Optical Fiber Nanotips Coated with Molecular Beacons for DNA Detection
Abstract
:1. Introduction
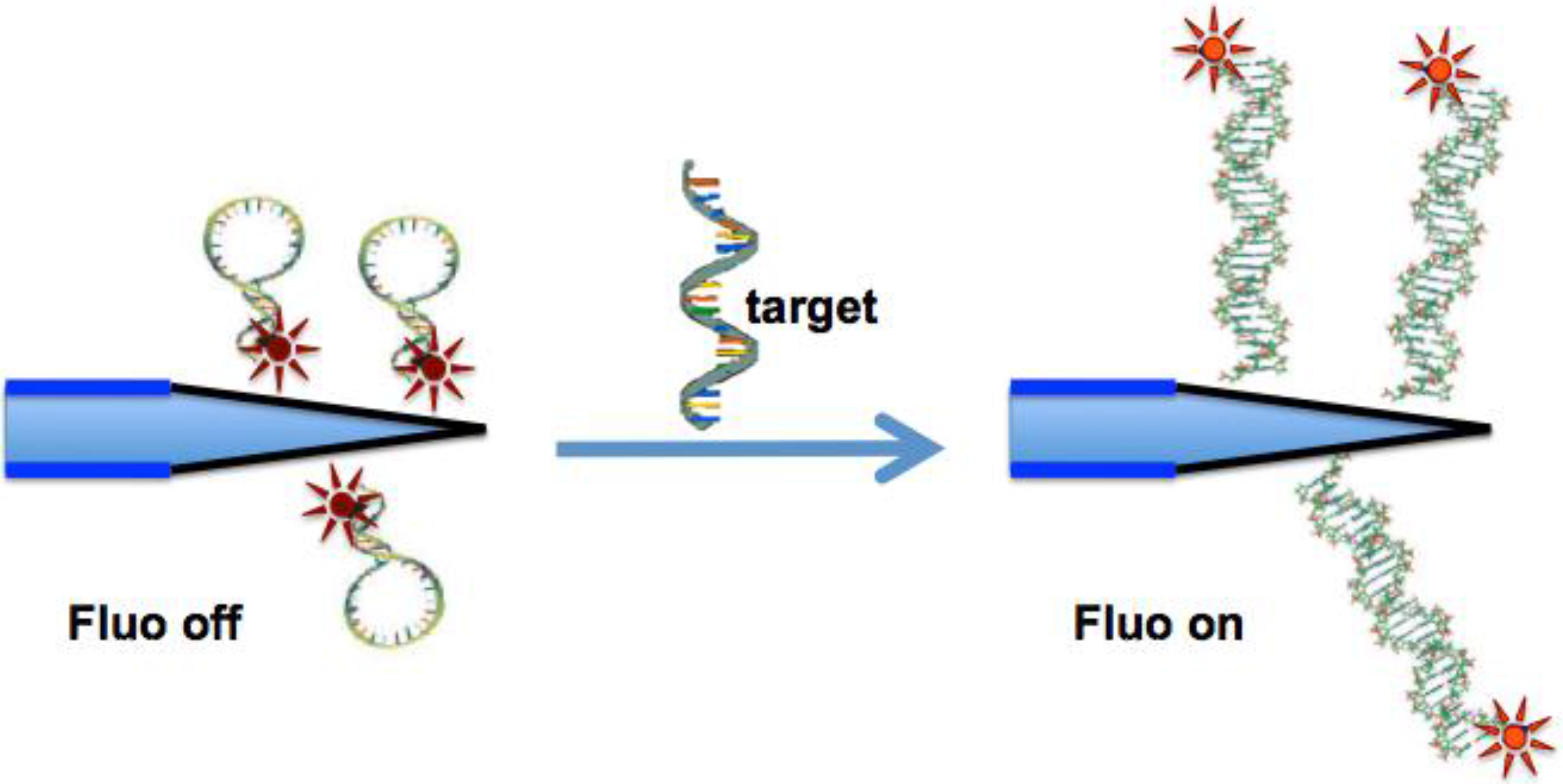
2. Experimental Section
2.1. Chemicals
2.2. Molecular Beacon and Target Sequences
- MB 5'-(ATTO647N)CGACGGAGAAAGGGCTGCCACGXCG(BBQ)-3' X=C6-dT Thio
- Target 5'-CCCCTGCCTGGCAGCCCTTTCTCAAGGACC-3'
- Random sequence 5'-ATCGGTGCGCTTGTCG-3'
- Linear probe 5'-(ATTO647N)GAGAAAGGGCTGCCA(Thiol)-3'
2.3. Optical Fiber Nanotip Preparation
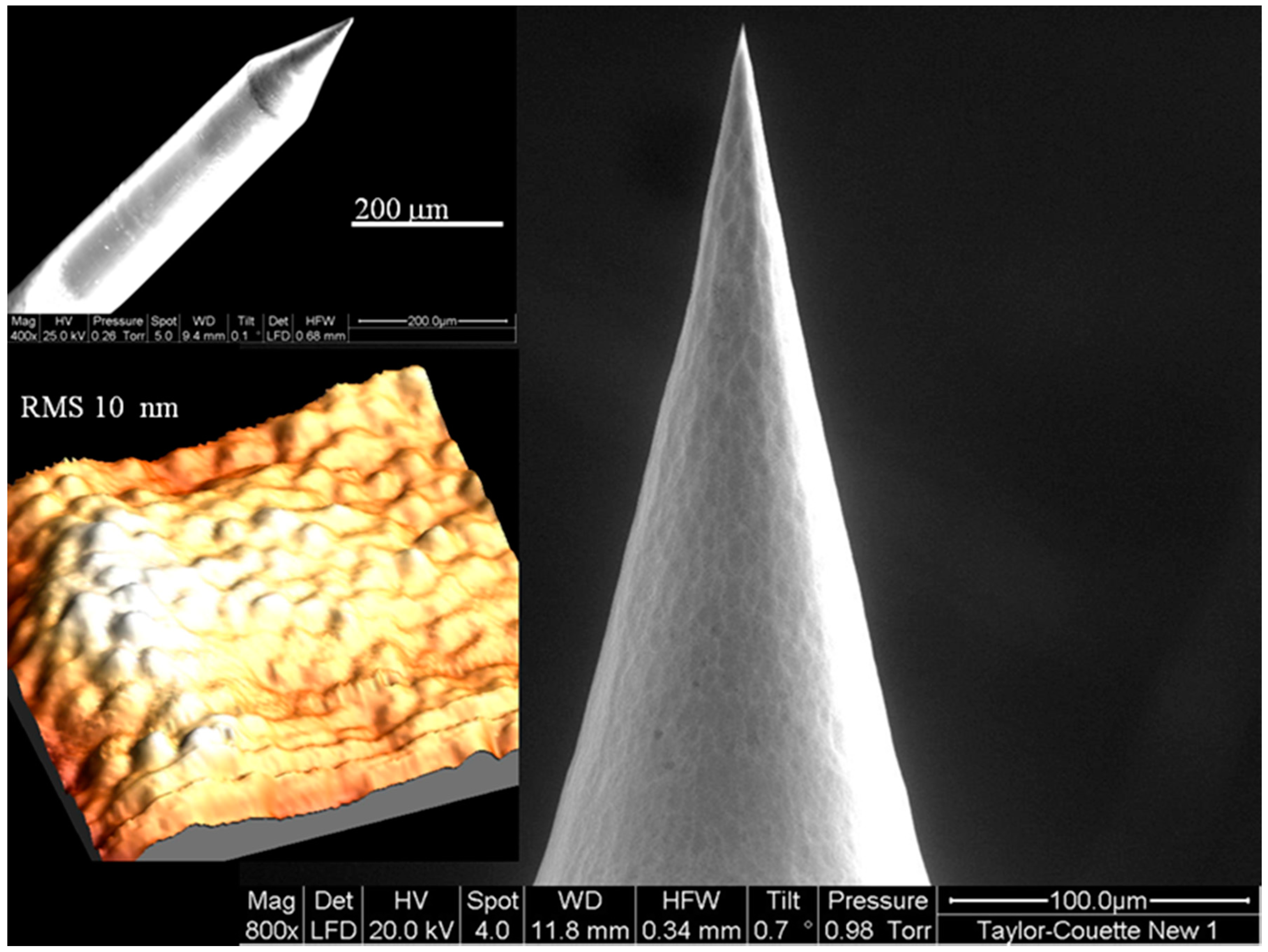

2.4. The Optical Measurements
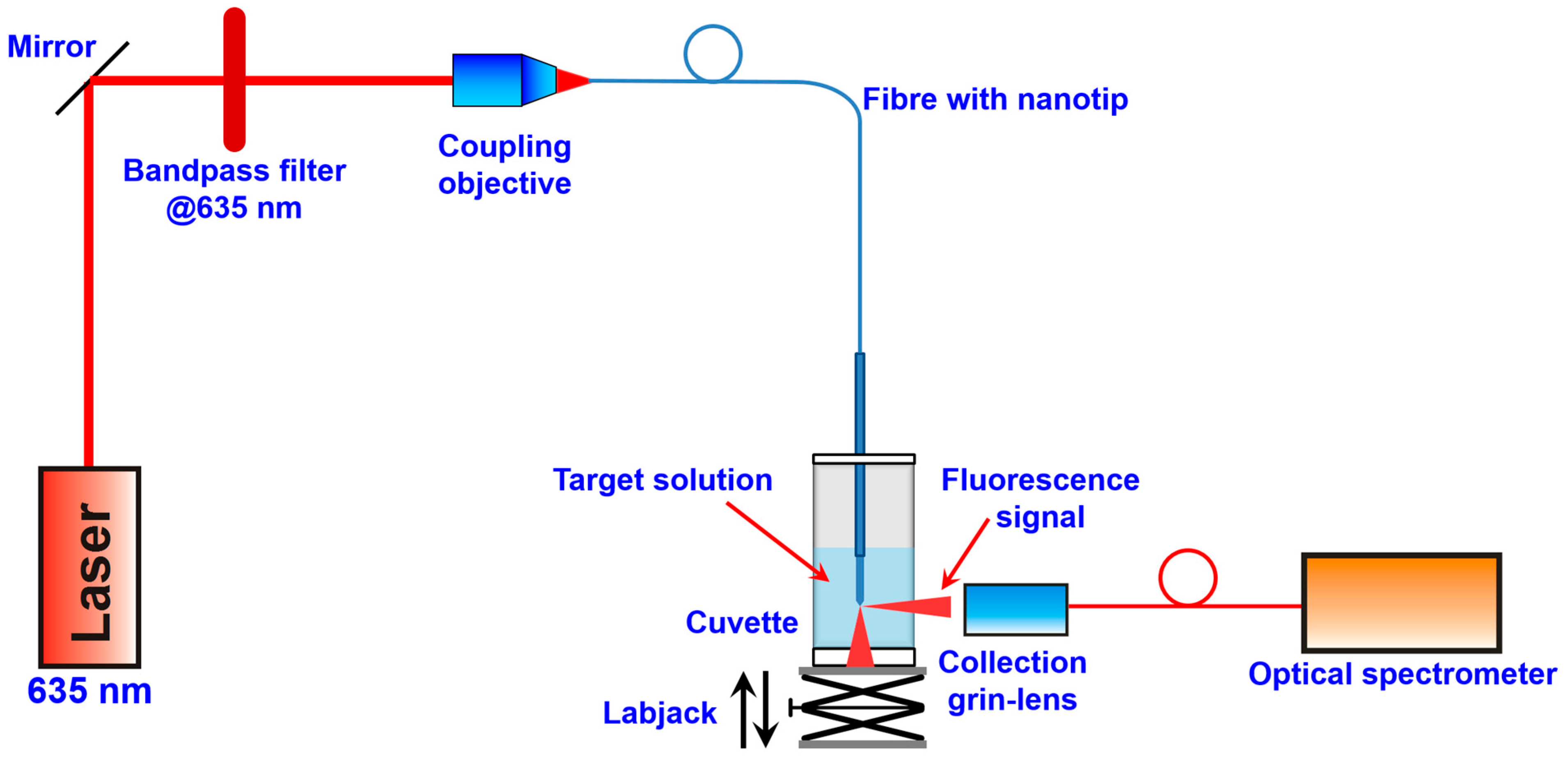
3. Results and Discussion
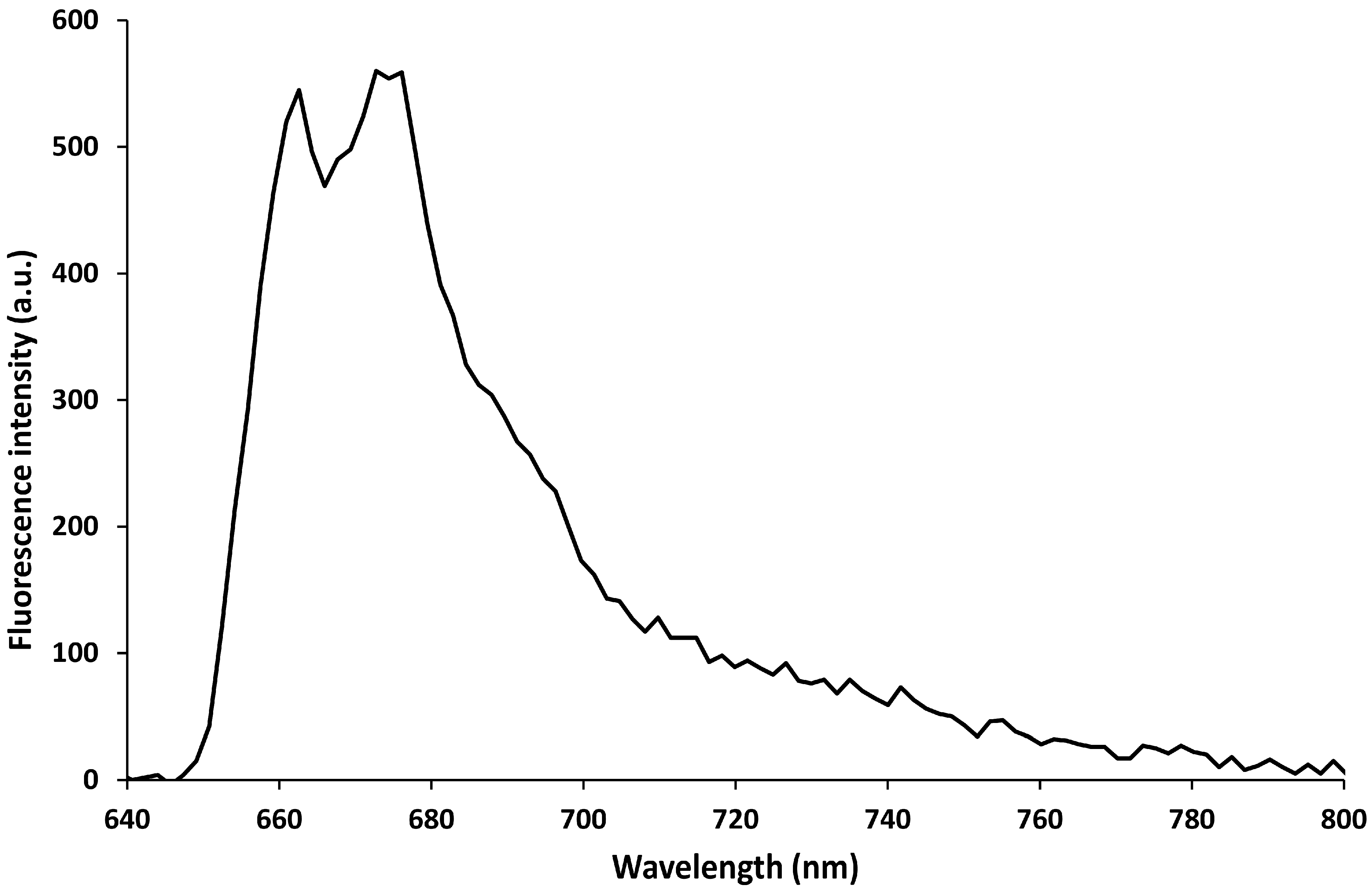
3.1. MB Characterization in Solution
3.2. MB and Linear Probe Characterization onto the Nanotip

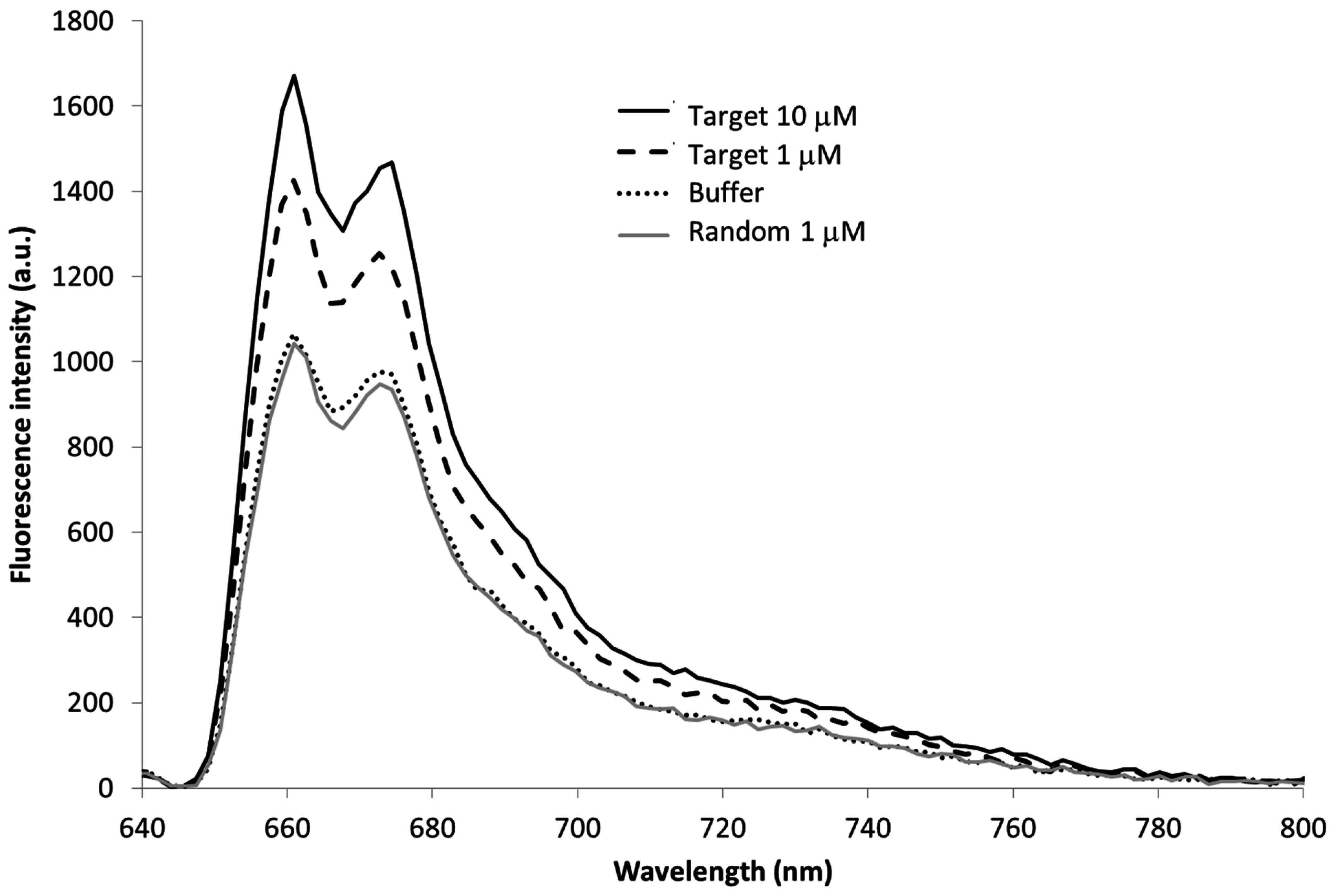
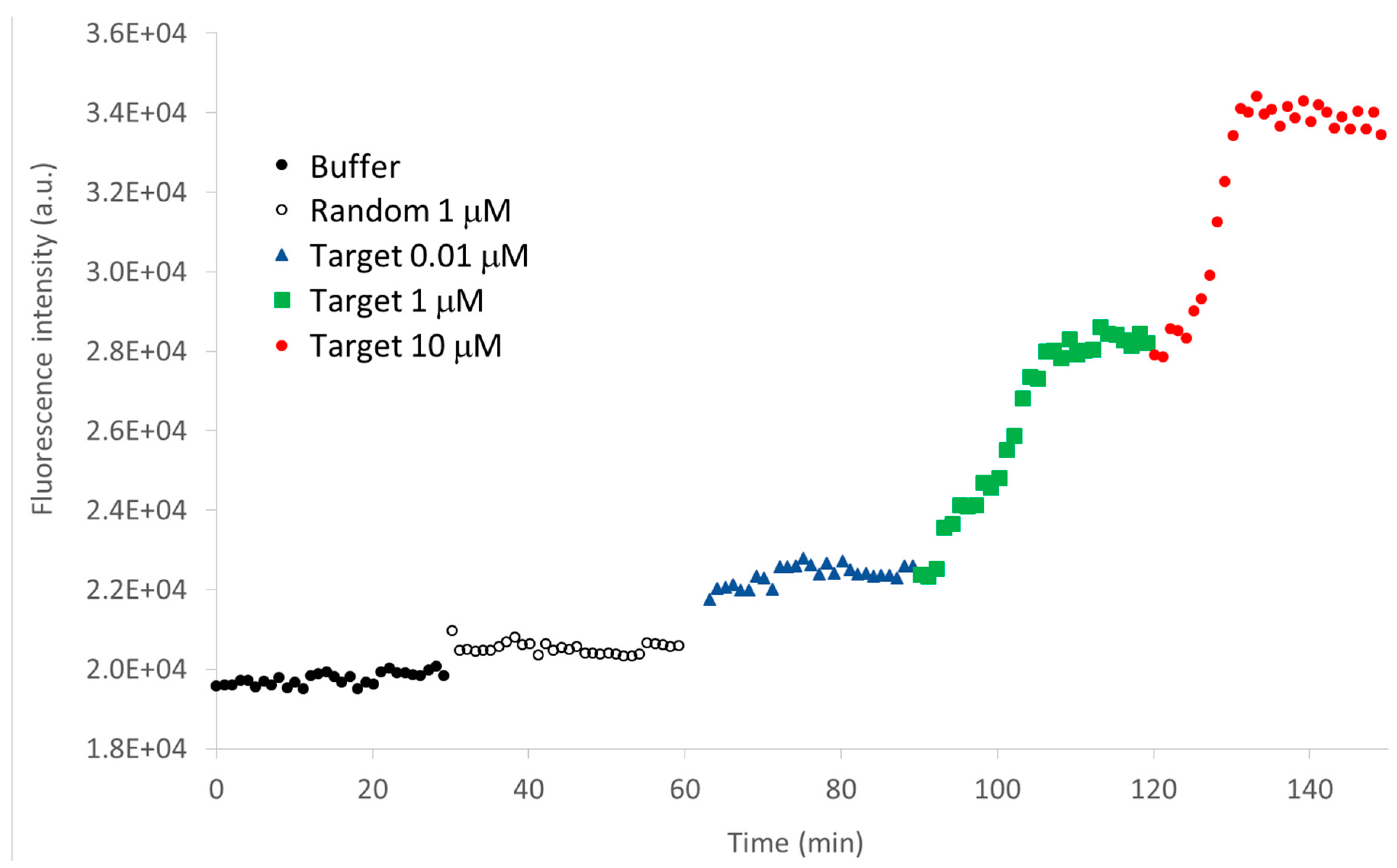
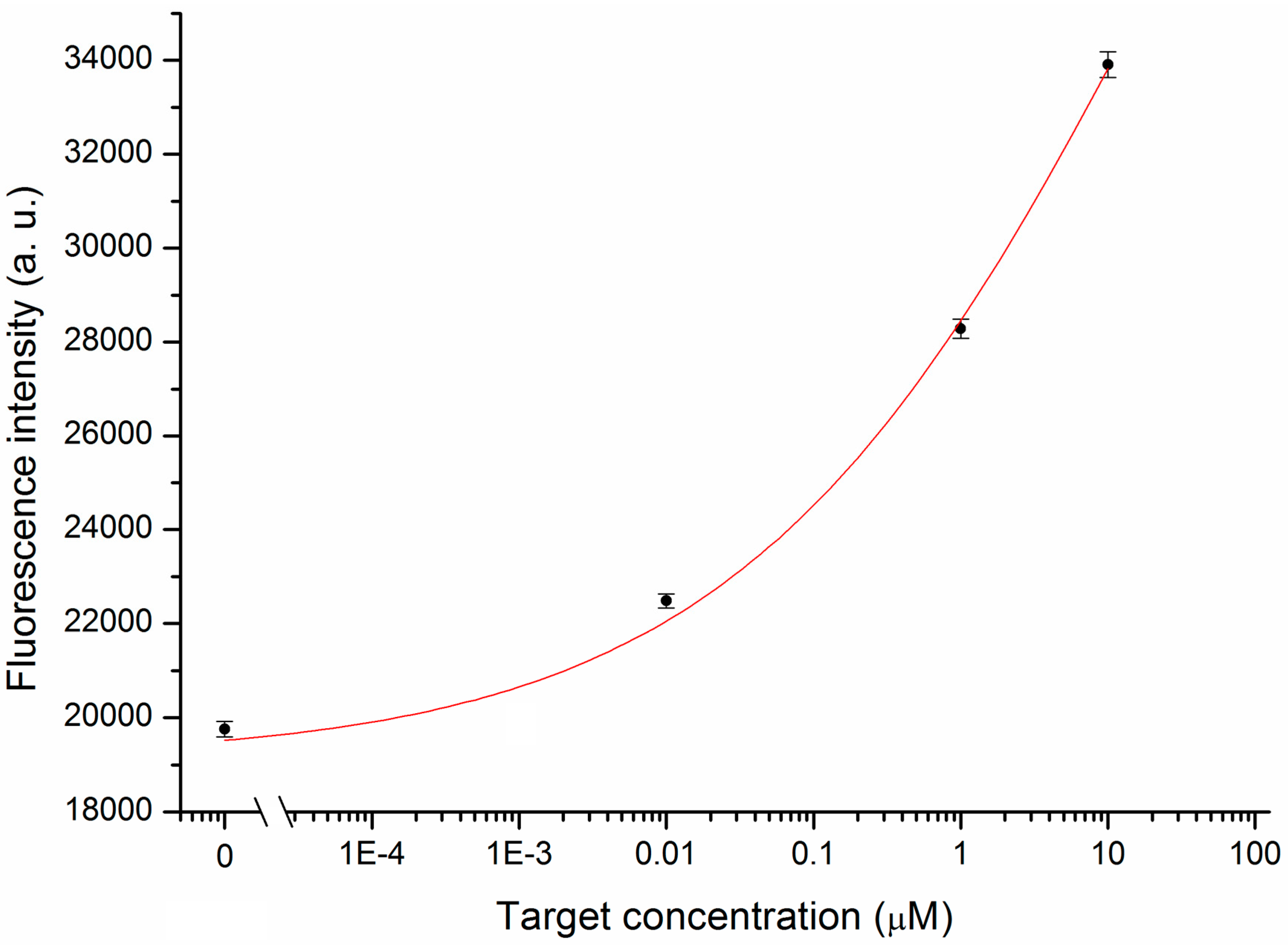
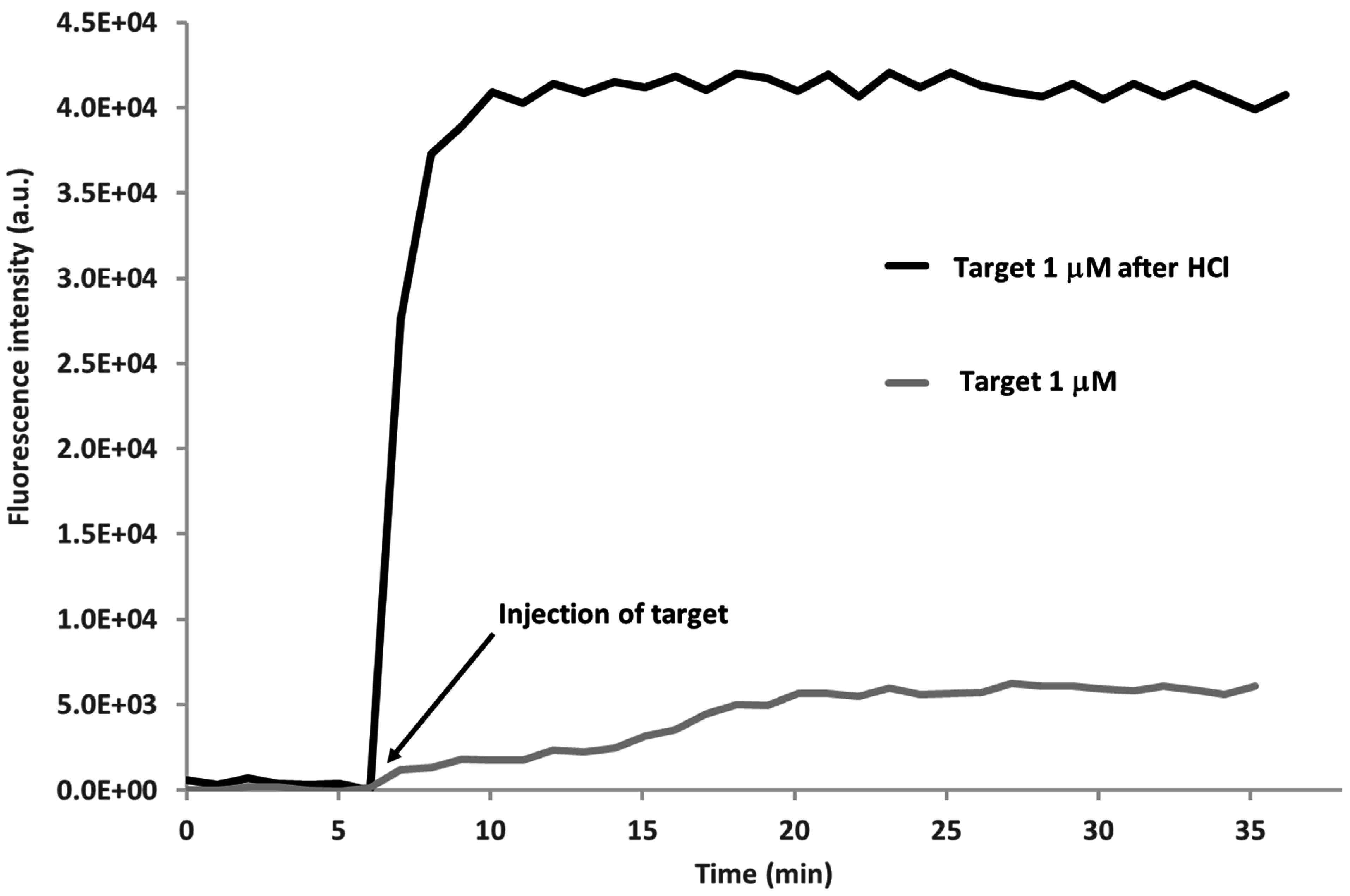
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Vo-Dinh, T. Nanosensing at the single cell level. Spectrochim. Acta Part B At. Spectrosc. 2008, 63, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.T.; Hu, W.; Wang, H.; Yang, H.; Zhou, W.; Li, C.M. Bifunctional electro-optical nanoprobe to real-time detect local biochemical processes in single cells. Biosens. Bioelectron. 2011, 26, 4484–4490. [Google Scholar] [CrossRef] [PubMed]
- Dahan, M.; Lévi, S.; Luccardini, C.; Rostaing, P.; Riveau, B.; Triller, A. Diffusion dynamics of glycine receptors revealed by single-quantum dot tracking. Science 2003, 302, 442–445. [Google Scholar] [CrossRef] [PubMed]
- Pathak, S.; Cao, E.; Davidson, M.C.; Jin, S.; Silva, G.A. Quantum Dot Applications to Neuroscience: New Tools for Probing Neurons and Glia. J. Neurosci. 2006, 26, 1893–1895. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.; Lin, Z.; Yang, N.; Li, Y.; Su, X.-G. A novel carboxymethyl chitosan–quantum dot-based intracellular probe for Zn2+ ion sensing in prostate cancer cells. Acta Biomater. 2014, 10, 868–874. [Google Scholar] [CrossRef] [PubMed]
- Duchi, S.; Sotgiu, G.; Lucarelli, E.; Ballestri, M.; Dozza, B.; Santi, S.; Guerrini, A.; Dambruoso, P.; Giannini, S.; Donati, D.; et al. Mesenchymal stem cells as delivery vehicle of porphyrin loaded nanoparticles: Effective photoinduced in vitro killing of osteosarcoma. J. Control Release 2013, 168, 225–237. [Google Scholar] [CrossRef] [PubMed]
- Giannetti, A.; Tombelli, S.; Baldini, F. Oligonucleotide optical switches for intracellular sensing. Anal. Bioanal. Chem. 2013, 405, 6181–6196. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, V.; Epple, M. Inorganic Nanoparticles as Carriers of Nucleic Acids into Cells. Angew. Chem. Int. Ed. 2008, 47, 1382–1395. [Google Scholar] [CrossRef]
- Ratto, F.; Matteini, P.; Centi, S.; Rossi, F.; Pini, R. Gold nanorods as new nanochromophores for photothermal therapies. J. Biophoton. 2010, 4, 64–73. [Google Scholar] [CrossRef]
- Kihara, T.; Yoshida, N.; Kitagawa, T.; Nakamura, C.; Nakamura, N.; Miyakea, J. Development of a novel method to detect intrinsic mRNA in a living cell by using a molecular beacon-immobilized nanoneedle. Biosens. Bioelectron. 2010, 26, 1449–1454. [Google Scholar] [CrossRef] [PubMed]
- Yum, K.; Wang, N.; Yu, M.-F. Nanoneedle: A multifunctional tool for biological studies in living cells. Nanoscale 2010, 2, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Dufrêne, Y.; Garcia-Parajo, M.F. Recent progress in cell surface nanoscopy: Light and force in the near-field. Nano Today 2012, 7, 390–403. [Google Scholar] [CrossRef]
- Zheng, X.T.; Li, C.M. Single living cell detection of telomerase over-expression for cancer detection by an optical fibre nanobiosensor. Biosen. Bioelectron. 2010, 25, 1548–1552. [Google Scholar] [CrossRef]
- Zheng, X.T.; Yang, H.B.; Li, C.M. Optical Detection of Single Cell Lactate Release for Cancer Metabolic Analysis. Anal. Chem. 2010, 82, 5082–5087. [Google Scholar] [CrossRef] [PubMed]
- Vitol, E.A.; Orynbayeva, Z.; Friedman, G.; Gogotsi, Y. Nanoprobes for intracellular and single cell surface-enhanced Raman spectroscopy (SERS). J. Raman Spectrosc. 2012, 43, 817–827. [Google Scholar] [CrossRef]
- Brasuel, M.; Kopelman, R.; Kasman, I.; Miller, T.J.; Philbert, M.A. Ion Concentrations in Live Cells from Highly Selective Ion Correlations Fluorescent Nano-Sensors for Sodium. IEEE Proc. 2002, 1, 288–292. [Google Scholar]
- McCulloch, S.; Uttamchandani, D. Development of a fibre optic micro-optrode for intracellular pH measurements. IEE Proc. Optoelectron. 1997, 144, 162–167. [Google Scholar] [CrossRef]
- Xu, H.; Aylott, J.W.; Kopelman, R.; Miller, T.J.; Philbert, M.A. A Real-Time Ratiometric Method for the Determination of Molecular Oxygen Inside Living Cells Using Sol−Gel-Based Spherical Optical Nanosensors with Applications to Rat C6 Glioma. Anal. Chem. 2001, 73, 4124–4133. [Google Scholar] [CrossRef] [PubMed]
- Vo-Dinh, T.; Alarie, J.P.; Cullum, B.M.; Griffin, G.D. Antibody-based nanoprobe for measurement of a fluorescent analyte in a single cell. Nat. Biotechnol. 2000, 18, 764–767. [Google Scholar] [CrossRef] [PubMed]
- Vo-Dinh, T.; Kasili, P. Fibre-optic nanosensors for single-cell monitoring. Anal. Bioanal. Chem. 2005, 382, 918–925. [Google Scholar] [CrossRef] [PubMed]
- Vo-Dinh, T.; Kasili, P.; Wabuyele, M. Nanoprobes and nanobiosensors for monitoring and imaging individual living cells. Nanomed. Nanotechnol. Biol. Med. 2006, 2, 22–30. [Google Scholar] [CrossRef]
- Zhang, Y.; Dhawan, A.; Vo-Dinh, T. Design and Fabrication of Fibre-Optic Nanoprobes for Optical Sensing. Nanoscale Res. Lett. 2011, 6. [Google Scholar] [CrossRef]
- Zhang, J.; Laiwalla, F.; Kim, J.A.; Urabe, H.; Van Wagenen, R.; Song, Y.-K.; Connors, B.W.; Zhang, F.; Deisseroth, K.; Nurmikko, A.V. Integrated device for optical stimulation and spatiotemporal electrical recording of neural activity in light-sensitized brain tissue. J. Neural Eng. 2009, 6. [Google Scholar] [CrossRef]
- Tyagi, S.; Kramer, F.R. Molecular beacons: Probes that fluoresce upon hybridization. Nat. Biotechnol. 1996, 14, 303–308. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Farmerie, W.; Schuster, S.; Tan, W. Molecular Beacons for DNA Biosensors with Micrometer to Submicrometer Dimensions. Anal. Biochem. 2000, 283, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Monroy-Contreras, R.; Vaca, L. Molecular Beacons: Powerful Tools for Imaging RNA in Living Cells. J. Nucleic Acids 2011, 2011. [Google Scholar] [CrossRef]
- Santangelo, P.J. Molecular beacons and related probes for intracellular RNA imaging. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2010, 2, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Chen, L.; Long, Y.; Tian, H.; Wu, J. Molecular Beacons of Xeno-Nucleic Acid for Detecting Nucleic Acid. Theranostics 2013, 3, 395–408. [Google Scholar] [CrossRef] [PubMed]
- Boutorine, A.S.; Novopashina, D.S.; Krasheninina, O.A.; Nozeret, K.; Venyaminova, A.H. Fluorescent Probes for Nucleic Acid Visualization in Fixed and Live Cells. Molecules 2013, 18, 15357–15397. [Google Scholar] [CrossRef] [PubMed]
- Hecht, B.; Sick, B.; Wild, U.P.; Deckert, V.; Zenobi, R. Scanning near-field optical microscopy with aperture probes: Fundamentals and applications. J. Chem. Phys. 2000, 112, 7761–7774. [Google Scholar] [CrossRef]
- Carpi, C.; Fogli, S.; Giannetti, A.; Adinolfi, B.; Tombelli, S.; Da Pozzo, E.; Vanni, A.; Martinotti, E.; Martini, C.; Breschi, M.C.; et al. Theranostic properties of a survivin-directed molecular beacon in human melanoma cells. PLoS ONE 2014, 11. [Google Scholar] [CrossRef]
- Adinolfi, B.; Carpi, S.; Giannetti, A.; Nieri, P.; Pellegrino, M.; Sotgiu, G.; Tombelli, S.; Trono, C.; Varchi, G.; Baldini, F. Complex nanostructures based on oligonucleotide optical switches and nanoparticles for intracellular mRNA sensing and silencing. Proc. Eng. 2014, 87, 751–754. [Google Scholar] [CrossRef]
- Nitin, N.; Santangelo, P.J.; Kim, G.; Nie, S.; Bao, G. Peptide-linked molecular beacons for efficient delivery and rapid mRNA detection in living cells. Nucleic Acids Res. 2004, 32. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, P.J.; Nix, B.; Tsourkas, A.; Bao, G. Dual FRET molecular beacons for mRNA detection in living cells. Nucleic Acids Res. 2004, 32. [Google Scholar] [CrossRef] [PubMed]
- Qiao, G.; Gao, Y.; Li, N.; Yu, Z.; Zhuo, L.; Tang, B. Simultaneous Detection of Intracellular Tumor mRNA with Bi-Color Imaging Based on a Gold Nanoparticle/Molecular Beacon. Chem. Eur. J. 2011, 17, 11210–11215. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; An, R.; Zhang, D.; Zhao, J.; Wang, X.; Yang, L.; He, D. Detection of survivin expression in cervical cancer cells using molecular beacon imaging: New strategy for the diagnosis of cervical cancer. Eur. J. Obstet. Gynecol. Reprod. Biol. 2011, 159, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Barucci, A.; Cosi, F.; Pelli, S.; Soria, S.; Nunzi Conti, G.; Giannetti, A.; Righini, G. Method of Fabricating Structures, Starting from Material Rods. Patent Pending PCT/EP2014/071743, 10 October 2014. [Google Scholar]
- Barucci, A.; Cosi, F.; Giannetti, A.; Pelli, S.; Griffini, D.; Insinna, M.; Salvadori, S.; Tiribilli, B.; Righini, G.C. Optical fibre nanotips fabricated by a dynamic chemical etching for sensing applications. J. Appl. Phys. 2015, 117. [Google Scholar] [CrossRef]
- Taylor, G.I. Stability of a Viscous Liquid Contained between Two Rotating Cylinders. Philosoph. Trans. R. Soc. London Ser. A Contain. Pap. Math. Phys. Charact. 1923, 223, 289–343. [Google Scholar] [CrossRef]
- Andereck, C.D.; Liu, S.S.; Swinney, H.L. Flow regimes in a circular Couette system with independently rotating cylinders. J. Fluid Mech. 1986, 164, 155–183. [Google Scholar] [CrossRef]
- Tombelli, S.; Ballestri, M.; Giambastiani, G.; Giannetti, A.; Guerrini, A.; Sotgiu, G.; Trono, C.; Tuci, G.; Varchi, G.; Baldini, F. Oligonucleotide switches and nanomaterials for intracellular mRNA sensing. Proc. SPIE 2013, 8798. [Google Scholar] [CrossRef]
- Pini, V.; Tiribilli, B.; Gambi, C.M.C.; Vassalli, M. Dynamical characterization of vibrating AFM cantilevers forced by photothermal excitation. Phys. Rev. B. 2010, 81. [Google Scholar] [CrossRef]
- Giannetti, A.; Baldini, F.; Ballestri, M.; Ghini, G.; Giambastiani, G.; Guerrini, A.; Sotgiu, G.; Tombelli, S.; Trono, C.; Tuci, G.; et al. Intracellular nanosensing and nanodelivery by PMMA nanoparticles. Lect. Notes Electri. Eng. 2014, 162, 69–75. [Google Scholar]
- Giannetti, A.; Tombelli, S.; Trono, C.; Ballestri, M.; Giambastiani, G.; Guerrini, A.; Sotgiu, G.; Tuci, G.; Varchi, G.; Baldini, F. Intracellular delivery of molecular beacons by PMMA nanoparticles and carbon nanotubes for mRNA sensing. Proc. SPIE. 2013, 8596. [Google Scholar] [CrossRef]
- Lechuga, L.M. New frontiers in optical biosensing. In Proceedings of the 13th European Conference on Integrated Optics (ECIO 2007), Copenhagen, Denmark, 25–27 April 2007.
- Mannelli, I.; Minunni, M.; Tombelli, S.; Wang, R.; Spiriti, M.M.; Mascini, M. Direct immobilization of DNA probes for the development of affinity biosensors. Bioelectrochemistry 2005, 66, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos Riccardi, C.; Yamanaka, H.; Josowicz, M.; Kowalik, J.; Mizaikoff, B.; Kranz, C. Label-free DNA detection based on modified conducting polypyrrole films at microelectrodes. Anal. Chem. 2006, 78, 1139–1145. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giannetti, A.; Barucci, A.; Cosi, F.; Pelli, S.; Tombelli, S.; Trono, C.; Baldini, F. Optical Fiber Nanotips Coated with Molecular Beacons for DNA Detection. Sensors 2015, 15, 9666-9680. https://doi.org/10.3390/s150509666
Giannetti A, Barucci A, Cosi F, Pelli S, Tombelli S, Trono C, Baldini F. Optical Fiber Nanotips Coated with Molecular Beacons for DNA Detection. Sensors. 2015; 15(5):9666-9680. https://doi.org/10.3390/s150509666
Chicago/Turabian StyleGiannetti, Ambra, Andrea Barucci, Franco Cosi, Stefano Pelli, Sara Tombelli, Cosimo Trono, and Francesco Baldini. 2015. "Optical Fiber Nanotips Coated with Molecular Beacons for DNA Detection" Sensors 15, no. 5: 9666-9680. https://doi.org/10.3390/s150509666
APA StyleGiannetti, A., Barucci, A., Cosi, F., Pelli, S., Tombelli, S., Trono, C., & Baldini, F. (2015). Optical Fiber Nanotips Coated with Molecular Beacons for DNA Detection. Sensors, 15(5), 9666-9680. https://doi.org/10.3390/s150509666











