3-D Image-Driven Morphological Crop Analysis: A Novel Method for Detection of Sunflower Broomrape Initial Subsoil Parasitism
Abstract
1. Introduction
2. Materials and Methods
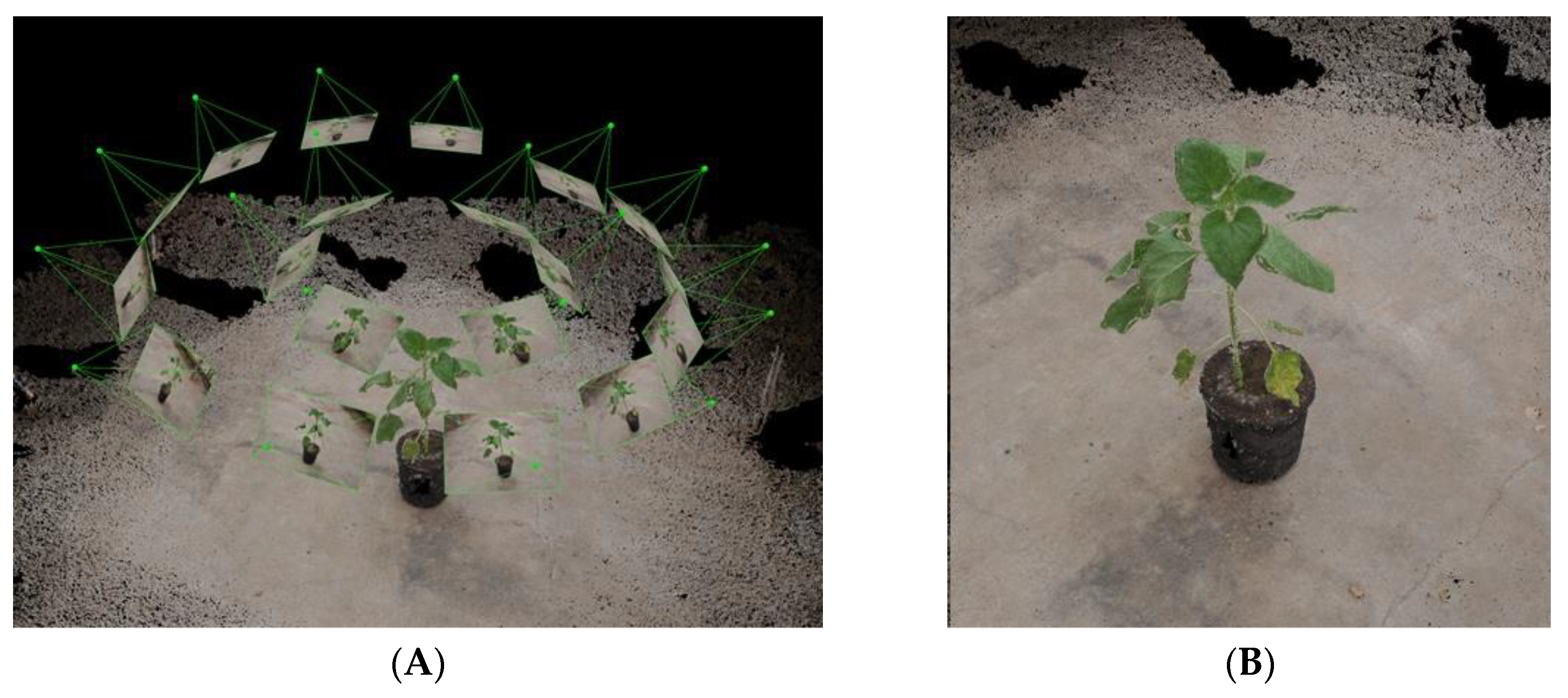
2.1. Image Acquisition and Analysis
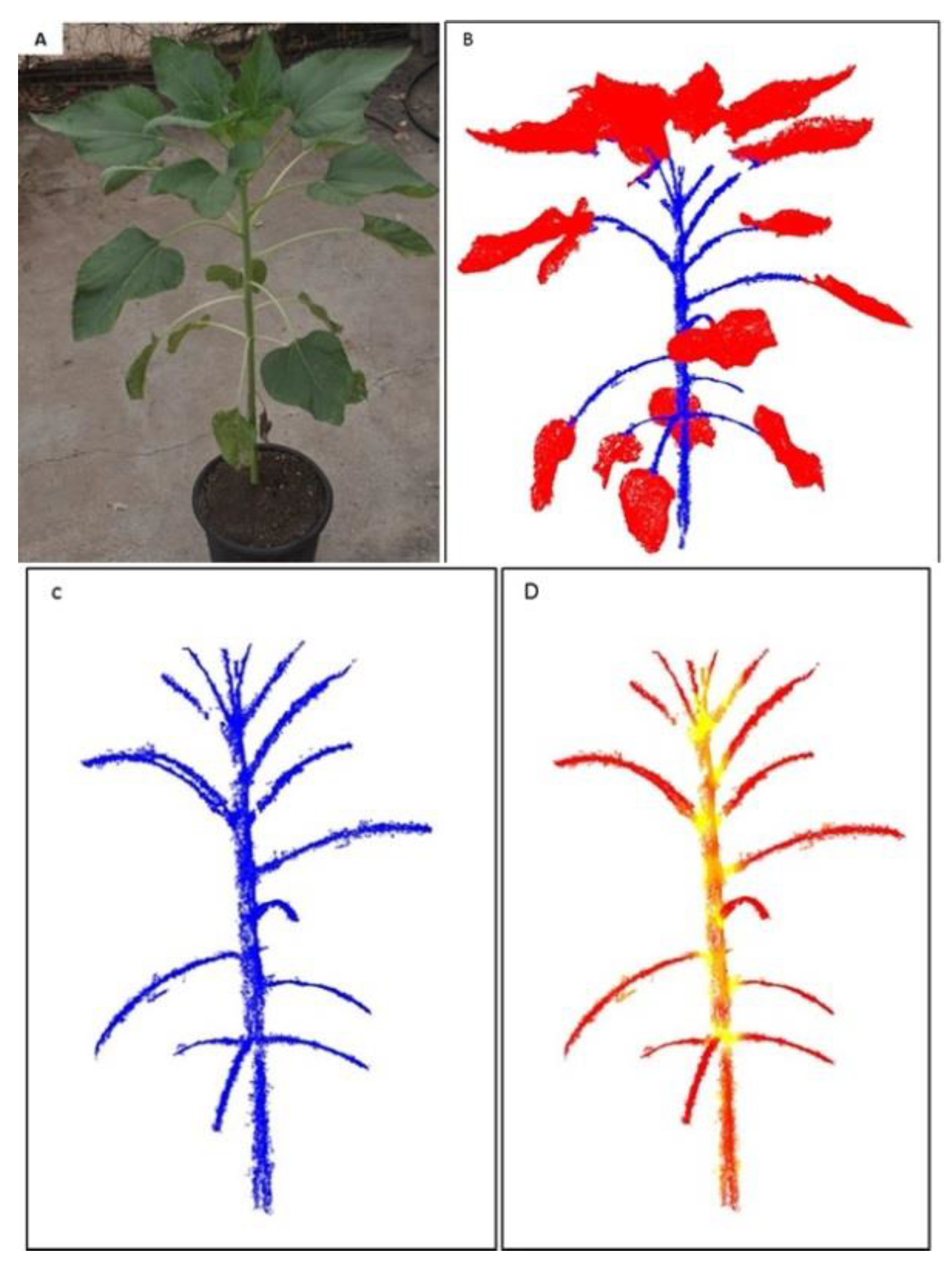
2.2. Plant Segmentation and Parameter Extraction
2.3. Minirhizotron Experiments
2.4. Statistical Analysis
3. Results
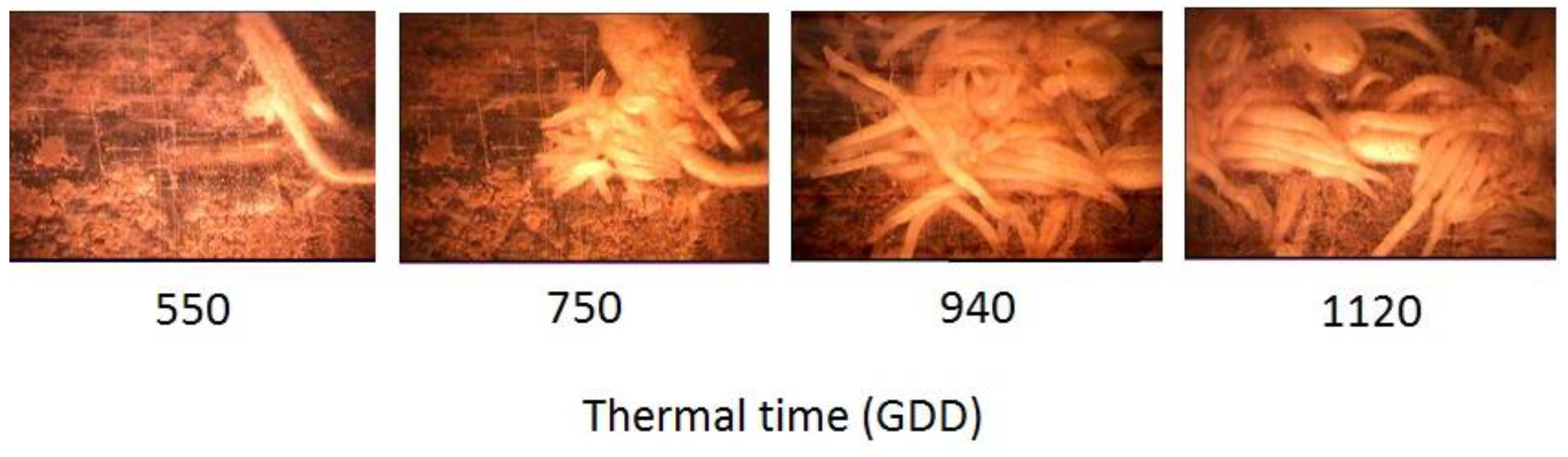
3.1. O. Cumana Parasitism Dynamics: Minirhizotron Experiments
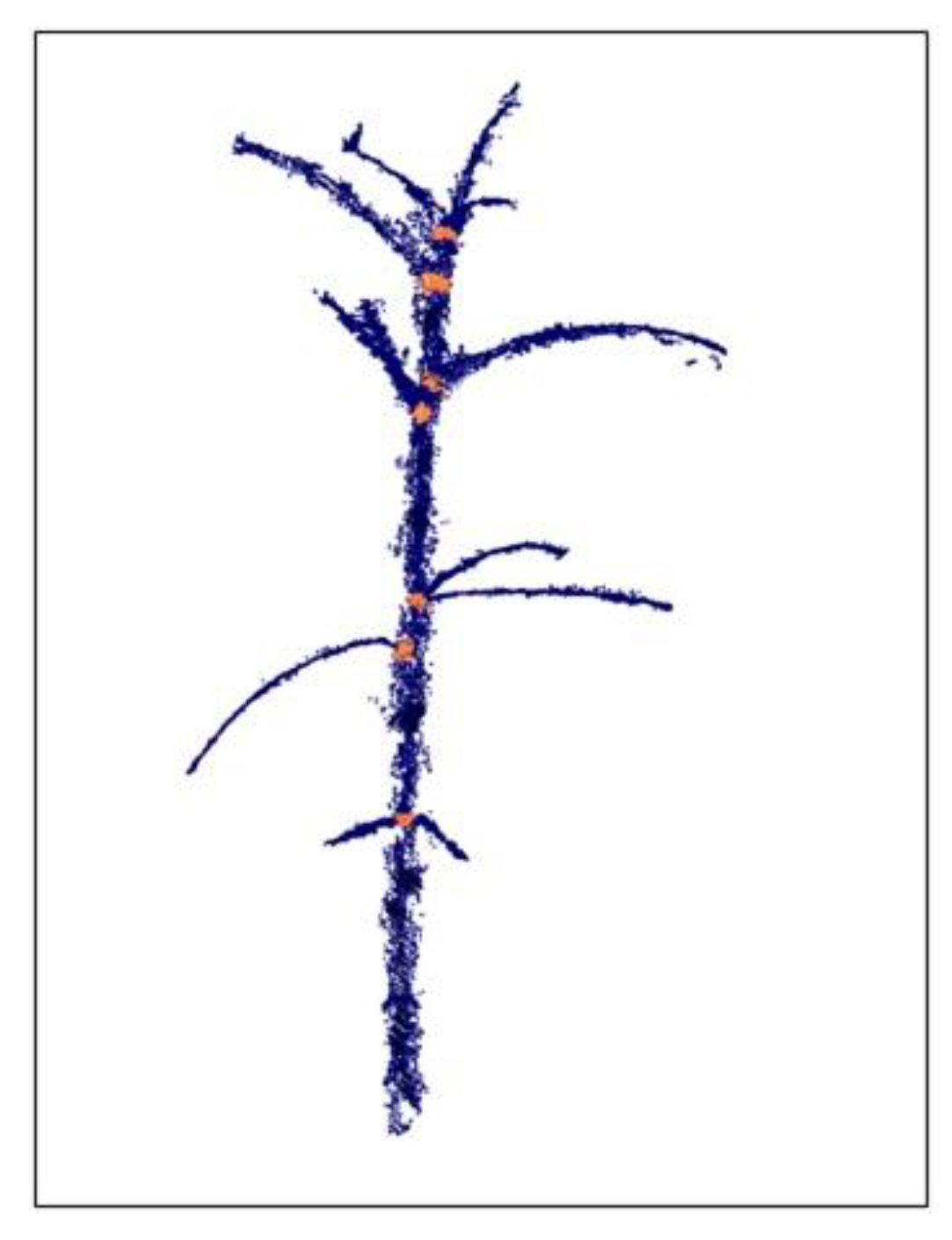
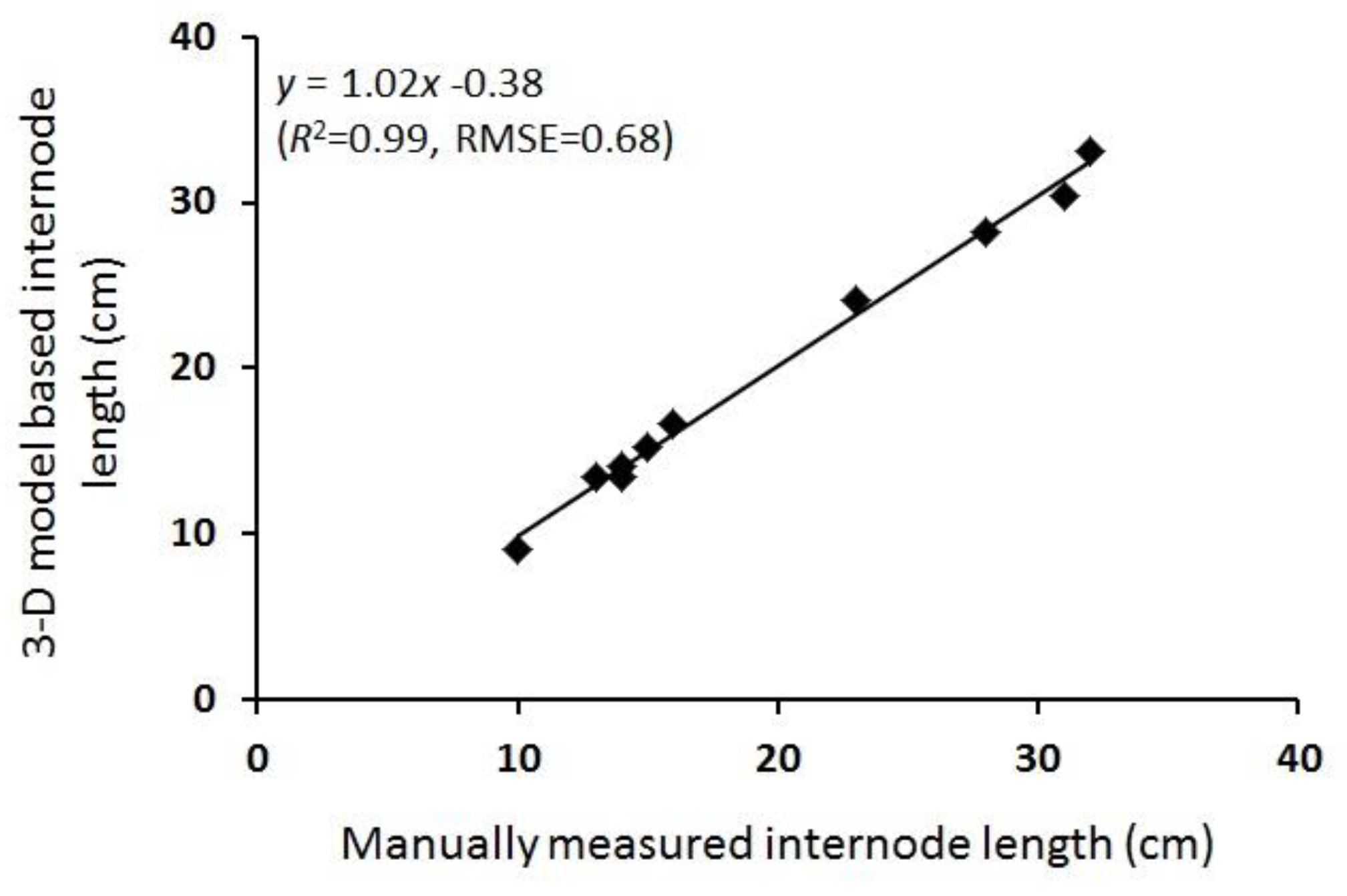
3.2. 3-D Reconstruction and Internode Estimation
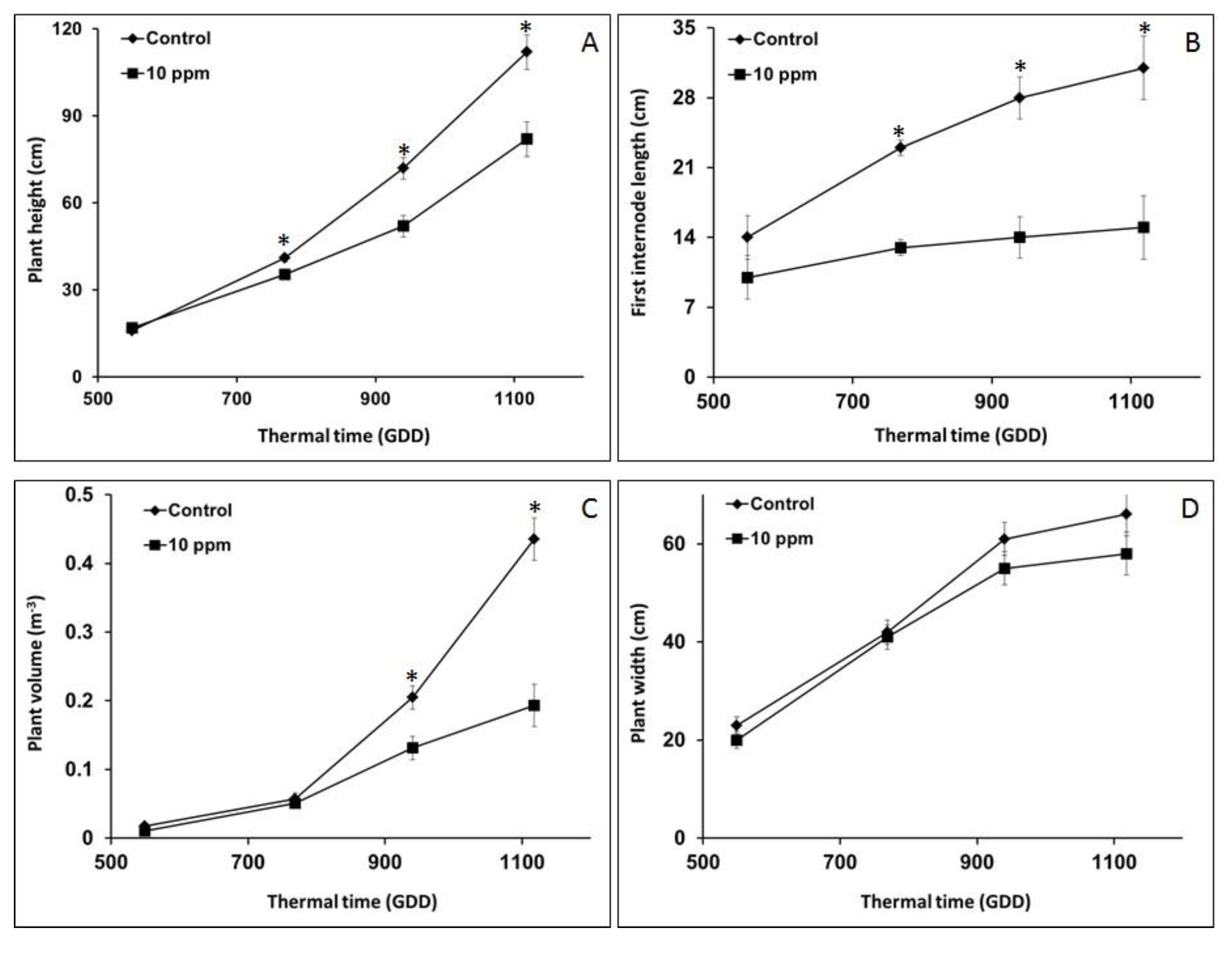
3.3. Morphological Analysis for O. Cumana Detection
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lati, R.N.; Filin, S.; Eizenberg, H. Plant growth parameter estimation from sparse 3D reconstruction based on highly-textured feature points. Precis. Agric. 2013, 14, 586–605. [Google Scholar] [CrossRef]
- Ephrath, J.E.; Eizenberg, H. Quantification of the dynamics of Orobanche cumana and Phelipanche aegyptiaca parasitism in confectionery sunflower. Weed Res. 2010, 50, 140–152. [Google Scholar] [CrossRef]
- Cochavi, A.; Achdari, G.; Smirnov, Y.; Rubin, B.; Eizenberg, H. Egyptian broomrape (Phelipanche aegyptiaca) management in carrot under field conditions. Weed Technol. 2015, 29, 519–528. [Google Scholar] [CrossRef]
- Aly, R.; Goldwasser, Y.; Eizenberg, H.; Hershenhorn, J.; Golan, S.; Kleifeld, Y. Broomrape (Orobanche cumana) control in sunflower (Helianthus annuus) with imazapic. Weed Technol. 2009, 15, 306–309. [Google Scholar] [CrossRef]
- Aly, R. Conventional and biotechnological approaches for control of parasitic weeds. In Vitro Cell. Dev. Biol. Plant 2007, 43, 304–317. [Google Scholar] [CrossRef]
- Eizenberg, H.; Hershenhorn, J.; Achdari, G.; Ephrath, J.E. A thermal time model for predicting parasitism of Orobanche cumana in irrigated sunflower-field validation. Field Crop. Res. 2012, 137, 49–55. [Google Scholar] [CrossRef]
- Eizenberg, H.; Aly, R.; Cohen, Y. Technologies for smart chemical control of broomrape (Orobanche spp. and Phelipanche spp.). Weed Sci. 2012, 60, 316–323. [Google Scholar] [CrossRef]
- Bajwa, A.A.; Mahajan, G.; Chauhan, B.S. Nonconventional weed management strategies for modern agriculture. Weed Sci. 2015, 63, 723–747. [Google Scholar] [CrossRef]
- Eizenberg, H.; Hershenhorn, J.; Ephrath, J.H.; Kanampiu, F. Chemical control. In Parasitic Orobanchaceae; Joel, D.M., Gressel, J., Musselman, L.J., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 415–432. [Google Scholar]
- Cochavi, A.; Rapaport, T.; Gendler, T.; Karnieli, A.; Eizenberg, H.; Rachmilevitch, S.; Ephrath, J.E. Recognition of orobanche cumana below-ground parasitism through physiological and hyper spectral measurements in sunflower (Helianthus annuus L.). Front. Plant Sci. 2017, 8, 909. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Bustos, C.M.; Pérez-Bueno, M.L.; Barón, M.; Molinero-Ruiz, L. Fluorescence imaging in the red and far-Rrd region during growth of sunflower plantlets. diagnosis of the early infection by the parasite Orobanche cumana. Front. Plant Sci. 2016, 7, 884. [Google Scholar] [CrossRef]
- Chaivivatrakul, S.; Tang, L.; Dailey, M.N.; Nakarmi, A.D. Automatic morphological trait characterization for corn plants via 3D holographic reconstruction. Comput. Electron. Agric. 2014, 109, 109–123. [Google Scholar] [CrossRef]
- Rose, C.J.; Paulus, S.; Kuhlmann, H. Accuracy analysis of a multi-view stereo approach for phenotyping of tomato plants at the organ level. Sensors 2015, 15, 9651–9665. [Google Scholar] [CrossRef]
- Paproki, A.; Sirault, X.; Berry, S.; Furbank, R.; Fripp, J. A novel mesh processing based technique for 3D plant analysis. BMC Plant Biol. 2012, 12, 63. [Google Scholar] [CrossRef]
- Paulus, S.; Dupuis, J.; Mahlein, A.-K.; Kuhlmann, H. Surface feature based classification of plant organs from 3D laserscanned point clouds for plant phenotyping. BMC Bioinform. 2013, 14, 238. [Google Scholar] [CrossRef]
- McCormick, R.F.; Truong, S.K.; Mullet, J.E. 3D sorghum reconstructions from depth images identify QTL regulating shoot architecture. Plant Physiol. 2016, 172, 823–834. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Slaughter, C.D.; Max, N.; Maloof, N.J.; Sinha, N. Structured light-based 3D reconstruction system for plants. Sensors 2015, 15, 18587–18612. [Google Scholar] [CrossRef]
- Salas Fernandez, M.G.; Bao, Y.; Tang, L.; Schnable, P.S. A high-throughput, field-based phenotyping technology for tall biomass crops. Plant Physiol. 2017, 174, 2008–2022. [Google Scholar] [CrossRef] [PubMed]
- Piron, A.; Leemans, V.; Lebeau, F.; Destain, M.F. Improving in-row weed detection in multispectral stereoscopic images. Comput. Electron. Agric. 2009, 69, 73–79. [Google Scholar] [CrossRef]
- Telem, G.; Filin, S. Photogrammetric modeling of the relative orientation in underwater environments. ISPRS J. Photogramm. Remote Sens. 2013, 86, 150–156. [Google Scholar] [CrossRef]
- Hosoi, F.; Omasa, K. Voxel-based 3-D modeling of individual trees for estimating leaf area density using high-resolution portable scanning lidar. IEEE Trans. Geosci. Remote Sens. 2006, 44, 3610–3618. [Google Scholar] [CrossRef]
- Elnashef, B.; Filin, S.; Lati, R.N. Tensor-based classification and segmentation of three-dimensional point clouds for organ-level plant phenotyping and growth analysis. Comput. Electron. Agric. 2019, 156, 51–61. [Google Scholar] [CrossRef]
- Eizenberg, H.; Shtienberg, D.; Silberbush, M.; Ephrath, J.E. A new method for in-situ monitoring of the underground development of Orobanche cumana in sunflower (Helianthus annuus) with a mini-rhizotron. Ann. Bot. 2005, 96, 1137–1140. [Google Scholar] [CrossRef] [PubMed]
- McMaster, G.S.; Wilhelm, W.W. Growing degree-days: One equation, two interpretations. Agric. For. Meteorol. 1997, 87, 291–300. [Google Scholar] [CrossRef]
- Simmons, A.M.; Yeargan, K.V.; Godfrey, L.D. Ovipositional sites of the potato leafhopper (Homoptera: Cicadellidae) on vegetative stage soybean plants. Environ. Entomol. 1985, 14, 165–169. [Google Scholar] [CrossRef]
- Peteinatos, G.G.; Weis, M.; Andújar, D.; Rueda Ayala, V.; Gerhards, R. Potential use of ground-based sensor technologies for weed detection. Pest Manag. Sci. 2014, 70, 190–199. [Google Scholar] [CrossRef] [PubMed]
- Hatfield, J.L.; Prueger, J.H. Temperature extremes: Effect on plant growth and development. Weather Clim. Extrem. 2015, 10, 4–10. [Google Scholar] [CrossRef]
- Liu, R.; Dai, M.; Wu, X.; Li, M.; Liu, X. Suppression of the root-knot nematode [Meloidogyne incognita (Kofoid & White) Chitwood] on tomato by dual inoculation with arbuscular mycorrhizal fungi and plant growth-promoting rhizobacteria. Mycorrhiza 2012, 22, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Guo, W.; Ninomiya, S. Node detection and internode length estimation of tomato seedlings based on image analysis and machine learning. Sensors 2016, 16, 1044. [Google Scholar] [CrossRef]
- Brichet, N.; Fournier, C.; Turc, O.; Strauss, O.; Artzet, S.; Pradal, C.; Welcker, C.; Tardieu, F.; Cabrera-Bosquet, L. A robot-assisted imaging pipeline for tracking the growths of maize ear and silks in a high-throughput phenotyping platform. Plant Methods 2017, 13, 96. [Google Scholar] [CrossRef]
- Surov, T.; Aviv, D.; Aly, R.; Joel, D.M.; Goldman-Guez, T.; Gressel, J. Generation of transgenic asulam-resistant potatoes to facilitate eradication of parasitic broomrapes (Orobanche spp.), with the sul gene as the selectable marker. Theor. Appl. Genet. 1998, 96, 132–137. [Google Scholar] [CrossRef]
- Tan, S.; Evans, R.R.; Dahmer, M.L.; Singh, B.K.; Shaner, D.L. Imidazolinone-tolerant crops: History, current status and future. Pest Manag. Sci. 2005, 61, 246–257. [Google Scholar] [CrossRef]
- Lin, Y. LiDAR: An important tool for next-generation phenotyping technology of high potential for plant phenomics? Comput. Electron. Agric. 2015, 119, 61–73. [Google Scholar] [CrossRef]
- Long, D.S.; McCallum, J.D. Mapping straw yield using on-combine light detection and ranging (lidar). Int. J. Remote Sens. 2013, 34, 6121–6134. [Google Scholar] [CrossRef]
- Zhang, L.; Grift, T.E. A LIDAR-based crop height measurement system for Miscanthus giganteus. Comput. Electron. Agric. 2012, 85, 70–76. [Google Scholar] [CrossRef]
- Fricke, T.; Wachendorf, M. Combining ultrasonic sward height and spectral signatures to assess the biomass of legume-grass swards. Comput. Electron. Agric. 2013, 99, 236–247. [Google Scholar] [CrossRef]
- Pittman, J.J.; Arnall, D.B.; Interrante, S.M.; Moffet, C.A.; Butler, T.J. Estimation of biomass and canopy height in Bermudagrass, Alfalfa, and wheat using ultrasonic, laser, and spectral sensors. Sensors 2015, 15, 2920–2943. [Google Scholar] [CrossRef]
- Paulus, S.; Dupuis, J.; Riedel, S.; Kuhlmann, H. Automated analysis of barley organs using 3D laser scanning: An approach for high throughput phenotyping. Sensors 2014, 14, 12670–12686. [Google Scholar] [CrossRef]
- Duan, T.; Chapman, S.C.; Holland, E.; Rebetzke, G.J.; Guo, Y.; Zheng, B. Dynamic quantification of canopy structure to characterize early plant vigour in wheat genotypes. J. Exp. Bot. 2016, 67, 4523–4534. [Google Scholar] [CrossRef]
- Paulus, S.; Schumann, H.; Kuhlmann, H.; Léon, J. High-precision laser scanning system for capturing 3D plant architecture and analysing growth of cereal plants. Biosyst. Eng. 2014, 121, 1–11. [Google Scholar] [CrossRef]
- Nguyen, T.T.; Slaughter, D.C.; Maloof, J.N.; Sinha, N. Plant phenotyping using multi-view stereo vision with structured lights. In Proceedings of the Autonomous Air and Ground Sensing Systems for Agricultural Optimization and Phenotyping (SPIE 9866), Baltimore, MD, USA, 17–21 April 2016; p. 986608. [Google Scholar] [CrossRef]
- Oveisi, M.; Yousefi, A.R.; Gonzalez-Andujar, J.L. Spatial distribution and temporal stability of crenate broomrape (Orobanche crenata Forsk) in faba bean (Vicia faba L.): A long-term study at two localities. Crop Prot. 2010, 29, 717–720. [Google Scholar] [CrossRef]
- Jay, S.; Rabatel, G.; Hadoux, X.; Moura, D.; Gorretta, N. In-field crop row phenotyping from 3D modeling performed using structure from motion. Comput. Electron. Agric. 2015, 110, 70–77. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, C.; Paterson, A.H. High throughput phenotyping of cotton plant height using depth images under field conditions. Comput. Electron. Agric. 2016, 130, 57–68. [Google Scholar] [CrossRef]
- Kise, M.; Zhang, Q. Development of a stereovision sensing system for 3D crop row structure mapping and tractor guidance. Biosyst. Eng. 2008, 101, 191–198. [Google Scholar] [CrossRef]
- Andújar, D.; Fernández-Quintanilla, C.; Dorado, J. Matching the best viewing angle in depth cameras for biomass estimation based on poplar seedling geometry. Sensors 2015, 15, 12999–13011. [Google Scholar] [CrossRef]
- Nakarmi, A.D.; Tang, L. Automatic inter-plant spacing sensing at early growth stages using a 3D vision sensor. Comput. Electron. Agric. 2012, 82, 23–31. [Google Scholar] [CrossRef]






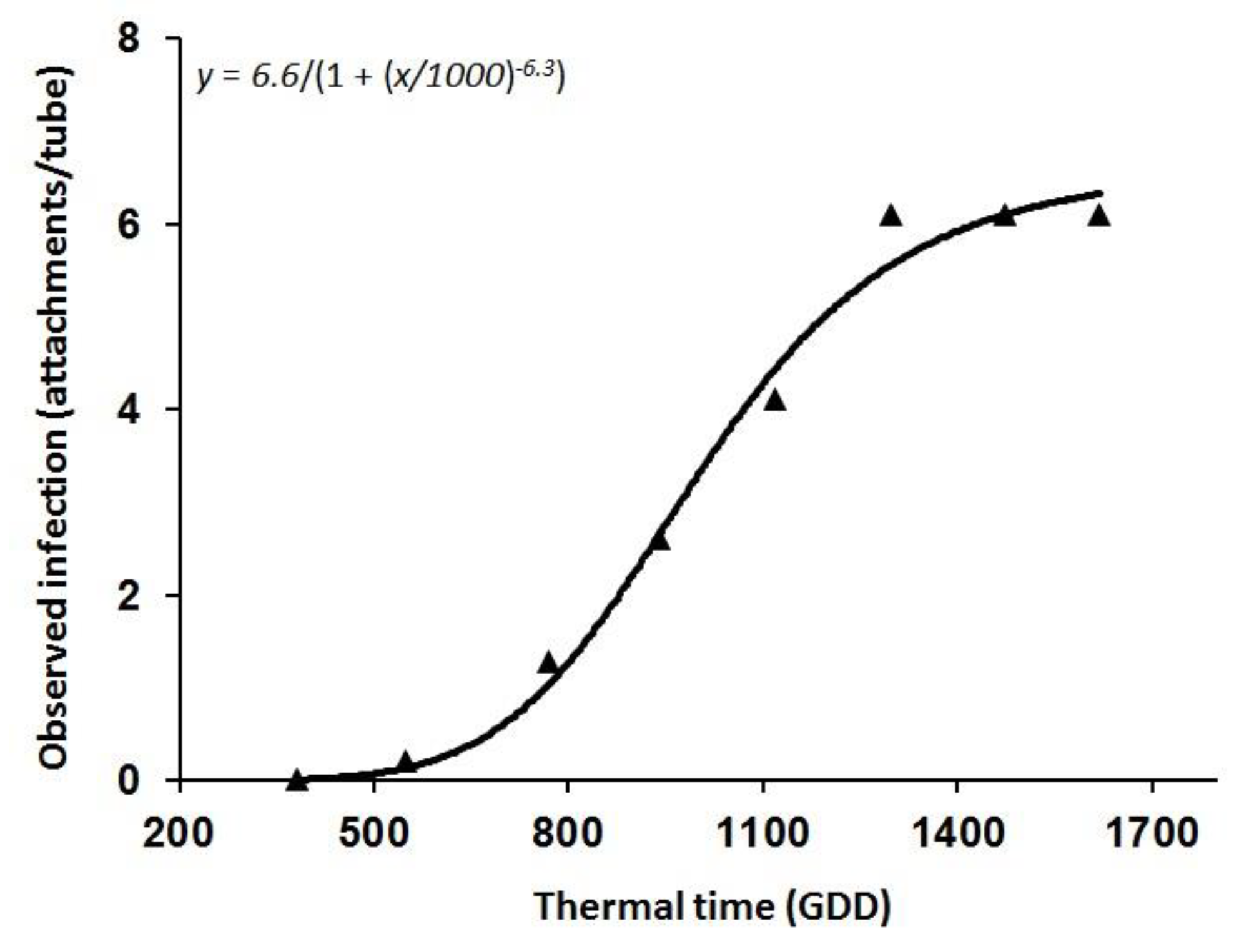

| Parameter | Estimate | SE |
|---|---|---|
| a | 6.62 | 0.42 |
| x0 | 1000.18 | 34.69 |
| b | −6.37 | 1.17 |
| R2 | 0.98 | |
| p | <0.0001 | |
| RMSE | 0.32 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lati, R.N.; Filin, S.; Elnashef, B.; Eizenberg, H. 3-D Image-Driven Morphological Crop Analysis: A Novel Method for Detection of Sunflower Broomrape Initial Subsoil Parasitism. Sensors 2019, 19, 1569. https://doi.org/10.3390/s19071569
Lati RN, Filin S, Elnashef B, Eizenberg H. 3-D Image-Driven Morphological Crop Analysis: A Novel Method for Detection of Sunflower Broomrape Initial Subsoil Parasitism. Sensors. 2019; 19(7):1569. https://doi.org/10.3390/s19071569
Chicago/Turabian StyleLati, Ran Nisim, Sagi Filin, Bashar Elnashef, and Hanan Eizenberg. 2019. "3-D Image-Driven Morphological Crop Analysis: A Novel Method for Detection of Sunflower Broomrape Initial Subsoil Parasitism" Sensors 19, no. 7: 1569. https://doi.org/10.3390/s19071569
APA StyleLati, R. N., Filin, S., Elnashef, B., & Eizenberg, H. (2019). 3-D Image-Driven Morphological Crop Analysis: A Novel Method for Detection of Sunflower Broomrape Initial Subsoil Parasitism. Sensors, 19(7), 1569. https://doi.org/10.3390/s19071569





