Single-Step Incubation Determination of miRNAs in Cancer Cells Using an Amperometric Biosensor Based on Competitive Hybridization onto Magnetic Beads
Abstract
:1. Introduction
2. Experimental
2.1. Apparatus and Electrodes
2.2. Reagents and Solutions
2.3. MBs Modification
2.4. Electrochemical Measurements
2.5. Cultured Cells and RNAt Extraction
3. Results and Discussion
- -
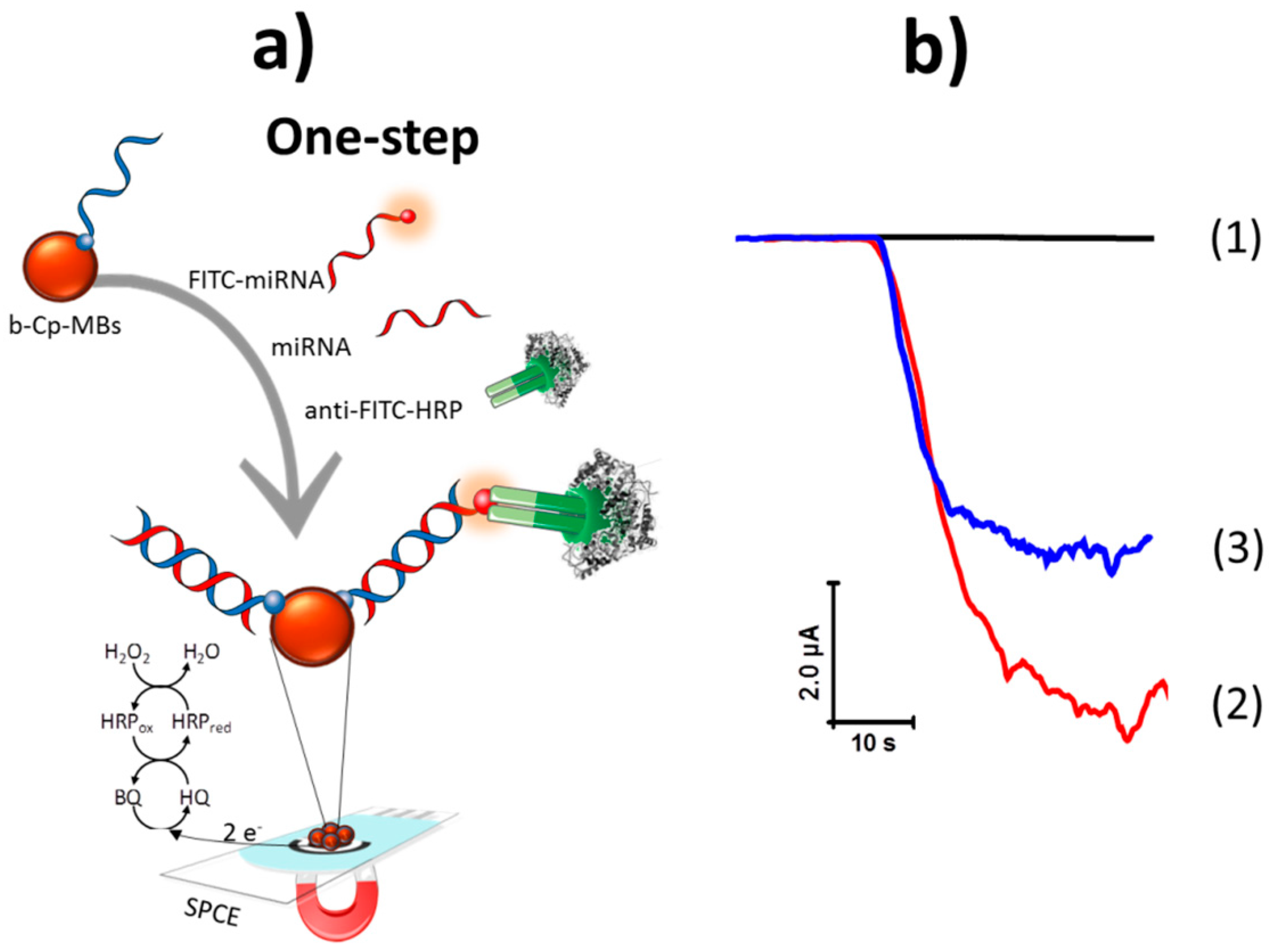
- Protocol 1, single step: incubating the Strep-MBs with a mixture solution containing b-Cp, target miRNA-21, FITC-miRNA, and anti-FITC-HRP.
- -
- Protocol 2, 2 steps: (1) immobilization of the b-Cp onto the Strep-MBs and (2) incubation of the b-Cp-MBs with a mixture solution containing target miRNA-21, FITC-miRNA, and anti-FITC-HRP.
- -
- Protocol 3A, 3 steps: (1) immobilization of the b-Cp onto the Strep-MBs, (2) hybridization of the miRNA-21 onto the b-Cp-MBs, and (3) incubation of the miRNA-21/b-Cp-MBs with a mixture solution containing FITC-miRNA and anti-FITC-HRP.
- -
- Protocol 3B, 3 steps: (1) immobilization of the b-Cp onto the Strep-MBs, (2) incubation of the b-Cp-MBs with a mixture solution containing target miRNA-21 and FITC-miRNA, and (3) labeling of the FITC-miRNA attached to the b-Cp-MBs with the anti-FITC-HRP.
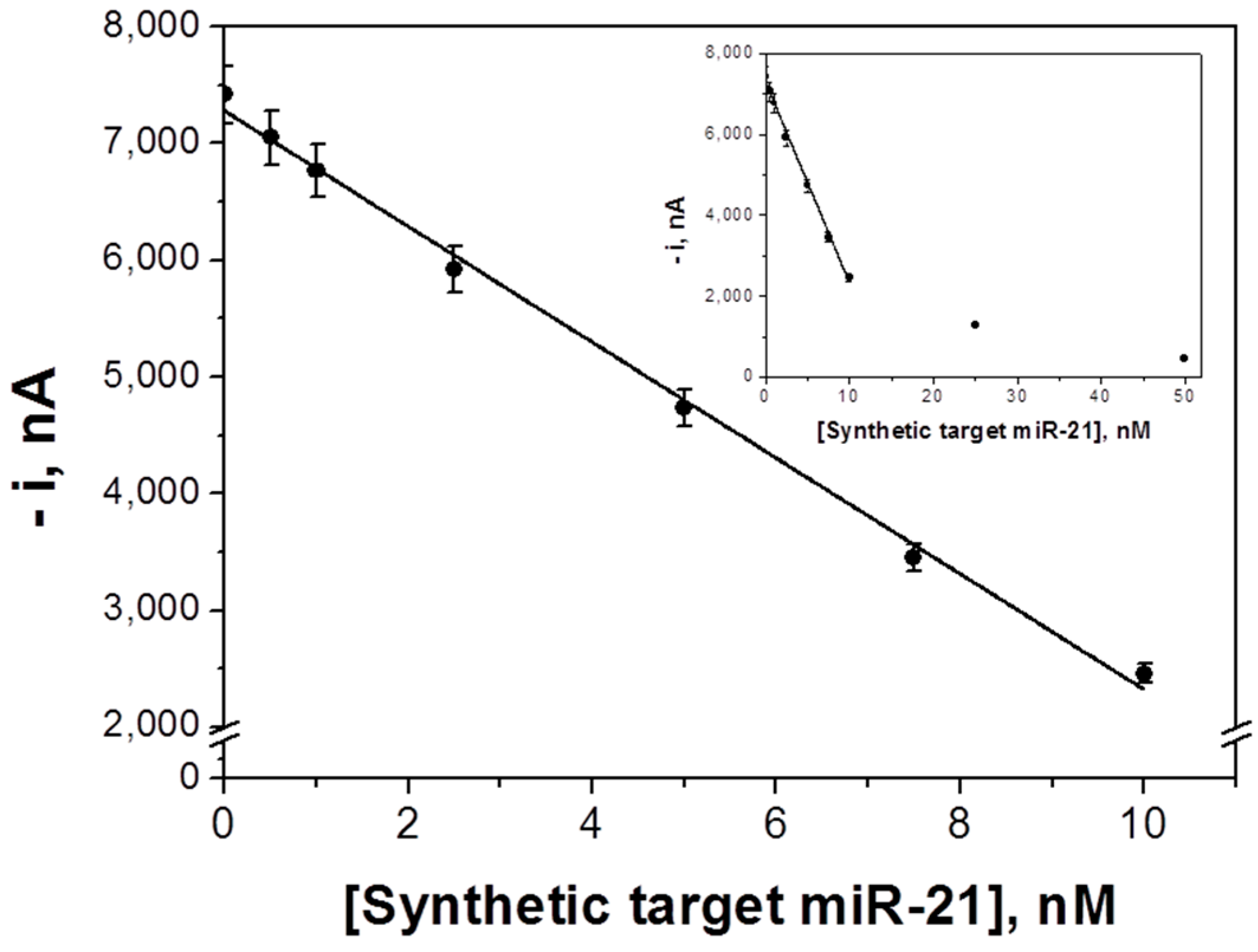
3.1. Analytical Characteristics
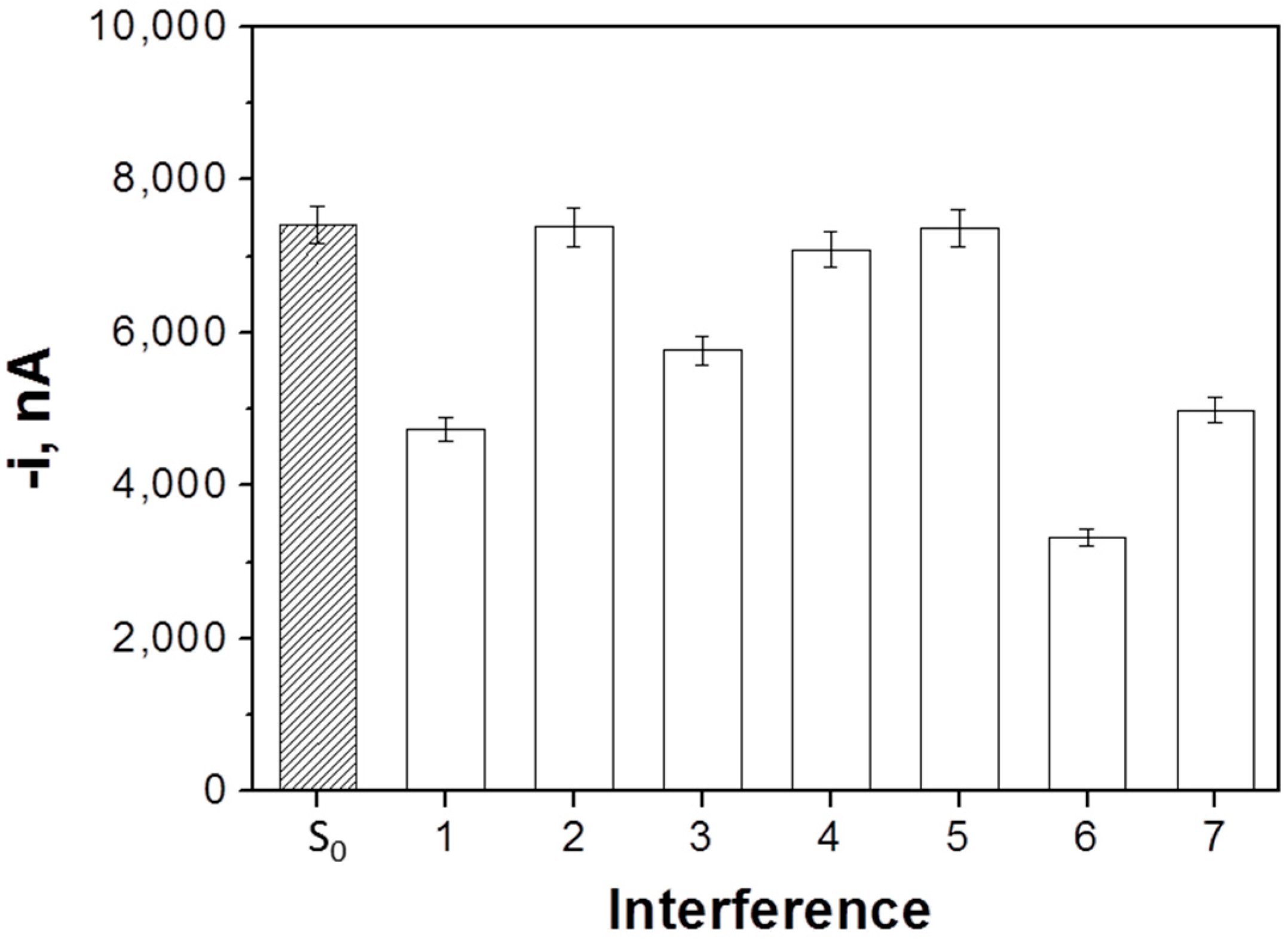
3.2. Selectivity
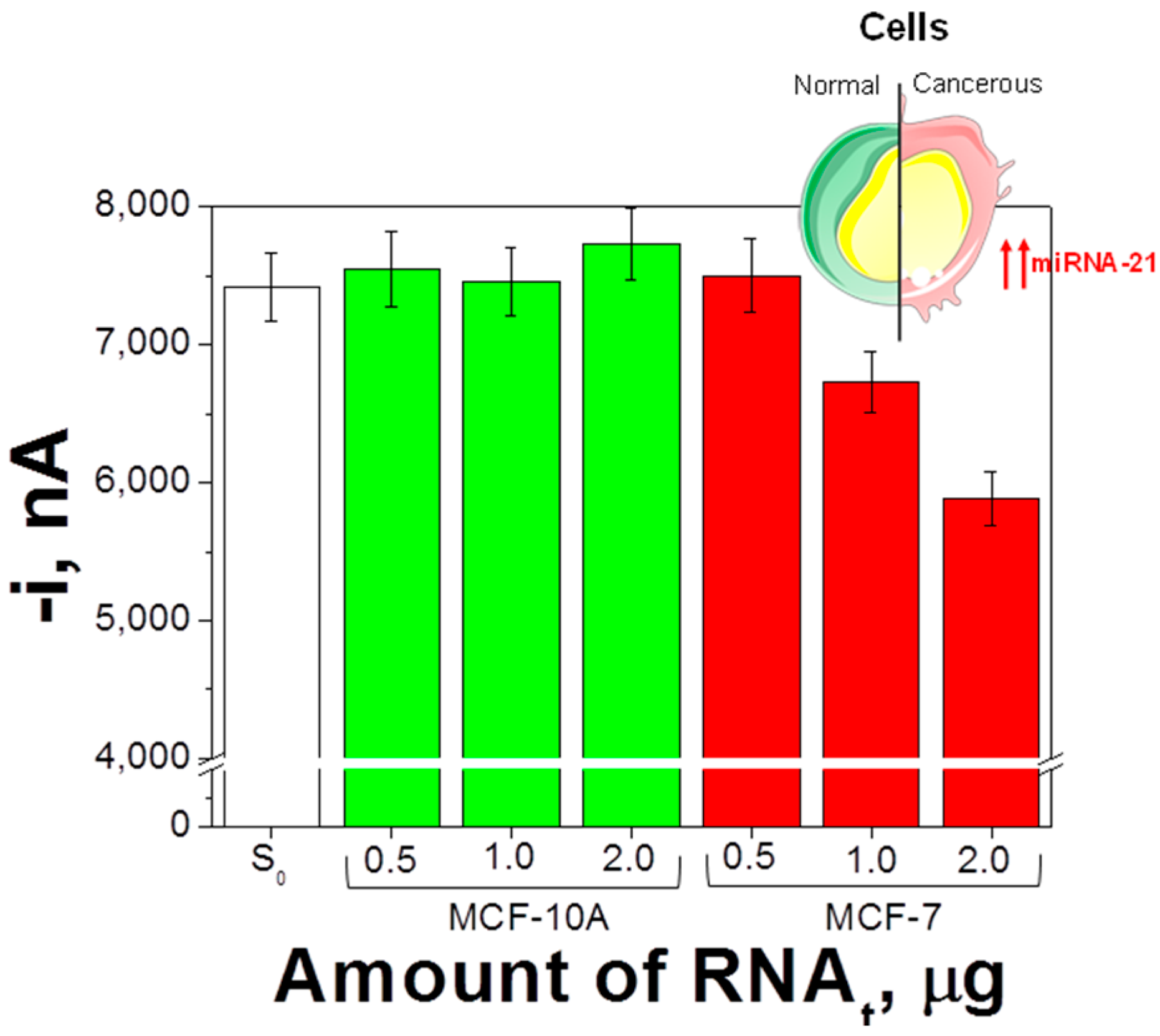
3.3. Determination of Mature miRNA-21 in RNAt Extracted from Cancer Cells
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hamam, R.; Hamam, D.; Alsaleh, K.A.; Kassem, M.; Zaher, W.; Alfayez, M.; Aldahmash, A.; Alajez, N.M. Circulating microRNAs in breast cancer: Novel diagnostic and prognostic biomarkers. Cell Death Dis. 2017, 8, e3045. [Google Scholar] [CrossRef] [PubMed]
- Campuzano, S.; Pedrero, M.; Pingarrón, J.M. Non-Invasive Breast Cancer Diagnosis through Electrochemical Biosensing at Different Molecular Levels. Sensors 2017, 17, 1993. [Google Scholar] [CrossRef] [PubMed]
- Nicolini, A.; Ferrari, P.; Duffy, M.J. Prognostic and predictive biomarkers in breast cancer: Past, present and future. Semin. Cancer Biol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-X.; Huang, K.-J.; Niu, K.-X. Recent advances in signal amplification strategy based on oligonucleotide and nanomaterials for microRNA detection-a review. Biosens. Bioelectron. 2018, 99, 612–624. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.W.; Chen, L.; Man, Y.-G. miRNA Biomarkers in Breast Cancer Detection and Management. J. Cancer 2011, 2, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Hasanzadeh, M.; Shadjou, N.; de la Guardia, M. Early stage screening of breast cancer using electrochemical biomarker detection. Trends Anal. Chem. 2017, 91, 67–76. [Google Scholar] [CrossRef]
- Cissell, K.A.; Rahimi, Y.; Shrestha, S.; Hunt, E.A.; Deo, S.K. Bioluminescence-Based Detection of MicroRNA, miR21 in Breast Cancer Cells. Anal. Chem. 2008, 80, 2319–2325. [Google Scholar] [CrossRef] [PubMed]
- Šípova, H.; Zhang, S.; Dudley, A.M.; Galas, D.; Wang, K.; Homola, J. Surface plasmon resonance biosensor for rapid label-free detection of microribonucleic acid at subfemtomole level. Anal. Chem. 2010, 82, 10110–10115. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, F. Label-free and enzyme-free homogeneous electrochemical biosensing strategy based on hybridization chain reaction: A facile, sensitive, and highly specific microRNA assay. Anal. Chem. 2015, 87, 11368–11374. [Google Scholar]
- Liao, Y.; Fu, Y.; Wu, Y.; Huang, R.; Zhou, X.; Xing, D. Ultrasensitive detection of microRNA in tumor cells and tissues via continuous assembly of DNA probe. Biomacromolecules 2015, 16, 3543–3551. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wu, D.-Z.; Cai, S.-X.; Chen, M.; Xia, Y.-K.; Wu, F.; Chen, J.-H. An immobilization-free electrochemical impedance biosensor based on duplex-specific nuclease assisted target recycling for amplified detection of microRNA. Biosens. Bioelectron. 2016, 75, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Kilic, T.; Erdem, A.; Ozsoz, M.; Carrara, S. MicroRNA biosensors: Opportunities and challenges among conventional and commercially available techniques. Biosens. Bioelectron. 2018, 99, 525–546. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.T.; Baldwin, D.A.; Scearce, L.M.; Oberholtzer, J.C.; Tobias, J.W.; Mourelatos, Z. Microarray-based, high-throughput gene expression profiling of microRNAs. Nat. Methods 2004, 1, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Liang, R.-Q.; Li, W.; Li, Y.; Tan, C.; Li, J.X.; Jin, Y.X.; Ruan, K.C. An oligonucleotide microarray for microRNA expression analysis based on labeling RNA with quantum dot and nanogold probe. Nucleic Acids Res. 2005, 33, e17. [Google Scholar] [CrossRef] [PubMed]
- Git, A.; Dvinge, H.; Salmon-Divon, M.; Osborne, M.; Kutter, C.; Hadfield, J.; Bertone, P.; Caldas, C. Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA 2010, 16, 991–1006. [Google Scholar] [CrossRef] [PubMed]
- Clancy, E.; Burke, M.; Arabkari, V.; Barry, T.; Kelly, H.; Dwyer, R.M.; Kerin, M.J.; Smith, T.J. Amplification-free detection of microRNAs via a rapid microarray-based sandwich assay. Anal. Bioanal. Chem. 2017, 409, 3497–3505. [Google Scholar] [CrossRef] [PubMed]
- Válóczi, A.; Hornyik, C.; Varga, N.; Burgyán, J.; Kauppinen, S.; Havelda, Z. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 2004, 32, e175. [Google Scholar] [CrossRef] [PubMed]
- Pall, G.S.; Codony-Servat, C.; Byrne, J.; Ritchie, L.; Hamilton, A. Carbodiimide-mediated cross-linking of RNA to nylon membranes improves the detection of siRNA, miRNA and piRNA by northern blot. Nucleic Acids Res. 2007, 35, e60. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yao, B.; Huang, H.; Wang, Z.; Sun, C.; Fan, Y.; Chang, Q.; Li, S.; Wang, X.; Xi, J. Real-time polymerase chain reaction microRNA detection based on enzymatic stem-loop probes ligation. Anal. Chem. 2009, 81, 5446–5451. [Google Scholar] [CrossRef] [PubMed]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Centi, S.; Laschi, S.; Frànek, M.; Mascini, M. A disposable immunomagnetic electrochemical sensor based on functionalized magnetic beads and carbon-based screen-printed electrodes (SPCEs) for the detection of polychlorinated biphenyls (PCBs). Anal. Chim. Acta 2005, 538, 205–212. [Google Scholar] [CrossRef]
- Zacco, E.; Adrian, J.; Galve, R.; Marco, M.P.; Alegret, S.; Pividori, M.I. Electrochemical magneto immunosensing of antibiotic residues in milk. Biosens. Bioelectron. 2007, 22, 2184–2191. [Google Scholar] [CrossRef] [PubMed]
- Ricci, F.; Volpe, G.; Micheli, L.; Palleschi, G. A review on novel developments and applications of immunosensors in food analysis. Anal. Chim. Acta 2007, 605, 111–129. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wang, E. Electrochemical biosensors based on magnetic micro/nano particles. Electrochim. Acta 2012, 84, 62–73. [Google Scholar] [CrossRef]
- Campuzano, S.; Torrente-Rodríguez, R.M.; López-Hernández, E.; Conzuelo, F.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Magnetobiosensors based on viral protein p19 for microRNA determination in cancer cells and tissues. Angew. Chem. Int. Ed. 2014, 53, 6168–6171. [Google Scholar] [CrossRef] [PubMed]
- Torrente-Rodríguez, R.M.; Campuzano, S.; López-Hernández, E.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Direct determination of miR-21 in total RNA extracted from breast cancer samples using magnetosensing platforms and the p19 viral protein as detector bioreceptor. Electroanalysis 2014, 26, 2080–2087. [Google Scholar] [CrossRef]
- Li, C.; Liu, Z.; Cai, S.; Wen, F.; Wu, D.; Liu, Y.; Wu, F.; Lan, J.; Han, Z.; Chen, J. An electrochemical microRNA biosensor based on protein p19 combining an acridone derivate as indicator and DNA concatamers for signal amplification. Electrochem. Commun. 2015, 60, 185–189. [Google Scholar] [CrossRef]
- Torrente-Rodríguez, R.M.; Ruiz-Valdepeñas Montiel, V.; Campuzano, S.; Fachardo-Dinia, M.; Barderas, R.; San Segundo-Acosta, P.; Montoya, J.J.; Pingarrón, J.M. Fast electrochemical miRNAs determination in cancer cells and tumor tissues with antibody-functionalized magnetic microcarriers. ACS Sens. 2016, 1, 896–903. [Google Scholar] [CrossRef]
- Torrente-Rodríguez, R.M.; Campuzano, S.; Ruiz-Valdepeñas Montiel, V.; Sagrera, A.; Domínguez-Cañete, J.J.; Vargas, E.; Montoya, J.J.; Granados, R.; Sánchez-Puelles, J.M.; Pingarrón, J.M. Electrochemical miRNAs Determination in Formalin-Fixed, Paraffin-Embedded Breast Tumor Tissues Association with HER2 Expression. JSM Biotechnol. Bioeng. 2016, 3, 1064. [Google Scholar]
- Vargas, E.; Torrente-Rodríguez, R.M.; Ruiz-Valdepeñas Montiel, V.; Povedano, E.; Pedrero, M.; Montoya, J.J.; Campuzano, S.; Pingarrón, J.M. Magnetic Beads-Based Sensor with tailored sensitivity for rapid and single-step amperometric determination of miRNAs. Int. J. Mol. Sci. 2017, 18, 2151. [Google Scholar] [CrossRef] [PubMed]
- Torrente-Rodríguez, R.M.; Campuzano, S.; Ruiz-Valdepeñas Montiel, V.; Montoya, J.J.; Pingarrón, J.M. Sensitive electrochemical determination of miRNAs based on a sandwich assay onto magnetic microcarriers and hybridization chain reaction amplification. Biosens. Bioelectron. 2016, 86, 516–521. [Google Scholar] [CrossRef] [PubMed]
- Gangwar, R.S.; Rajagopalan, S.; Natarajan, R.; Deiuliis, J.A. Noncoding RNAs in Cardiovascular Disease: Pathological Relevance and Emerging Role as Biomarkers and Therapeutics. Am. J. Hypertens. 2018, 31, 150–165. [Google Scholar] [CrossRef] [PubMed]
- Gamella, M.; Campuzano, S.; Conzuelo, F.; Reviejo, A.J.; Pingarrón, J.M. Amperometric Magnetoimmunosensors for Direct Determination of D-Dimer in Human Serum. Electroanalysis 2012, 24, 2235–2243. [Google Scholar] [CrossRef]
- Wang, W.; Kong, T.; Zhang, D.; Zhang, J.; Cheng, G. Label-Free MicroRNA Detection Based on Fluorescence Quenching of Gold Nanoparticles with a Competitive Hybridization. Anal. Chem. 2015, 87, 10822–10829. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Peng, J.; Wang, J.; Tang, H.; Tan, L.; Xie, Q.; Yao, S. Carbon nanotube-based label-free electrochemical biosensor for sensitive detection of miRNA-24. Biosens. Bioelectron. 2014, 54, 158–164. [Google Scholar] [CrossRef] [PubMed]
- Markou, A.; Yousef, G.M.; Stathopoulos, E.; Georgoulias, V.; Lianidou, E. Prognostic significance of metastasis-related microRNAs in early breast cancer patients with a long follow-up. Clin. Chem. 2014, 60, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, B.S.; Balslev, E.; Poulsen, T.S.; Nielsen, D.; Møller, T.; Mortensen, C.E.; Holmstrøm, K.; Høgdall, E. miR-21 expression in cancer cells may not predict resistance to adjuvant trastuzumab in primary breast cancer. Front. Oncol. 2014, 4, 207. [Google Scholar] [CrossRef] [PubMed]
- De Mattos-Arruda, L.; Bottai, G.; Nuciforo, P.G.; Di Tommaso, L.; Giovannetti, E.; Peg, V.; Losurdo, A.; Pérez-Garcia, J.; Masci, G.; Corsi, F.; et al. MicroRNA-21 links epithelial-to-mesenchymal transition and inflammatory signals to confer resistance to neoadjuvant trastuzumab and chemotherapy in HER2-positive breast cancer patients. Oncotarget 2015, 6, 37269–37280. [Google Scholar] [CrossRef] [PubMed]
- Kurozumi, S.; Yamaguchi, Y.; Kurosumi, M.; Ohira, M.; Matsumoto, H.; Horiguchi, J. Recent trends in microRNA research into breast cancer with particular focus on the associations between microRNAs and intrinsic subtypes. J. Hum. Genet. 2017, 62, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.X.; Wu, Q.N.; Zhang, Y.; Li, Y.Y.; Liao, D.Z.; Hou, J.H.; Fu, J.; Zeng, M.S.; Yun, J.P.; Wu, Q.L.; et al. Knockdown of miR-21 in human breast cancer cell lines inhibits proliferation, In Vitro migration and In Vivo tumor growth. Breast Cancer Res. 2011, 13, R2. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Yang, T.; Chen, Y. Quantification of microRNA by DNA−peptide probe and liquid chromatography−tandem mass spectrometry-based quasi-targeted proteomics. Anal. Chem. 2016, 88, 754–763. [Google Scholar] [CrossRef] [PubMed]
- Novara, C.; Chiadò, A.; Paccotti, N.; Catuogno, S.; Esposito, C.L.; Condorelli, G.; De Franciscis, V.; Geobaldo, F.; Rivolo, P.; Giorgis, F. SERS-active metal-dielectric nanostructures integrated in microfluidic devices for label-free quantitative detection of miRNA. Faraday Discuss. 2017, 205, 271–289. [Google Scholar] [CrossRef] [PubMed]

| Oligonucleotide | Sequence (5′→3′) |
|---|---|
| Biotinylated capture probe (b-Cp) | TCAACATCAGTCTGATAAGCTA-Biotin |
| Target miRNA-21 | UAGCUUAUCAGACUGAUGUUGA |
| FITC-modified miRNA (FITC-miRNA) | UAGCUUAUCAGACUGAUGUUGA-FITC |
| 1-mismatched in central position (1-m(c)) | UAGCUUAUCAAACUGAUGUUGA |
| 1-mismatched in terminal position (1-m(t)) | UAGCUUAUCAGACUGAUGUUGG |
| Non-complementary 1 (NC1, miRNA-205) | UCCUUCAUUCCACCGGAGUCU |
| Non-complementary 2 (NC2, miRNA-122) | UGGAGUGUGACAAUGGUGUUUG |
| Experimental Variable | Tested Range | Selected Value |
|---|---|---|
| Strep-MBs, μL | 0.25–5.0 | 0.5 |
| (b-Cp), nM | 0.5–50.0 | 2.5 |
| Incubation time with b-Cp, min | 15–60 | 15 |
| (FITC-miRNA), nM | 0.25–5.0 | 2.5 |
| anti-FITC-HRP | 1/5000–1/250 | 1/500 |
| Number of steps | 1–4 | 2 |
| Incubation time with mixture (miRNA-21 + FITC-miRNA + anti-FITC-HRP), min | 15–270 | 120 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vargas, E.; Povedano, E.; Montiel, V.R.-V.; Torrente-Rodríguez, R.M.; Zouari, M.; Montoya, J.J.; Raouafi, N.; Campuzano, S.; Pingarrón, J.M. Single-Step Incubation Determination of miRNAs in Cancer Cells Using an Amperometric Biosensor Based on Competitive Hybridization onto Magnetic Beads. Sensors 2018, 18, 863. https://doi.org/10.3390/s18030863
Vargas E, Povedano E, Montiel VR-V, Torrente-Rodríguez RM, Zouari M, Montoya JJ, Raouafi N, Campuzano S, Pingarrón JM. Single-Step Incubation Determination of miRNAs in Cancer Cells Using an Amperometric Biosensor Based on Competitive Hybridization onto Magnetic Beads. Sensors. 2018; 18(3):863. https://doi.org/10.3390/s18030863
Chicago/Turabian StyleVargas, Eva, Eloy Povedano, Víctor Ruiz-Valdepeñas Montiel, Rebeca M. Torrente-Rodríguez, Mohamed Zouari, Juan José Montoya, Noureddine Raouafi, Susana Campuzano, and José M. Pingarrón. 2018. "Single-Step Incubation Determination of miRNAs in Cancer Cells Using an Amperometric Biosensor Based on Competitive Hybridization onto Magnetic Beads" Sensors 18, no. 3: 863. https://doi.org/10.3390/s18030863
APA StyleVargas, E., Povedano, E., Montiel, V. R.-V., Torrente-Rodríguez, R. M., Zouari, M., Montoya, J. J., Raouafi, N., Campuzano, S., & Pingarrón, J. M. (2018). Single-Step Incubation Determination of miRNAs in Cancer Cells Using an Amperometric Biosensor Based on Competitive Hybridization onto Magnetic Beads. Sensors, 18(3), 863. https://doi.org/10.3390/s18030863






