Noise Sources and Requirements for Confocal Raman Spectrometers in Biosensor Applications
Abstract
:1. Introduction
2. Materials and Methods
2.1. Instrumentation
- a low-cost, small home-build spectrometer with CMOS camera (mini CMOS Raman spectrometer);
- a middle price-class small Czerny-Turner spectrometer with CCD detector (mini CCD Raman spectrometer);
- a commercial, laboratory grade, confocal Raman spectrometer with a deeply cooled CCD detector (reference Raman spectrometer).
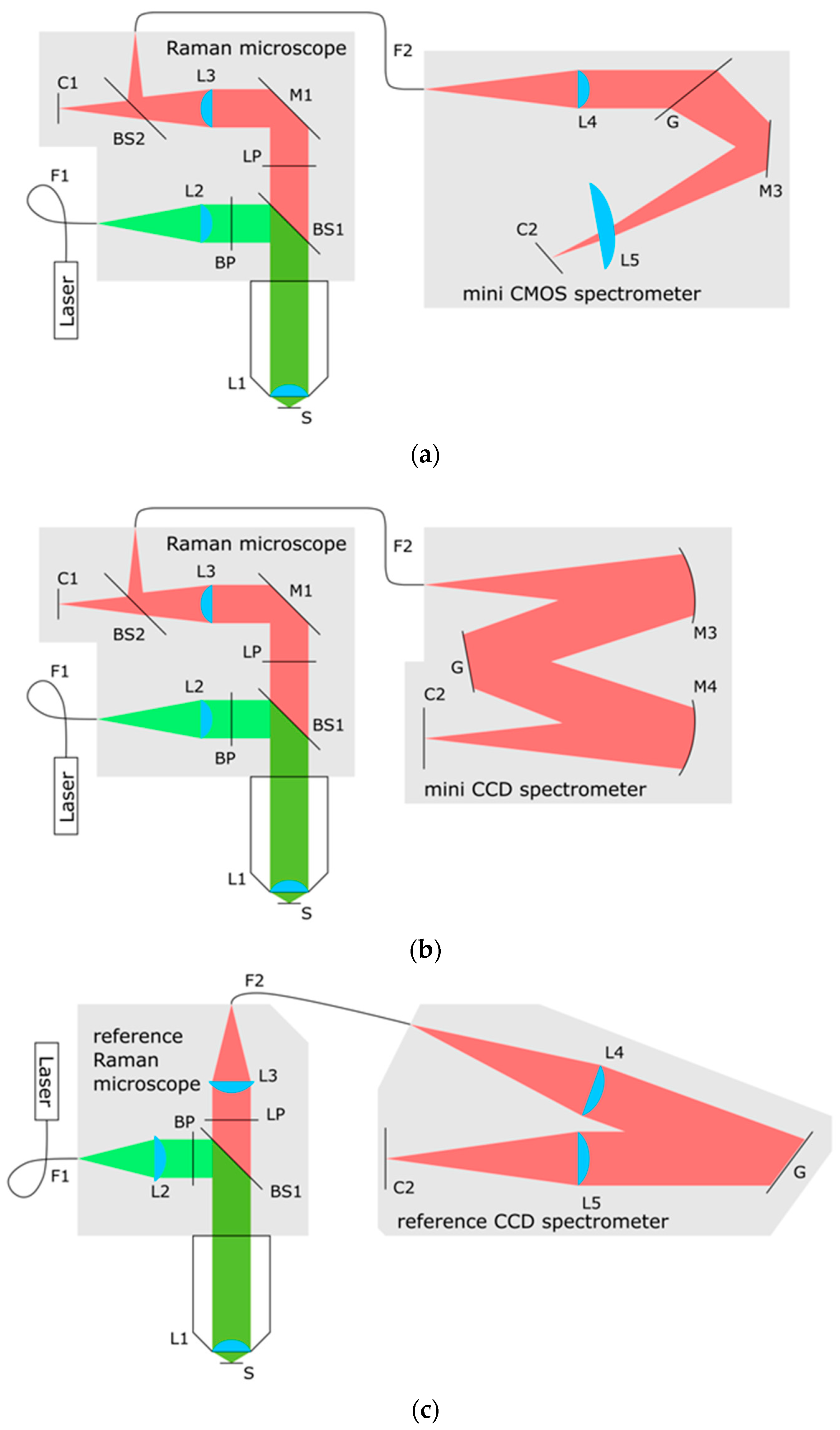
2.1.1. Raman Microscope Components
2.1.2. The Mini CMOS Spectrometer
2.1.3. The Mini CCD Spectrometer
2.1.4. The Reference Raman Spectrometer
2.2. Bacterial Strains and Culturing Conditions
2.3. Raman Measuremnts
2.4. Signal-to-Noise Ratio (SNR) Considerations
- variance of signal photon noise (shot noise of photons),
- variance of background noise (shot noise of fluorescence),
- variance of dark current noise (shot noise of dark current) and
- variance of readout noise.
2.5. Data Pre-Processing and Chemometrical Analysis
3. Results and Discussion
3.1. Photon Budget of Instruments
3.2. Signal-to-Noise Ratio of Spectra
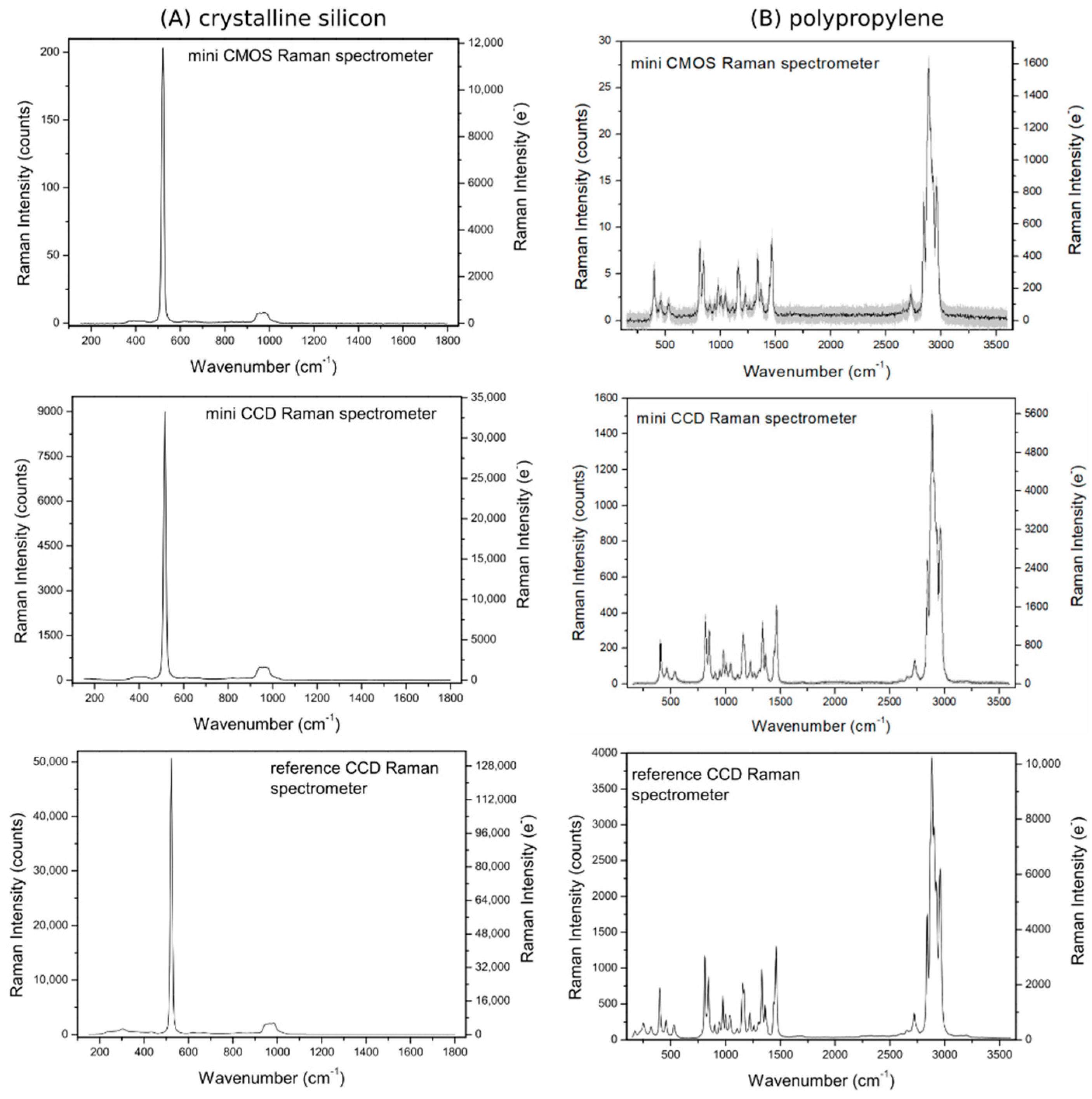
3.2.1. Samples
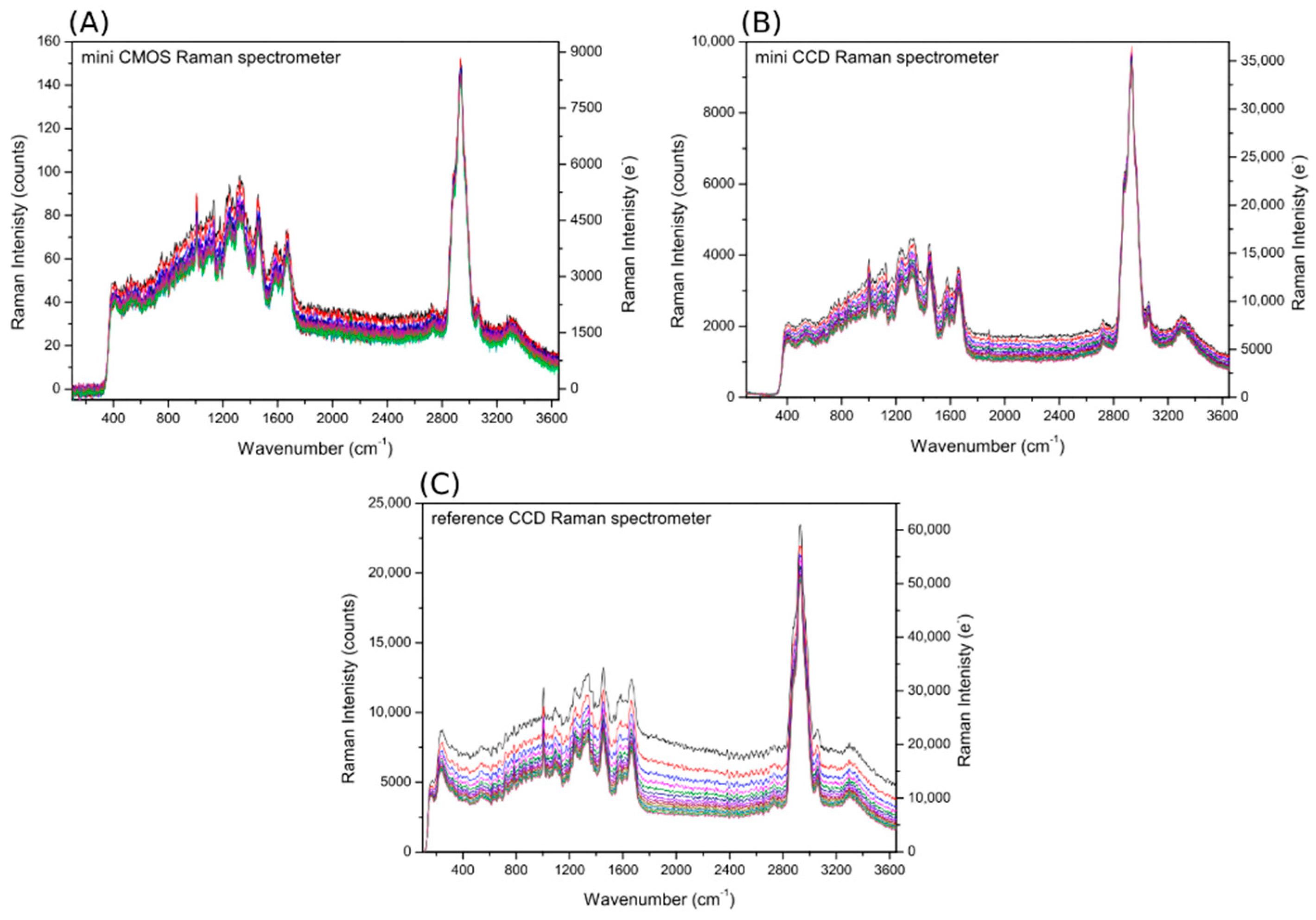
3.2.2. Raman Spectra
3.2.3. The Role of Confocality
3.2.4. Raman Signal Independent Noise Sources
3.2.5. Calculated vs. Experimental SNR—The Case of Non-Fluorescing Samples
3.2.6. Calculated SNR—The Case of Fluorescing Samples
3.2.7. The Influence of Acquisition Time on the SNR
3.2.8. Other Noise Sources
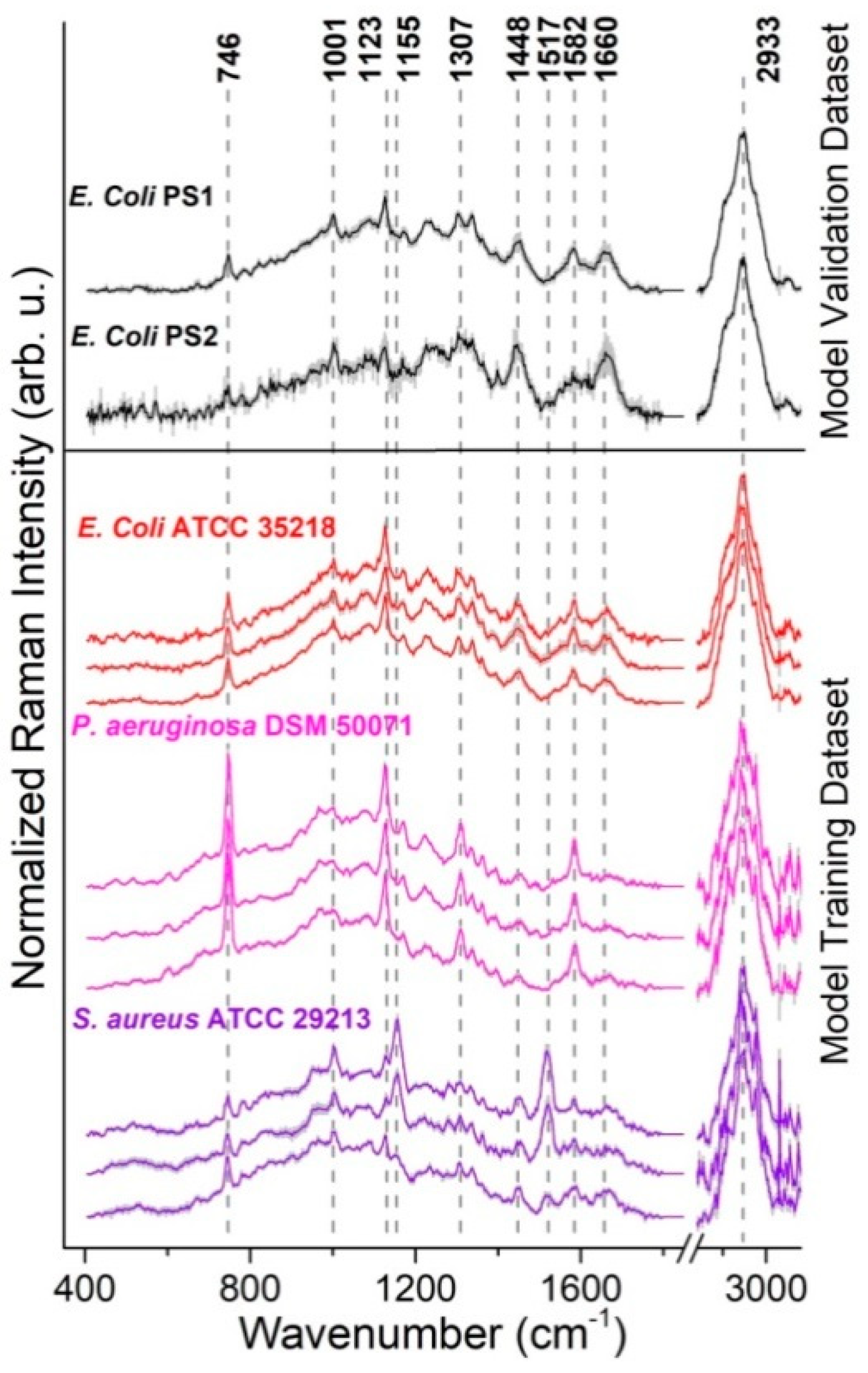
3.3. Pathogen Identification
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, X.; Liu, Y.; Huang, J.; Liu, W.; Huang, J.; Zhang, Y.; Fu, W. Label-free techniques for laboratory medicine applications. Front. Lab. Med. 2017, 1, 82–85. [Google Scholar] [CrossRef]
- Krafft, C.; Popp, J. Raman Spectroscopy to Solve Unmet Needs in Histopathology. In Proceedings of the International Photonics and Optoelectronics Meeting 2017, Wuhan, China, 3 November 2017; Optical Society of America: Wuhan, China, 2017; p. ASu1A.1. [Google Scholar]
- Orringer, D.A.; Pandian, B.; Niknafs, Y.S.; Hollon, T.C.; Boyle, J.; Lewis, S.; Garrard, M.; Hervey-Jumper, S.L.; Garton, H.J.L.; Maher, C.O.; et al. Rapid intraoperative histology of unprocessed surgical specimens via fibre-laser-based stimulated Raman scattering microscopy. Nat. Biomed. Eng. 2017, 1, 27. [Google Scholar] [CrossRef] [PubMed]
- Hubbard, T.J.E.; Shore, A.; Stone, N. Raman spectroscopy for rapid intra-operative margin analysis of surgically excised tumour specimens. Analyst 2019, 144, 6479–6496. [Google Scholar] [CrossRef]
- Kloß, S.; Rösch, P.; Pfister, W.; Kiehntopf, M.; Popp, J. Toward Culture-Free Raman Spectroscopic Identification of Pathogens in Ascitic Fluid. Anal. Chem. 2015, 87, 937–943. [Google Scholar] [CrossRef]
- Stöckel, S.; Kirchhoff, J.; Neugebauer, U.; Rösch, P.; Popp, J. The application of Raman spectroscopy for the detection and identification of microorganisms. J. Raman Spectrosc. 2016, 47, 89–109. [Google Scholar] [CrossRef]
- Kumar, S.; Gopinathan, R.; Chandra, G.K.; Umapathy, S.; Saini, D.K. Rapid detection of bacterial infection and viability assessment with high specificity and sensitivity using Raman microspectroscopy. Anal. Bioanal. Chem. 2020, 412, 2505–2516. [Google Scholar] [CrossRef] [PubMed]
- Vanden-Hehir, S.; Tipping, W.J.; Lee, M.; Brunton, V.G.; Williams, A.; Hulme, A.N. Raman Imaging of Nanocarriers for Drug Delivery. Nanomaterials 2019, 9, 341. [Google Scholar] [CrossRef] [Green Version]
- Jung, N.; Windbergs, M. Raman spectroscopy in pharmaceutical research and industry. Phys. Sci. Rev. 2018, 3, 8. [Google Scholar] [CrossRef]
- Krishnan, K.S. The Raman Effect in Crystals. Nature 1928, 122, 477–478. [Google Scholar] [CrossRef]
- Schmitt, M.; Mayerhöfer, T.; Popp, J.; Kleppe, I.; Weisshart, K. Light–Matter Interaction. In Handbook of Biophotonics; Wiley-VCH: Berlin, Germany, 2012; pp. 87–261. [Google Scholar]
- Sellar, R.G.; Boreman, G.D. Comparison of relative signal-to-noise ratios of different classes of imaging spectrometer. Appl. Opt. 2005, 44, 1614–1624. [Google Scholar] [CrossRef]
- Hauswald, W.; Förster, R.; Popp, J.; Heintzmann, R. Thermal illumination limits in 3D Raman microscopy: A comparison of different sample illumination strategies to obtain maximum imaging speed. PLoS ONE 2019, 14, e0220824. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Hu, C.; Chu, K.; Smith, Z.J. Low resolution Raman: The impact of spectral resolution on limit of detection and imaging speed in hyperspectral imaging. Analyst 2020, 145, 6607–6616. [Google Scholar] [CrossRef]
- Schie, I.W.; Krafft, C.; Popp, J. Cell classification with low-resolution Raman spectroscopy (LRRS). J. Biophotonics 2016, 9, 994–1000. [Google Scholar] [CrossRef]
- Shipp, D.W.; Sinjab, F.; Notingher, I. Raman spectroscopy: Techniques and applications in the life sciences. Adv. Opt. Photonics 2017, 9, 315–428. [Google Scholar] [CrossRef] [Green Version]
- Krafft, C.; Schmitt, M.; Schie, I.W.; Cialla-May, D.; Matthäus, C.; Bocklitz, T.; Popp, J. Label-Free Molecular Imaging of Biological Cells and Tissues by Linear and Nonlinear Raman Spectroscopic Approaches. Angew. Chem. Int. Ed. 2017, 56, 4392–4430. [Google Scholar] [CrossRef] [PubMed]
- Pahlow, S.; Kloß, S.; Blättel, V.; Kirsch, K.; Hübner, U.; Cialla, D.; Rösch, P.; Weber, K.; Popp, J. Isolation and Enrichment of Pathogens with a Surface-Modified Aluminium Chip for Raman Spectroscopic Applications. ChemPhysChem 2013, 14, 3600–3605. [Google Scholar] [CrossRef]
- Ramoji, A.; Galler, K.; Glaser, U.; Henkel, T.; Mayer, G.; Dellith, J.; Bauer, M.; Popp, J.; Neugebauer, U. Characterization of different substrates for Raman spectroscopic imaging of eukaryotic cells. J. Raman Spectrosc. 2016, 47, 773–786. [Google Scholar] [CrossRef]
- McCreery, R.L. Signal-to-Noise in Raman Spectroscopy. In Raman Spectroscopy for Chemical Analysis; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2000; pp. 49–71. [Google Scholar]
- Team, R.C. R: A Language and Environment for Statistical Computing. Available online: https://www.R-project.org/ (accessed on 29 June 2021).
- Ryabchykov, O.; Guo, S.; Bocklitz, T. Micro-raman spectroscopy. In 4. Analyzing Raman Spectroscopic Data; Jürgen, P., Thomas, M., Eds.; De Gruyter: Berlin, Germany, 2020; pp. 81–106. [Google Scholar]
- Storozhuk, D.; Ryabchykov, O. Online Software Platform for Raman Spectroscopic Data Analysis. Available online: https://ramanmetrix.eu (accessed on 29 June 2021).
- Ryabchykov, O.; Bocklitz, T.; Ramoji, A.; Neugebauer, U.; Foerster, M.; Kroegel, C.; Bauer, M.; Kiehntopf, M.; Popp, J. Automatization of spike correction in Raman spectra of biological samples. Chemom. Intell. Lab. Syst. 2016, 155, 1–6. [Google Scholar] [CrossRef]
- Bocklitz, T.W.; Dörfer, T.; Heinke, R.; Schmitt, M.; Popp, J. Spectrometer calibration protocol for Raman spectra recorded with different excitation wavelengths. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2015, 149, 544–549. [Google Scholar] [CrossRef]
- Ryan, C.G.; Clayton, E.; Griffin, W.L.; Sie, S.H.; Cousens, D.R. SNIP, a statistics-sensitive background treatment for the quantitative analysis of PIXE spectra in geoscience applications. Nucl. Instrum. Methods Phys. Res. Sect. B Beam Interact. Mater. At. 1988, 34, 396–402. [Google Scholar] [CrossRef]
- Guo, S.; Bocklitz, T.; Neugebauer, U.; Popp, J. Common mistakes in cross-validating classification models. Anal. Methods 2017, 9, 4410–4417. [Google Scholar] [CrossRef]
- van Vliet, L.J.; Sudar, D.; Young, I.T. Digital fluorescence imaging using cooled CCD array cameras. In Cell Biology; Celis, J., Ed.; Academic Press: New York, NY, USA, 1998; Volume III, pp. 109–120. [Google Scholar]
- Aggarwal, R.L.; Farrar, L.W.; Saikin, S.K.; Aspuru-Guzik, A.; Stopa, M.; Polla, D.L. Measurement of the absolute Raman cross section of the optical phonon in silicon. Solid State Commun. 2011, 151, 553–556. [Google Scholar] [CrossRef]
- Wendel, H. Theoretical study of the Raman cross section and its pressure dependence in silicon. Solid State Commun. 1979, 31, 423–426. [Google Scholar] [CrossRef]
- Widenhorn, R.; Blouke, M.; Weber, A.; Rest, A.; Bodegom, E. Temperature Dependence of Dark Current in a CCD; SPIE: Bellingham, WA, USA, 2002; Volume 4669. [Google Scholar]
- Ho, C.-S.; Jean, N.; Hogan, C.A.; Blackmon, L.; Jeffrey, S.S.; Holodniy, M.; Banaei, N.; Saleh, A.A.E.; Ermon, S.; Dionne, J. Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat. Commun. 2019, 10, 4927. [Google Scholar] [CrossRef] [PubMed]
- Pahlow, S.; Meisel, S.; Cialla-May, D.; Weber, K.; Rösch, P.; Popp, J. Isolation and identification of bacteria by means of Raman spectroscopy. Adv. Drug Deliv. Rev. 2015, 89, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, B.; Ali, N.; Bocklitz, T.; Rösch, P.; Popp, J. Discrimination between pathogenic and non-pathogenic E. coli strains by means of Raman microspectroscopy. Anal. Bioanal. Chem. 2020, 412, 8241–8247. [Google Scholar] [CrossRef]
- Guo, S.; Rösch, P.; Popp, J.; Bocklitz, T. Modified PCA and PLS: Towards a better classification in Raman spectroscopy–based biological applications. J. Chemom. 2020, 34, e3202. [Google Scholar] [CrossRef] [Green Version]
- Macernis, M.; Galzerano, D.; Sulskus, J.; Kish, E.; Kim, Y.-H.; Koo, S.; Valkunas, L.; Robert, B. Resonance Raman Spectra of Carotenoid Molecules: Influence of Methyl Substitutions. J. Phys. Chem. A 2015, 119, 56–66. [Google Scholar] [CrossRef]

| Raman Spectrometer | Mini CMOS | Mini CCD | Reference | Units |
|---|---|---|---|---|
| Excitation laser wavelength | 532 | 532 | 514 | nm |
| Laser power at the sample | 3.8 | 3.8 | 3.8 | mW |
| Conversion factor (datasheet) | - | 4 | 2 | e−/ADU |
| System magnification | 13.9 | 13.9 | 45.8 | x |
| Collection fiber diameter | 25 | 25 | 50 | µm |
| Integrated c-Si signal 519 cm−1 | 816 | 28,065 | 119,948 | ADU/s |
| Conversion factor measured | 58 (a) | 3.7 | 2.6 | e−/ADU |
| Photoelectrons at detector | 0.047 × 106 | 0.104 × 106 | 0.312 × 106 | e−/s |
| Photons ideally expected at the sample (b) | (5.2 ± 3.5) × 106 | (5.2 ± 3.5) × 106 | (4.2 ± 2.7) × 106 | 1/s |
| Theoretical efficiency (c) | 0.92% | 2.01% | 7.42% | |
| Optical efficiency (d) | 7.0% | 10.4% | 27% | |
| Remaining efficiency (e) | 13% | 19% | 27% |
| Raman Spectrometer | Mini CMOS | Mini CCD | Reference | Units | |
|---|---|---|---|---|---|
| Signal Intensity c-Si (519 cm−1) | 11,714 | 33,253 | 131,542 | e− | |
| Normalized signal intensity Si | 0.09 | 0.25 | 1 | ref | |
| Signal free noise sum a (1 s) | 57.23 | 12.81 | 7.04 | e−/px | |
| Signal shot noise | 122 | 183 | 363 | e−/px | |
| SNRc-Si | Calculated b | 96 | 182 | 363 | |
| Experimental c | 87 | 169 | 505 | ||
| Signal Intensity PP (2888 cm−1) | 1569 | 5590 | 10,129 | e− | |
| Normalized signal intensity PP | 0.15 | 0.55 | 1 | ref | |
| Signal free noise sum a (1 s) | 57.23 | 12.81 | 7.04 | e−/px | |
| Signal shot noise | 40 | 75 | 101 | e−/px | |
| SNRPP | Calculated b | 23 | 77 | 100 | |
| Experimental c | 24 | 71 | 141 | ||
| Signal Intensity (S) E. coli (2932 cm−1) | 6960 | 29,711 | 42,185 | e− | |
| Normalized signal intensity | 0.16 | 0.7 | 1 | ref | |
| Raman signal free noise sum a (15 s) | 95.03 | 74.54 | 86.40 | e−/px | |
| Signal shot noise | 83 | 172 | 205 | e−/px | |
| SNRE. coli | Calculated b | 55 | 158 | 185 | |
| Experimental c | N.A. d | N.A. d | N.A. d | ||
| (A) | |||||
| Predicted | Sensitivity (%) | Specificity (%) | |||
| True | E. coli | P. aeruginosa | S. aureus | ||
| E. coli | 75 | 0 | 0 | 100 | 99 |
| P. aeruginosa | 0 | 75 | 0 | 100 | 100 |
| S. aureus | 2 | 0 | 73 | 97.3 | 100 |
| (B) | |||||
| Predicted | Sensitivity (%) | Sensitivity (%) | |||
| True | E. coli | P. aeruginosa | S. aureus | ||
| E. coli | 55 | 0 | 0 | 100 | n/a |
| P. aeruginosa | 0 | 0 | 0 | n/a | 100 |
| S. aureus | 0 | 0 | 0 | n/a | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jahn, I.J.; Grjasnow, A.; John, H.; Weber, K.; Popp, J.; Hauswald, W. Noise Sources and Requirements for Confocal Raman Spectrometers in Biosensor Applications. Sensors 2021, 21, 5067. https://doi.org/10.3390/s21155067
Jahn IJ, Grjasnow A, John H, Weber K, Popp J, Hauswald W. Noise Sources and Requirements for Confocal Raman Spectrometers in Biosensor Applications. Sensors. 2021; 21(15):5067. https://doi.org/10.3390/s21155067
Chicago/Turabian StyleJahn, Izabella J., Alexej Grjasnow, Henry John, Karina Weber, Jürgen Popp, and Walter Hauswald. 2021. "Noise Sources and Requirements for Confocal Raman Spectrometers in Biosensor Applications" Sensors 21, no. 15: 5067. https://doi.org/10.3390/s21155067
APA StyleJahn, I. J., Grjasnow, A., John, H., Weber, K., Popp, J., & Hauswald, W. (2021). Noise Sources and Requirements for Confocal Raman Spectrometers in Biosensor Applications. Sensors, 21(15), 5067. https://doi.org/10.3390/s21155067






