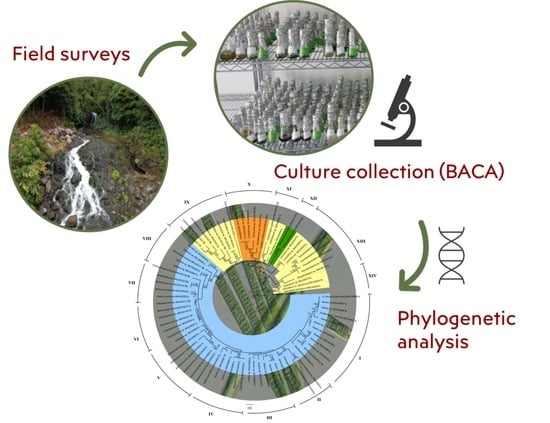
Cyanobacteria Phylogenetic Studies Reveal Evidence for Polyphyletic Genera from Thermal and Freshwater Habitats
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Site and Sample Collection
2.2. Strains Isolation, Morphological Characterization and Culture Conditions
2.3. DNA Extraction, Polymerase Chain Reaction (PCR) Amplification, and Sequencing
2.4. Phylogenetic Analysis
3. Results
3.1. Sampling Site Features
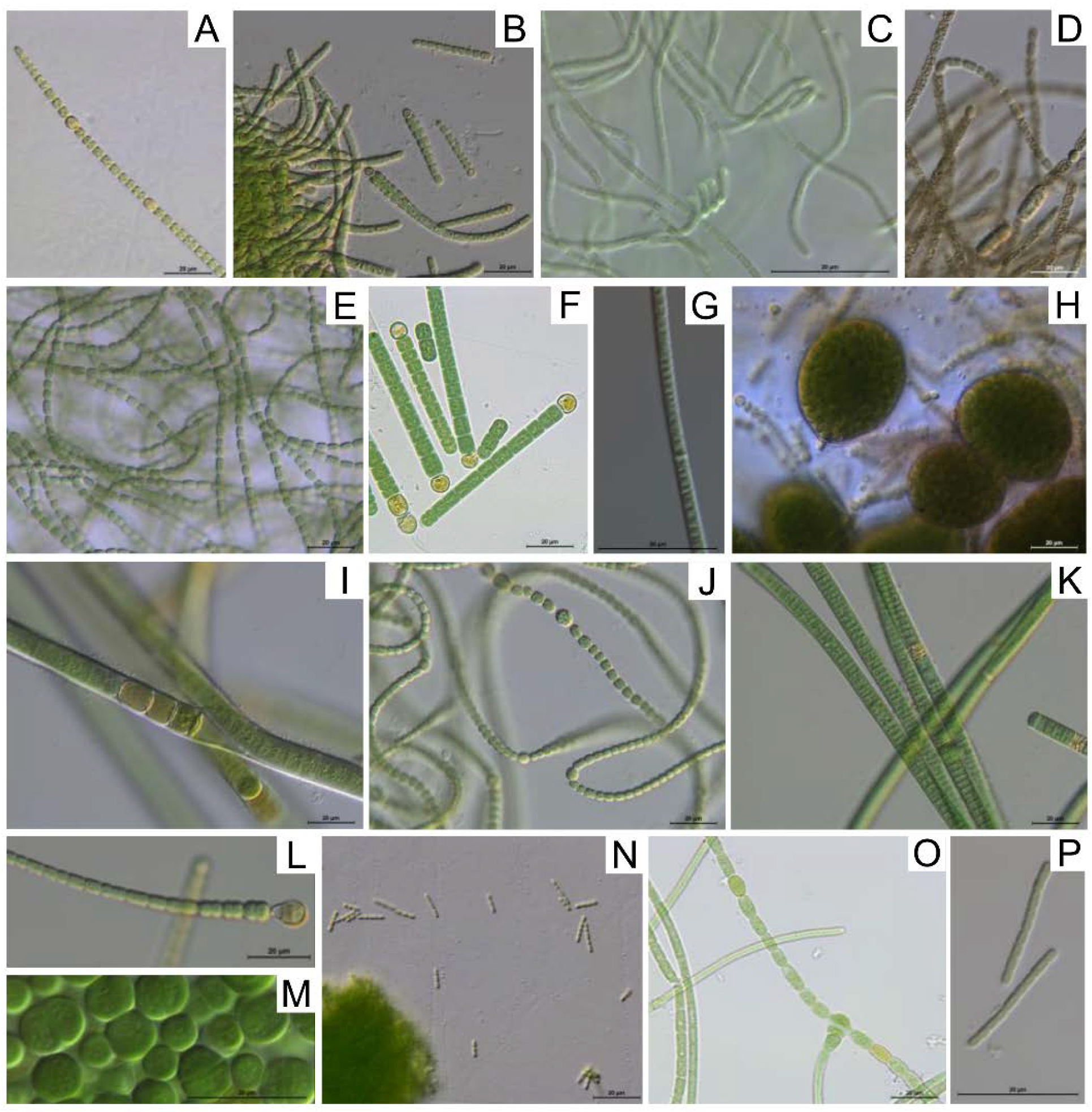
3.2. Cyanobacterial Isolation and Morphological Identification
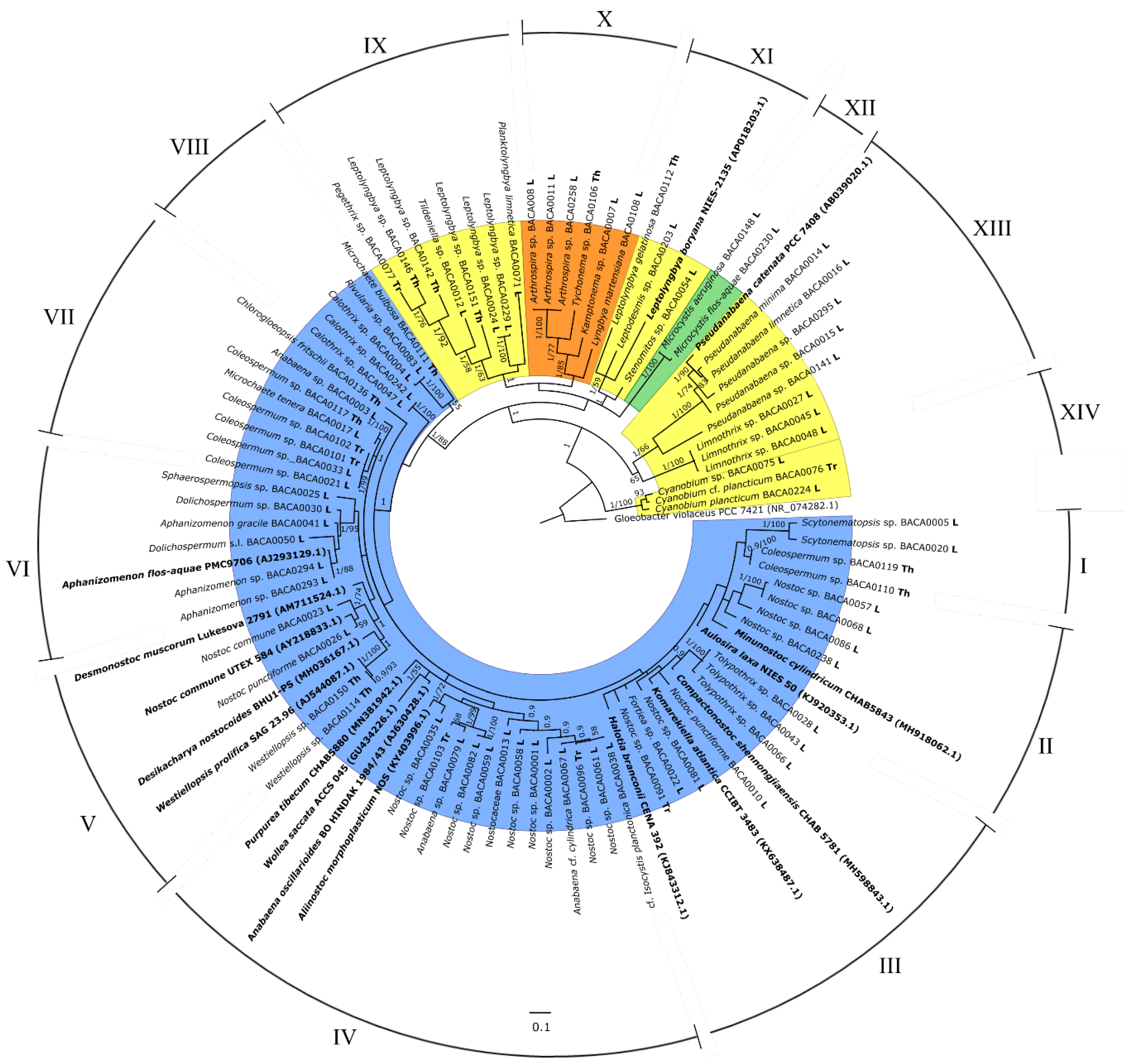
3.3. Phylogeny Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Codd, G.A.; Meriluoto, J.; Metcalf, J.S. Introduction: Cyanobacteria, Cyanotoxins, Their Human Impact, and Risk Management. In Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis; Codd, G.A., Meriluoto, J., Metcalf, J.S., Eds.; John Wiley & Sons, Ltd.: Chichester, UK, 2017; pp. 1–8. [Google Scholar]
- Paerl, H.W.; Otten, T.G. Harmful cyanobacterial blooms: Causes, consequences, and controls. Microb. Ecol. 2013, 65, 995–1010. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. Cyanobacteria: Biology, ecology and evolution. In Cyanobacteria an Economic Perspective; Sharma, N., Rai, A., Stal, L., Eds.; John Wiley & Sons, Ltd.: Oxford, UK, 2014; pp. 3–20. [Google Scholar]
- Cirés, S.; Casero, M.C.; Quesada, A. Toxicity at the edge of life: A review on cyanobacterial toxins from extreme environments. Mar. Drugs 2017, 15, 233. [Google Scholar] [CrossRef] [PubMed]
- Archer, W. Notes on some collections made from Furnas Lake, Azores, containing algae and a few other organisms. Bot. J. Linn. Soc. 1874, 14, 328–340. [Google Scholar] [CrossRef]
- Luz, R.; Cordeiro, R.; Vilaverde, J.; Raposeiro, P.; Fonseca, A.; Gonçalves, V. Cyanobacteria from freshwater lakes in the Azores archipelago, Portugal: Data from long term phytoplankton monitoring. Biodivers. Data J. 2020, 8, e51928. [Google Scholar] [CrossRef]
- Johansson, C. Freshwater algal vegetation in the Azores. Bol. Soc. Brot. 1976, 50, 117–141. [Google Scholar]
- Santos, M.C.R.; Muelle, H.; Pacheco, D.M.D. Cyanobacteria and microcystins in lake Furnas (S. Miguel island-Azores). Limnetica 2012, 31, 107–118. [Google Scholar]
- Moreira, C.; Martins, A.; Moreira, C.; Vasconcelos, V. Toxigenic cyanobacteria in volcanic lakes and hot springs of a North Atlantic island (S. Miguel, Azores, Portugal). Fresenius Environ. Bull. 2011, 20, 420–426. [Google Scholar]
- Porteiro, J.M.M. Lagoas dos Açores. Elementos de Suporte ao Planeamento Integrado. Ph.D. Thesis, University of Azores, Azores, Portugal, 2000. [Google Scholar]
- Santos, M.C.R.; Pacheco, D.M.D.; Santana, F.; Muelle, H. Cyanobacteria blooms in Sete-Cidades lake (S. Miguel Island—Azores). Arch. Hydrobiol. Suppl. Algol. Stud. 2005, 117, 393–406. [Google Scholar] [CrossRef]
- Cruz, J.V.; França, Z. Hydrogeochemistry of thermal and mineral water springs of the Azores archipelago (Portugal). J. Volcanol. Geotherm. Res. 2006, 151, 382–398. [Google Scholar] [CrossRef]
- Luz, R.F.S. Biological Activity Screening of Isolated Freshwater and Thermal Water Cyanobacteria from the Azores. Master’s Thesis, University of Azores, Azores, Portugal, 2018. [Google Scholar]
- Komárek, J.; Kastovsky, J.; Mares, J.; Johansen, J.R. Taxonomic classification of cyanoprokaryotes (cyanobacterial genera) 2014, using a polyphasic approach. Preslia 2014, 86, 295–335. [Google Scholar]
- Komárek, J. A polyphasic approach for the taxonomy of cyanobacteria: Principles and applications. Eur. J. Phycol. 2016, 51, 346–353. [Google Scholar] [CrossRef]
- Hrouzek, P.; Lukešová, A.; Mareš, J.; Ventura, S. Description of the cyanobacterial genus Desmonostoc gen. nov. including D. muscorum comb. nov. as a distinct, phylogenetically coherent taxon related to the genus Nostoc. Fottea 2013, 13, 201–213. [Google Scholar]
- Mai, T.; Johansen, J.R.; Pietrasiak, N.; Bohunická, M.; Martin, M.P. Revision of the Synechococcales (Cyanobacteria) through recognition of four families including Oculatellaceae fam. nov. and Trichocoleaceae fam. nov. and six new genera containing 14 species. Phytotaxa 2018, 365, 1–59. [Google Scholar] [CrossRef]
- Genuário, D.B.; Andreote, A.P.D.; Vaz, M.G.; Fiore, M.F. Heterocyte-forming cyanobacteria from Brazilian saline-alkaline lakes. Mol. Phylogenet. Evol. 2017, 109, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Genuário, D.B.; Vaz, M.G.; Melo, I.S. Phylogenetic insights into the diversity of homocytous cyanobacteria from Amazonian rivers. Mol. Phylogenet. Evol. 2017, 116, 120–135. [Google Scholar] [CrossRef] [PubMed]
- Genuário, D.B.; Vaz, M.G.; Hentschke, G.S.; Sant’Anna, C.L.; Fiore, M.F. Halotia gen. nov., a phylogenetically and physiologically coherent cyanobacterial genus isolated from marine coastal environments. Int. J. Syst. Evol. Microbiol. 2015, 65, 663–675. [Google Scholar] [CrossRef]
- Dvořák, P.; Poulíčková, A.; Hašler, P.; Belli, M.; Casamatta, D.A.; Papini, A. Species concepts and speciation factors in cyanobacteria, with connection to the problems of diversity and classification. Biodivers. Conserv. 2015, 24, 739–757. [Google Scholar] [CrossRef]
- Komarek, J. Cyanobacterial Taxonomy: Current problems and prospects for the integration of traditional and molecular approaches. ALGAE 2006, 21, 349–375. [Google Scholar] [CrossRef]
- Engene, N.; Rottacker, E.C.; Kastovsky, J.; Byrum, T.; Choi, H.; Ellisman, M.H.; Komarek, J.; Gerwick, W.H. Moorea producens gen. nov., sp. nov. and Moorea bouillonii comb. nov., tropical marine cyanobacteria rich in bioactive secondary metabolites. Int. J. Syst. Evol. Microbiol. 2012, 62, 1171–1178. [Google Scholar] [CrossRef]
- Saraf, A.; Suradkar, A.; Dawda, H.G.; Gaysina, L.A.; Gabidullin, Y.; Kumat, A.; Behere, I.; Kotulkar, M.; Batule, P.; Singh, P. Phylogenetic complexities of the members of Rivulariaceae with the re-creation of the family Calotrichaceae and description of Dulcicalothrix necridiiformans gen nov., sp nov., and reclassification of Calothrix desertica. FEMS Microbiol. Lett. 2019, 366, fnz219. [Google Scholar] [CrossRef]
- Bagchi, S.N.; Singh, P. Importance of Cyanobacterial Taxonomy in Biotechnological Applications. In Microbial Diversity in Ecosystem Sustainability and Biotechnological Applications; Satyanarayana, T., Johri, B.N., Das, S.K., Eds.; Springer: Singapore, 2019; pp. 387–414. ISBN 978-981-13-8314-4. [Google Scholar]
- Ramos, V.; Morais, J.; Castelo-Branco, R.; Pinheiro, Â.; Martins, J.; Regueiras, A.; Pereira, A.L.; Lopes, V.R.; Frazão, B.; Gomes, D.; et al. Cyanobacterial diversity held in microbial biological resource centers as a biotechnological asset: The case study of the newly established LEGE culture collection. J. Appl. Phycol. 2018, 30, 1437–1451. [Google Scholar] [CrossRef] [PubMed]
- Lourenço, S.O.; Vieira, A.A.H. Culture collections of microalgae in Brazil: Progress and constraints. Nov. Hedwigia 2004, 79, 149–173. [Google Scholar] [CrossRef]
- Bryant, D.A. A Brief History of Cyanobacterial Research: Past, Present, and Future Prospects. In The Cell Biology of Cyanobacteria; Flores, E., Herrero, A., Eds.; Caister Academic Press: Norfolk, UK, 2014; pp. 1–6. [Google Scholar]
- Rippka, R.; Deruelles, J.; Waterbury, J.B.; Herdman, M.; Stanier, R.Y. Generic Assignments, Strain Histories and Properties of Pure Cultures of Cyanobacteria. J. Gen. Microbiol. 1979, 111, 1–61. [Google Scholar] [CrossRef]
- Hauer, T.; Bohunická, M.; Johansen, J.R.; Mareš, J.; Berrendero-Gomez, E. Reassessment of the cyanobacterial family Microchaetaceae and establishment of new families Tolypothrichaceae and Godleyaceae. J. Phycol. 2014, 50, 1089–1100. [Google Scholar] [CrossRef]
- Raabová, L.; Kovacik, L.; Elster, J.; Strunecký, O. Review of the genus Phormidesmis (Cyanobacteria) based on environmental, morphological, and molecular data with description of a new genus Leptodesmia. Phytotaxa 2019, 395, 1–16. [Google Scholar] [CrossRef]
- Rippka, R. Isolation and purification of cyanobacteria. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 1988; Volume 167, pp. 3–27. [Google Scholar]
- Rippka, R.; Waterbury, J.; Stanier, R. Isolation and Purification of Cyanobacteria: Some General Principles. In The Prokaryotes: A Handbook on Habitats, Isolation and Identification of Bacteria; Starr, M.P., Stolp, H., Trüper, H.G., Balows, A., Schlegel, H.G., Eds.; Springer: Berlin/Heidelberg, Germany, 1981; ISBN 978-3-662-13189-3. [Google Scholar]
- Castenholz, R.W. Culturing methods for cyanobacteria. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 1988; pp. 68–93. [Google Scholar]
- Komárek, J.; Anagnostidis, K. Cyanoprokaryota 2. Teil/Part 2: Oscillatoriales; Spektrum Akademischer Verlag: Münche, Germany, 2005. [Google Scholar]
- Komárek, J.; Anagnostidis, K. Cyanoprokaryota, Teil 1/Part 1: Chroococcales; Spektrum Akademischer Verlag: Berlin, Germany, 2008. [Google Scholar]
- Komárek, J. Cyanoprokaryota 3. Teil/Part 3: Heterocytous Genera; Spektrum Akademischer Verlag: Berlin, Germany, 2013. [Google Scholar]
- Xavier, E.D.; Gonçalves, V.; Reis, A.; Azevedo, J.M.N.; Neto, A.I. Culture collection of freshwater microalgae from the Azores archipelago: Resource for taxonomic and phycoprospecting research. Cryptogam. Algol. 2018, 39, 227–237. [Google Scholar] [CrossRef]
- Nübel, U.; Garcia-Pichel, F.; Muyzer, G. PCR primers to amplify 16S rRNA genes from cyanobacteria. Appl. Environ. Microbiol. 1997, 63, 3327–3332. [Google Scholar] [CrossRef] [PubMed]
- Mühling, M.; Woolven-Allen, J.; Murrell, J.C.; Joint, I. Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J. 2008, 2, 379–392. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. Raxml version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Edler, D.; Klein, J.; Antonelli, A.; Silvestro, D. raxmlGUI 2.0 beta: A graphical interface and toolkit for phylogenetic analyses using RAxML. bioRxiv 2019. [Google Scholar]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef]
- Moseley, H.N. Notes on Fresh-water Algæ obtained at the Boiling Springs at Furnas, St. Michael’s, Azores, and their neighbourhood. Bot. J. Linn. Soc. 1874, 14, 321–325. [Google Scholar] [CrossRef]
- Papaefthimiou, D.; Hrouzek, P.; Mugnai, M.A.; Lukesova, A.; Turicchia, S.; Rasmussen, U.; Ventura, S. Differential patterns of evolution and distribution of the symbiotic behaviour in nostocacean cyanobacteria. Int. J. Syst. Evol. Microbiol. 2008, 58, 553–564. [Google Scholar] [CrossRef]
- Svenning, M.M.; Eriksson, T.; Rasmussen, U. Phylogeny of symbiotic cyanobacteria within the genus Nostoc based on 16S rDNA sequence analyses. Arch. Microbiol. 2005, 183, 19–26. [Google Scholar] [CrossRef]
- Hrouzek, P.; Ventura, S.; Lukešová, A.; Mugnai, M.; Angela Turicchia, S.; Komárek, J. Diversity of soil Nostoc strains: Phylogenetic and phenotypic variability. Algol. Stud. Hydrobiol. Suppl. Vol. 2005, 117, 251–264. [Google Scholar] [CrossRef]
- Silva, C.S.P.; Genuário, D.B.; Vaz, M.G.; Fiore, M.F. Phylogeny of culturable cyanobacteria from Brazilian mangroves. Syst. Appl. Microbiol. 2014, 37, 100–112. [Google Scholar] [CrossRef]
- Bravakos, P.; Kotoulas, G.; Skaraki, K.; Pantazidou, A.; Economou-Amilli, A. A polyphasic taxonomic approach in isolated strains of Cyanobacteria from thermal springs of Greece. Mol. Phylogenet. Evol. 2016, 98, 147–160. [Google Scholar] [CrossRef]
- Bagchi, S.N.; Dubey, N.; Singh, P. Phylogenetically distant clade of Nostoc-like taxa with the description of Aliinostoc gen. nov. and Aliinostoc morphoplasticum sp. nov. Int. J. Syst. Evol. Microbiol. 2017, 67, 3329–3338. [Google Scholar] [CrossRef] [PubMed]
- Řeháková, K.; Johansen, J.R.; Casamatta, D.A.; Xuesong, L.; Vincent, J. Morphological and molecular characterization of selected desert soil cyanobacteria: Three species new to science including Mojavia pulchra gen. et sp. nov. Phycologia 2007, 46, 481–502. [Google Scholar] [CrossRef]
- Lyra, C.; Suomalainen, S.; Gugger, M.; Vezie, C.; Sundman, P.; Paulin, L.; Sivonen, K. Molecular characterization of planktic cyanobacteria of Anabaena, Aphanizomenon, Microcystis and Planktothrix genera. Int. J. Syst. Evol. Microbiol. 2001, 51, 513–526. [Google Scholar] [CrossRef] [PubMed]
- Gugger, M.F.; Hoffmann, L. Polyphyly of true branching cyanobacteria (Stigonematales). Int. J. Syst. Bacteriol. Evol. Microbiol. 2004, 54, 349–357. [Google Scholar] [CrossRef]
- Neilan, B.A.; Jacobs, D.; Goodman, A.E. Genetic diversity and phylogeny of toxic cyanobacteria determined by DNA polymorphisms within the phycocyanin locus. Appl. Environ. Microbiol. 1995, 61, 3875–3883. [Google Scholar] [CrossRef]
- Janet, M. Westiellopsis prolifica, gen. et sp. nov., a new member of the Stigonemataceae. Ann. Bot. 1941, 5, 167–170. [Google Scholar] [CrossRef]
- Finsinger, K.; Scholz, I.; Serrano, A.; Morales, S.; Uribe-Lorio, L.; Mora, M.; Sittenfeld, A.; Weckesser, J.; Hess, W.R. Characterization of true-branching cyanobacteria from geothermal sites and hot springs of Costa Rica. Environ. Microbiol. 2008, 10, 460–473. [Google Scholar] [CrossRef]
- Saber, A.A.; Cantonati, M.; Mareš, J.; Anesi, A.; Guella, G. Polyphasic characterization of Westiellopsis prolifica (Hapalosiphonaceae, Cyanobacteria) from the El-Farafra Oasis (Western Desert, Egypt). Phycologia 2017, 56, 697–709. [Google Scholar] [CrossRef]
- Vaccarino, M.A.; Johansen, J.R. Scytonematopsis contorta sp. nov. (Nostocales), a new species from the Hawaiian Islands. Fottea 2011, 11, 149–161. [Google Scholar] [CrossRef]
- Komárek, J. Recent changes (2008) in cyanobacteria taxonomy based on a combination of molecular background with phenotype and ecological consequences (genus and species concept). Hydrobiologia 2010, 639, 245–259. [Google Scholar] [CrossRef]
- Kling, H.J.; Dail Laughinghouse IV, H.; Marda, J.; Komarek, J.; Acreman, J.; Bruun, K.; Watson, S.B.; Chen, F. A new red colonial Pseudanabaena (Cyanoprokaryota, Oscillatoriales) from North American large lakes. Fottea 2012, 12, 327–339. [Google Scholar] [CrossRef]
| Order | Family | Genera | No. 1 | Ecosystem |
|---|---|---|---|---|
| Chroococcales | Chroococcaceae | Gloeocapsopsis | 1 | Th |
| Chroococcales | Cyanobacteriaceae | Cyanobacterium | 1 | L |
| Chroococcales | Microcystaceae | Microcystis | 2 | L |
| Nostocales | Aphanizomenonaceae | Anabaenopsis | 1 | L |
| Nostocales | Aphanizomenonaceae | Aphanizomenon | 4 | L |
| Nostocales | Aphanizomenonaceae | Dolichospermum | 3 | L |
| Nostocales | Aphanizomenonaceae | Sphaerospermopsis | 1 | L |
| Nostocales | Chlorogloeopsidaceae | Chlorogloeopsis | 5 | Th |
| Nostocales | Fortieaceae | Fortiea | 2 | L |
| Nostocales | Hapalosiphonaceae | Fischerella | 1 | Th |
| Nostocales | Hapalosiphonaceae | Hapalosiphon | 1 | Tr |
| Nostocales | Hapalosiphonaceae | Mastigocladus | 8 | Th |
| Nostocales | Hapalosiphonaceae | Westiellopsis | 2 | Th |
| Nostocales | Microchaetaceae | Coleospermum | 19 | L, Th, Tr |
| Nostocales | Microchaetaceae | Goleter | 1 | L |
| Nostocales | Microchaetaceae | Microchaete | 2 | L, Th |
| Nostocales | Nostocaceae | - | 1 | L |
| Nostocales | Nostocaceae | Anabaena | 5 | L |
| Nostocales | Nostocaceae | Cylindrospermum | 2 | S, Tr |
| Nostocales | Nostocaceae | Hydrocoryne | 1 | L |
| Nostocales | Nostocaceae | Isocystis | 1 | L |
| Nostocales | Nostocaceae | Nostoc | 32 | L, S, Tr |
| Nostocales | Rivulariaceae | Calothrix | 11 | L, S |
| Nostocales | Rivulariaceae | Rivularia | 3 | L |
| Nostocales | Scytonemataceae | Scytonematopsis | 2 | L |
| Nostocales | Tolypothrichaceae | Tolypothrix | 9 | L, S |
| Oscillatoriales | Oscillatoriaceae | Kamptonema | 1 | L |
| Oscillatoriales | Oscillatoriaceae | Lyngbya | 1 | Tr |
| Oscillatoriales | Oscillatoriaceae | Phormidium | 2 | L, Th |
| Oscillatoriales | Oscillatoriaceae | Tychonema | 1 | L |
| Oscillatoriales | Microcoleaceae | Arthrospira | 3 | L |
| Synechococcales | Leptolyngbyaceae | Leptolyngbya | 12 | L, Th |
| Synechococcales | Leptolyngbyaceae | Stenomitos | 1 | L |
| Synechococcales | Oculatellaceae | Pegethrix | 1 | Tr |
| Synechococcales | Oculatellaceae | Tildeniella | 1 | L |
| Synechococcales | Pseudanabaenaceae | Pseudanabaena | 5 | L |
| Synechococcales | Pseudanabaenaceae | Limnothrix | 3 | L |
| Synechococcales | Synechococcaceae | Cyanobium | 3 | L, Tr |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cordeiro, R.; Luz, R.; Vasconcelos, V.; Gonçalves, V.; Fonseca, A. Cyanobacteria Phylogenetic Studies Reveal Evidence for Polyphyletic Genera from Thermal and Freshwater Habitats. Diversity 2020, 12, 298. https://doi.org/10.3390/d12080298
Cordeiro R, Luz R, Vasconcelos V, Gonçalves V, Fonseca A. Cyanobacteria Phylogenetic Studies Reveal Evidence for Polyphyletic Genera from Thermal and Freshwater Habitats. Diversity. 2020; 12(8):298. https://doi.org/10.3390/d12080298
Chicago/Turabian StyleCordeiro, Rita, Rúben Luz, Vitor Vasconcelos, Vítor Gonçalves, and Amélia Fonseca. 2020. "Cyanobacteria Phylogenetic Studies Reveal Evidence for Polyphyletic Genera from Thermal and Freshwater Habitats" Diversity 12, no. 8: 298. https://doi.org/10.3390/d12080298
APA StyleCordeiro, R., Luz, R., Vasconcelos, V., Gonçalves, V., & Fonseca, A. (2020). Cyanobacteria Phylogenetic Studies Reveal Evidence for Polyphyletic Genera from Thermal and Freshwater Habitats. Diversity, 12(8), 298. https://doi.org/10.3390/d12080298