An Improved Method for Assessing Simple Sequence Repeat (SSR) Variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass)
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. DNA Extraction and Quality Analysis
2.3. Molecular Characterization of Species
2.4. SSR Loci Amplification and Protocol Optimization
2.5. DNA Fingerprinting Analysis
2.6. Statistical Analysis
3. Results
3.1. Molecular Characterization of Species
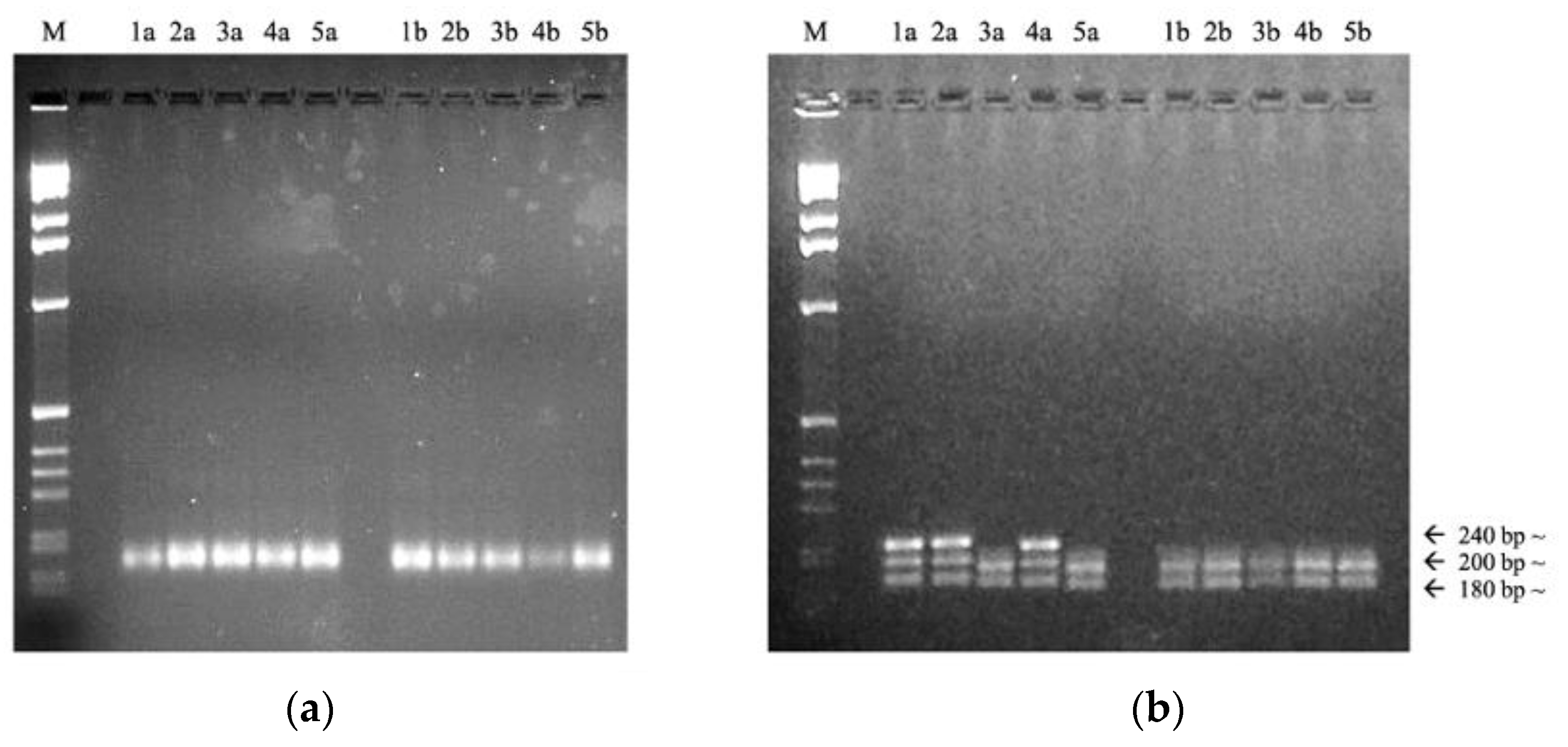
3.2. SSR Protocol Optimization
3.3. Genetic Richness and Diversity Analysis
3.4. Analysis of Molecular Variance (AMOVA)
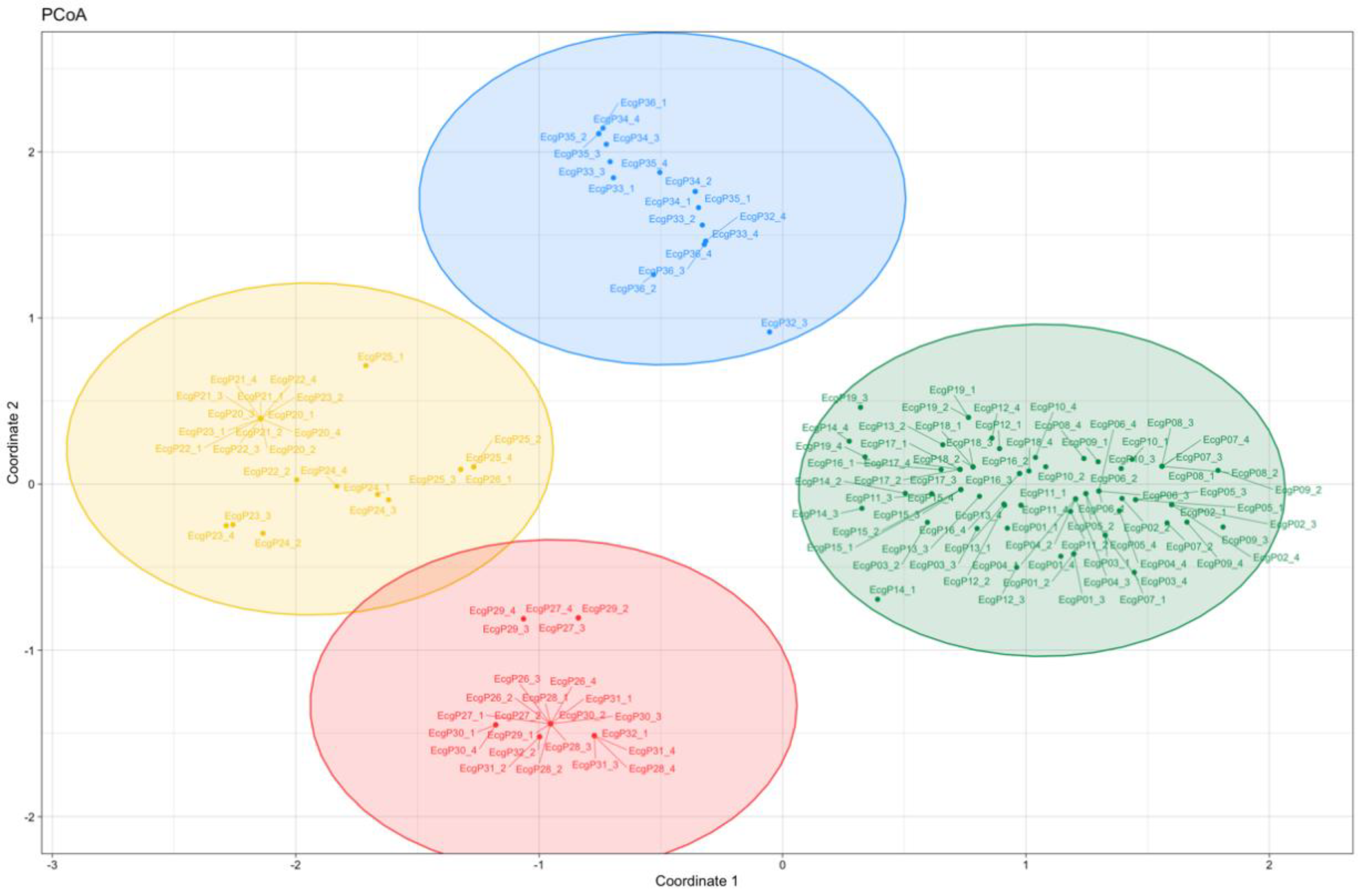
3.5. Hierarchical Clustering and Principal Coordinates Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Storkey, J.; Neve, P. What good is weed diversity? Weed Res. 2018, 58, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.C.; Röder, M.S.; Fahima, T.; Kirzhner, V.M.; Beiles, A.; Korol, A.B.; Nevo, E. Natural selection causing microsatellite divergence in wild emmer wheat at the ecologically variable microsite at Ammiad, Israel. Theor. Appl. Genet. 2000, 100, 985–999. [Google Scholar] [CrossRef]
- Green, J.M.; Barker, J.H.A.; Marshall, E.J.P.; Froud-Williams, R.J.; Peters, N.C.B.; Arnold, G.M.; Dawson, K.; Karp, A. Microsatellite analysis of the inbreeding grass weed Barren Brome (Anisantha sterilis) reveals genetic diversity at the within- and between-farm scales. Mol. Ecol. 2001, 10, 1035–1045. [Google Scholar] [CrossRef] [PubMed]
- Tabacchi, M.; Mantegazza, R.; Spada, A.; Ferrero, A. Morphological traits and molecular markers for classification of Echinochloa species from Italian rice fields. Weed Sci. 2006, 54, 1086–1093. [Google Scholar] [CrossRef]
- Sparacino, A.C.; Santin, C.; Ditto, D.; Tano, F. Morphological characteristics of the most spread Echinochloa spp. in Italian rice-fields. In In Proceedings of the 4th International Temperate Rice Conference, Novara, Italy, 25–28 June 2007. [Google Scholar]
- Pignatti, S. Flora d’Italia; Edagricole: Bologna, Italy, 1982; pp. 607–609. [Google Scholar]
- Ye, C.-Y.; Lin, Z.; Li, G.; Wang, Y.-Y.; Qiu, J.; Fu, F.; Zhang, H.; Chen, L.; Ye, S.; Song, W.; et al. Echinochloa Chloroplast Genomes: Insights into the Evolution and Taxonomic Identification of Two Weedy Species. PLoS ONE 2014, 9, e113657. [Google Scholar] [CrossRef]
- Ye, C.-Y.; Wu, D.; Mao, L.; Jia, L.; Qiu, J.; Lao, S.; Chen, M.; Jiang, B.; Tang, W.; Peng, Q.; et al. The Genomes of the Allohexaploid Echinochloa crus-galli and Its Progenitors Provide Insights into Polyploidization-Driven Adaptation. Mol. Plant 2020, 13, 1298–1310. [Google Scholar] [CrossRef]
- Altop, E.K.; Mennan, H. Genetic and morphologic diversity of Echinochloa crus-galli populations from different origins. Phytoparasitica 2010, 39, 93–102. [Google Scholar] [CrossRef]
- Brusoni, M. Studio sulla variabilità intraspecifica di Echinochloa crus-galli (L.) P. Beauv. Atti Ist. Bot. Lab. Crittogam. 1991, 10, 39–88. [Google Scholar]
- Norris, R.F. Morphological and Phenological Variation in Barnyardgrass (Echinochloa crus-galli) in California. Weed Sci. 1996, 44, 804–814. [Google Scholar] [CrossRef]
- Rutledge, J.; Talbert, R.; Sneller, C. RAPD analysis of genetic variation among propanil-resistant and -susceptible Echinochloa crus-galli populations in Arkansas. Weed Sci. 2000, 48, 669–674. [Google Scholar] [CrossRef]
- Tasrif, A.; Juraimi, A.S.; Kadir, J.; Sastroutomo, S.S.; Napis, S. Genetic diversity of Echinochloa crus-galli var. crus-galli (L.) Beauv (Barnyardgrass: Poaceae) ecotypes in Malaysia and Indonesia as revealed by RAPD markers. Asian J. Plant Sci. 2004, 3, 231–238. [Google Scholar] [CrossRef]
- Iwakami, S.; Uchino, A.; Watanabe, H.; Yamasue, Y.; Inamura, T. Isolation and expression of genes for acetolactate synthase and acetyl-CoA carboxylase in Echinochloa phyllopogon, a polyploid weed species. Pest Manag. Sci. 2012, 68, 1098–1106. [Google Scholar] [CrossRef]
- Kaloumenos, N.S.; Chatzilazaridou, S.L.; Mylona, P.V.; Polidoros, A.N.; Eleftherohorinos, I.G. Target-site mutation associated with cross-resistance to ALS-inhibiting herbicides in late watergrass (Echinochloa oryzicola Vasing.). Pest Manag. Sci. 2013, 9, 865–873. [Google Scholar] [CrossRef]
- Panozzo, S.; Scarabel, L.; Tranel, P.J.; Sattin, M. Target-site resistance to ALS inhibitors in the polyploid species Echinochloa crus-galli. Pestic. Biochem. Physiol. 2013, 105, 93–101. [Google Scholar] [CrossRef]
- Mascanzoni, E.; Perego, A.; Marchi, N.; Scarabel, L.; Panozzo, S.; Ferrero, A.; Acutis, M.; Sattin, M. Epidemiology and agronomic predictors of herbicide resistance in rice at a large scale. Agron. Sustain. Dev. 2018, 38, 68. [Google Scholar] [CrossRef]
- Claerhout, S.; Dewaele, K.; De Riek, J.; Reheul, D.; De Cauwer, B. Morphological and genetic variability of local Echinochloa accessions and the link with herbicide sensitivity. Weed Res. 2015, 56, 137–148. [Google Scholar] [CrossRef]
- Ruiz-Santaella, J.P.; Bastida, F.; Franco, A.R.; De Prado, R. Morphological and molecular characterization of different Echinochloa spp. and Oryza sativa populations. J. Agric. Food Chem. 2006, 54, 1166–1172. [Google Scholar] [CrossRef]
- Le, D.; Nguyen, C.M.; Mann, R.K.; Yerkes, C.N.; Kumar, B.V.N. Genetic diversity and herbicide resistance of 15 Echinochloa crus-galli populations to quinclorac in Mekong Delta of Vietnam and Arkansas of United States. J. Plant Biotechnol. 2017, 44, 472–477. [Google Scholar] [CrossRef]
- Nozawa, H.; Takahashi, M.; Nakai, H.; Sato, Y. Difference in SSR Variations Between Japanese Barnyard Millet (Echinochloa esculenta) and its Wild Relative E. crus-galli. Breed. Sci. 2006, 56, 335–340. [Google Scholar] [CrossRef][Green Version]
- Chen, G.; Zhang, W.; Fang, J.; Dong, L. Restriction site-associated DNA sequencing allows for the rapid identification of simple sequence repeat markers in Echinochloa crus-galli. Weed Biol. Manag. 2017, 17, 68–76. [Google Scholar] [CrossRef]
- Scialpi, A.; Mengoni, A. La PCR e le sue varianti. In Quaderno di Laboratorio; Firenze University Press: Firenze, Italy, 2008. [Google Scholar]
- Lee, J.; Park, K.W.; Lee, I.Y.; Kim, C.S.; Kown, O.D.; Park, T.S. Simple sequence repeat analysis of genetic diversity among Acetyl-CoA carboxylase inhibitor-resistant and inhibitor-susceptible Echinochloa crus-galli and E. oryzicola populations in Korea. Weed Res. 2015, 55, 90–100. [Google Scholar] [CrossRef]
- Lee, E.; Nah, G.; Yook, M.; Lim, S.; Park, T.; Lee, D.; Kim, D. Phylogenetic Relationship of Echinochloa Species Based on Simple Sequence Repeat and Phenotypic Marker Analyses. Weed Sci. 2016, 64, 441–454. [Google Scholar] [CrossRef]
- Mondini, L.; Noorani, A.; Pagnotta, M.A. Assessing Plant Genetic Diversity by Molecular Tools. Diversity 2009, 1, 19–35. [Google Scholar] [CrossRef]
- Manco, R.; Chiaiese, P.; Basile, B.; Corrado, G. Comparative analysis of genomic- and EST-SSRs in European plum (Prunus domestica L.): Implications for the diversity analysis of polyploids. 3 Biotech 2020, 10, 543. [Google Scholar] [CrossRef]
- Stift, M.; Kolář, F.; Meirmans, P.G. STRUCTURE is more robust than other clustering methods in simulated mixed-ploidy populations. Heredity 2019, 123, 429–441. [Google Scholar] [CrossRef]
- Meena, R.; Bhandari, M.; Ginwal, H. Usage of microsatellite markers for characterization of polyploids: A case study in reference to hexaploid bamboo species. Silvae Genet. 2020, 69, 94–97. [Google Scholar] [CrossRef]
- Rajput, S.G.; Wable, K.J.; Sharma, K.M.; Kubde, P.D.; Mulay, S.A. Reproducibility testing of RAPD and SSR markers in tomato. Afr. J. Biotechnol. 2006, 5, 108–112. [Google Scholar]
- Manimekalai, M.; Dhasarathan, M.; Karthikeyan, A.; Murukarthick, J.; Renganathan, V.G.; Thangaraj, K.; Vellaikumar, S.; Vanniarajan, C.; Senthil, N.N. Genetic diversity in the barnyard millet (Echinochola frumentacea) germplasms revealed by morphological traits and simple sequence repeat markers. Curr. Plant Biol. 2018, 14, 71–78. [Google Scholar]
- Teixeira, H.; Rodríguez-Echeverría, S.; Nabais, C. Correction: Genetic Diversity and Differentiation of Juniperus thurifera in Spain and Morocco as Determined by SSR. PLoS ONE 2015, 10, e0126042. [Google Scholar] [CrossRef]
- Altop, E.K.; Jabran, K.; Mennan, H. Determination of morphological and genetic diversity of ALS (acetolactate synthase)-herbicide-resistant Echinochloa oryzoides biotypes in rice. Int. J. Agric. Biol. 2018, 20, 628–636. [Google Scholar] [CrossRef]
- Ferchichi, Y.; Sakhraoui, A.; Ltaeif, H.B.; Ben Mhara, Y.; Elimem, M.; Ben Naceur, M.; Ghrabi-Gammar, Z.; Rouz, S. Eco-Geographical, Morphological and Molecular Characterization of a Collection of the Perennial Endemic Species Medicago tunetana (Murb.) A.W. Hill (Fabaceae) from Tunisia. Plants 2021, 10, 1923. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Pan, Y.B.; Wang, Q.N.; Wang, J.D.; Chen, J.L.; Gao, S.J. Genetic diversity and population structure analysis of Saccharum and Erianthus genera using microsatellite (SSR) markers. Sci. Rep. 2019, 9, 395. [Google Scholar] [CrossRef] [PubMed]
- Assoumane, A.; Zoubeirou, A.M.; Rodier-Goud, M.; Favreau, B.; Bezançon, G.; Verhaegen, D. Highlighting the occurrence of tetraploidy in Acacia senegal (L.) Willd. and genetic variation patterns in its natural range revealed by DNA microsatellite markers. Tree Genet. Genomes 2013, 9, 93–106. [Google Scholar] [CrossRef]
- Kashyap, P.L.; Kumar, S.; Kumar, R.S.; Tripathi, R.; Sharma, P.; Sharma, A.; Jasrotia, P.; Singh, G.P. Identification of Novel Microsatellite Markers to Assess the Population Structure and Genetic Differentiation of Ustilago hordei Causing Covered Smut of Barley. Front. Microbiol. 2020, 10, 2929. [Google Scholar] [CrossRef]
- Li, Q.; Su, X.; Ma, H.; Du, K.; Yang, M.; Chen, B.; Fu, S.; Fu, T.; Xiang, C.; Zhao, Q.; et al. Development of genic SSR marker resources from RNA-seq data in Camellia japonica and their application in the genus Camellia. Sci. Rep. 2021, 11, 9919. [Google Scholar] [CrossRef]
- Pagnotta, M.A. Comparison among Methods and Statistical Software Packages to Analyze Germplasm Genetic Diversity by Means of Codominant Markers. J 2018, 1, 197–215. [Google Scholar] [CrossRef]
- Meirmans, P.G.; Liu, S. Analysis of Molecular Variance (AMOVA) for Autopolyploids. Front. Ecol. Evol. 2018, 6, 66. [Google Scholar] [CrossRef]
- Lee, J.; Kim, J.-W.; Lee, I.-Y. Development of Cross-specific Microsatellite Markers for Echinochloa spp. (Poaceae). Weed Turfgrass Sci. 2019, 8, 21–29. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissues. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Amaro-Blanco, I.; Romano, Y.; Palmerin, J.A.; Gordo, R.; Palma-Bautista, C.; De Prado, R.; Osuna, M.D. Different Mutations Providing Target Site Resistance to ALS- and ACCase-Inhibiting Herbicides in Echinochloa spp. from Rice Fields. Agriculture 2021, 11, 382. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef]
- Kamvar, Z.N.; Brooks, J.C.; Grunwald, N.J. Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front. Genet. 2015, 6, 208. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Que, Y.; Pan, Y.B. Highly Polymorphic Microsatellite DNA Markers for Sugarcane Germplasm Evaluation and Variety Identity Testing. Sugar Tech. 2011, 13, 129–136. [Google Scholar] [CrossRef]
- Prevost, A.; Wilkinson, M.J. A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor. Appl. Genet. 1999, 98, 107–112. [Google Scholar] [CrossRef]
- Tiwari, G.; Singh, R.; Singh, N.; Choudhury, D.R.; Paliwal, R.; Kumar, A.; Gupta, V. Study of arbitrarily amplified (RAPD and ISSR) and gene targeted (SCoT and CBDP) markers for genetic diversity and population structure in Kalmegh [Andrographis paniculata (Burm. f.) Nees]. Ind. Crop. Prod. 2016, 86, 1–11. [Google Scholar] [CrossRef]
- Shannon, C.E. A mathematical theory of communication. ACM SIGMOBILE Mob. Comput. Commun. Rev. 2001, 5, 3–55. [Google Scholar] [CrossRef]
- Stoddart, J.A.; Taylor, J.F. Genotypic diversity: Estimation and prediction in samples. Genetics 1988, 118, 705–711. [Google Scholar] [CrossRef]
- Simpson, E. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Pielou, E.C. Ecological Diversity; Wiley: New York, NY, USA, 1975; p. 165. [Google Scholar]
- Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [CrossRef]
- Paradis, E. Pegas: An R package for population genetics with an integrated-modular approach. Bioinformatics 2010, 26, 419–420. [Google Scholar] [CrossRef]
- Pembleton, L.W.; Cogan, N.O.I.; Forster, J.W. StAMPP: An R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol. Ecol. Resour. 2013, 13, 946–952. [Google Scholar] [CrossRef] [PubMed]
- Clark, L.; Jasieniuk, M. Polysat: An R package for polyploid microsatellite analysis. Mol. Ecol. Resour. 2011, 11, 562–566. [Google Scholar] [CrossRef] [PubMed]
- Clark, L.; Schreier, A.D. Resolving microsatellite genotype ambiguity in populations of allopolyploid and diploidized autopolyploid organisms using negative correlations between allelic variables. Mol. Ecol. Resour. 2017, 17, 1090–1103. [Google Scholar] [CrossRef]
- Galili, T. Dendextend: An R package for visualizing, adjusting, and comparing trees of hierarchical clustering. Bioinformatics 2015, 31, 3718–3720. [Google Scholar] [CrossRef]
- Lê, S.; Josse, J.; Husson, F. FactoMineR: An R Package for Multivariate Analysis. J. Stat. Softw. 2008, 25, 1–18. [Google Scholar] [CrossRef]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Khamassi, K.; Khoufi, S.; Chaabane, R.; Da Silva, J.A.T.; Naceur, M.B. Optimization of conditions forassessment of genetic diversity in chickpea (Cicer arietinum L.) using SSR markers. Int. J. Plant Breed. 2011, 5, 141–145. [Google Scholar]
- Doğrar, N.; Akkaya, M.S. Optimization of PCR Amplification of Wheat Simple Sequence Repeat DNA Markers. Turk. J. Biol. 2001, 25, 153–158. [Google Scholar]
- Narina, S.S.; D’Orgeix, C.A.; Sayre, B.L. Optimization of PCR conditions to amplify microsatellite loci in the bunchgrass lizard (Sceloporus slevini) genomic DNA. BMC Res. Notes 2011, 4, 26. [Google Scholar] [CrossRef] [PubMed]
- Ur-rahman, M.; Malik, T.A.; Aslam, N.; Asif, M.; Ahmad, R.; Khan, I.A.; Zafar, Y. Optimization of PCR Conditions to Amplify Microsatellite Loci in Cotton (Gossypium hirsutum L.) Genomic DNA. Int. J. Agric. Biol. 2004, 2, 282–284. [Google Scholar]
- Shinde, D.; Lai, Y.; Sun, F.; Arnheim, N. Taq DNA polymerase slippage mutation rates measured by PCR and quasi-likelihood analysis: (CA/GT)n and (A/T)n microsatellites. Nucleic Acids Res. 2003, 31, 974–980. [Google Scholar] [CrossRef]
- Blanchard, M.M.; Tailon-Miller, P.; Nowotny, P.; Nowotny, V. PCR buffer optimization with a uniform temperature regimen to facilitate automation. Genome Res. 1993, 2, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Meirmans, P.G.; Liu, S.; Van Tienderen, P.H. The Analysis of Polyploid Genetic Data. J. Hered. 2018, 109, 283–296. [Google Scholar] [CrossRef]
- Wright, S. The Interpretation of Population Structure by F-Statistics with Special Regard to Systems of Mating. Evolution 1965, 19, 395–420. [Google Scholar] [CrossRef]
- Montiel Castelán, P.; Cortés-Cruz, M.; Mendoza-Castillo, M.d.C.; Cruz-Izquierdo, S.; López-Upton, J.; Sandoval Padilla, I.; Guerra de la Cruz, V. Diversity and Genetic Structure Inferred with Microsatellites in Natural Populations of Pseudotsuga menziesii (Mirb.) Franco (Pinaceae) in the Central Region of Mexico. Forests 2019, 10, 101. [Google Scholar] [CrossRef]
- Nybom, H. Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol. Ecol. 2004, 13, 1143–1155. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Pan, T.; Qian, H.; Zhang, M.; Yang, G.; Wang, X. Genetic Diversity and Population Structure Revealed by SSR Markers on Endemic Species Osmanthus serrulatus Rehder from Southwestern Sichuan Basin, China. Forests 2021, 12, 1365. [Google Scholar] [CrossRef]


| Locus Name | SSR Motif | Primer Sequences (5′—3′) |
|---|---|---|
| EG1 | (TG)7 | F: GCTCCTGAACTGTGTACATTCTTGC |
| R: TCGATTCACCCTTCAGCTTCTC | ||
| EG2 | (TA)6 | F: CATCGGATTCAGATTGAAAGGG |
| R: GGTCGTAGGTCTATAGTCCGTAGAGTCA | ||
| EG301 | (AT)5 | F: GCGTCGTCAAGTCGTTCTTCTA |
| R: TGTATTCAGCTGTCGTGCATGT | ||
| EG302 | (ATTT)8 | F: ATTCGAACACCCATCAACCAAC |
| R: GAAACAGAAGGGAGGTGTGCTG | ||
| EG305 | (AT)4 | F: AGCCGTTCCTCTAGTCGGATTTCT |
| R: TATTCAGCTGCCGTGCATGTAGTA | ||
| EG306 | (CT)8 | F: TAAAACAAAACGACCGGCGTAA |
| R: TCAATCATTTCAGCCTTCGGAT | ||
| EG307 | (ATC)11 | F: AACATTGTCATCACAAATATCATCATCA |
| R: AATCAAGGAAGCCCCTTCACTC | ||
| EG320 | (TA)5 | F: CAACTCATAAGACAATTCAAAGGGTTT |
| R: GCATCATTTAAGCATCAAAATGACA |
|
|
|---|---|
| PCR Mixture (in a Total Volume = 10 µL) | PCR Mixture (in a Total Volume = 10 µL) |
| 0.2 µL of crude DNA (6–8 ng) extract | 2 µL of diluted DNA from crude extract (10 ng/µL) |
| 0.4 µL of each primer (0.4 µM) | 1 µL of each primer (10 µM) |
| 5 µL of Taq polymerase Ready Mix (0.27 UI) (Dongsheng Biotech) | 5.3 µL of Taq polymerase Ready Mix (0.4 UI) KAPA 2X Taq Extra Hot Start Ready-mix PCR Kit (Resnova S.r.l.) |
| (MgCl2 total concentration = 1.6 mM) | Addition of 0.5 µL of MgCl2 (MgCl2 total concentration = 2.5 mM) |
| nuclease-free H2O—ad volume | nuclease-free H2O—ad volume |
| PCR program | PCR program |
| initial denaturation step at 94 °C for 4 min | initial denaturation step at 95 °C for 5 min |
| 35 cycles of: 94 °C for 30 s relative annealing temperatures for 30 s 72°C for 1 min | 35 cycles of: 95 °C for 30 s relative annealing temperatures for 30 s 72°C for 1 min |
| final extension step at 72 °C for 10 min | final extension step at 72 °C for 10 min |
| Locus Name | AT According to et al. Chen et al. (2017) | AT According Post-Gradient PCR Results |
|---|---|---|
| EG1 | 49 °C | 40.6 °C |
| EG2 | 51.5 °C | 50 °C |
| EG301 | 57 °C | 43.3 °C |
| EG302 | 57 °C | 48 °C |
| EG305 | 57 °C | 55 °C |
| EG306 | 57 °C | 43.2 °C |
| EG307 | 57 °C | 55.6 °C |
| EG320 | 57 °C | 46.5 °C |
| Locus | Na | PIC |
|---|---|---|
| EG1 | 5 | 0.93 |
| EG2 | 3 | 0.88 |
| EG302 | 8 | 0.97 |
| EG305 | 5 | 0.96 |
| EG306 | 4 | 0.94 |
| EG307 | 2 | 0.76 |
| EG320 | 9 | 0.98 |
| EG301 | 12 | 0.98 |
| Mean | 6 | 0.92 |
| Population ID | N | %P | MLG | H | G | Lambda | E.5 | He | Ho |
|---|---|---|---|---|---|---|---|---|---|
| EcgP01 | 4 | 41.15 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.33 | 0.67 |
| EcgP02 | 4 | 39.06 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP03 | 4 | 41.15 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP04 | 4 | 39.06 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP05 | 4 | 36.46 | 2 | 0.69 | 2.00 | 0.50 | 1.00 | 0.01 | 0.01 |
| EcgP06 | 4 | 33.33 | 1 | 0.00 | 1.00 | 0.00 | -- | 0.01 | 0.00 |
| EcgP07 | 4 | 38.02 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP08 | 4 | 38.54 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP09 | 4 | 34.90 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP10 | 4 | 32.29 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP11 | 4 | 37.50 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP12 | 4 | 33.33 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP13 | 4 | 38.54 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP14 | 4 | 38.54 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP15 | 4 | 35.42 | 1 | 0.00 | 1.00 | 0.00 | -- | 0.01 | 0.00 |
| EcgP16 | 4 | 35.94 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP17 | 4 | 35.42 | 1 | 0.00 | 1.00 | 0.00 | -- | 0.01 | 0.00 |
| EcgP18 | 4 | 32.81 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP19 | 4 | 32.81 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP20 | 4 | 33.33 | 1 | 0.00 | 1.00 | 0.00 | -- | 0.01 | 0.00 |
| EcgP21 | 4 | 33.33 | 1 | 0.00 | 1.00 | 0.00 | -- | 0.01 | 0.00 |
| EcgP22 | 4 | 31.77 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP23 | 4 | 31.77 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP24 | 4 | 36.98 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP25 | 4 | 28.13 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP26 | 4 | 29.17 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP27 | 4 | 31.25 | 2 | 0.69 | 2.00 | 0.50 | 1.00 | 0.01 | 0.01 |
| EcgP28 | 4 | 28.65 | 2 | 0.56 | 1.60 | 0.38 | 0.79 | 0.01 | 0.01 |
| EcgP29 | 4 | 31.25 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP30 | 4 | 28.13 | 2 | 0.69 | 2.00 | 0.50 | 1.00 | 0.01 | 0.01 |
| EcgP31 | 4 | 27.08 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP32 | 4 | 30.73 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP33 | 4 | 34.38 | 4 | 1.39 | 4.00 | 0.75 | 1.00 | 0.03 | 0.02 |
| EcgP34 | 4 | 35.94 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP35 | 4 | 39.06 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| EcgP36 | 4 | 35.42 | 3 | 1.04 | 2.67 | 0.63 | 0.91 | 0.02 | 0.01 |
| Total | 144 | ---- | 78 | 29.8 | 36.64 | 0.97 | 28.21 | 0.96 | 1.10 |
| Mean | 4 | 34.46 | 2.61 | 0.82 | 2.40 | 0.50 | 0.91 | 0.03 | 0.03 |
| Source | DF | SS | MS | Est. Var. | % | p |
|---|---|---|---|---|---|---|
| Between agricultural managements | 1 | 8.20 | 8.20 | 0.11 | 47.23% | <0.001 |
| Among populations | 34 | 18.66 | 0.54 | 0.12 | 37.01% | <0.001 |
| Within populations | 108 | 4.24 | 0.03 | 0.04 | 15.74% | <0.001 |
| Total | 143 | 31.10 | 0.21 | 0.27 | 100% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cusaro, C.M.; Grazioli, C.; Zambuto, F.; Capelli, E.; Brusoni, M. An Improved Method for Assessing Simple Sequence Repeat (SSR) Variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass). Diversity 2022, 14, 3. https://doi.org/10.3390/d14010003
Cusaro CM, Grazioli C, Zambuto F, Capelli E, Brusoni M. An Improved Method for Assessing Simple Sequence Repeat (SSR) Variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass). Diversity. 2022; 14(1):3. https://doi.org/10.3390/d14010003
Chicago/Turabian StyleCusaro, Carlo Maria, Carolina Grazioli, Francesco Zambuto, Enrica Capelli, and Maura Brusoni. 2022. "An Improved Method for Assessing Simple Sequence Repeat (SSR) Variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass)" Diversity 14, no. 1: 3. https://doi.org/10.3390/d14010003
APA StyleCusaro, C. M., Grazioli, C., Zambuto, F., Capelli, E., & Brusoni, M. (2022). An Improved Method for Assessing Simple Sequence Repeat (SSR) Variation in Echinochloa crus-galli (L.) P. Beauv (Barnyardgrass). Diversity, 14(1), 3. https://doi.org/10.3390/d14010003







