3D Laser Triangulation for Plant Phenotyping in Challenging Environments
Abstract
:1. Introduction
2. Material and Methods
2.1. The 3D Laser Triangulation Scanner
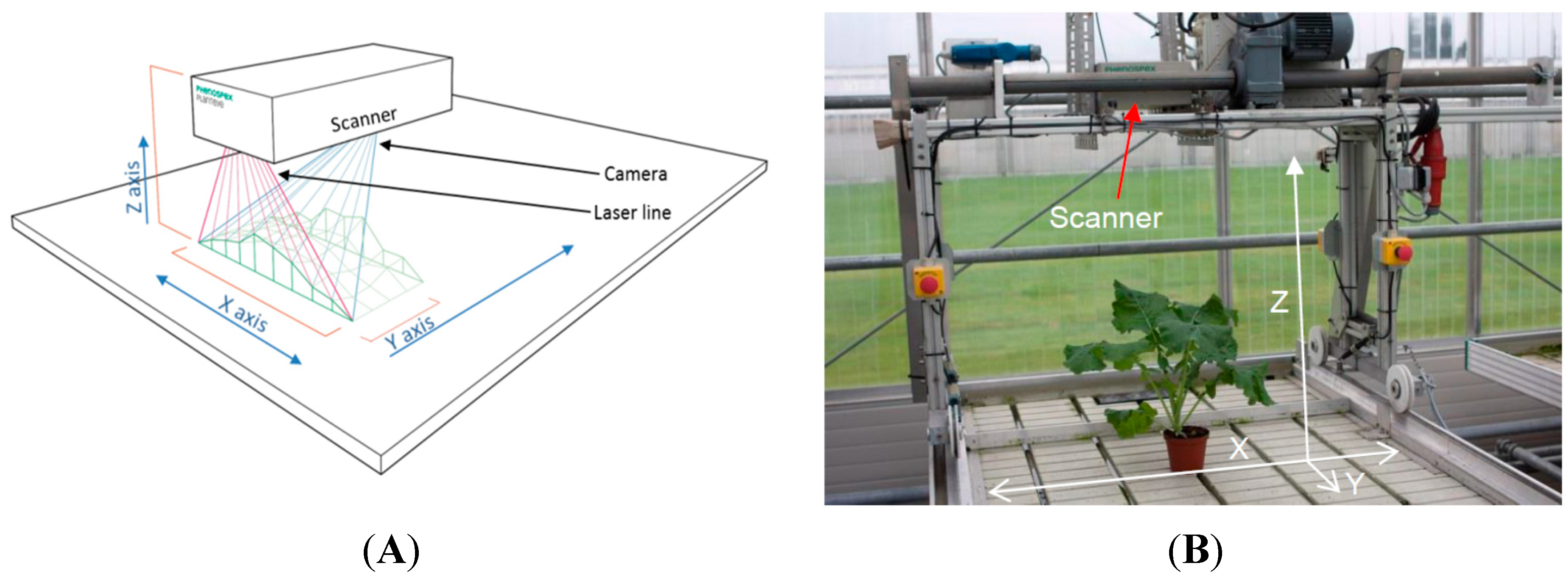
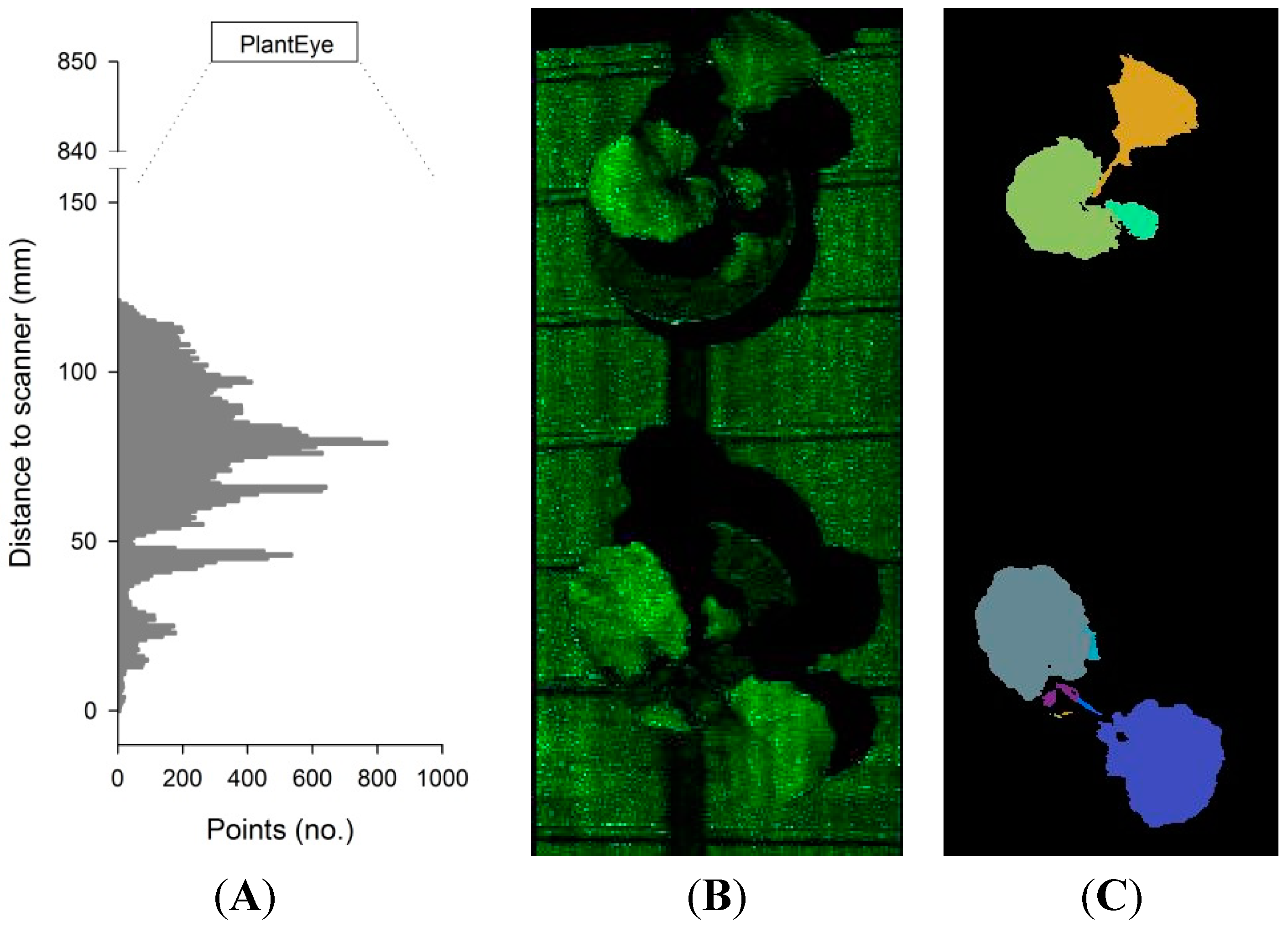
2.2. Experiment 1: Testing a Potential Influence of the Projected Laser Line on Photosynthetic Activity
2.3. Experiment 2: Predicting Growth Parameters of Rapeseed by 3D Laser Triangulation
2.4. Experiment 3: Validating the Model on Ten Rapeseed Genotypes
2.5. Data Analysis
3. Results and Discussion
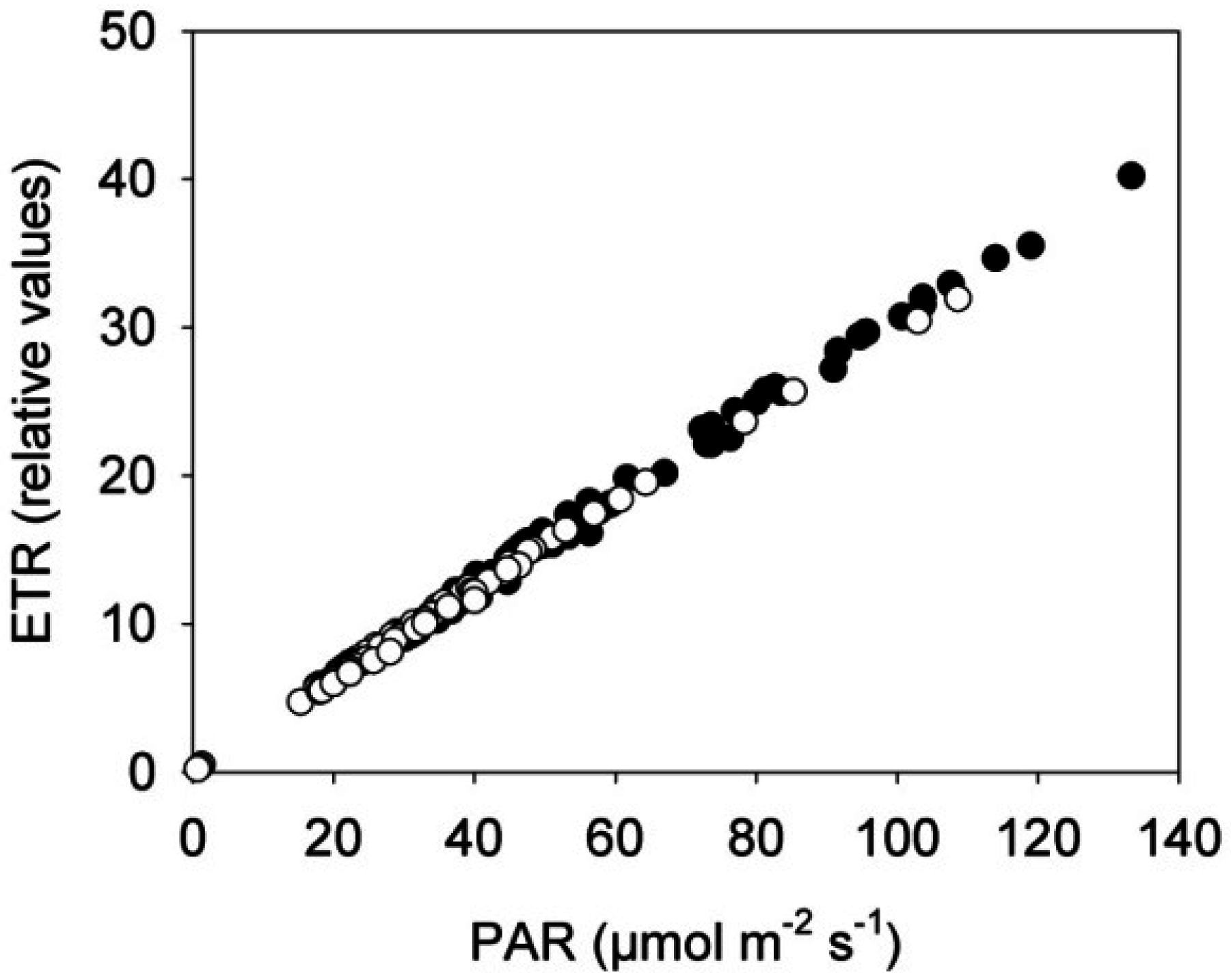
3.1. Photosystem II Efficiency of Rapeseed Was Not Affected by the Near-Infrared (NIR) Laser Line
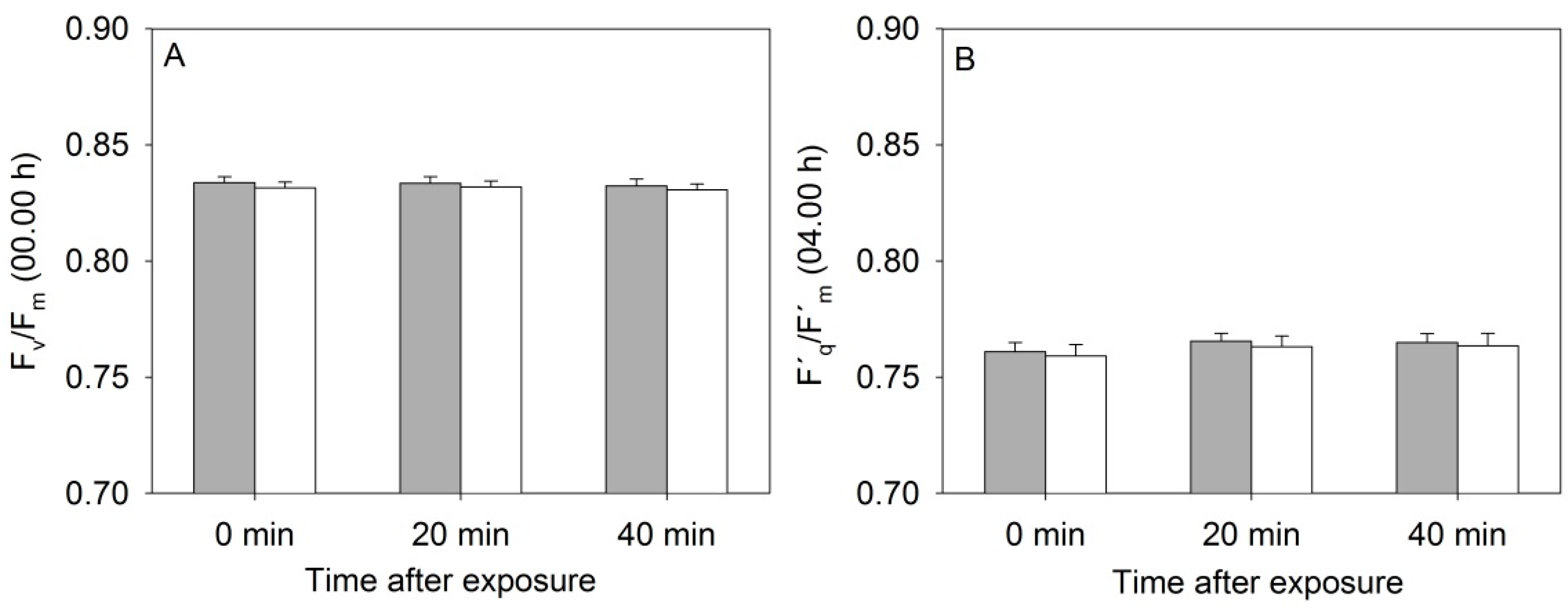
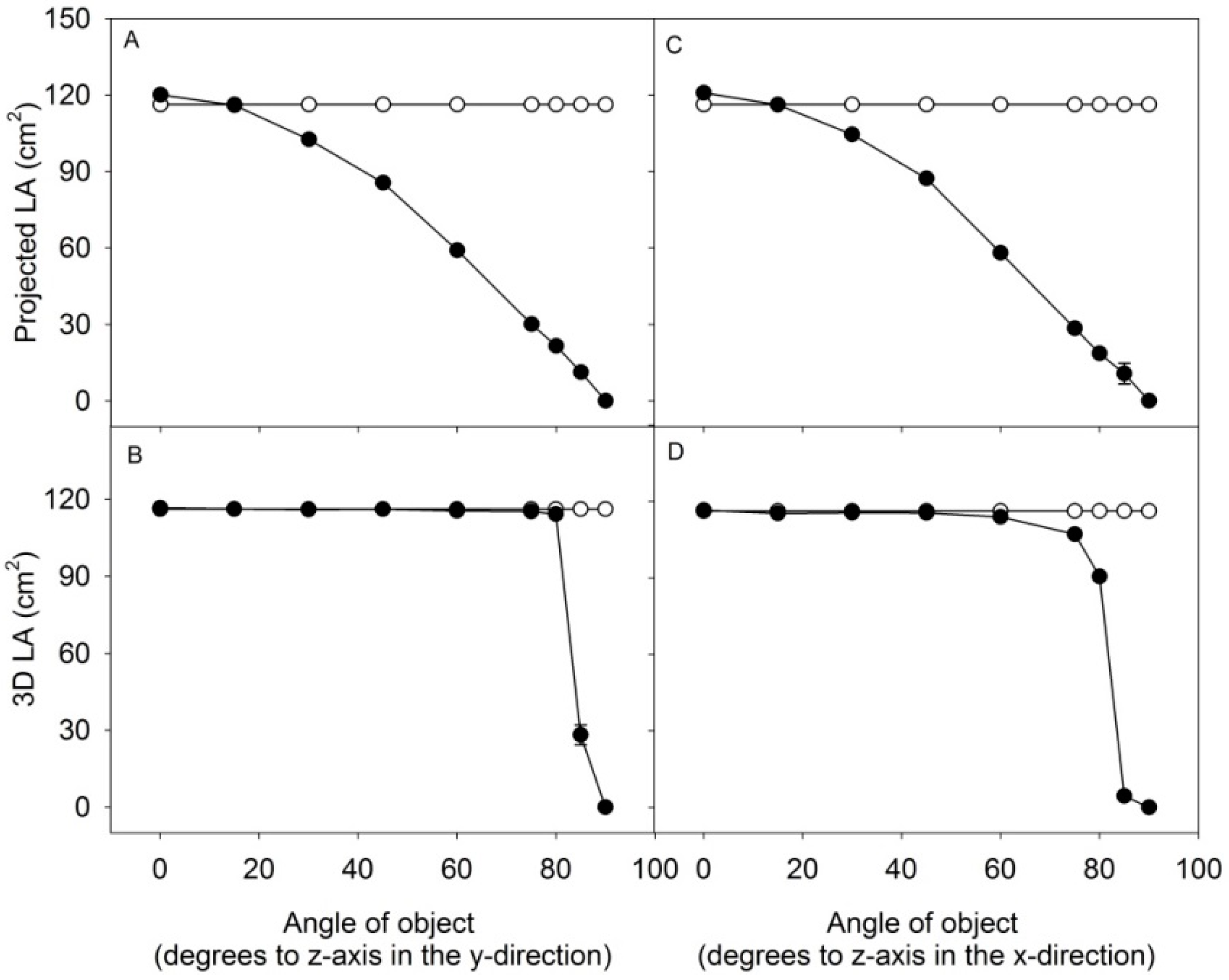
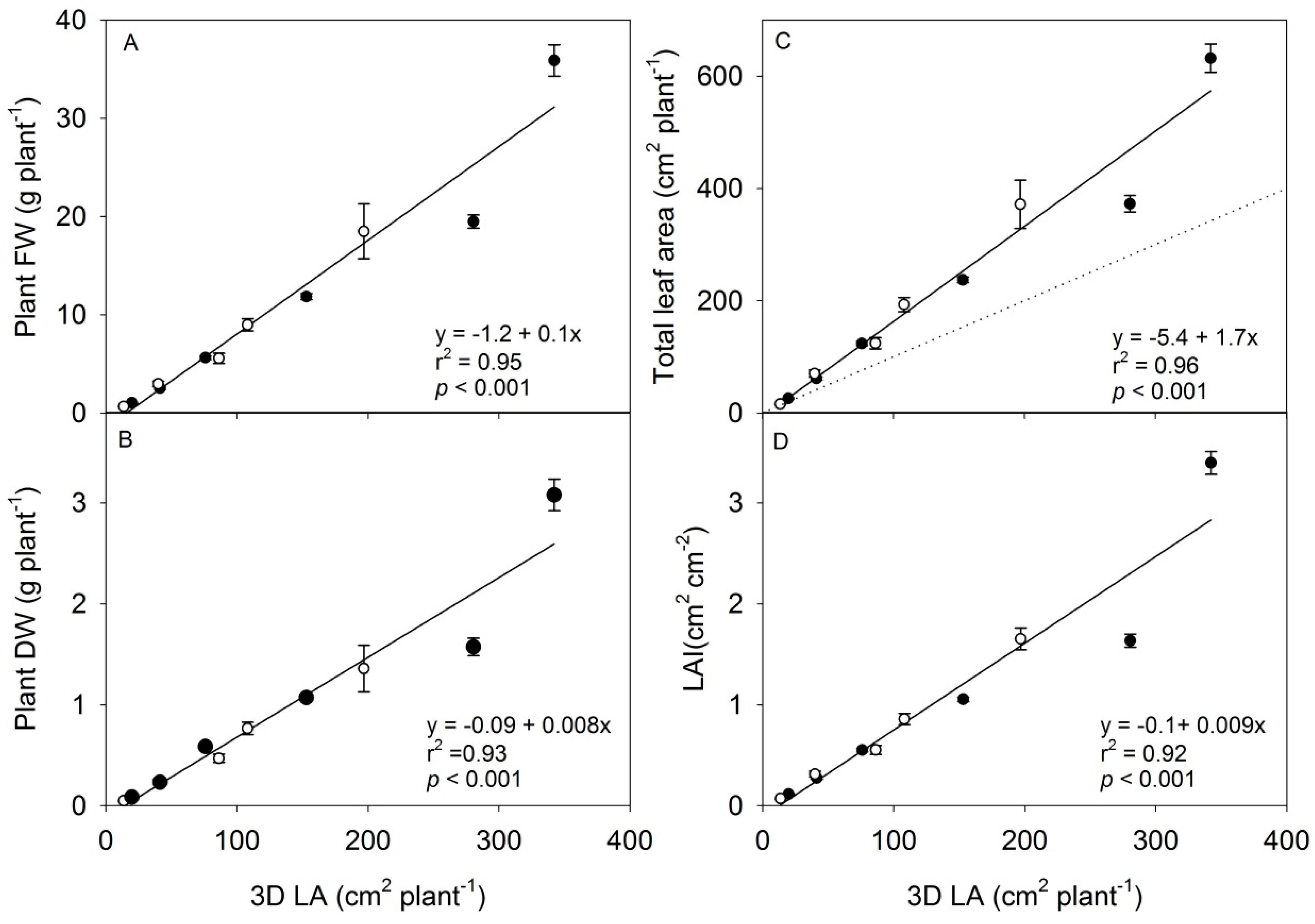
3.2. The Scanning Parameters Correlate with Growth of Rapeseed, but Total Leaf Area Is Underestimated
| LAI | FW | DW | LN | LA | ||
|---|---|---|---|---|---|---|
| Height | r2 | 0.86 | 0.93 | 0.93 | 0.89 | 0.94 |
| p | <0.01 | <0.001 | <0.001 | <0.01 | <0.001 | |
| 3D LA | r2 | 0.93 | 0.97 | 0.97 | 0.88 | 0.97 |
| p | <0.001 | <0.001 | <0.001 | <0.01 | <0.001 | |
| Proj LA | r2 | 0.93 | 0.97 | 0.97 | 0.88 | 0.97 |
| p | <0.001 | <0.001 | <0.001 | <0.01 | <0.001 |
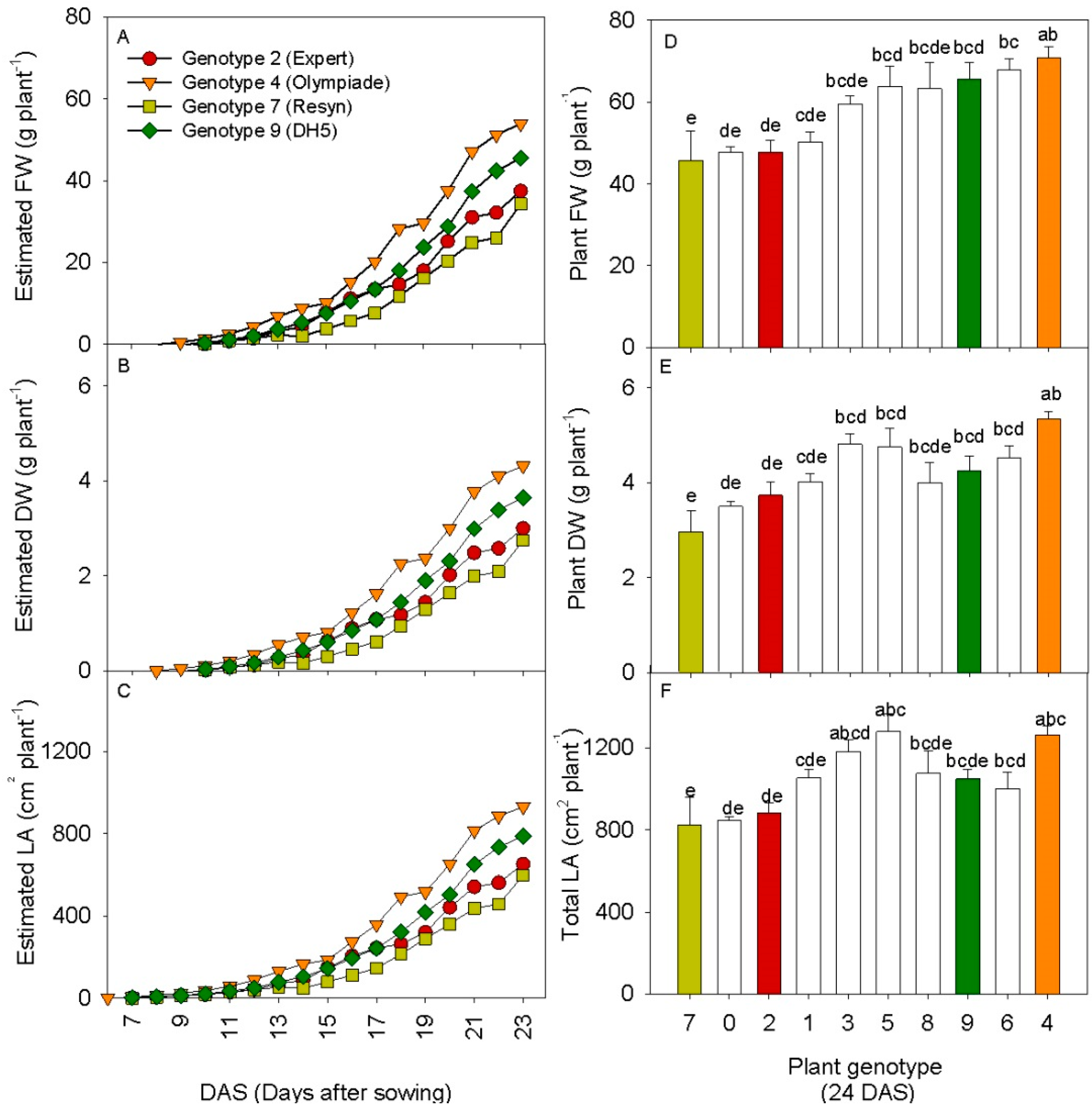
3.3. 3D Laser Triangulation Identified Phenotypic Variation in Rapeseed
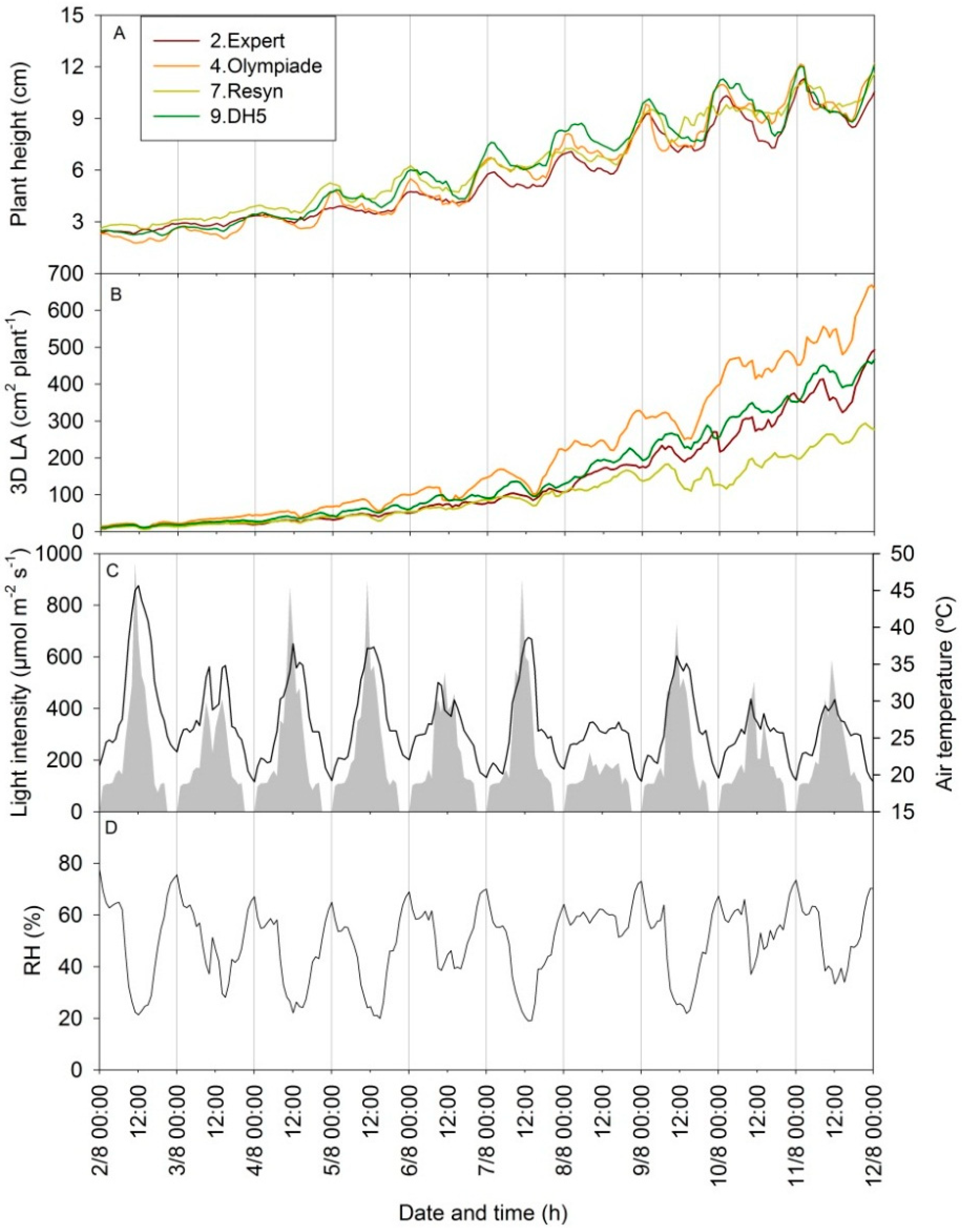
3.4. Diurnal Patterns in Scanning Parameters Contain Information on Changes in 3D Plant Structure
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Pereyra-Irujo, G.A.; Gasco, E.D.; Peirone, L.S.; Aguirrezabal, L.A. GlyPh: A low-cost platform for phenotyping plant growth and water use. Funct. Plant. Biol. 2012, 39, 905–913. [Google Scholar] [CrossRef]
- Golzarian, M.R.; Frick, R.A.; Rajendran, K.; Berger, B.; Roy, S.; Tester, M.; Lun, D.S. Accurate inference of shoot biomass from high-throughput images of cereal plants. Plant Methods 2011, 7. [Google Scholar] [CrossRef] [PubMed]
- Brien, C.J.; Berger, B.; Rabie, H.; Tester, M. Accounting for variation in designing greenhouse experiments with special reference to greenhouse containing plants on conveyor belts. Plant Methods 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Fanourakis, D.; Briese, C.; Max, J.F.; Kleinen, S.; Putz, A.; Fiorani, F.; Ulbrich, A.; Schurr, U. Rapid determination of leaf area and plant height by using light curtain arrays in four species with contrasting shoot architecture. Plant Methods 2014, 10. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Czauderna, T.; Hoffmann, R.; Stein, N.; Schreiber, F. HTPheno: An image analysis pipeline for high-throughput phenotyping. BMC Bioinform. 2011, 12. [Google Scholar] [CrossRef]
- Van Zanten, M.; Pons, T.L.; Janssen, J.A.M.; Voesenek, L.A.C.J.; Peeters, A.J.M. On the relevance and control of leaf angle. Crit. Rev. Plant Sci. 2010, 29, 300–316. [Google Scholar] [CrossRef]
- Chéné, Y.; Rousseau, D.; Lucidarme, P.; Bertheloot, J.; Caffier, V.; Morel, P.; Belin, E.; Chapeau-Blondeau, F. On the use of depth camera for 3D phenotyping of entire plants. Comput. Electron. Agric. 2012, 82, 122–127. [Google Scholar] [CrossRef]
- Biskup, B.; Scharr, H.; Schurr, U.; Rascher, U. A stereo imaging system for measuring structural parameters of plant canopies. Plant Cell Environ. 2007, 30, 1299–1308. [Google Scholar] [CrossRef] [PubMed]
- Lefsky, M.A.; Cohen, W.B.; Parker, G.G.; Harding, D.J. Lidar remote sensing for ecosystem studies. BioScience 2002, 52, 19–30. [Google Scholar] [CrossRef]
- Omasa, K.; Hosoi, F.; Konishi, A. 3D lidar imaging for detecting and understanding plant responses and canopy structure. J. Exp. Bot. 2007, 58, 881–899. [Google Scholar] [CrossRef] [PubMed]
- Vadez, V.; Kholová, J.; Hummel, G.; Zhockhavets, U.; Gupta, S.K.; Tom Hash, C. LeasyScan: A novel concept combining 3D imaging and lysimetry for high-troughput phenotyping of traits of traits controlling plant water budget. J. Exp. Bot. 2015. [Google Scholar] [CrossRef] [PubMed]
- Boussac, A.; Sugiura, M.; Kirilowsky, D.; Rutherford, W. Near-infrared-induced transitions in the manganese cluster of photosystem II. Action spectra for the S2 and S3 redox state. Plant Cell Physiol. 2005, 46, 837–842. [Google Scholar] [CrossRef] [PubMed]
- Thapper, A.; Mamedov, F.; Mokvist, F.; Hammarström, L.; Styring, S. Defining the far-red limit of photosystem II in Spinach. Plant Cell 2009, 21, 2391–2401. [Google Scholar] [CrossRef] [PubMed]
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef] [PubMed]
- The R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing; R Core Team: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 8 June 2015).
- Seidel, D.; Beyer, F.; Hertel, D.; Fleck, S.; Leuschner, C. 3D-laser scanning: A non-destructive method for studying above-ground biomass and growth of juvenile trees. Agric. For. Meteorol. 2011, 151, 1305–1311. [Google Scholar] [CrossRef]
- Bunce, J.A. Growth-Rate, photosynthesis and respiration in relation to leaf-area index. Ann. Bot. 1989, 63, 459–463. [Google Scholar]
- Poiré, R.; Chochois, V.; Sirault, X.R.R.; Vogel, J.P.; Watt, M.; Furbank, R.T. Digital imaging approaches for phenotyping whole plant nitrogen and phosphorus response in Brachypodium distachyon. J. Int. Plant Biol. 2014, 8, 781–796. [Google Scholar] [CrossRef] [PubMed]
- Ruts, T.; Matsubara, S.; Wiese-Klinkenberg, A.; Walter, A. Aberrant temporal growth pattern and morphology of root and shoot caused by a defective circadian clock in Arabidopsis thaliana. Plant J. 2012, 72, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Satter, R. Leaf movements and tendril curling. In Encyclopedia of Plant Physiology, New Series, Volume 7. Physiology of Movements; Haupt, W., Feinleib, M.E., Eds.; Springer Verlag: Berlin, Germany, 1979; pp. 442–484. [Google Scholar]
- Liu, C.C.; Welham, C.V.; Zhang, X.Q.; Wang, R.Q. Leaflet movement of Robinia pseudoacacia in response to a changing light environment. J. Int. Plant Biol. 2007, 49, 419–424. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kjaer, K.H.; Ottosen, C.-O. 3D Laser Triangulation for Plant Phenotyping in Challenging Environments. Sensors 2015, 15, 13533-13547. https://doi.org/10.3390/s150613533
Kjaer KH, Ottosen C-O. 3D Laser Triangulation for Plant Phenotyping in Challenging Environments. Sensors. 2015; 15(6):13533-13547. https://doi.org/10.3390/s150613533
Chicago/Turabian StyleKjaer, Katrine Heinsvig, and Carl-Otto Ottosen. 2015. "3D Laser Triangulation for Plant Phenotyping in Challenging Environments" Sensors 15, no. 6: 13533-13547. https://doi.org/10.3390/s150613533
APA StyleKjaer, K. H., & Ottosen, C.-O. (2015). 3D Laser Triangulation for Plant Phenotyping in Challenging Environments. Sensors, 15(6), 13533-13547. https://doi.org/10.3390/s150613533







