], in which the dataset was first randomly split into a training set (80%) and a test (20%) set. Validation datasets were then generated by further dividing the training dataset into K subsets, or “folds”, where each fold is a group of test subjects, and each of the K folds is used once as a validation dataset (“leave one out”) while the remaining K-1 folds are combined together as the training dataset. This procedure guarantees that every test subject will be in a validation set exactly once and in a training set K-1 times. The error estimate is averaged over all K trials to derive the performance of each model. As is common practice, we use K = 5 to balance bias and variance of test error estimates [85]. Cross-validation was applied multiple times for different values of the hyperparameters, and the parameters that optimized each model were selected by maximizing the concussion classification F1-score across each of the selected validation folds (F1 ± 2 standard deviations). In this manner, cross-validation addresses the problem of overfitting [82], since cross-validated models that perform well over the test data and give good accuracy have not overfitted the training data and can be used for prediction. The hyperparameters that optimized each model are listed in Appendix A.
- The classification performance of the four different ML models was ranked based on the F1 scores when applied to the testing set. The F1-score represents a balanced approach for conveying a model’s performance in terms of its correct and incorrect classifications. Specifically, F1 weighs both false negatives (FN) and false positives (FP) in conveying a model’s accuracy and is prioritized for ranking the performance of the current ML models for both binary (Use Case 1) and multiclass (Use Case 2) classification experiments. All metric descriptions and formal calculations are provided in Appendix A.
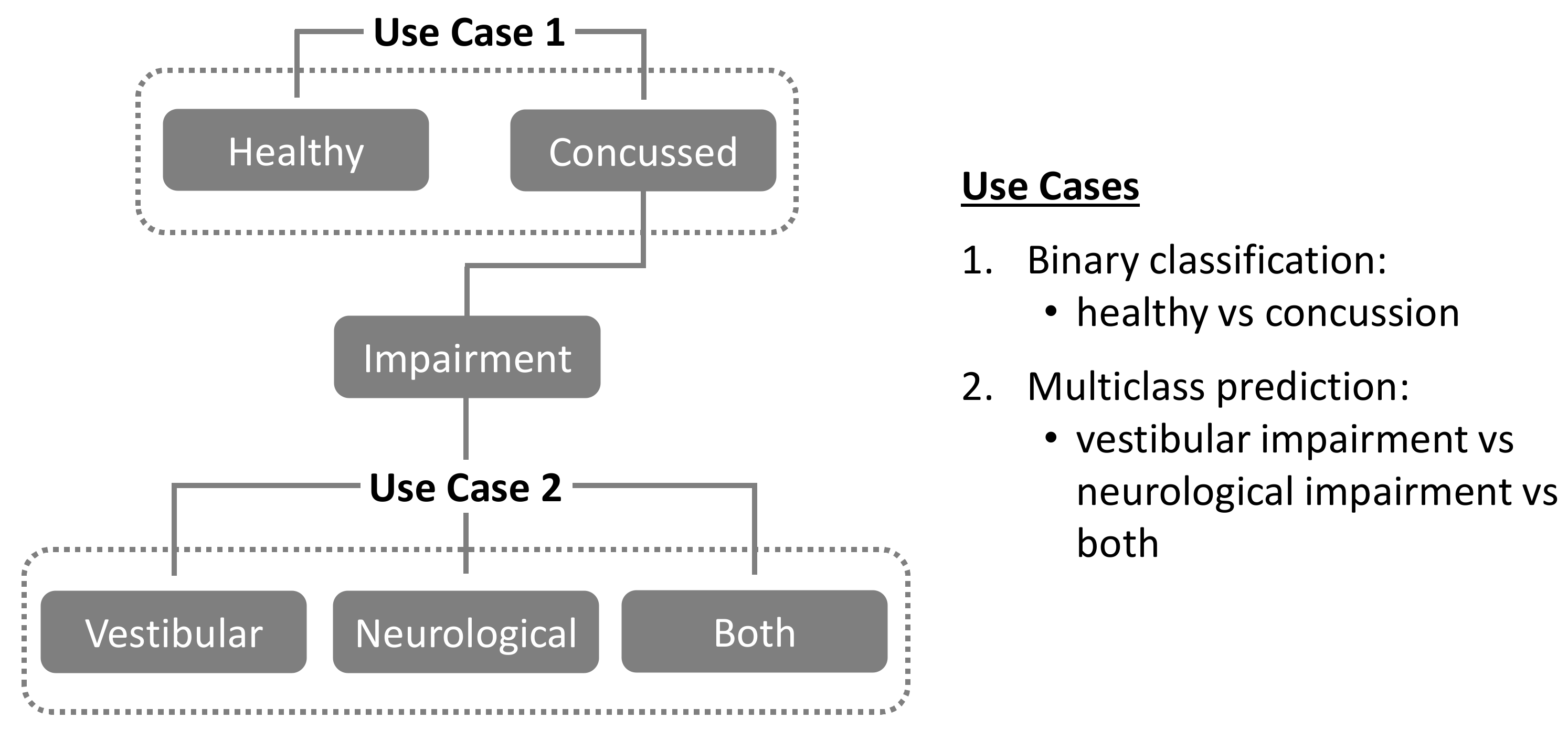
- In Use Case 1, the random assignment of “healthy” and “concussed” individuals into training, validation, and testing datasets maintained the original proportional balance in each dataset.
- In Use Case 2, the random assignment of the concussed individuals into training, validation, and testing datasets for multiclass prediction (“vestibular” vs. “neurological” vs. “both”) also maintained the original proportional balance in each dataset.
3. Results
3.1. Use Case 1: Classifying “Healthy” vs. “Concussion”
3.1.1. Comparison of TSA and NTS Data Preprocessing Pipelines
3.1.2. Machine Learning Model Comparisons
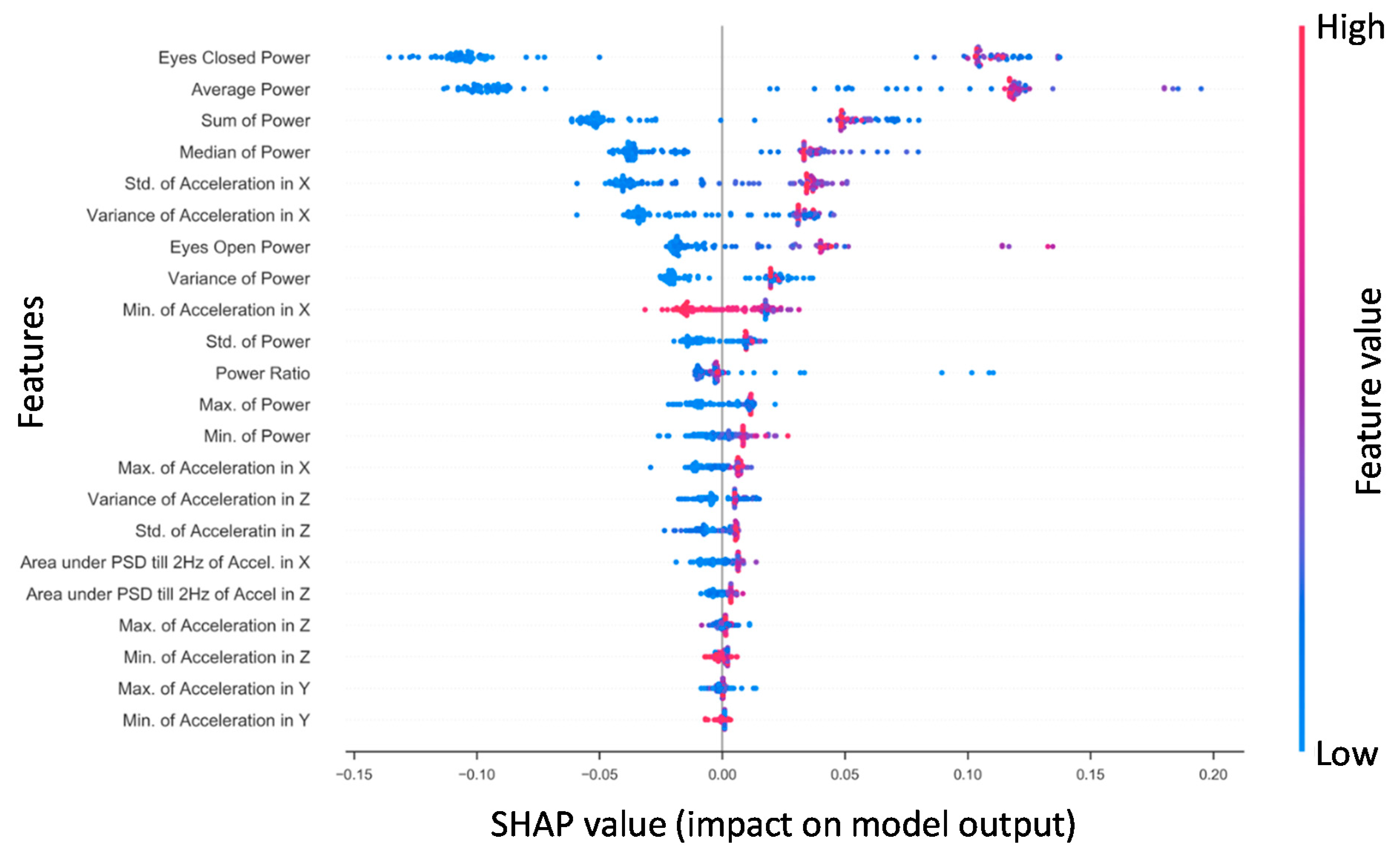
3.1.3. Concussed vs. Healthy SHAP for RF NTS Model
3.2. Use Case 2: Concussion Impairment Classification
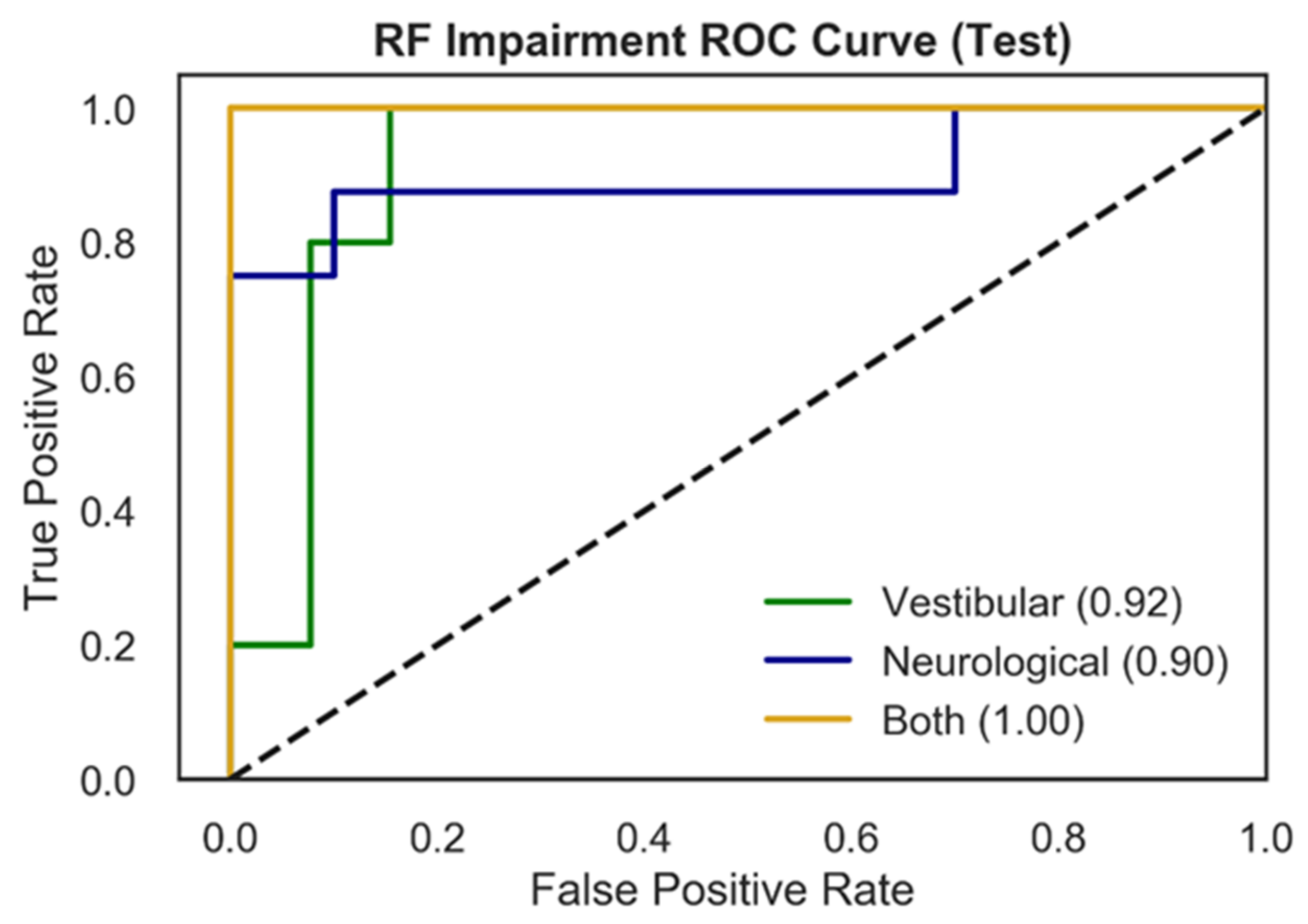
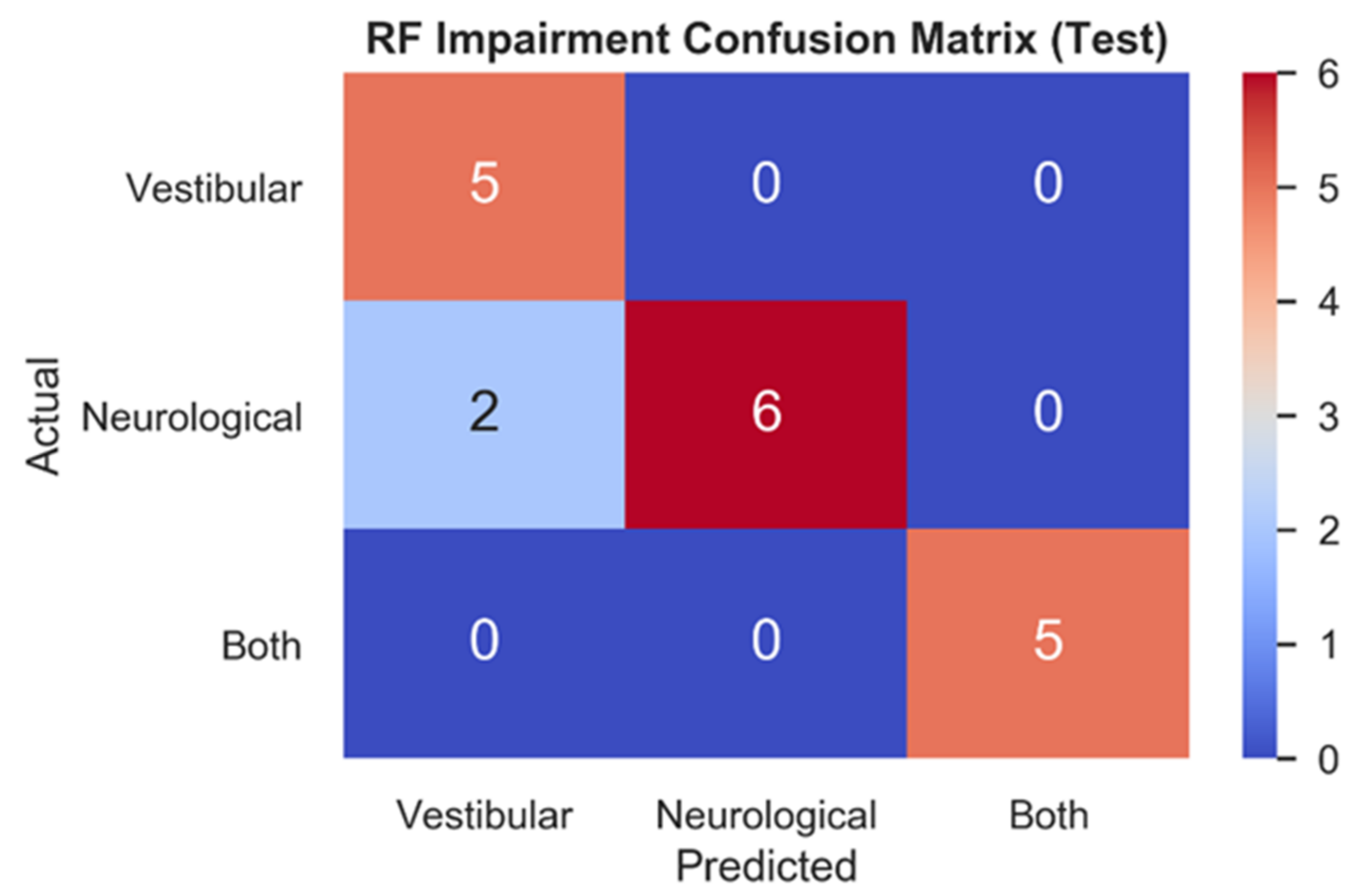
3.2.1. Comparison of Model Performance
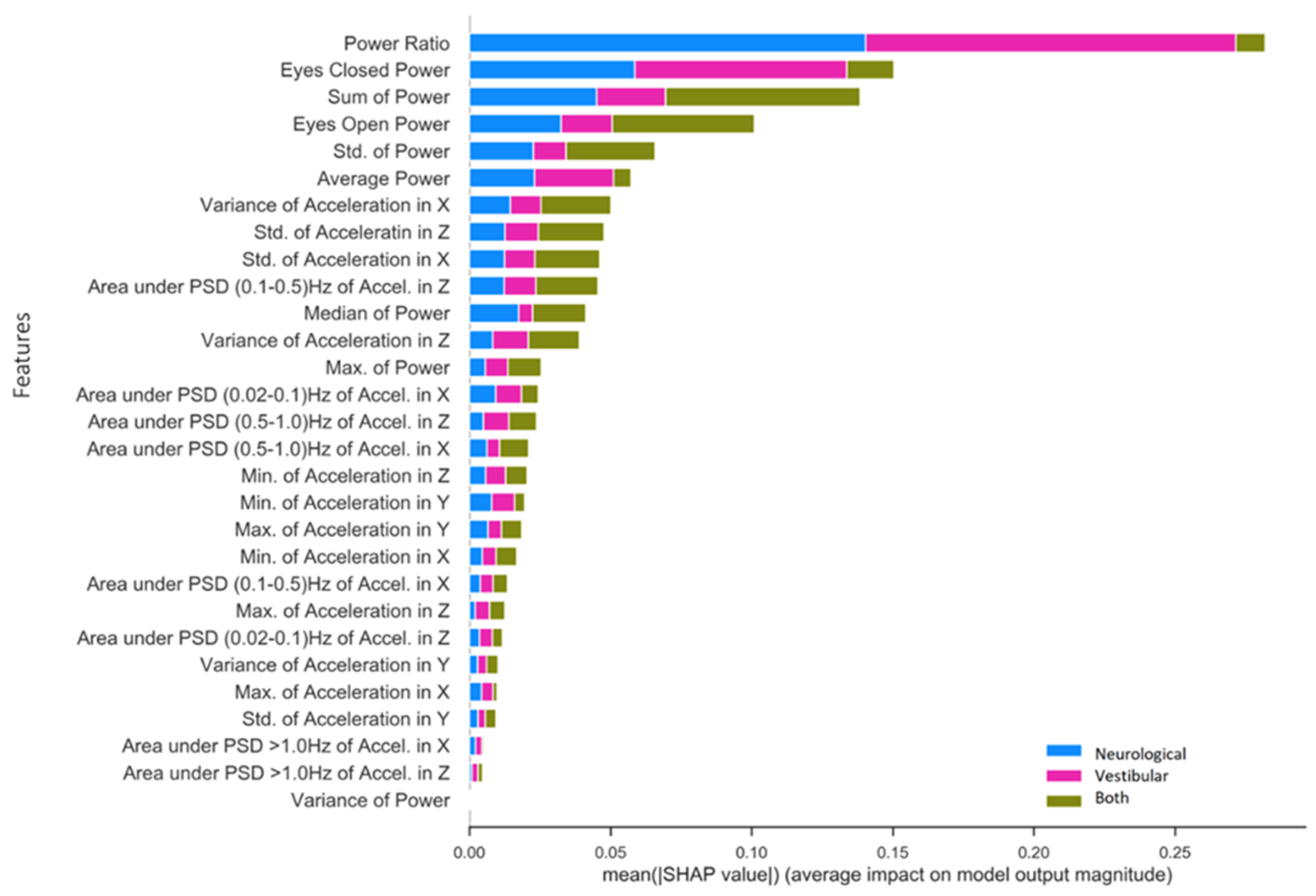
3.2.2. Specific Impairment SHAP for NTS RF Model
4. Discussion
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
- C = 20
- decision_function_shape = “ovr”
- kernel = “rbf”
- degree = 3
- probability = True
- class_weight = “balanced”
- oob_score = True
- n_estimators = 150
- class_weight = “balanced”
- min_samples_leaf = 2
- criterion = “gini”
- max_depth = None
- max_features = squareroot(n_features)
- n_estimators = 15
- max_depth = 6
- gamma = 0
- learning_rate = 0.3
- min_child_weight = 1
- subsample = 1
- C = 15
- decision_function_shape = “ovr”
- kernel = “rbf”
- degree = 5
- probability = True
- class_weight = “balanced”
- oob_score = True
- n_estimators = 150
- class_weight = “balanced”
- min_samples_leaf = 2
- criterion = “gini”
- max_depth = None
- max_features = squareroot(n_features)
- n_estimators = 50
- max_depth = 8
- gamma = 0
- learning_rate = 0.4
- min_child_weight = 1
- subsample = 1
- input_channels = 4
- output_channels = 64
- kernel_size = 6
- stride = 1
- activation_function = “ReLU”
- input_channels = 64
- output_channels = 128
- kernel_size = 6
- stride = 2
- activation_function = “ReLU”
- input_channels = 640
- output_channels = 100
- activation_layer = “ReLU”
- Linear Layer
- input_channels = 100
- output_channels = 1
References
- Memar, M.H.; Seidi, M.; Margulies, S. Head rotational kinematics, tissue deformations, and their relationships to the acute traumatic axonal injury. ASME J. Biomech. Eng. 2020, 142, 31006. [Google Scholar] [CrossRef]
- Chen, J.; Kouts, J.; Rippee, M.A.; Lauer, S.; Smith, D.; McDonald, T.; Kurylo, M.; Filardi, T. Developing a Comprehensive, Interdisciplinary Concussion Program. Health Serv. Insights 2020, 13, 1178632920938674. [Google Scholar] [CrossRef]
- Kelly, J.C.; Amerson, E.H.; Barth, J.T. Mild traumatic brain injury: Lessons learned from clinical, sports, and combat concussions. Rehabil. Res. Pract. 2012, 2012, 371970. [Google Scholar] [CrossRef] [Green Version]
- Yaramothu, C.; Goodman, A.M.; Alvarez, T.L. Epidemiology and Incidence of Pediatric Concussions in General Aspects of Life. Brain Sci. 2019, 9, 257. [Google Scholar] [CrossRef] [Green Version]
- Jacob, L.; Azouvi, P.; Kostev, K. Age-Related Changes in the Association between Traumatic Brain Injury and Dementia in Older Men and Women. J. Head Trauma Rehabil. 2021, 36, E139–E146. [Google Scholar] [CrossRef] [PubMed]
- Rădoi, A.; Poca, M.A.; Ga’ndara, D.; Castro, L.; Cevallos, M.; Pacios, M.E.; Sahuquillo, J. The Sport Concussion Assessment Tool (SCAT2) for evaluating civilian mild traumatic brain injury. A pilot normative study. PLoS ONE 2019, 14, e0212541. [Google Scholar] [CrossRef]
- Galea, O.A.; Cottrell, M.A.; Treleaven, J.M.; O’Leary, S.P. Sensorimotor and physiological indicators of impairment in mild traumatic brain injury: A meta-analysis. Neurorehabil. Neural Repair 2018, 32, 115–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKee, A.C. The Neuropathology of Chronic Traumatic Encephalopathy: The Status of the Literature. Semin Neurol. 2020, 40, 359–369. [Google Scholar] [CrossRef]
- Cavanaugh, J.T.; Guskiewicz, K.M.; Giuliani, C.; Marshall, S.; Mercer, V.; Stergiou, N. Detecting altered postural control after cerebral concussion in athletes with normal postural stability. Br. J. Sports Med. 2005, 39, 805–811. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dubose, D.F.; Herman, D.C.; Jones, D.L.; Tillman, S.M.; Clugston, J.R.; Pass, A.; Hernandez, J.A.; Vasilopoulos, T.; Horodyski, M.; Chmielewski, T.L. Lower Extremity Stiffness Changes after Concussion in Collegiate Football Players. Med. Sci. Sports Exerc. 2017, 49, 167–172. [Google Scholar] [CrossRef] [Green Version]
- Mang, C.S.; Whitten, T.A.; Cosh, M.S.; Scott, S.H.; Wiley, J.P.; Debert, C.T.; Dukelow, S.P.; Benson, B.W. Robotic Assessment of Motor, Sensory, and Cognitive Function in Acute Sport-Related Concussion and Recovery. J. Neurotrauma 2019, 36, 308–321. [Google Scholar] [CrossRef]
- Zhu, D.C.; Covassin, T.; Nogle, S.; Doyle, S.; Russell, D.; Pearson, R.L.; Monroe, J.; Liszewski, C.M.; Demarco, J.K.; Kaufman, D.I. A Potential Biomarker in Sports-Related Concussion: Brain Functional Connectivity Alteration of the Default-Mode Network Measured with Longitudinal Resting-State fMRI over Thirty Days. J. Neurotrauma 2015, 32, 327–341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garcia, G.P.; Yang, J.; Lavieri, M.S.; McAllister, T.W.; McCrea, M.A.; Broglio, S.P. Optimizing components of the sport concussion assessment tool for acute concussion assessment. Neurosurgery 2020, 87, 971–981. [Google Scholar] [CrossRef]
- Barlow, M.; Schlabach, D.; Peiffer, J.; Cook, C. Differences in change scores and the predictive validity of three commonly used measures following concussion in the middle school and high school aged population. Int. J. Sports Phys. Ther. 2011, 6, 150–157. [Google Scholar]
- Mann, A.; Tator, C.H.; Carson, J.D. Concussion diagnosis and management: Knowledge and attitudes of family medicine residents. Can. Fam. Physician 2017, 63, 460–466. [Google Scholar] [PubMed]
- Rowe, B.H.; Eliyahu, L.; Lowes, J.; Gaudet, L.A.; Beach, J.; Mrazik, M.; Cummings, G.; Voaklander, D. Concussion diagnoses among adults presenting to three Canadian emergency departments: Missed opportunities. Am. J. Emerg. Med. 2018, 36, 2144–2151. [Google Scholar] [CrossRef]
- Burke, M.J.; Fralick, M.; Nejatbakhsh, N.; Tartaglia, M.C.; Tator, C.H. In search of evidence-based treatment for concussion: Characteristics of current clinical trials. Brain Inj. 2015, 29, 300–305. [Google Scholar] [CrossRef]
- Cheng, B.; Knaack, C.; Forkert, N.D.; Schnabel, R.; Gerloff, C.; Thomalla, G. Stroke subtype classification by geometrical descriptors of lesion shape. PLoS ONE 2017, 12, e0185063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khezrian, M.; Myint, P.K.; McNeil, C.; Murray, A.D. A review of frailty syndrome and its physical, cognitive and emotional domains in the elderly. Geriatrics 2017, 2, 36. [Google Scholar] [CrossRef] [Green Version]
- Vellinga, M.; Geurts, J.; Rostrup, E.; Uitdehaag, B.; Polman, C.; Barkhof, F.; Vrenken, H. Clinical correlations of brain lesion distribution in multiple sclerosis. J. Magn. Reson. Imaging 2009, 29, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Kouli, A.; Torsney, K.M.; Kuan, W.L. Parkinson’s disease: Etiology, neuropathology, and pathogenesis. In Parkinson’s Disease: Pathogenesis and Clinical Aspects; Stoker, T.B., Greenland, J.C., Eds.; Codon Publications: Singapore, 2018. [Google Scholar] [CrossRef]
- Meier, T.B.; Giraldo-Chica, M.; España, L.Y.; Mayer, A.R.; Harezlak, J.; Nencka, A.S.; Wang, Y.; Koch, K.M.; Wu, Y.-C.; Saykin, A.J.; et al. Resting-State fMRI Metrics in Acute Sport-Related Concussion and Their Association with Clinical Recovery: A Study from the NCAA-DOD CARE Consortium. J. Neurotrauma 2020, 37, 152–162. [Google Scholar] [CrossRef] [PubMed]
- Toledo, E.; Lebel, A.; Becerra, L.; Minster, A.; Linnman, C.; Maleki, N.; Dodick, D.W.; Borsook, D. The young brain and concussion: Imaging as a biomarker for diagnosis and prognosis. Neurosci. Biobehav. Rev. 2012, 36, 1510–1531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrader, H.; Mickeviciene, D.; Gleizniene, R.; Jakstiene, S.; Surkiene, D.; Stovner, L.J.; Obelieniene, D. Magnetic resonance imaging after most common form of concussion. BMC Med. Imaging 2009, 9, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cavanaugh, J.T.; Guskiewicz, K.M.; Stergiou, N. A nonlinear dynamic approach for evaluating postural control: New directions for the management of sport-related cerebral concussion. Sports Med. 2005, 35, 935–950. [Google Scholar] [CrossRef] [PubMed]
- Buster, T.W.; Chernyavskiy, P.; Harms, N.R.; Kaste, E.G.; Burnfield, J.M. Computerized dynamic posturography detects balance deficits in individuals with a history of chronic severe traumatic brain injury. Brain Inj. 2016, 30, 1249–1255. [Google Scholar] [CrossRef]
- Martini, D.N.; Sabin, M.J.; DePesa, S.A.; Leal, E.W.; Negrete, T.N.; Sosnoff, J.J.; Broglio, S.P. The Chronic Effects of Concussion on Gait. Arch. Phys. Med. Rehabil. 2011, 92, 585–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kawata, K.; Liu, C.Y.; Merkel, S.F.; Ramirez, S.H.; Tierney, R.T.; Langford, D. Blood biomarkers for brain injury: What are we measuring? Neurosci. Biobehav. Rev. 2016, 68, 460–473. [Google Scholar] [CrossRef] [Green Version]
- Broglio, S.P.; CARE Consortium Investigators; Katz, B.P.; Zhao, S.; McCrea, M.; McAllister, T. Test-Retest Reliability and Interpretation of Common Concussion Assessment Tools: Findings from the NCAA-DoD CARE Consortium. Sports Med. 2018, 48, 1255–1268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keith, J.; Williams, M.; Taravath, S.; Lecci, L. A Clinician’s Guide to Machine Learning in Neuropsychological Research and Practice. J. Pediatr. Neuropsychol. 2019, 5, 177–187. [Google Scholar] [CrossRef]
- Vergara, V.M.; Mayer, A.R.; Damaraju, E.; Kiehl, K.A.; Calhoun, V. Detection of Mild Traumatic Brain Injury by Machine Learning Classification Using Resting State Functional Network Connectivity and Fractional Anisotropy. J. Neurotrauma. 2017, 34, 1045–1053. [Google Scholar] [CrossRef] [Green Version]
- Jacquin, A.; Kanakia, S.; Oberly, D.; Prichep, L.S. A multimodal biomarker for concussion identification, prognosis, and management. Comput. Biol. Med. 2018, 102, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Schmid, W.; Fan, Y.; Chi, T.; Golanov, E.; Regnier-Golanov, A.S.; Austerman, R.J.; Podell, K.; Cherukuri, P.; Bentley, T.; Steele, C.T.; et al. Review of wearable technologies and machine learning methodologies for systematic detection of mild traumatic brain injuries. J. Neural Eng. 2021, 18, 41006. [Google Scholar] [CrossRef]
- Boshra, R.; Dhindsa, K.; Boursalie, O.; Ruiter, K.I.; Sonnadara, R.; Samavi, R.; Doyle, T.E.; Reilly, J.P.; Connolly, J.F. From Group-Level Statistics to Single-Subject Prediction: Machine Learning Detection of Concussion in Retired Athletes. IEEE Trans. Neural Syst. Rehabil. Eng. 2019, 27, 1492–1501. [Google Scholar] [CrossRef]
- McNerney, M.W.; Hobday, T.; Cole, B.; Ganong, R.; Winans, N.; Matthews, D.; Hood, J.; Lane, S. Objective Classification of mTBI Using Machine Learning on a Combination of Frontopolar Electroencephalography Measurements and Self-reported Symptoms. Sports Med. Open 2019, 5, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prichep, L.S.; Ghosh-Dastidar, S.; Jacquin, A.; Koppes, W.; Miller, J.; Radman, T.; O’Neil, B.; Naunheim, R.; Huff, J.S. Classification algorithms for the identification of structural injury in TBI using brain electrical activity. Comput. Biol. Med. 2014, 53, 125–133. [Google Scholar] [CrossRef]
- Visscher, R.M.S.; Feddermann-Demont, N.; Romano, F.; Straumann, D.; Bertolini, G. Artificial intelligence for understanding concussion: Retrospective cluster analysis on the balance and vestibular diagnostic data of concussion patients. PLoS ONE 2019, 14, e0214525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Subhash, S.; Obafemi-Ajayi, T.; Goodman, D.; Wunsch, D.; Olbricht, G.R. Predictive Modeling of Sports-Related Concussions using Clinical Assessment Metrics. In Proceedings of the 2020 IEEE Symposium Series on Computational Intelligence (SSCI), Canberra, Australia, 1–4 December 2020; pp. 513–520. [Google Scholar] [CrossRef]
- Castellanos, J.; Phoo, C.P.; Eckner, J.T.; Franco, L.; Broglio, S.P.; McCrea, M.; McAllister, T.; Wiens, J.; The CARE Consortium Investigators. Predicting Risk of Sport-Related Concussion in Collegiate Athletes and Military Cadets: A Machine Learning Approach Using Baseline Data from the CARE Consortium Study. Sports Med. 2021, 51, 567–579. [Google Scholar] [CrossRef]
- Talkar, T.; Yuditskaya, S.; Williamson, J.R.; Lammert, A.C.; Rao, H.; Hannon, D.J.; O’Brien, A.; Vergara-Diaz, G.; DeLaura, R.; Quatieri, T.F.; et al. Detection of Subclinical Mild Traumatic Brain Injury (mTBI) Through Speech and Gait. In Proceedings of the INTERSPEECH 2020, Shanghai, China, 25–29 October 2020; pp. 135–139. [Google Scholar] [CrossRef]
- Landry, A.P.; Ting, W.K.C.; Zador, Z.; Sadeghian, A.; Cusimano, M.D. Using artificial neural networks to identify patients with concussion and postconcussion syndrome based on antisaccades. J. Neurosurg. 2019, 131, 1235–1242. [Google Scholar] [CrossRef] [Green Version]
- Peacock, W.F.; Van Meter, T.E.; Nazanin, M.; Ferber, K.; Gerwien, R.; Rao, V.; Sair, H.I.; Diaz-Arrastia, R.; Korley, F.K. Derivation of a Three Biomarker Panel to Improve Diagnosis in Patients with Mild Traumatic Brain Injury. Front. Neurol. 2017, 8, 641. [Google Scholar] [CrossRef] [Green Version]
- Rooks, T.F.; Dargie, A.S.; Chancey, V.C. Machine Learning Classification of Head Impact Sensor Data. In Proceedings of the ASME International Mechanical Engineering Congress and Exposition, Salt Lake City, UT, USA, 11–14 November 2019. Paper No: IMECE2019-12173. [Google Scholar] [CrossRef]
- Shim, V.B.; Holdsworth, S.; Champagne, A.A.; Coverdale, N.S.; Cook, D.J.; Lee, T.-R.; Wang, A.D.; Li, S.; Fernandez, J.W. Rapid Prediction of Brain Injury Pattern in mTBI by Combining FE Analysis with a Machine-Learning Based Approach. IEEE Access 2020, 8, 179457–179465. [Google Scholar] [CrossRef]
- Cai, Y.; Wu, S.; Zhao, W.; Li, Z.; Wu, Z.; Ji, S. Concussion classification via deep learning using whole-brain white matter fiber strains. PLoS ONE 2018, 13, e0197992. [Google Scholar] [CrossRef] [Green Version]
- Howell, D.R.; Kirkwood, M.W.; Provance, A.; Iverson, G.L.; Meehan, W.P., III. Using concurrent gait and cognitive assessments to identify impairments after concussion: A narrative review. Concussion 2018, 3, 54. [Google Scholar] [CrossRef] [Green Version]
- Fjeldstad, C.; Pardo, G.; Bemben, D.; Bemben, M. Decreased postural balance in multiple sclerosis patients with low disability. Int. J. Rehabil. Res. 2011, 34, 53–58. [Google Scholar] [CrossRef]
- Sutliff, M.H. Contribution of impaired mobility to patient burden in multiple sclerosis. Curr. Med. Res. Opin. 2010, 26, 109–119. [Google Scholar] [CrossRef]
- Hebert, J.R.; Manago, M.M. Reliability and Validity of the Computerized Dynamic Posturography Sensory Organization Test in People with Multiple Sclerosis. Int. J. MS Care 2017, 151–157. [Google Scholar] [CrossRef]
- Filli, L.; Sutter, T.; Easthope, C.S.; Killeen, T.; Meyer, C.; Reuter, K.; Lörincz, L.; Bolliger, M.; Weller, M.; Curt, A.; et al. Profiling walking dysfunction in multiple sclerosis: Characterisation, classification and progression over time. Sci. Rep. 2018, 8, 4984. [Google Scholar] [CrossRef]
- Lynch, S.G.; Parmenter, B.A.; Denney, D.R. The association between cognitive impairment and physical disability in multiple sclerosis. Mult. Scler. 2005, 11, 469–476. [Google Scholar] [CrossRef]
- Howell, D.; Osternig, L.; Chou, L.-S. Monitoring recovery of gait balance control following concussion using an accelerometer. J. Biomech. 2015, 48, 3364–3368. [Google Scholar] [CrossRef]
- King, L.A.; Mancini, M.; Fino, P.C.; Chestnutt, J.; Swanson, C.W.; Markwardt, S.; Chapman, J.C. Sensor-Based Balance Measures Outperform Modified Balance Error Scoring System in Identifying Acute Concussion. Ann. Biomed. Eng. 2017. [Google Scholar] [CrossRef]
- Wile, D.J.; Ranawaya, R.; Kiss, Z.H.T. Smart watch accelerometry for analysis and diagnosis of tremor. J. Neurosci. Methods 2014, 230, 1–4. [Google Scholar] [CrossRef]
- Rand, D.; Eng, J.J.; Tang, P.F.; Jeng, J.S.; Hung, C. How active are people with stroke?: Use of accelerometers to assess physical activity. Stroke 2009, 40, 163–168. [Google Scholar] [CrossRef] [Green Version]
- Moe-Nilssen, R. A new method for evaluating motor control in gait under real-life environmental conditions. Part 1: The instrument. Clin. Biomech. 1998, 13, 320–327. [Google Scholar] [CrossRef]
- Vienne-Jumeau, A.; Quijoux, F.; Vidal, P.-P.; Ricard, D. Wearable inertial sensors provide reliable biomarkers of disease severity in multiple sclerosis: A systematic review and meta-analysis. Ann. Phys. Rehabil. Med. 2020, 63, 138–147. [Google Scholar] [CrossRef] [PubMed]
- Ralston, J.D.; Raina, A.; Benson, B.W.; Peters, R.M.; Roper, J.M.; Ralston, A.B. Physiological Vibration Acceleration (Phybrata) Sensor Assessment of Multi-System Physiological Impairments and Sensory Reweighting Following Concussion. Med. Devices Evid. Res. 2020, 13, 411–438. [Google Scholar] [CrossRef]
- Godfrey, A.; Bourke, A.K.; Ólaighin, G.M.; van de Ven, P.; Nelson, J. Activity classification using a single chest mounted tri-axial accelerometer. Med. Eng. Phys. 2011, 33, 1127–1135. [Google Scholar] [CrossRef]
- Fortino, G.; Giannantonio, R.; Gravina, R.; Kuryloski, P.; Jafari, R. Enabling effective programming and flexible management of efficient body sensor network applications. IEEE Trans. Hum. Mach. Syst. 2013, 43, 115–133. [Google Scholar] [CrossRef]
- Dief, T.N.; Abdollah, V.; Ralston, J.D.; Ho, C.; Rouhani, H. Investigating the Validity of a Single Tri-axial Accelerometer Mounted on the head for Monitoring the Activities of Daily Living and the Timed-Up and Go Test. Gait Posture 2021, 90, 137–140. [Google Scholar]
- Nazarahari, M.; Rouhani, H. Detection of daily postures and walking modalities using a single chest-mounted tri-axial accelerometer. Med. Eng. Phys. 2018, 57, 75–81. [Google Scholar] [CrossRef]
- Paraschiv-Ionescu, A.; Buchser, E.E.; Rutschmann, B.; Najafi, B.; Aminian, K. Ambulatory system for the quantitative and qualitative analysis of gait and posture in chronic pain patients treated with spinal cord stimulation. Gait Posture 2004, 20, 113–125. [Google Scholar] [CrossRef] [PubMed]
- Oba, N.; Sasagawa, S.; Yamamoto, A.; Nakazawa, K. Difference in postural control during quiet standing between young children and adults: Assessment with center of mass acceleration. PLoS ONE 2015, 10, e0140235. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, I.-S.; Lai, D.-M.; Ding, J.-J.; Chien, A.; Cheng, C.-H.; Wang, S.-F.; Wang, J.-L.; Kuo, C.-L.; Hsu, W.-L. Reweighting of the sensory inputs for postural control in patients with cervical spondylotic myelopathy after surgery. J. Neuroeng. Rehabil. 2019, 16, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Assländer, L.; Peterka, R.J. Sensory reweighting dynamics in human postural control. J. Neurophysiol. 2014, 111, 1852–1864. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Diener, H.C.; Dichgans, J.; Bacher, M.; Gompf, B. Quantification of postural sway in normals and patients with cerebellar diseases. Electroencephalogr. Clin. Neurophysiol. 1984, 57, 134–142. [Google Scholar] [CrossRef] [Green Version]
- Kanekar, N.; Lee, Y.-J.; Aruin, A.S. Frequency analysis approach to study balance control in individuals with multiple sclerosis. J. Neurosci. Methods 2014, 222, 91–96. [Google Scholar] [CrossRef]
- Grafton, S.T.; Ralston, A.B.; Ralston, J.D. Monitoring of postural sway with a head-mounted wearable device: Effects of gender, participant state, and concussion. Med. Devices Evid. Res. 2019, 12, 151–164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, W.G.; Tierney, R.T.; McDevitt, J. Visual-vestibular processing deficits in mild traumatic brain injury. J. Vest. Res. 2017, 27, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Kolev, O.I.; Sergeeva, M. Vestibular disorders following different types of head and neck trauma. Funct. Neurol. 2016, 31, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Banman, C.J.; Schneider, K.J.; Cluff, T.; Peters, R.M. Altered Vestibular Balance Function in Combat Sport Athletes. J. Neurotrauma 2021, 38, 2291–2300. [Google Scholar] [CrossRef] [PubMed]
- Wallace, B.; Lifshitz, J. Traumatic brain injury and vestibulo-ocular function: Current challenges and future prospects. Eye Brain 2016, 8, 153–164. [Google Scholar] [CrossRef] [Green Version]
- Baruch, M.; Barth, J.T.; Cifu, D.; Leibman, M. Utility of a multimodal neurophysiological assessment tool in distinguishing between individuals with and without a history of mild traumatic brain injury. J. Rehabil. Res. Dev. 2016, 53, 959–972. [Google Scholar] [CrossRef]
- Resch, J.E.; Brown, C.N.; Schmidt, J.; Macciocchi, S.N.; Blueitt, D.; Cullum, C.M.; Ferrara, M.S. The sensitivity and specificity of clinical measures of sport concussion: Three tests are better than one. BMJ Open Sport Exerc. Med. 2016, 2, e000012. [Google Scholar] [CrossRef] [Green Version]
- Balaban, C.; Hoffer, M.E.; Szczupak, M.; Snapp, H.; Crawford, J.; Murphy, S.; Marshall, K.; Pelusso, C.; Knowles, S.; Kiderman, A. Oculomotor, Vestibular, and Reaction Time Tests in Mild Traumatic Brain Injury. PLoS ONE 2016, 11, e0162168. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hauenstein, A.; Roper, J.M.; Ralston, A.B.; Ralston, J.D. Signal classification of wearable inertial motion sensor data using a convolutional neural network. In Proceedings of the IEEE-EMBS 2019 International Conference Biomed & Health Informatics, Chicago, IL, USA, 19–22 May 2019. [Google Scholar]
- Welch, P. The use of fast Fourier transform for the estimation of power spectra: A method based on time averaging over short, modified periodograms. IEEE Trans. Audio Electroacoust. 1967, 15, 70–73. [Google Scholar] [CrossRef] [Green Version]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A scalable tree boosting system. In Proceedings of the KDD ‘16 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar] [CrossRef] [Green Version]
- Paszke, A.; Gross, S.; Chintala, S.; Chanan, G.; Yang, E.; DeVito, Z.; Lerer, A. Automatic differentiation in pytorch. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Berrar, D. Cross-validation. In Encyclopedia of Bioinformatics and Computational Biology; Elsevier: Amsterdam, The Netherlands, 2018; Volume 1, pp. 542–545. [Google Scholar] [CrossRef]
- Arlot, S.; Celisse, A. A survey of cross-validation procedures for model selection. Stat. Surv. 2010, 4, 40–79. [Google Scholar] [CrossRef]
- Jung, Y. Multiple predicting K-fold cross-validation for model selection. J. Nonparametr. Stat. 2018, 30, 197–215. [Google Scholar] [CrossRef]
- Burman, P. A comparative study of ordinary cross-validation, v-fold cross-validation and the repeated learning-testing methods. Biometrika 1989, 76, 503–514. [Google Scholar] [CrossRef] [Green Version]
- Lundberg, S.M.; Lee, S.I. A unified approach to interpreting model predictions. In Proceedings of the 31st International Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 4768–4777. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Faul, F.; Erdfelder, E.; Lang, A.-G.; Buchner, A. G*Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav. Res. Methods 2007, 39, 175–191. [Google Scholar] [CrossRef]
- Corwin, D.J.; Wiebe, D.J.; Zonfrillo, M.; Grady, M.F.; Robinson, R.L.; Goodman, A.M.; Master, C.L. Vestibular Deficits following Youth Concussion. J. Pediatr. 2015, 166, 1221–1225. [Google Scholar] [CrossRef] [Green Version]
- Christy, J.B.; Cochrane, G.; Almutairi, A.; Busettini, C.; Swanson, M.W.; Weise, K.K. Peripheral Vestibular and Balance Function in Athletes with and Without Concussion. J. Neurol. Phys. Ther. 2019, 43, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Schwab, N.; Grenier, K.; Hazrati, L.-N. DNA repair deficiency and senescence in concussed professional athletes involved in contact sports. Acta Neuropathol. Commun. 2019, 7, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Chancellor, S.E.; Franz, E.S.; Minaeva, O.V.; Goldstein, L.E. Pathophysiology of concussion. Semin. Pediatr. Neurol. 2019, 30, 14–25. [Google Scholar] [CrossRef] [PubMed]
| Use-Case | Model | Preprocessing Pipeline | Specificity | Sensitivity | F1 |
|---|---|---|---|---|---|
| Concussed vs. Healthy | RF | TSA | 0.94 | 0.94 | 0.94 |
| NTS | 0.88 | 0.99 | 0.94 | ||
| SVM | TSA | 0.94 | 0.47 | 0.62 | |
| NTS | 0.88 | 0.94 | 0.91 | ||
| XGB | TSA | 0.88 | 0.94 | 0.91 | |
| NTS | 0.88 | 0.99 | 0.94 | ||
| CNN | TSA | 0.88 | 0.94 | 0.91 |
| Use-Case | Model | Preprocessing Pipeline | Specificity | Sensitivity | F1 |
|---|---|---|---|---|---|
| Vestibular vs. Neurological vs. Both | RF | NTS | 0.93 | 0.89 | 0.90 |
| SVM | NTS | 0.83 | 0.72 | 0.73 | |
| XGB | NTS | 0.93 | 0.83 | 0.85 |
| Data Source | ML Model(s) | Sensitivity | Specificity | F1 | AUC | Reference |
|---|---|---|---|---|---|---|
| Phybrata sensor | RF, SVM, XGB, CNN | 0.94 | 0.94 | 0.94 | 0.98 | present work |
| Multimodal: Neurocognitive tests, clinical scales, symptoms checklists, balance and gait testing | CNN | nr | nr | 0.85 | 0.95 | [30] |
| MRI | SVM | 0.89 | 0.79 | 0.84 | 0.84 | [31] |
| Multimodal: EEG, neurocognitive tests, standard concussion assessment tools | Genetic Algorithm (GA) classifier | 0.92 | 0.75 | 0.81 | 0.92 | [32] |
| EEG | SVM | 0.82 | 0.80 | 0.81 | nr | [34] |
| EEG | Genetic Algorithm (GA) classifier | 0.98 | 0.60 | nr | 0.90 | [36] |
| Clinical scales and assessment metrics: retrospective analysis | C5.0 Decision Tree, Recursive Partitioning, Random Forest, XGB | 0.97–0.99 | 0.43–0.58 | 0.71–0.78 | nr | [38] |
| Eye tracking | CNN | 0.63 | 0.74 | 0.67 | 0.75 | [41] |
| 3 blood biomarkers | Random Forest | 0.98 | 0.72 | nr | 0.91 | [42] |
| Head impact data | SVM, Random Forest, CNN | 0.84 | 0.88 | 0.86 | 0.9 | [45] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hope, A.J.; Vashisth, U.; Parker, M.J.; Ralston, A.B.; Roper, J.M.; Ralston, J.D. Phybrata Sensors and Machine Learning for Enhanced Neurophysiological Diagnosis and Treatment. Sensors 2021, 21, 7417. https://doi.org/10.3390/s21217417
Hope AJ, Vashisth U, Parker MJ, Ralston AB, Roper JM, Ralston JD. Phybrata Sensors and Machine Learning for Enhanced Neurophysiological Diagnosis and Treatment. Sensors. 2021; 21(21):7417. https://doi.org/10.3390/s21217417
Chicago/Turabian StyleHope, Alex J., Utkarsh Vashisth, Matthew J. Parker, Andreas B. Ralston, Joshua M. Roper, and John D. Ralston. 2021. "Phybrata Sensors and Machine Learning for Enhanced Neurophysiological Diagnosis and Treatment" Sensors 21, no. 21: 7417. https://doi.org/10.3390/s21217417
APA StyleHope, A. J., Vashisth, U., Parker, M. J., Ralston, A. B., Roper, J. M., & Ralston, J. D. (2021). Phybrata Sensors and Machine Learning for Enhanced Neurophysiological Diagnosis and Treatment. Sensors, 21(21), 7417. https://doi.org/10.3390/s21217417






