Electrochemical Biosensors for Rapid Detection of Foodborne Salmonella: A Critical Overview
Abstract
1. Introduction
2. Nano- and Micro-Sized Materials for Improving Detection
3. Electrochemical Immunosensors for Salmonella Detection
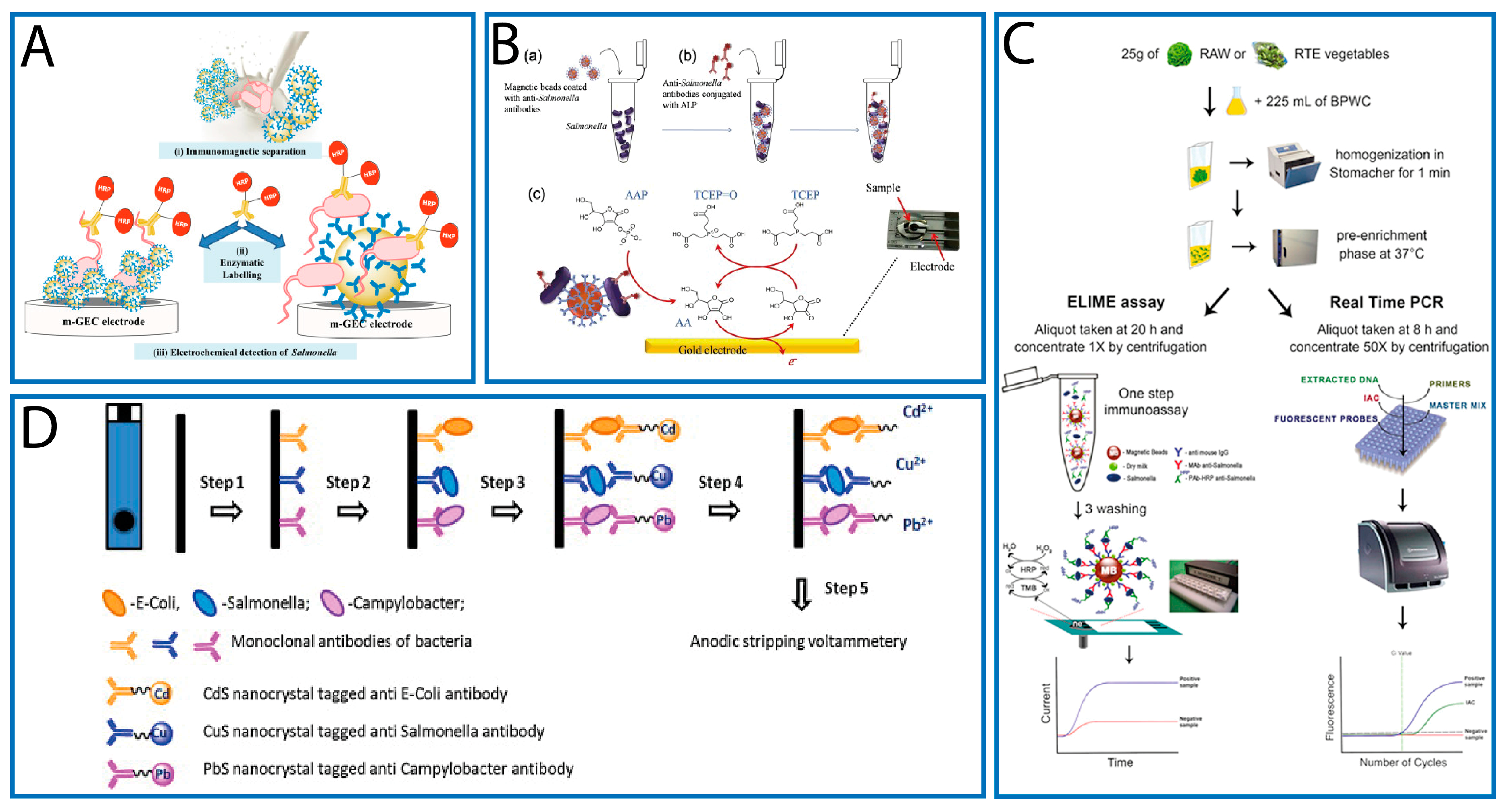
3.1. Label-Based Electrochemical Immunosensors
3.1.1. GNPs
3.1.2. MBs
3.1.3. QDs
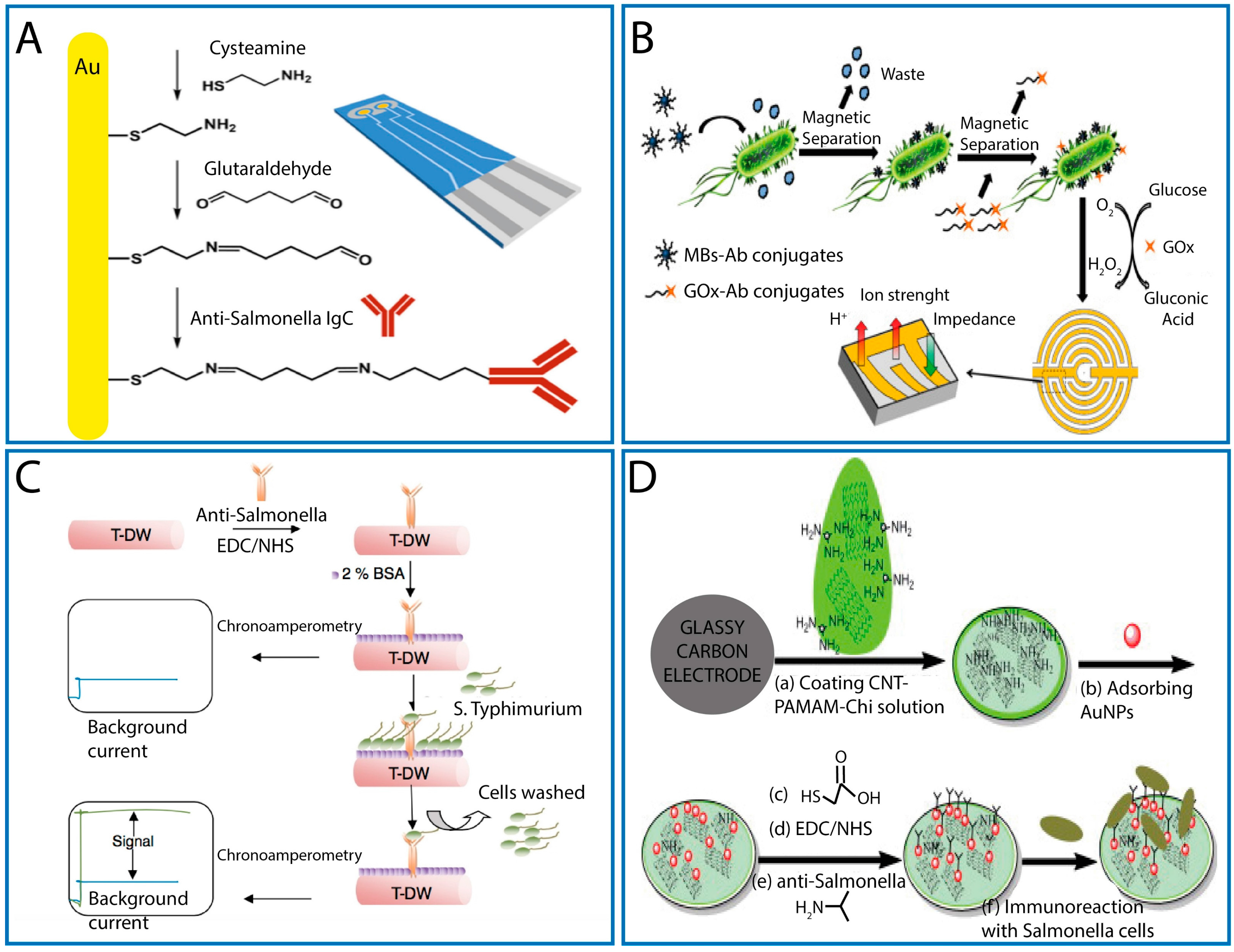
3.2. Label Free Electrochemical Immunosensors
3.2.1. MBs
3.2.2. CNTs
3.2.3. GR
4. Electrochemical Genosensors, Phagosensors and Aptasensors for Salmonella Detection
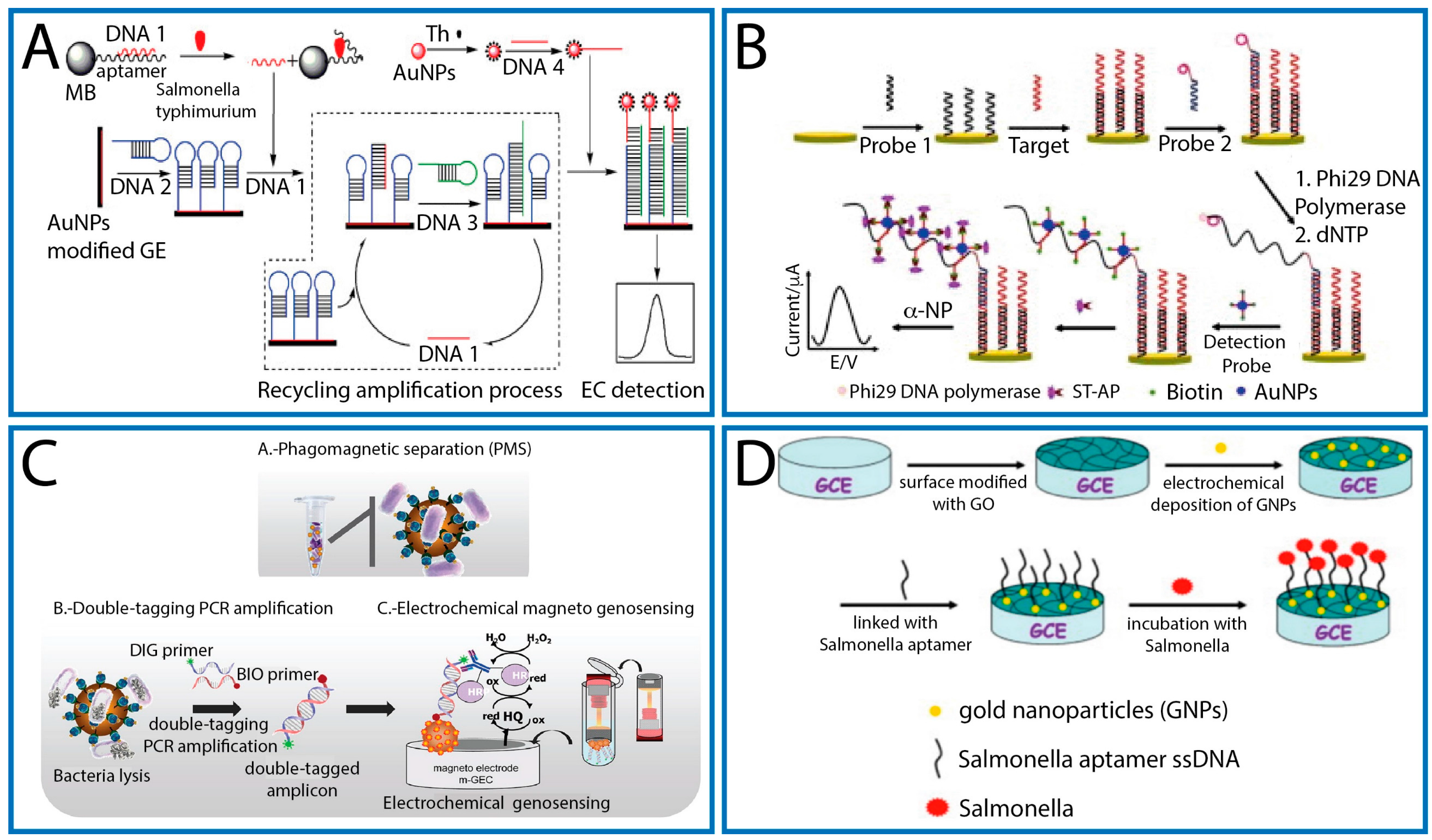
4.1. Label-Based Approaches
4.1.1. Genosensors
GNPs
MBs
4.1.2. Phagosensors
MBs
4.1.3. Aptasensors
GNPs
MBs
4.2. Label-Free Approaches
4.2.1. Genosensors
GNPs
4.2.2. Aptasensors
GNPs
Conductive Polymers
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Park, S.; Szonyi, B.; Gautam, R.; Nightingale, K.; Anciso, J.; Ivanek, R. Risk factors for microbial contamination in fruits and vegetables at the preharvest level: A systematic review. J. Food Prot. 2012, 75, 2055–2081. [Google Scholar] [CrossRef] [PubMed]
- Hanning, I.B.; Nutt, J.D.; Ricke, S.C. Salmonellosis outbreaks in the United States due to fresh produce: Sources and potential intervention measures. Foodborne Pathog. Dis. 2009, 6, 635–648. [Google Scholar] [CrossRef] [PubMed]
- Franz, E.; van Bruggen, A.H.C. Ecology of E. coli O157:H7 and Salmonella enterica in the Primary Vegetable Production Chain. Crit. Rev. Microbiol. 2008, 34, 143–161. [Google Scholar] [CrossRef] [PubMed]
- Sivapalasingam, S.; Friedman, C.R.; Cohenand, L.; Tauxe, R.V. Fresh produce: A growing cause of outbreaks of foodborne illness in the United States 1973 through 1997. J. Food Protect. 2004, 67, 2342–2353. [Google Scholar] [CrossRef]
- Commission Regulation (EC) No 2073/2005 on Microbiological Criteria for Foodstuffs. 2005. Available online: http://eur-lex.europa.eu/legal-content/IT/TXT/?uri=celex:32005R2073 (accessed on 17 August 2017).
- Melo, A.M.A.; Alexandre, D.L.; Furtado, R.F.; Borges, M.F.; Figueiredo, E.A.T.; Biswas, A.; Cheng, H.N. Electrochemical immunosensors for Salmonella detection in food. Appl. Microbiol. Biotechnol. 2016, 100, 5301–5312. [Google Scholar] [CrossRef] [PubMed]
- Labib, M.; Zamay, A.S.; Kolovskaya, O.S.; Reshetneva, I.T.; Zamay, G.S.; Kibbee, R.J.; Sattar, A.A.; Zamay, T.N.; Berezovski, M.V. Aptamer-based viability impedimetric sensor for bacteria. Anal. Chem. 2012, 84, 8966–8969. [Google Scholar] [CrossRef] [PubMed]
- Irwin, P.; Gehring, A.; Tu, S.; Chen, C. Blocking nonspecific adsorption of native food-borne microorganisms by immunomagnetic beads with Ι-carrageenan. Carbohydr. Res. 2004, 339, 613–621. [Google Scholar] [CrossRef] [PubMed]
- Bard, A.J.; Faulkner, L.R. Electrochemical Method Fundamentals and Applications; Wiley: New York, NY, USA, 1980. [Google Scholar]
- Wang, J. Analytical Electrochemistry; John Wiley & Sons: New York, NY, USA, 2006. [Google Scholar]
- Arduini, F.; Cinti, S.; Scognamiglio, V.; Moscone, D. Nanomaterials in electrochemical biosensors for pesticide detection: Advances and challenges in food analysis. Microchim. Acta 2016, 183, 2063–2083. [Google Scholar] [CrossRef]
- Wang, J. Nanomaterial-based electrochemical biosensors. Analyst 2005, 130, 421–426. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.; Chatterjee, S. Nanomaterials based electrochemical sensors for biomedical applications. Chem. Soc. Rev. 2013, 42, 5425–5438. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Santos, D.; González-García, M.B.; García, A.C. Metal-nanoparticles based electroanalysis. Electroanalysis 2002, 14, 1225–1235. [Google Scholar] [CrossRef]
- Cinti, S.; Santella, F.; Moscone, D.; Arduini, F. Hg2+ detection using a disposable and miniaturized screen-printed electrode modified with nanocomposite carbon black and gold nanoparticles. Environ. Sci. Pollut. Res. Int. 2016, 23, 8192–8199. [Google Scholar] [CrossRef] [PubMed]
- Cinti, S.; Arduini, F. Graphene-based screen-printed electrochemical (bio)sensors and their applications: Efforts and criticisms. Biosens. Bioelectron. 2016, 89, 107–122. [Google Scholar] [CrossRef] [PubMed]
- Arduini, F.; Cinti, S.; Scognamiglio, V.; Moscone, D.; Palleschi, G. How cutting-edge technologies impact the design of electrochemical (bio)sensors for environmental analysis. Anal. Chim. Acta 2017, 959, 15–42. [Google Scholar] [CrossRef] [PubMed]
- Renedo, O.D.; Alonso-Lomillo, M.A.; Martínez, M.A. Recent developments in the field of screen-printed electrodes and their related applications. Talanta 2007, 73, 202–219. [Google Scholar] [CrossRef] [PubMed]
- Albareda-Sirvent, M.; Merkoci, A.; Alegret, S. Configurations used in the design of screen-printed enzymatic biosensors. A review. Sens. Actuators B 2000, 69, 153–163. [Google Scholar] [CrossRef]
- Metters, J.P.; Kadara, R.O.; Banks, C.E. New directions in screen printed electroanalytical sensors: An overview of recent developments. Analyst 2011, 136, 1067–1076. [Google Scholar] [CrossRef] [PubMed]
- Cinti, S.; Minotti, C.; Moscone, D.; Palleschi, G.; Arduini, F. Fully integrated ready-to-use paper-based electrochemical biosensor to detect nerve agents. Biosens. Bioelectron. 2017, 93, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Soreta, T.R.; Strutwolf, J.; Henry, O.; O’Sullivan, C.K. Electrochemical surface structuring with palladium nanoparticles for signal enhancement. Langmuir 2010, 26, 12293–12299. [Google Scholar] [CrossRef] [PubMed]
- De la Escosura-Muñiz, A.; Ambrosi, A.; Merkoçi, A. Electrochemical analysis with nanoparticle-based biosystems. Trends Anal. Chem. 2008, 27, 568–584. [Google Scholar] [CrossRef]
- Cinti, S.; Neagu, D.; Carbone, M.; Cacciotti, I.; Moscone, D.; Arduini, F. Novel carbon black-cobalt phthalocyanine nanocomposite as sensing platform to detect organophosphorus pollutants at screen-printed electrode. Electrochim. Acta 2016, 188, 574–581. [Google Scholar] [CrossRef]
- Cinti, S.; Basso, M.; Moscone, D.; Arduini, F. A paper based-nanomodified electrochemical biosensor for ethanol detection in beers. Anal. Chim. Acta 2017, 960, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Cinti, S.; Talarico, D.; Palleschi, G.; Moscone, D.; Arduini, F. Novel reagentless paper-based screen-printed electrochemical sensor to detect phosphate. Anal. Chim. Acta 2016, 919, 78–84. [Google Scholar] [CrossRef] [PubMed]
- De la Escosura-Muñiz, A.; Parolo, C.; Merkoçi, A. Immunosensing using nanoparticles. Mater. Today 2010, 13, 24–34. [Google Scholar] [CrossRef]
- Saha, K.; Agasti, S.S.; Kim, C.; Li, X.; Rotello, V.M. Gold nanoparticles in chemical and biological sensing. Chem. Rev. 2012, 112, 2739–2779. [Google Scholar] [CrossRef] [PubMed]
- Merkoçi, A.; Pumera, M.; Llopis, X.; Pérez, B.; del Valle, M.; Alegret, S. New materials for electrochemical sensing VI: Carbon nanotubes. Trends Anal. Chem. 2005, 24, 826–838. [Google Scholar] [CrossRef]
- Cinti, S.; Arduini, F.; Carbone, M.; Sansone, L.; Cacciotti, I.; Moscone, D.; Palleschi, G. Screen-Printed Electrodes Modified with Carbon Nanomaterials: A Comparison among Carbon Black, Carbon Nanotubes and Graphene. Electroanalysis 2015, 27, 2230–2238. [Google Scholar] [CrossRef]
- Nambiar, S.; Yeow, J.T. Conductive polymer-based sensors for biomedical applications. Biosens. Bioelectron. 2011, 26, 1825–1832. [Google Scholar] [CrossRef] [PubMed]
- Golabi, M.; Turner, A.P.; Jager, E.W. Tunable conjugated polymers for bacterial differentiation. Sens. Actuators B 2016, 222, 839–848. [Google Scholar] [CrossRef]
- Goicoechea, J.; Arregui, F.J.; Matias, I.R. Quantum dots for sensing. In Sensors Based on Nanostructured Materials; Springer: New York, NY, USA, 2009; pp. 131–181. [Google Scholar]
- Wang, J.; Liu, G.; Merkoçi, A. Electrochemical coding technology for simultaneous detection of multiple DNA targets. J. Am. Chem. Soc. 2003, 125, 3214–3215. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.A.; Wang, J.; Kawde, A.N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar] [CrossRef] [PubMed]
- Haukanes, B.I.; Kvam, C. Application of magnetic beads in bioassays. Nat. Biotechnol. 1993, 11, 60–63. [Google Scholar] [CrossRef]
- Yáñez-Sedeño, P.; Campuzano, S.; Pingarrón, J.M. Magnetic particles coupled to disposable screen printed transducers for electrochemical biosensing. Sensors 2016, 16, 1585. [Google Scholar] [CrossRef] [PubMed]
- Catanante, G.; Rhouati, A.; Hayat, A.; Marty, J.L. An overview of recent electrochemical immunosensing strategy for mycotoxin detection. Electroanalysis 2016, 28, 1750–1753. [Google Scholar] [CrossRef]
- Arduini, F.; Micheli, L.; Moscone, D.; Palleschi, G.; Piermarini, S.; Ricci, F.; Volpe, G. Electrochemical biosensors based on nanomodified sceen-printed electrodes: Recent applications in clinical analysis. Trends Anal. Chem. 2016, 79, 114–126. [Google Scholar] [CrossRef]
- Pandiaraj, M.; Sethy, N.K.; Bhargava, K.; Kameswararao, V.; Karunakaran, C. Designing label free electrochemical immunosensors for cytochrome c using nanocomposites functionalized screen-printed electrodes. Biosens. Bioelectron. 2014, 54, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Haab, B.B. Methods and applications of antibody microarrays in cancer research. Proteomics 2003, 3, 2116–2122. [Google Scholar] [CrossRef] [PubMed]
- Xiang, C.; Li, R.; Adhikari, B.; She, Z.; Li, Y. Sensitive electrochemical detection of Salmonella with chitosan-gold nanoparticles composite film. Talanta 2015, 140, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Fei, J.; Dou, W.; Zhao, G. A sandwich electrochemical immunosensors for Salmonella pollurum and Salmonella gallinarum based on a screen-printe carbone electrode modified with an ionic liquid and electrodeposited gold nanoparticles. Microchim. Acta 2015, 182, 2267–2275. [Google Scholar] [CrossRef]
- Afonso, A.S.; Lopez, B.P.; Faria, R.C.; Mattoso, L.H.C.; Hernandez-Herrero, M.; Roig-Sagues, A.X.; Maltez-da Costa, M.; Merkoci, A. Electrochemical detection of Salmonella using gold nanoparticles. Biosens. Bioelectron. 2013, 40, 121–126. [Google Scholar] [CrossRef] [PubMed]
- Brandao, D.; Liebana, S.; Campoy, S.; Alegret, S. Immunomagnetic separation of Salmonella with tailored magnetic micro and nanocarriers. A comparative study. Talanta 2015, 143, 198–204. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Wang, Z.; Chen, J.; Kinchla, A.J.; Nugen, S.R. Rapid detection of Salmonella using a redox cycling-based electrochemical method. Food Control 2016, 62, 81–88. [Google Scholar] [CrossRef]
- Fabiani, L.; Pucci, E.; Delibato, E.; Volpe, G.; Piermarini, S.; De Medici, D.; Capuano, F.; Palleschi, G. ELIME assay vs. Real-Time PCR and conventional culture method for an effective detection of Salmonella in fresh leafy green vegetables. Talanta 2017, 166, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, S.; Rani, C.; Ho, J.A. Electrochemical immunosensors for multiplexed detection of food-borne pathogens using nanocrystal bioconjugares and MWCNT scree-printed electrode. Talanta 2012, 94, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Volpe, G.; Delibato, E.; Fabiani, L.; Pucci, E.; Piermarini, S.; D’Angelo, A.; Capuano, F.; De Medici, D.; Palleschi, G. Development and evaluation of an ELIME assay to reveal the presence of Salmonella in irrigation water: Comparison with Real-Time PCR and the Standard Culture Method. Talanta 2016, 149, 202–210. [Google Scholar] [CrossRef] [PubMed]
- Pratiwi, F.W.; Rijiravanich, P.; Somasundrum, M.; Surareungchai, W. Electrochemical immunoassay for Salmonella Typhimurium based on magnetically collected Ag-enhanced DNA biobarcode labels. Analyst 2013, 138, 5011–5018. [Google Scholar] [CrossRef] [PubMed]
- Nam, J.M.; Thaxton, C.S.; Mirkin, C.A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 2003, 301, 1884–1886. [Google Scholar] [CrossRef] [PubMed]
- Freitas, M.; Viswanathan, S.; Nouws, H.P.A.; Oliveira, M.B.P.P.; Delerue-Matos, C. Iron oxide/gold core/shell nanomagnetic probes and CdS biolabels for amplified electrochemical immunosensing of Salmonella Typhimurium. Biosens. Bioelectron. 2014, 51, 195–200. [Google Scholar] [CrossRef] [PubMed]
- Farka, Z.; Juřík, T.; Pastucha, M.; Kovář, D.; Lacina, K.; Skládal, P. Rapid immunosensing of Salmonella Typhimurium using electrochemical impedance spectroscopy: The effect of sample treatment. Electroanalysis 2016, 28, 1–8. [Google Scholar] [CrossRef]
- Xu, M.; Wang, R.; Li, Y. Rapid detection of Escherichia coli O157:H7 and Salmonella Typhimurium in foods using an electrochemical immunosensors based on screen-printed interdigitated microelectrode and immunomagnetic separation. Talanta 2016, 148, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Punbusayakul, N.; Talapatra, S.; Ajayan, P.M.; Surareungchai, W. Label-free as-grown double wall carbon nanotubes bundles for Salmonella Typhimurium immunoassay. Chem. Cent. J. 2013, 7, 102–108. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Zhao, H.M.; Xu, Q.; Ma, S.; Ai, A. label-free electrochemical impedance immunosensors based on AuNPs/PAMAM-MWCNT-Chi nanocomposite modified glassy carbon electrode for detection of Salmonella Typhimurium in milk. Food Chem. 2013, 141, 1980–1986. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Chai, C.; Yao, B. Rapid evaluation of Salmonella pullorum contamination in chicken based on a portable amperometric sensor. Biosens. Bioelectron. 2013, 4, 137–143. [Google Scholar]
- Mutreja, R.; Jariyal, M.; Pathania, P.; Sharma, A.; Sahoo, D.K.; Suri, C.R. Novel surface antigen based impedimetric immunosensors for detection of Salmonella Typhimurium in water and juice samples. Biosens. Bioelectron. 2016, 85, 707–713. [Google Scholar] [CrossRef] [PubMed]
- García, T.; Revenga-Parra, M.; Añorga, L.; Arana, S.; Pariente, F.; Lorenzo, E. Disposable DNA biosensor based on thin-film gold electrodes for selective Salmonella detection. Sens. Actuators B 2012, 161, 1030–1037. [Google Scholar] [CrossRef]
- Tabrizi, M.A.; Shamsipur, M. A label-free electrochemical DNA biosensor based on covalent immobilization of Salmonella DNA sequences on the nanoporous glassy carbon electrode. Biosens. Bioelectron. 2015, 69, 100–105. [Google Scholar] [CrossRef] [PubMed]
- Das, R.; Sharma, M.K.; Rao, V.K.; Bhattacharya, B.K.; Garg, I.; Venkatesh, V.; Upadhyay, S. An electrochemical genosensor for Salmonella typhi on gold nanoparticles-mercaptosilane modified screen printed electrode. J. Biotechnol. 2014, 188, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Smartt, A.E.; Xu, T.; Jegier, P.; Carswell, J.J.; Blount, S.A.; Sayler, G.S.; Ripp, S. Pathogen detection using engineered bacteriophages. Anal. Bioanal. Chem. 2012, 402, 3127–3146. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Poshtiban, S.; Evoy, S. Recent advances in bacteriophage based biosensors for food-borne pathogen detection. Sensors 2013, 13, 1763–1786. [Google Scholar] [CrossRef] [PubMed]
- Peltomaa, R.; López-Perolio, I.; Benito-Peña, E.; Barderas, R.; Moreno-Bondi, M.C. Application of bacteriophages in sensor development. Anal. Bioanal. Chem. 2016, 408, 1805–1828. [Google Scholar] [CrossRef] [PubMed]
- Liébana, S.; Spricigo, D.A.; Cortés, M.P.; Barbé, J.; Llagostera, M.; Alegret, S.; Pividori, M.I. Phagomagnetic separation and electrochemical magneto-genosensing of pathogenic bacteria. Anal. Chem. 2013, 85, 3079–3086. [Google Scholar] [CrossRef] [PubMed]
- Sheikhzadeh, E.; Chamsaz, M.; Turner, A.P.F.; Jager, E.W.H.; Beni, V. Label-free impedimetric biosensor for Salmonella Typhimurium detection based on poly[pyrrole-co-3-carboxyl-pyrrole] copolymer supported aptamer. Biosens. Bioelectron. 2016, 80, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Cheng, W.; Zhang, D.; Yu, T.; Yin, Y.; Ju, H.; Ding, S. Rapid and sensitive strategy for Salmonella detection using an InvA gene-based electrochemical DNA sensor. Int. J. Electrochem. Sci. 2012, 7, 844–856. [Google Scholar]
- Zong, Y.; Liu, F.; Zhang, Y.; Zhan, T.; He, Y.; Hun, X. Signal amplification technology based on entropy-driven molecular switch for ultrasensitive electrochemical determination of DNA and Salmonella Typhimurium. Sens. Actuators B 2016, 225, 420–427. [Google Scholar] [CrossRef]
- Zhu, D.; Yan, Y.; Lei, P.; Shen, B.; Cheng, W.; Ju, H.; Ding, S. A novel electrochemical sensing strategy for rapid and ultrasensitive detection of Salmonella by rolling circle amplification and DNA-AuNPs probe. Anal. Chim. Acta 2014, 846, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Jiang, Y.; Jia, F.; Yu, Y.; Chen, J.; Wang, Z. An aptamer-based electrochemical biosensor for the detection of Salmonella. J. Microbiol. Methods 2014, 98, 94–98. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Fu, H.; He, Y.; Zhai, Q.; Guo, J.; Qing, K.; Yi, G. Electrochemical Aptasensor for Rapid and Sensitive Determination of Salmonella Based on Target-Induced Strand Displacement and Gold Nanoparticle Amplification. Anal. Lett. 2016, 49, 2405–2417. [Google Scholar] [CrossRef]
- Liébana, S.; Brandão, D.; Cortés, P.; Campoy, S.; Alegret, S.; Pividori, M.I. Electrochemical genosensing of Salmonella, Listeria and Escherichia coli on silica magnetic particles. Anal. Chim. Acta 2016, 904, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Yang, H.; Lakshmanan, R.S.; Johnson, M.L.; Wan, J.; Chen, I.-H.; Wikle, H.C.; Petrenko, V.A.; Barbaree, J.M.; Chin, B.A. Sequential detection of Salmonella Typhimurium and Bacillus anthracis spores using magnetoelastic biosensors. Biosens. Bioelectron. 2009, 24, 1730–1736. [Google Scholar] [CrossRef] [PubMed]
- Chai, Y.; Li, S.; Horikawa, S.; Park, M.-K.; Vodyanoy, V.; Chin, B.A. Rapid and sensitive detection of Salmonella Typhimurium on eggshells by using wireless biosensors. J. Food Prot. 2012, 75, 631–636. [Google Scholar] [CrossRef] [PubMed]
- Bagheryan, Z.; Raoof, J.B.; Golabi, M.; Turner, A.P.; Beni, V. Diazonium-based impedimetric aptasensor for the rapid label-free detection of Salmonella Typhimurium in food sample. Biosens. Bioelectron. 2016, 80, 566–573. [Google Scholar] [CrossRef] [PubMed]
| Platform | Nano- and Micro-Sized Materials | Electrochemical Technique | Detection Time | LOD in Broth Cultures CFU/mL | Food Analysis | Reference |
|---|---|---|---|---|---|---|
| GCE | GNPs | DPV | 4 h | 5 | - | [42] |
| SPE | GNPs | CV | 1 h 20 min | 3000 | Eggs and chicken meat Accuracy 80–100% | [43] |
| SPE | MBs/GNPs | DPV | 1 h 30 min | 143 | Milk Recovery 83–94% (1.5 × 103–1.5 × 105 CFU/mL) | [44] |
| GCE | NMBs | Amperometry | 1 h | 462 | Milk LOD 291 CFU/mL LOD 1 CFU/25 mL (8 h pre-enrichment) Milk LOD 538 CFU/mL | [45] |
| MMBs | 835 | |||||
| G-SPE | MBs | Amperometry | 1 h 30 min | 760 | Agricultural water LOD 6 × 102 CFU/mL LOD 10 CFU/mL (4 h of pre-enrichment) | [46] |
| 8-SPE strip | MBs | Amperometry | 1 h | 1000 | Irrigation water LOD 1 CFU/L (10 h of pre-enrichment) | [49] |
| 8-SPE strip | MBs | Amperometry | 1 h | 1000 | Fresh leafy green vegetables LOD 1 CFU/25 g (20 h of pre-enrichment) | [47] |
| SPE | MBs | DPASV | 1 h | 12 | Green bean sprouts, egg, milk LOD 13–26 CFU/mL | [50] |
| SPE | QDs/CNTs | SWASV | 4 h | 400 | Milk Recovery 95% (104 CFU/mL) | [51] |
| SPE | MBs/QDs | SWASV | <1 h | 13 | Milk Recovery 77.6–77.8% (103–104 CFU/mL) | [48] |
| G-SPE | - | EIS | 20 min | <1000 | Milk LOD 6 × 102 CFU/mL | [52] |
| SP-IDME | MBs | EIS | 2 h | 1660 | Chicken carcass rinse water LOD 103 CFU/mL | [53] |
| T-DW | CNTs | Amperometry | 2 h | 8.9 | - | [54] |
| SPE | CNTs | Amperometry | 30 min | 100 | Chicken meat LOD 60–100 CFU/mL (1.5–2 h of pre-enrichment) | [57] |
| GCE | GNPs/CNTs | EIS | 1 h | 500 | Milk Recovery 94.5–106.6% (103–107 CFU/mL) | [56] |
| SPE | rG-GO | EIS | 3 h | 10 | Water and fruit juices LOD 10 CFU/mL | [58] |
| Platform | Nano- and Micro-Sized Materials | Electrochemical Technique | Detection Time | LOD in Broth Cultures | Food Analysis | Reference |
|---|---|---|---|---|---|---|
| GE A | - | DPV | 3 h 30 min | 10 CFU/mL (after PCR) | - | [67] |
| GE A | GNPs, thionine/GNPs | DPV | 5 h | 0.3 fM | - | [68] |
| GE B | MBs | DPV | 4 h 25 min | 13 CFU/mL | Milk Recovery 96.1–103.0% (57–1093 CFU/mL) | [68] |
| GE A | GNPs | DPV | 1 h | 6 CFU/mL (after PCR) | Milk LOD 6 CFU/mL | [69] |
| m-GEC A | Silica MBs | Amperometry | 3 h | 0.04 μg/mL (after PCR) | - | [72] |
| m-GEC C | MBs | Amperometry | 4 h | 3 CFU/mL (after PCR) | - | [65] |
| GE B | GNPs | DPV | 3 h 30 min | 20 CFU/mL D | Milk LOD 200 CFU/mL | [71] |
| Thin-film GE A | DPV | 1 h | 0.208 μM D | - | [59] | |
| GCE A | - | DPV | 1 h | 2.1 pM D | - | [60] |
| EIS | 0.15 pM D | |||||
| SPE A | GNPs | DPV | 1 h 5 min | 50 pM D | - | [61] |
| GCE B | GO/GNPs | EIS | 1 h 30 min | 3 CFU/mL | Pork meat Recovery 97.3–105% (10–1000 CFU/mL) | [70] |
| SPE B | - | EIS | 45 min | 6 CFU/mL | Apple juice Recovery 300–440% (102–106 CFU/mL) | [75] |
| GE B | PPy | EIS | 1 h | 3 CFU/mL | Apple juice Recovery 140–410% (102–106 CFU/mL) | [66] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cinti, S.; Volpe, G.; Piermarini, S.; Delibato, E.; Palleschi, G. Electrochemical Biosensors for Rapid Detection of Foodborne Salmonella: A Critical Overview. Sensors 2017, 17, 1910. https://doi.org/10.3390/s17081910
Cinti S, Volpe G, Piermarini S, Delibato E, Palleschi G. Electrochemical Biosensors for Rapid Detection of Foodborne Salmonella: A Critical Overview. Sensors. 2017; 17(8):1910. https://doi.org/10.3390/s17081910
Chicago/Turabian StyleCinti, Stefano, Giulia Volpe, Silvia Piermarini, Elisabetta Delibato, and Giuseppe Palleschi. 2017. "Electrochemical Biosensors for Rapid Detection of Foodborne Salmonella: A Critical Overview" Sensors 17, no. 8: 1910. https://doi.org/10.3390/s17081910
APA StyleCinti, S., Volpe, G., Piermarini, S., Delibato, E., & Palleschi, G. (2017). Electrochemical Biosensors for Rapid Detection of Foodborne Salmonella: A Critical Overview. Sensors, 17(8), 1910. https://doi.org/10.3390/s17081910





