Phyllopshere Bacterial Community Structure of Spinach (Spinacia oleracea) as Affected by Cultivar and Environmental Conditions at Time of Harvest
Abstract
: Modern molecular ecology techniques were used to demonstrate the effects of plant genotype and environmental conditions prior to harvest on the spinach epiphytic bacterial community. Three cultivars of spinach with different leaf topographies were collected at three different periods during the fall growing season. Leaf surface topography had an effect on diversity and number of culturable bacteria on the phylloepiphtyic community of spinach. Savoy cultivars, which had larger surface area and more stomata and glandular trichomes, where bacterial aggregates were observed, featured more diverse communities with increased richness and larger bacterial populations compared to flat-leaved cultivars. Bacterial community richness was compared using denaturant gradient gel electrophoresis (DGGE), while abundance was quantified using 16s rRNA primers for major phyla. The most diverse communities, both in richness and abundance, were observed during the first sampling period, immediately following a period of rapid spinach growth. Exposure to lower air and soil temperatures and decreased precipitation resulted in significantly reduced bacterial population size and bacterial community richness in November and December. This study describes the effect of the plant characteristics and environmental conditions that affect spinach microbiota population size and diversity, which might have implications in the survival of food and plant bacterial pathogens.1. Introduction
Phyllosphere microbial communities are diverse, consisting of bacteria, fungi, algae, protozoa and, infrequently, nematodes. Phylloepiphytic bacteria that successfully colonize the leaf surface must survive adverse conditions including nutrient limitation, low water availability, and UV irradiation which all affect bacterial fitness and diversity [1,2]. The membership of these phyllosphere communities is dominated by members of a few phylogenetic groups, chiefly α-proteobacteria, γ-proteobacteria and Bacteroidetes. β-proteobacteria and Firmicutes also are a large part of the bacterial community, while Acidobacteria, Actinobacteria and Cyanobacteria occur infrequently [3]. Recent culture-independent analyses of leaf washings of several plants indicated that phylloepiphytic bacterial communities are more complex than had been reported by culture-based methods. Novel bacterial species have been recently reported from leaf surfaces of several crop species, vegetables and trees [4].
Epiphytic bacterial population sizes and diversity are influenced by environmental conditions [5], plant species [6], plant cultivar [7,8], and stage of growth [9,10]. Differences in temperature, rainfall and UV exposure, which typically fluctuate with season, are associated with total culturable bacteria on leaves of maize, sugar beets and olive trees [11-13]. In relation to cultivar, differences in leaf surface topography and nutrients are important factors that affect the microbial community of the phyllosphere [14,15]. Bacteria are associated in larger numbers with leaf structures including trichomes, veins, stomata and epidermal cell wall surfaces [6,16,17]. Nutrient compositions, water availability and plant secondary metabolites influence total microbial populations [7,18]. It is possible that changes in leaf blade topography might be related to changes on the leaf surface, influencing the microbial diversity and population size on plants.
In recent years, spinach consumption was implicated in an E. coli O157:H7 outbreak [19]. Study of the phyllosphere of spinach leaves would improve the understanding of how pathogenic microorganisms are able to populate edible plants during pre- and postharvest handling since possible interactions with the members of the bacterial populations on plants might play a role on their survival. The purpose of this work was to determine how the bacterial diversity of the spinach leaf epiphytic community changes in relation to environmental conditions on savoy, semi-savoy and flat-leaf spinach cultivars. Characterization of the membership of the epiphytic microbial community and changes in its structure will provide a better understanding of the ecology of the spinach phyllosphere that could be applied to elucidate mechanisms of survival for plant or food pathogens.
2. Experimental Section
2.1. Spinach Production and Harvest
Three spinach (Spinacia oleracea) cultivars with different leaf topographies, ‘Monza’ (flat), ‘Menorca’ (savoy) and ‘Unipak’ (semi-savoy) (Seedway LLC. USA), were seeded on September 19, 2007 at the Virginia Tech Kentland Research Farm. One plot of 6 × 3.75 m was equally subdivided into four subplots, approximately 1.5 × 6 m. Each subplot contained four rows on raised beds, each seeded with a different cultivar. Organic agricultural practices were followed with nutrients applied using composted vegetable waste and no pesticides were used long-term. Drip irrigation was applied every two days from seeding until irrigation was terminated on November 21, 2007. Two hundred grams of spinach leaves were harvested when the leaf measured 10 cm long measured from petiole base of the leaf to the leaf tip (“baby” leaf stage) from plots randomly selected using a round plastic hoop, covering approximately 0.30 m2. The leaves were harvested with scissors at the base of the petioles and stored in self-sealing sterile plastic bags. Scissors were wiped with ethanol before use and between harvests of different cultivars. Samples were immediately taken to a laboratory, stored at approximately 4 °C, and processed within 3 h. Leaves were harvested 36 days after seeding on October 25, 2007 (October harvest), after 58 days on November 17, 2007 (November harvest) and after 78 days on December 7, 2007 (December harvest).
Environmental conditions, maximum and minimum daily air and soil temperatures and precipitation were recorded at a weather station ( http://www.vaes.vt.edu/colleges/kentland/weather/) approximately 500 m from the field plot at the Virginia Tech Kentland Research Farm (Table 1). The growing degree units (GDUs) were calculated as: Σ[(daily maximum temperature + daily minimum temperature)/2] − Base temperature.
2.2. Microbial Counts and DNA Isolation from Epiphytic Community on Spinach Leaves
For each cultivar, 10 g of spinach leaves were placed in a filtered bag with 90 mL of 1% (wt/vol) peptone water (Sigma-Aldrich Co., USA) and processed in a Pulsifier® (Microbiology International, Frederick, MD, USA) for 5 min. The pulsifier removed bacteria attached to the surface of the spinach leaves, without disrupting the tissue, and reduced the chlorophyll and spinach DNA in the sample [20]. All extractions were replicated in triplicate. Ten milliliters of the suspension were serially diluted and plated onto R2A media (Difco, MI, USA). Bacterial cells were harvested from the remaining volume by centrifugation, resuspended in 1X PBS (phosphate buffered saline; 1.37 mM NaCl, 1 mM Phosphate, 0.27 mM KCl, pH 7.4) and stored at −20 °C. DNA was extracted using the ZR soil microbe DNA kit™ (Zymo Research Co., Orange, CA, USA) per manufacturer's instructions.
2.3. Measurement of Plant Anatomical Features
The leaf area for each cultivar at the “baby” leaf stage was measured using a leaf area meter (model LI3000 LI-COR Biosciences, Lincoln, NE). Three replicates of 10, 10 cm long immature leaves each were selected and each leaf measurement was repeated three times. Five individual 10cm long leaves of each cultivar were collected from each subplot to determine the stomata density using differential interference contrast (DIC) microscopy (Axiophot compound microscope Zeiss, Thornwood, NY, USA) with 200× magnification. The numbers of stomata were manually counted on both the abaxial and adaxial surfaces of each leaf in five fields of approximately 0.051 mm2 each. Leaf surfaces were covered with clear nitrocellulose cotton solution that was removed along with the cuticle after drying with clear tape (Scotch, 3M, Minneapolis, MN, USA) to obtain stomatal imprints for counting. Stomata density was converted to stomata per cm2. The density of glandular trichomes was assessed using a dissection microscope (Cambridge Instruments, Watertown, MA, USA). Five leaves of each cultivar were selected and trichomes were counted in 1 cm2 on abaxial and adaxial surfaces of each leaf.
Microbiota surrounding stomata were observed on gold-coated abaxial surface of mature leaves using environmental scanning electronic microscopy (FEI quanta 600 FEG) under high vacuum (approximately 7 × 10−5 Torr).
2.4. DGGE Analysis of PCR Amplified 16S rRNA Gene Fragments
The 16S rRNA gene was amplified from the total epiphytic DNA (50 ng/μL) to generate a 566 bp fragment using the primers 341-F (5′-CCT ACG GGA GGC AGC AG-3′) and 907R (5′-CCG TCA ATT CMT TTG AGT TT-3′) [21]. The forward primer was modified to add a 40 nucleotide GC clamp (314GC) at the 5′ end (5′-CGC CCG CCG CGC GCG GCG GGC GGG GCG GGG GCA CGG GGG G-3′) [21]. Each 25 μL reaction contained 1.5 mM of MgCl2, 50 mM of KCl, 0.2 mM of each dinucleotide, 1% of DMSO (dimethylsulfoxide), 25 mM of Tris-HCl (pH 8), 1U/μL of HotStart-IT FideliTaq DNA polymerase (USB 71156, Cleveland, OH), 0.5 μM of primers 341GC and 907R, and 50 ng of DNA. The size and intensity of PCR products were confirmed using a 0.9% agarose gels (Fisher-Scientific, Atlanta, GA).
The PCR products were run on a 8% polyacrylamide gel in a 30-60% denaturant gradient of urea and formamide (100% denaturant corresponds to 7 M urea plus 40% (vol/vol) of deionized formamide) using the Bio-Rad DCode™ Universal Detection System (Bio-Rad, Hercules, CA, USA). Twenty two microliters of PCR products were separated at constant 85 V and 60 °C for 17 h. The DNA amplicons were visualized by staining with ethidium bromide (5 ug/mL) and photographed using the Molecular Imager ® GelDoc™ XR (Bio-Rad). The amplicons were analyzed using Quantity One®-1D analysis software (Bio-Rad) and the DGGE profiles were analyzed by clustering using the unweighted pair group method with mathematical averages (UPGMA; Dice coefficient of similarity) using the same software producing dendrograms for the 3 cultivars at each harvest time. Two different gels were analyzed for each sample.
Amplicons consistently present in all three cultivars and at all harvest times were excised and suspended in 100 μL of water, and incubated at 4 °C for 24 h to allow passive DNA diffusion. Diffused DNA was re-amplified using 341f and 907r primers as described above. The PCR products were purified from a 1% (w/v) agarose gel and cloned into pCR4-TOPO® VECTOR (Invitrogen) to transform One Shot® Mach1 ™ chemically competent E. coli (Invitrogen). Plasmids were isolated using Quick lyse miniprep (Qiagen) and inserts were sequenced. Sequences were compared to the NCBI database using BLASTN and identities assigned based on similarity.
2.5. Phylum Analysis Using Real Time PCR
Microbial community abundance was assessed by amplification of phyla specific regions of the 16S RNA gene as described by Blackwood et al. 2005 [22]. Phylum-specific abundance was determined targeting the following phylogenetic groups: α–proteobacteria, β-proteobacteria, Firmicutes, Actinobacteria and Bacteroidetes (Table 1).
Standard curves for real-time PCR analysis were constructed for each phylum using 16S rDNA universal primers for bacteria previously isolated from spinach leaves and identified at the genus level using 16S rDNA universal primers (Table 1). Serial dilutions of purified DNA were made to create 10-fold serial dilutions from 0.1 to 100 ng/μL. Standard curves using real time PCR amplification were prepared using the phylum-specific primers listed in Table 2 using: 100, 10, 1, and 0.1 ng. Each 25 μL reaction contained respective amounts of DNA template, 12.5 μL of HotSart-IT™SYBR®Green qPCR Master Mix 2X that contains 5 mM and 0.4 mM of MgCl2 and nucleotides, respectively (USB® 75770 Cleveland, OH, USA), 10nM of fluorescein as passive reference dye (USB® 75767 Cleveland, OH, USA) and 0.5 μM of forward and reverse primers. PCR conditions were denaturation at 95 °C for 2 min, followed by 40 cycles of denaturation at 95 °C for 30 s, 30 s at the annealing temperature, and 72 °C for 1 min. Annealing temperatures were: 60 °C for α-proteobacteria, β-proteobacteria, Actinobacteria, Firmicutes and 65 °C for Bacteroidetes [23]. Each concentration in the standard curve was done in triplicate using separated plates. Melting curve analysis of the PCR products was conducted following each assay to confirm that the fluorescence signal originated from a specific PCR product. Amplification was carried out with an iQ™ Optical system Real-time PCR detection system (Bio-Rad). Primers specificity was confirmed by testing primers with DNA of pure cultures of previously isolated spinach epiphytes belonging to the phylum analyzed (Table 1). All Ct values for bacteria that did not belong to the specific genera being tested ranged from 35 to 40, and a Ct value of 30 or less for spinach samples was considered for analysis (data not shown).
After confirmation of linearity with the specific primers was addressed, DNA samples from the three spinach cultivars and from the three times of sampling were adjusted to 50 ng/μL and real-time amplification was performed in duplicate as described for the standard curves using 50 ng of template.
2.6. Statistical Analysis
Bacterial enumeration experiments were repeated three times for each cultivar and sampling period with duplicate plate counts. Data was normalized by conversion to log10 CFU/g−1 of spinach. For phyla analysis results were performed on DNA isolated from two different sets of spinach leaves with a duplicate measurement. Statistical comparison among treatments was conducted using the MIX procedure and means were separated using Tukey's multiple comparison. All statistical calculations were assessed with Statistical Analysis System 9.2 (SAS Institute, Cary, NC, USA). Statistical significance was reported when p < 0.05.
3. Results
3.1. Environmental Conditions
From planting to first harvest, temperatures were warmest (Table 2). During the first harvest period, spinach plants were exposed to the most days over 25 °C, and temperatures never exceeded 25 °C after first harvest. All spinach cultivars were harvested as immature “baby” spinach after accumulating about 1000 GDUs (Table 2). Only 190 GDUs were accumulated in November and 85 in December so little crop development occurred after first harvest. Subfreezing temperatures were not reached until after first harvest. Average minimum temperatures fell from 10 degrees during the first harvest period to below zero for the remaining two periods. Soil temperatures declined gradually during each period, but remained above freezing. Precipitation for both September/October and November were very similar, 11.5 cm for both periods, while December was much drier (Table 2).
3.2. Aerobic Total Microbial Counts
Significantly larger microbial counts (p < 0.05) were recorded for savoy ‘Menorca’ and ‘Unipak’ than for the flat-leaf cultivar Monza at all sampling periods. The total numbers of aerobic culturable bacteria were significantly larger during the months of October and December (Table 3) for the three cultivars and harvest periods (p < 0.05) than those recorded in November (Table 3). After the population decline in November, the microbial counts recovered in December but only populations of ‘Monza’ recovered to the level observed in October.
3.3. Spinach Anatomical Features
The mean leaf surface area of cultivars differed by approximately 0.2 cm2 each. The surface area of the savoy ‘Menorca’ was largest, while the flat ‘Monza’ had the smallest mean leaf surface area (Table 4). Stomata densities on abaxial versus adaxial leaf surfaces were significantly different for each cultivar, however only ‘Monza’ possessed fewer stomata on the lower surface compared to the other cultivars (Table 4). Glandular trichomes were observed only on the mature leaves of ‘Menorca’ and ‘Unipak’ (Table 4).
Microbial aggregates were observed surrounding stomata on all three cultivars (Figure 1), but only on ‘Unipak’ were aggregates directly observed on guard cells. The largest microbial density was observed on ‘Menorca’ leaves.
3.4. DGGE Analysis
The V3 region of 16S rRNA gene was amplified, and the resulting amplicons were analyzed using DGGE. The number of amplicons (bands) corresponded to the species richness of the microbial community for each cultivar harvested during different time periods. Differences in number and position of amplicons were noted among the harvest periods and among three cultivars (results not shown). Seven amplicons, which were excised and 16S rDNA amplicons sequenced, were present in electropherograms of samples harvested in October and November for all three cultivars. Amplicons from DGGE were identified as belonging to the phyla Proteobacteria, Bacteroidetes and Actinobacteria (Table 5). The greatest species richness was observed during October. A few amplicons were absent from all cultivars in November (Table 5).
The majority of sequences were identified as nonculturable bacteria. Some bands were represented by more than one microorganism.
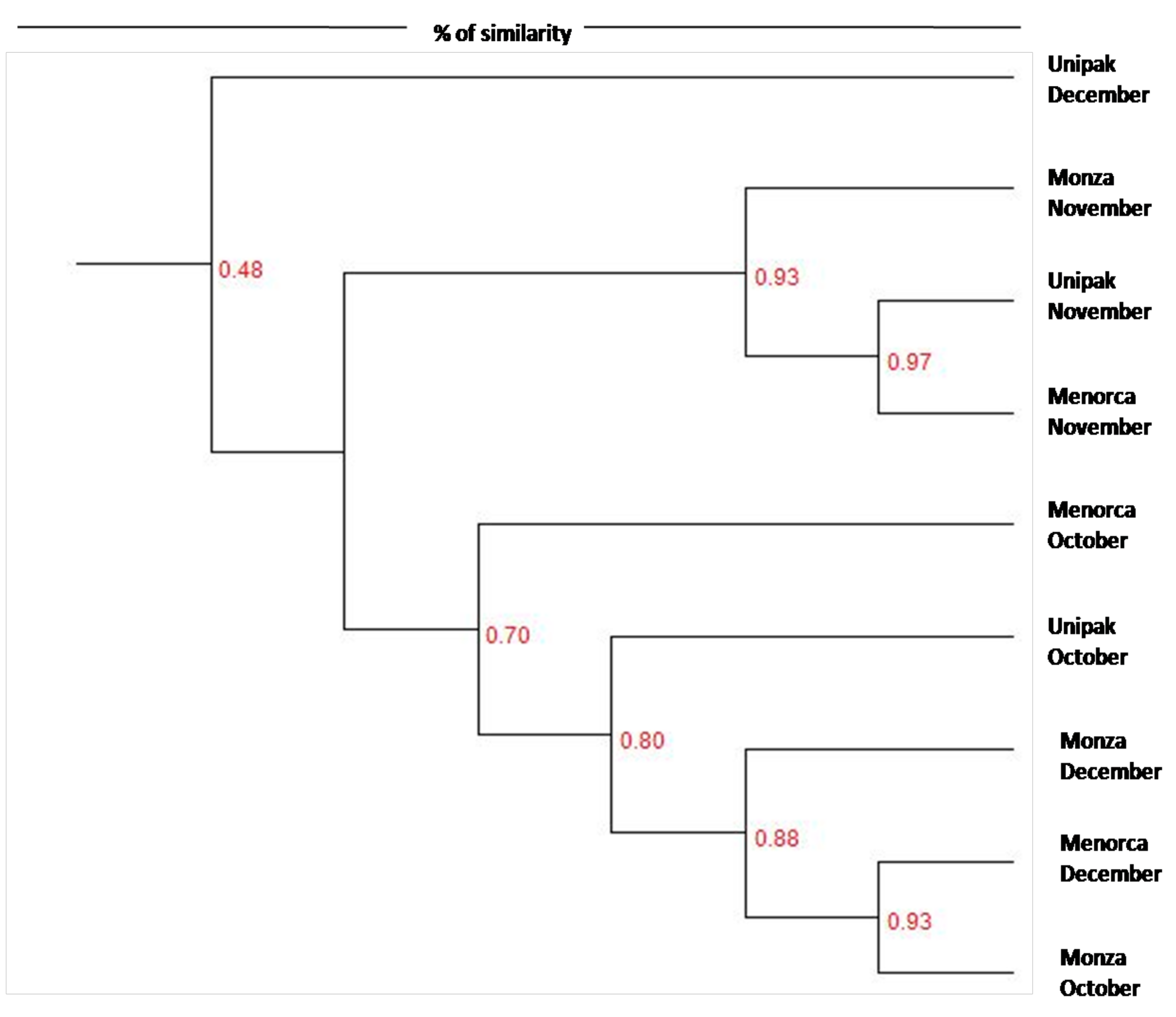
Dendrograms were constructed to distinguish similarities between community profiles (Figure 2). Two distinct clusters were identified. The first cluster is represented by ‘Unipak’ during December and the second cluster is further divided into two clusters; one contains bacterial community members from the three cultivars during November, while the other cluster consists of bacterial community members from cultivars harvested during October and December. The clustering indicates more similarity between communities harvested from spinach in October and December compared to communities harvested from spinach in November.
3.5. Determination of Specific Phyla Abundance and Diversity Indices
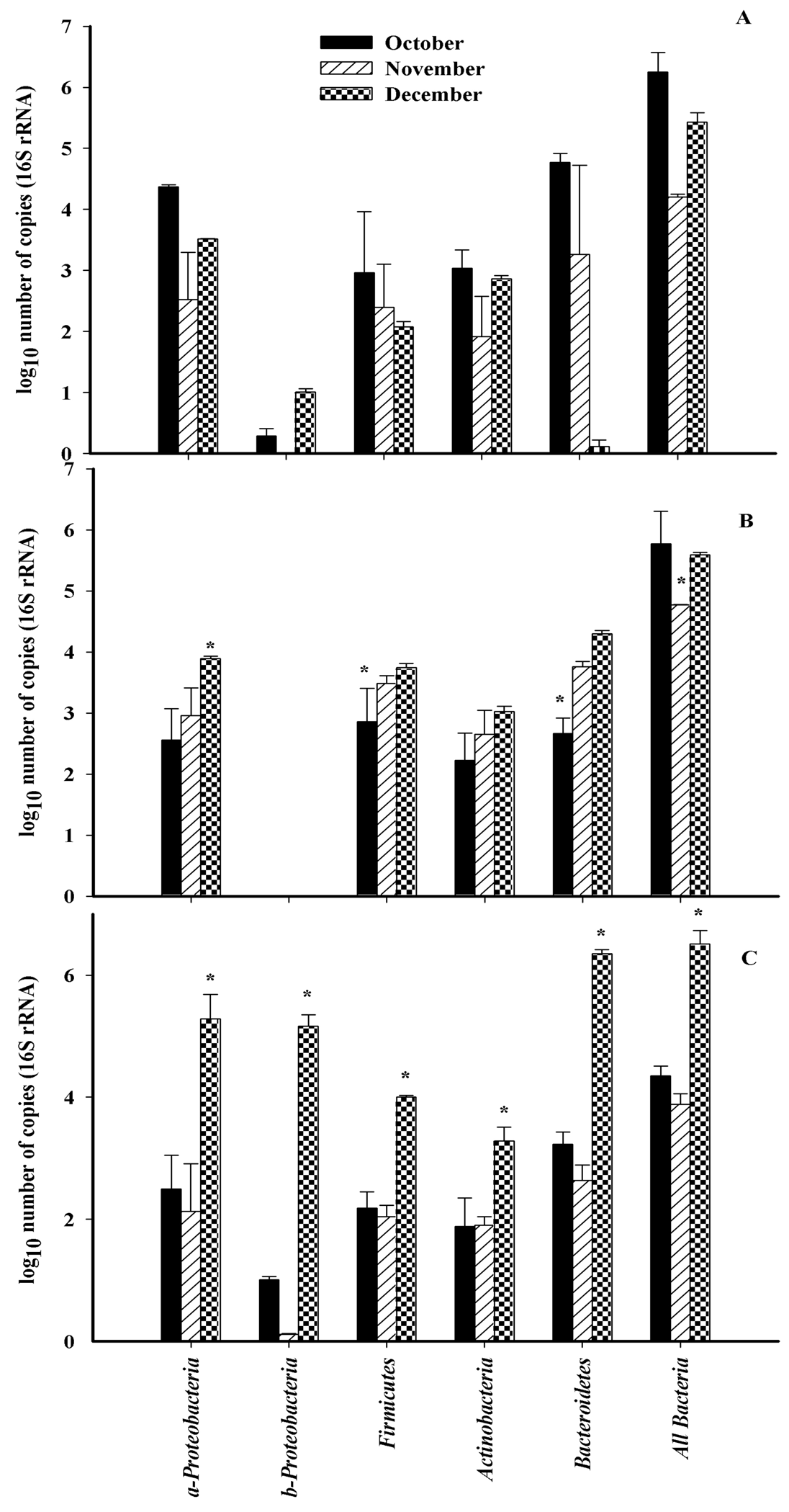
The abundance of all bacteria was determined using qRT-PCR using universal 16S rRNA primers. Community structure at the phylum level was determined using primers designed to specific regions of the 16S rDNA targeting all bacteria, Actinobacteria, Firmicutes, α-protoebacteria, β-proteobacteria and Bacteroidetes (Figure 3). The total number of bacterial 16S rDNA sequences detected in October and December were significantly higher (p < 0.05) than bacterial sequences detected in November for all three cultivars. The most abundant 16S rDNA sequences belonged to the α-proteobacteria, and Bacteroidetes (Figure 3). Members of the β-proteobacteria were the least abundant. The total number of bacterial sequences for α-proteobacteria and Bacteroidetes were significantly less abundant in November for all three cultivars. The abundance of sequences increased in December with the exception of Bacteroidetes on ‘Monza’. Actinobacteria and Firmicutes, were the most stable members of the community.
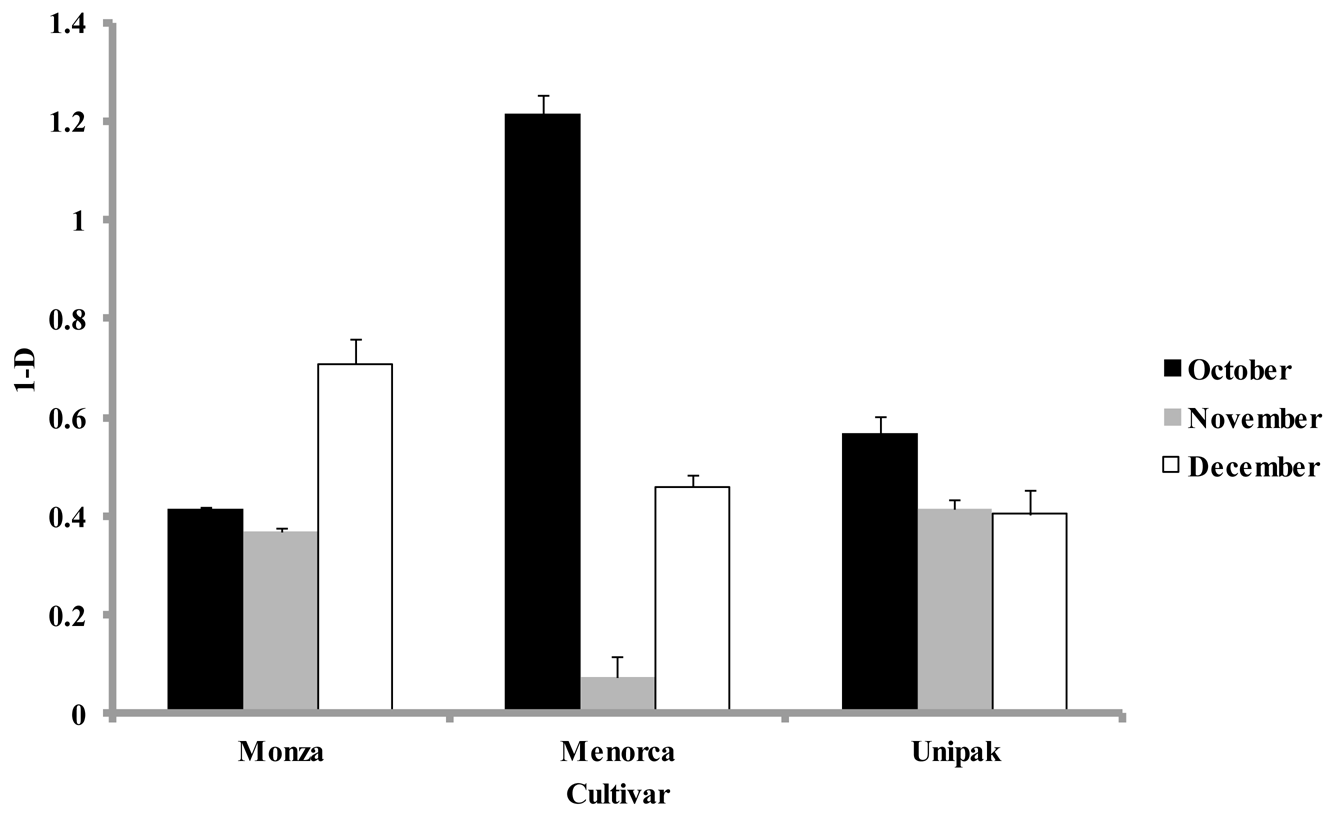
Simpson's diversity index, calculated at each harvest time for all cultivars using the abundance of individual groups, were significantly lower (p < 0.05) for samples collected during November than for October and December (Figure 4). The diversity indices for ‘Monza’ were not significantly different in samples from October and November (Figure 4). However, samples collected in December showed a significant increase in their diversity index. The largest diversity index during October was for ‘Menorca’, yet during November this cultivar had the lowest diversity index. ‘Unipak’ had a larger diversity index in October, but declined in November and December without significant differences between these months (p > 0.05). Real-time PCR showed no significant differences in phylum composition among cultivars collected during the same harvest periods, which was also observed in DGGE patterns obtained for all three cultivars.
4. Discussion
The phyllopshere bacterial community of spinach is influenced by environmental conditions prior to harvest (temperature and precipitation) and the host genotype (cultivar). In this study, the largest numbers of culturable bacteria, greatest species richness and abundances were seen at the first harvest in October. In November, the numbers of culturable bacteria, species richness and abundance declined. In December a small but significant increase in number of total and culturable bacteria were observed, with the cultivar Monza recovering culturable bacteria to numbers seen in October (Table 2 and Figure 3). In addition the samples from the majority of cultivars clustered together between the months of October and December (Figure 2). In December plants were harvested the day after a rainfall event that consisted of a combination of liquid and frozen rain, which may explain these differences. Intense rainfall events have been shown to precede the onset of rapid growth of Pseudomonas syringae on snap bean leaves under field conditions [14]. Additionally, the reduced air temperatures may have weakened the plant cells, allowing force from the raining ice crystals to damage the plant cells, releasing nutrients for bacterial growth. Alternatively, the increase in numbers of bacteria in December compared to November may reflect increases in psychrotrophic bacteria on spinach leaves. Increased populations and reduced bacterial counts have been observed on spinach produced in winter months in Italy [26] as seen in this study. The changes to microbial abundance and richness seen in November and December are similar to that seen on packaged leaves held at refrigeration temperatures [27].
The plant stage of development and leaf age may impact the bacterial community on the phyllopshere [9,11,28]. Previous studies indicate that microbial diversity patterns differed between immature and mature plants, with increased species richness seen in younger leaves of olive trees and on sugar beet leaves [11,13]. Additionally, the richness and diversity of the spinach phyllosphere were influenced by the stage of plant development [9], but was only minimally affected by time from leaf emergence [29]. Depletion of available nitrogen and carbon sources is associated with decreased survival of E.coli O157:H7 on older leaves of Romaine lettuce in comparison to younger inner leaves (30). In this study, all cultivars developed to the immature “baby” stage, like those typically harvested for modified atmosphere packaged salads, after accumulating approximately 1000 GDUs from planting to first harvest in October. Plants entered a non-growing, vegetative state after October because the amount of GDUs accumulated were below that necessary for spinach growth [31] (Table 1). Therefore, plants at all three sampling periods were at the same stage of development (6–8 leaf stage). To reduce the bias associated with leaf age, all leaves from each plant were combined, therefore presenting the possibility to examine changes to bacterial richness and abundance associated with environmental conditions and harvest period (time from seeding until harvest).
Variations in the numbers of bacteria with leaf blade topography have been previously reported [3]. Broad leaf plant species, such as cucumber, lettuce and beans typically have larger aerobic microbial counts than grasses [5]. Cultivar differences affect the ability of Pseudomonas syringae pv. syringae to colonize snap beans [10], and for Salmonella enterica to colonize lettuce [17]. We selected three spinach cultivars; Monza, Menorca and Unipak, that differed in leaf surface area (Table 2). In this study, the savoy “Menorca” and semi-savoy “Unipak” had significantly higher diversity indices and larger numbers of culturable bacteria, than the flat leaf “Monza” during the first sampling period. The Diversity indices varied between cultivars as well (Figure 4). In particular the diversity index was highest for the flat cultivar “Monza” in the month of December, while the other cultivars showed highest diversity in the warmer October month. This suggests that leaf blade topography influenced the number and metabolic strategies of microorganisms harbored on the plant surface. Changes attributed to cultivar may be related to differences in total surface area that serves as niches for microbial establishment and development. In this study significantly larger mean leaf area was measured for “Menorca”. Savoy cultivars such as Menorca and Unipak have leaf blades that are curly between leaf veins, increasing the leaf surface area available for colonization. “Menorca” and “Unipak” had more anatomical leaf features such as glandular trichomes and stomata (Table 3, Figure 1), where microbial aggregates and bacteria were observed. Veins, hooked trichomes and glandular trichomes, have preferential distribution associated with bacterial aggregates on leaf surfaces of beans [32] and strawberry leaves [28]. Foodborne pathogens such as Listeria monocytogenes and E. coli O157:H7 associated with leaves of Arabidopsis thaliana and spinach, respectively, have also shown to be associated with these plant structures [7,33]. DGGE profiles of the three cultivars are most similar during the month of November (Figure 3). However, the numbers of bacteria in different phyla were significantly different among cultivars harvested in November and December (Figure 4), indicating that the changes associated with spinach cultivars are due to differences in abundance but not in richness.
Overall, the microbial community of spinach is conserved and stable, as indicated by the presence of the same bands for all cultivars and at all harvest periods (Table 5). These bands likely represent bacteria that are robust, even in stressful conditions like low temperature. Actinobacteria and Firmicutes populations were maintained without significant variations during the season for the three spinach cultivars. This indicates that members of these two phyla maintain fitness regardless of changes in environmental conditions and therefore contribute to the overall stability of the community. In contrast, numbers of bacteria belonging to the phylogenetic group β-Proteobacteria (Figure 3) fluctuated greatly between October and December harvests and were the least stable members of the community. In this study the majority of the members of the epiphytic community belonged to the phylogenetic group α-Proteobacteria and Bacteroidetes. The same phylogenetic groups are dominant on spinach and other plant leaf surfaces including Zea mays and Capsicum annuum [4,34]. The dominant phylogenetic group on spinach and other leaf surfaces is the γ-Proteobacteria [3,34]. In this study we tried several available primer sets to examine changes to this class of bacteria, however non-specific amplication of phyllopshere bacteria belonging to other Proteobacteria and Actinobacteria occurred. Therefore the changes in this dominant bacterial class are not presented, but are likely to be biologically relevant to the phyllopshere community structure. Disadvantages related with the PCR reaction include different affinities of the designed oligos to the DNA template, different copy numbers of the target genes and the preferential amplification towards the more abundant members of the community as well as different amplification related with the GC content of the template, resulting in so called PCR bias [35,36] which can prevent amplification of other rRNA genes that will result in underestimation of the abundance. Microbial richness is underestimated using DGGE approaches due to PCR bias and co-migration of amplicons from different bacteria that have similar GC contents [21,37].
The microbial community structure of the spinach phyllosphere is impacted by the cultivar and environmental conditions during spinach development. Community members vary in fitness to decreased environmental temperatures and rainfall but after a period of adaptation, the phyllosphere community successfully recovered its abundance but not its richness. Time of harvest and weather conditions before harvest may have important implications for maintaining product quality [26] and potentially influence the establishment of human pathogens on the leaf surface. Recent review articles describe the frequent isolation of pathogens and bacteria closely related to Salmonella and E.coli from plant tissues and the rhizosphere, and describe mechanisms used by human pathogens to persist or even colonize plant systems [38,39]. Bacteria identified on the surface of spinach and other leafy greens have been shown to promote the growth of E.coli O157:H7 on leaf surfaces; however other bacteria have been identified that may reduce the growth of human pathogens [29,40,41]. Results of this study and others suggest that bio-control may offer a strategy to reduce persistence of human pathogens, however additional research is needed to produce pathogen- free crops. Research should work to identify bacteria that not only compete with human pathogens but also stimulate plant defenses to clear human pathogens. Future research should consider the cultivar, time of harvest and the post-harvest processing conditions on survival of human pathogens as strategies to maintain product quality, while minimizing risk of human illness.

| Phylogenetic target | Isolated bacterium used as positive control (# 16s rDNA copies)a | Forward primer (5′-3′) | Reverse primer (5′-3′) | Source of primer sequences |
|---|---|---|---|---|
| α-proteobacteria | Brevundimonas sp. (5) | -ACT CCT ACG GGA GGC AGC AG- | -TCT ACG RAT TTC ACC YCT AC- | [23] |
| Actinobacteria | Microbacterium sp. (7) | -CGC CGC CTA TCA GCT TGT TG- | -ATT ACC GCG GCT GCT GG- | [23] |
| Firmicutes | Exiguobacterium sp. (9) | -GCA GTA GGG AAT CTT CCG- | -ATT ACC GCG GCT GCT GG- | [23,24] |
| Bacteroidetes | Flavobacterium sp. (6) | -GTA CTG AGA CAC GGA CCA- | -ATT ACC GCG GCT GCT GG- | [23,25] |
| β-proteobacteria | Acidovorax sp. (3) | -ACT CCT ACG GGA GGC AGC AG- | -TCA CTG CTA CAC GYG- | [23,25] |
| All bacteria | Escherichia coli (7) | -ACT CCT ACG GGA GGC AGC AG- | -ATT ACC GCG GCT GCT GG- | [23] |
aCopy numbers obtained from http://rrndb.mmg.msu.edu/search.php.
| From day of seeding (September 19, 2007) to first harvest (October 25 2007) | From first harvest to second harvest (November 17 2007) | From second harvest to third harvest (December 7 2007) | |
|---|---|---|---|
| Sampling time | December | November | December |
| Accumulated GDUsc | 995 | 190 | 86 |
| Average of maximum air temperatures (°C) | 16.7 ± 3.7 | 6.8 ± 3.8 | 4.7 ± 4.4 |
| Average of minimum air temperatures (°C) | 9.8 ± 4.5 | –0.12 ± 4.8 | –2.0 ± 4.6 |
| Average of soil temperatures (°C) | 17.4 ± 2.0 | 10.4 ± 2.5 | 7.1 ± 2.0 |
| Total precipitation (cm) | 11.6 | 11.5 | 1.8 |
| Number of days at 25 °C or above | 13 | 0 | 0 |
| Number of days at 0 °C or below | 0 | 0 | 4 |
bData obtained from preliminary local climatic data at www.weather.gov/data/GSP, WS Forms F-6 for Blacksburg station for the months of September, October, November and December 2007.cAccumulated GDUs are the number of temperature degrees or heat units above 4.7 °C accumulated during the growing season, 5 °C is the base temperature to which plant growth is expected.
| Cultivar | log of CFU/g spinach* |
|---|---|
| Monza | |
| October Harvest | 5.93 ± 0.0331,C |
| November Harvest | 5.02 ± 0.0362,A |
| December Harvest | 5.84 ± 0.0601,A |
| Menorca | |
| October Harvest | 6.47 ± 0.0481,B |
| November Harvest | 4.58 ± 0.0652,B |
| December Harvest | 5.14 ± 0.0372,A |
| Unipak | |
| October Harvest | 7.47 ± 0.0951,A |
| November Harvest | 4.13 ± 0.0383,B |
| December Harvest | 5.49 ± 1.0902,A |
*Results are represented by the mean of three independent replicates ± standard deviation. Different numbers denote significant difference (p < 0.05) within each cultivar. Different letters denote significant difference (p < 0.05) within each month among the three cultivars.
| Leaf area Normalized area (leaf area/leaf length) | Stomata (no.of stomata/cm2) | Glandular trichomes (no.of glandular trichomes/cm2) | |||
|---|---|---|---|---|---|
| Cultivar | Entire leaf | Abaxial surface | Adaxial surface | Abaxial surface | Adaxial surface |
| Monza | 2.82 ± 0.37b | 2.63 x 104 ± 6370a | 3.69 × 104 ± 3368a | N/D | N/D |
| Menorca | 3.27 ± 0.12a | 3.28 x 104 ± 5590a | 4.79 × 104 ± 8207b | 9.15 ± 1.50b | 31.3 ± 6.79a |
| Unipak | 3.02 ± 0.11b | 2.63 x 104± 3069a | 5.35 × 104 ± 1375b | 3.00 ± 2.12b | 18.66 ± 9.81a |
dData represent means ± standard deviations of each measurement of 5 spinach leaves. Mean with different letter within a column are significantly different (p < 0.05). (N/D) glandular trichomes were not detected for ‘Monza’.
| Amplicon identification: closest match to NCBI database | % similarity | |
|---|---|---|
| Amplicons shared between all spinach cultivars at each harvest | Kineococcus spp. -FJ214365 | 97 |
| Oryzihumus spp. -GQ355280 | 99 | |
| Terrabacter spp. -FJ772036 | 99 | |
| Uncultured Actinobacteria- EF651015 | 89 | |
| Uncultured Oxalobacteraceae FJ037285 | 97 | |
| Uncultured Proteobacterium clone -EF602176 | 97 | |
| Uncultured β- Proteobacterium -EFO74569 | 98 | |
| Amplicons absent from November harvest | Flavobacterium spp. -AY599661 | 98 |
| Uncultured α– Proteobacteria -CU926667 | 95 | |
| Uncultured Baceroidetes bacterium -AM116743 | 96 | |
Acknowledgments
We acknowledge the assistance of A. Grant for manuscript formatting, Brinkley Benson for his contribution in the cultivation of spinach and Glenda Gillaspy for her expertise and assistance in the utilization of DIC microscopy. ESEM was carried out using instruments in the Nanoscale Characterization and Fabrication Laboratory, a Virginia Tech facility operated by the Institute for Critical Technology and Applied Science.
References and Notes
- Gnanamanickam, S.; Immanuel, J. Epiphytic bacteria, their ecology and functions. In Plant-Associated Bacteria; Gnanamanickam, S., Ed.; Springer: Dordrecht, The Netherlands, 2006; pp. 131–154. [Google Scholar]
- Wilson, M.; Lindow, S.E. Inoculum density-dependent mortality and colonization of the phyllosphere by Pseudomonas syringae. Appl. Environ. Microb. 1994, 60, 2232–2237. [Google Scholar]
- Whipps, J.M.; Hand, P.; Pink, D.; Bending, G.D. Phyllosphere microbiology with special reference to diversity and plant genotype. J. Appl. Microbiol. 2008, 105, 1744–1755. [Google Scholar]
- Yang, C.H.; Crowley, D.E.; Borneman, J.; Kenn, N.T. Microbial phyllosphere populations are more complex than previously realized. PNAS 2001, 98, 3889–3894. [Google Scholar]
- Kinkel, L.L.; Wilson, M.; Lindow, S.E. Plant species and plant incubation conditions influence variability in epiphytic bacterial population size. Microb. Ecol. 2000, 39, 1–11. [Google Scholar]
- Borges-Baldotto, L.E.; Lopes-Olivares, F. Phylloepiphytic interaction between bacteria and different plant species in a tropical agricultural system. Can. J. Microbiol. 2008, 54, 918–931. [Google Scholar]
- Mitra, R.; Cuesta-Alonso, E.; Wayadande, A.C.; Talley, J.; Gilliland, S.; Fletcher, J. Effect of route of introduction and host cultivar on the colonization, internalization, and movement of the human pathogen Escherichia coli O157:H7 in spinach. J. Food Prot. 2009, 72, 1521–1530. [Google Scholar]
- Rasche, F.; Trondl, R.; Naglreiter, C.; Reichenauer, T.G.; Sessitsch, A. Chilling and cultivar type affect the diversity of bacterial endophytes colonizing sweet pepper (Capsicum anuum L.). Can. J. Microbiol. 2006, 52, 1036–1045. [Google Scholar]
- Carder, P. A Study of the Microbial Community of Spinach at Various Stages of Plant Growth from Seed to Maturity. MS Thesis, Virginia Polytechnic Institute and State University, Blacksburg, VA, USA, 2010; pp. 33–56. [Google Scholar]
- Jacques, M.; Kinkel, L.L.; Morris, C.E. Population sizes, immigration, and growth of epiphytic bacteria on leaves of different ages and positions of field-grown endive (Cichorium endivia var. latifolia). Appl. Environ. Microb. 1995, 61, 899–906. [Google Scholar]
- Ercolani, G.L. Distribution of epiphytic bacteria on olive leaves and the influence of leaf age and sampling time. Microb. Ecol. 1991, 21, 35–48. [Google Scholar]
- Kadivar, H.; Stapleton, A.E. Ultraviolet radiation alters maize phyllosphere bacterial diversity. Microb. Ecol. 2003, 45, 353–361. [Google Scholar]
- Thompson, I.P.; Bailey, M.J.; Fenlon, J.S.; Fermor, T.R.; Lilley, A.K.; Lynch, J.M.; McCormack, P.J.; McQuilken, M.P.; Purdy, K.J. Quantitative and qualitative seasonal changes in the microbial community from the phyllosphere of sugar beet. Plant Soil. 1993, 150, 177–191. [Google Scholar]
- Hirano, S.S.; Baker, L.S.; Upper, C.D. Raindrop momentum triggers growth of leaf-associated populations of Pseudomonas syringae on field-grown snap bean plants. Appl. Environ. Microbiol. 1996, 62, 2560–2566. [Google Scholar]
- Kinkel, L.L. Microbial population dynamics on Leaves. Annu. Rev. Phytopathol. 1997, 35, 327–347. [Google Scholar]
- Klerks, M.; Franz, E.; Zijlstra, G.; Bruggen, A. Differential interaction of Salmonella enterica serovars with lettuce cultivars and plant-microbe factors influencing colonization efficiency. ISME J. 2007, 1, 620–631. [Google Scholar]
- Ruppel, S.; Krumbein, A.; Schreiner, M. Composition of the phyllospheric microbial populations on vegetable plants with different glucosinolate and carotenoid compositions. Microb. Ecol. 2008, 56, 364–372. [Google Scholar]
- Yadav, R.K.P.; Karamanoli, K.; Vokou, D. Bacterial colonization of the phyllosphere of mediterranean perennial species as influenced by leaf structure and chemical features. Microb. Ecol. 2005, 50, 185–196. [Google Scholar]
- Jay, M.T.; Cooley, M.; Carychao, D.; Wiscomb, G.W.; Sweitzer, R.A.; Crawford-Miksza, L.; Farrar, J.A.; Lau, D.K.; O'Connell, J.; Millington, A.; et al. Escherichia coli O157:H7 in feral swine near spinach fields and cattle central california coast. Emerg. Infect. Dis. 2007, 13, 1908–1911. [Google Scholar]
- Fung, D.Y.C.; Sharpe, A.N.; Hart, B.C.; Liu, Y. The Pulsifier: A new instrument for preparing food suspensions for microbiological analysis. J. Rapid Meth. Aut. Mic. 1998, 6, 43–49. [Google Scholar]
- Muyzer, G.; Smalla, K. Application of Denaturing Gradient Gel Electrophoresis (DGGE) and Temperature Gradient Gel Electrophoresis (TGGE) in microbial ecology. Anton. Van Leeuw. 1998, 73, 127–141. [Google Scholar]
- Blackwood, C.B.; Oaks, A.; Buyer, J.S. Phylum- and Class-specific PCR primers for general microbial community analysis. Appl. Environ. Microb. 2005, 71, 6193–6198. [Google Scholar]
- Fierer, N.; Jackson, J.A.; Vilgalys, R.; Jackson, R.B. Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl. Environ. Microb. 2005, 71, 4117–4120. [Google Scholar]
- Lane, D. 16S/23S rRNA sequencing. In Nucleic Acid Techniques in Bacterial Systematics; Stackebrandt, E., Goodfellow, M., Eds.; John Wiley & Sons: West Sussex, UK, 1991; pp. 115–175. [Google Scholar]
- Manz, W.; Amann, R.; W. Ludwig, W.; Vancanneyt, M.; Schleifer, K.H. Application of a suite of 16S rRNA-specific oligonucleotide probes designed to investigate bacteria of the phylum cytophaga-flavobacter-bacteroides in the natural environment. Microbiology 1996, 142, 1097–1106. [Google Scholar]
- Conte, A.; Conversa, G.; Scrocco, C.; Brescia, I.; Laverse, J.; Elia, A.; Del Nobile, M.A. Influence of growing periods on the quality of baby spinach leaves at harvest and during storage as minimally processed produce. Postharvest Biol. Tec. 2008, 50, 190–196. [Google Scholar]
- Lopez-Velasco, G.; Welbaum, G.; Boyer, R.; Mane, S.; Ponder, M. Changes in spinach phylloepiphytic bacteria communities following minimal processing and refrigerated storage described using pyrosequencing of 16S rRNA amplicons. J. Appl. Microbiol. 2011, 110, 1203–1214. [Google Scholar]
- Krimm, U.; Abanda-Nkpwatt, D.; Schwab, W.; Schreiber, L. Epiphytic microorganisms on strawberry plants (Fragaria ananassa cv. Elsanta): Identification of bacterial isolates and analysis of their interaction with leaf surfaces. FEMS Microbiol. Ecol. 2005, 53, 483–92. [Google Scholar]
- Tydings, H. Identification and Optimization of the Antagonistic Potential of Native Spinach Microbiota towards Escherichia coli O157:H7. MS Thesis, Virginia Polytechnic Institute and State University, Blacksburg, VA, USA, 2010. [Google Scholar]
- Brandl, M.T.; Amundson, R. Leaf age as a risk factor in the contamination of lettuce with Escherichia coli O157:H7 and Salmonella enterica. Appl. Environ. Microbiol. 2008, 74, 2298–2306. [Google Scholar]
- Rubatzky, V.; Yamaguchi, M. Spinach, table beets, and other vegetable chenopods. In World Vegetables, 2nd ed.; Rubatzky, V., Yamaguchi, M., Eds.; International Thompson Publishing: New York, NY, USA, 2007; pp. 457–463. [Google Scholar]
- Monier, J.M.; Lindow, S.E. Frequency, size, and localization of bacterial aggregates on bean leaf surfaces. Appl. Environ. Microbiol. 2004, 70, 346–55. [Google Scholar]
- Milillo, S.R.; Badamo, J.M.; Boor, K.J.; Wiedmann, M. Growth and persistence of Listeria monocytogenes isolates on the plant model Arabidopsis thaliana. Food Microbiol. 2008, 25, 698–704. [Google Scholar]
- Lopez-Velasco, G.; Davis, M.; Boyer, R.; Williams, R.; Ponder, M. Alterations of the phylloepiphytic bacterial community associated with interactions of Escherichia coli O157:H7 during storage of packaged spinach at refrigeration temperatures. Food Microbiol. 2010, 27, 476–486. [Google Scholar]
- Kirk, J.; Beaudette, L.; Hart, M.; Moutoglis, P.; Klironomos, J.; Lee, H.; Trevors, J. Methods of studying soil microbial diversity. J. Microbiol. Methods. 2004, 58, 169–188. [Google Scholar]
- Morales, S.; Cosart, T.; Johnson, J.; Holben, W. Extensive phylogenetic analysis of a soil bacterial community illustrates extreme taxon evenness and the effects of amplicon length, degree of coverage, and DNA fractionation on classification and ecological parameters. Appl. Environ. Microbiol. 2009, 75, 668–75. [Google Scholar]
- Handschur, M.; Pinar, G.; Gallist, B.; Lubitz, W.; Haslberger, A. Culture free DGGE and cloning based monitoring of changes in bacterial communities of salad due to processing. Food Chem. Toxicol. 2005, 43, 1595–1605. [Google Scholar]
- Teplitski, M.; Warriner, K.; Bartz, J.; Schneider, K. Untangling metabolic and communication networks: Interaction of enterics with phytobacteria and their implications in produce safety. Trends Microbiol. 2011, 19, 121–127. [Google Scholar]
- Critzer, F.; Doyle, M. Microbial ecology of foodborne pathogens associated with produce. Curr. Opin. Biotechnol. 2010, 21, 125–130. [Google Scholar]
- Cooley, M.; Chao, D.; Mandrell, R. Escherichia coli O157:H7 survival and growth on lettuce is altered by the presence of epiphytic bacteria. J. Food Prot. 2006, 69, 2329–2335. [Google Scholar]
- Johnston, M.; Harrison, M.; Morrow, R. Microbial antagonists of Escherichia coli O157:H7 on fresh-cut lettuce and spinach. J. Food Prot. 2009, 72, 1569–1575. [Google Scholar]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lopez-Velasco, G.; Welbaum, G.E.; Falkinham III, J.O.; Ponder, M.A. Phyllopshere Bacterial Community Structure of Spinach (Spinacia oleracea) as Affected by Cultivar and Environmental Conditions at Time of Harvest. Diversity 2011, 3, 721-738. https://doi.org/10.3390/d3040721
Lopez-Velasco G, Welbaum GE, Falkinham III JO, Ponder MA. Phyllopshere Bacterial Community Structure of Spinach (Spinacia oleracea) as Affected by Cultivar and Environmental Conditions at Time of Harvest. Diversity. 2011; 3(4):721-738. https://doi.org/10.3390/d3040721
Chicago/Turabian StyleLopez-Velasco, Gabriela, Gregory E. Welbaum, Joseph O. Falkinham III, and Monica A. Ponder. 2011. "Phyllopshere Bacterial Community Structure of Spinach (Spinacia oleracea) as Affected by Cultivar and Environmental Conditions at Time of Harvest" Diversity 3, no. 4: 721-738. https://doi.org/10.3390/d3040721
APA StyleLopez-Velasco, G., Welbaum, G. E., Falkinham III, J. O., & Ponder, M. A. (2011). Phyllopshere Bacterial Community Structure of Spinach (Spinacia oleracea) as Affected by Cultivar and Environmental Conditions at Time of Harvest. Diversity, 3(4), 721-738. https://doi.org/10.3390/d3040721






