Diversity of NC10 Bacteria and ANME-2d Archaea in Sediments of Fault Zones at Lake Baikal
Abstract
1. Introduction
2. Materials and Methods
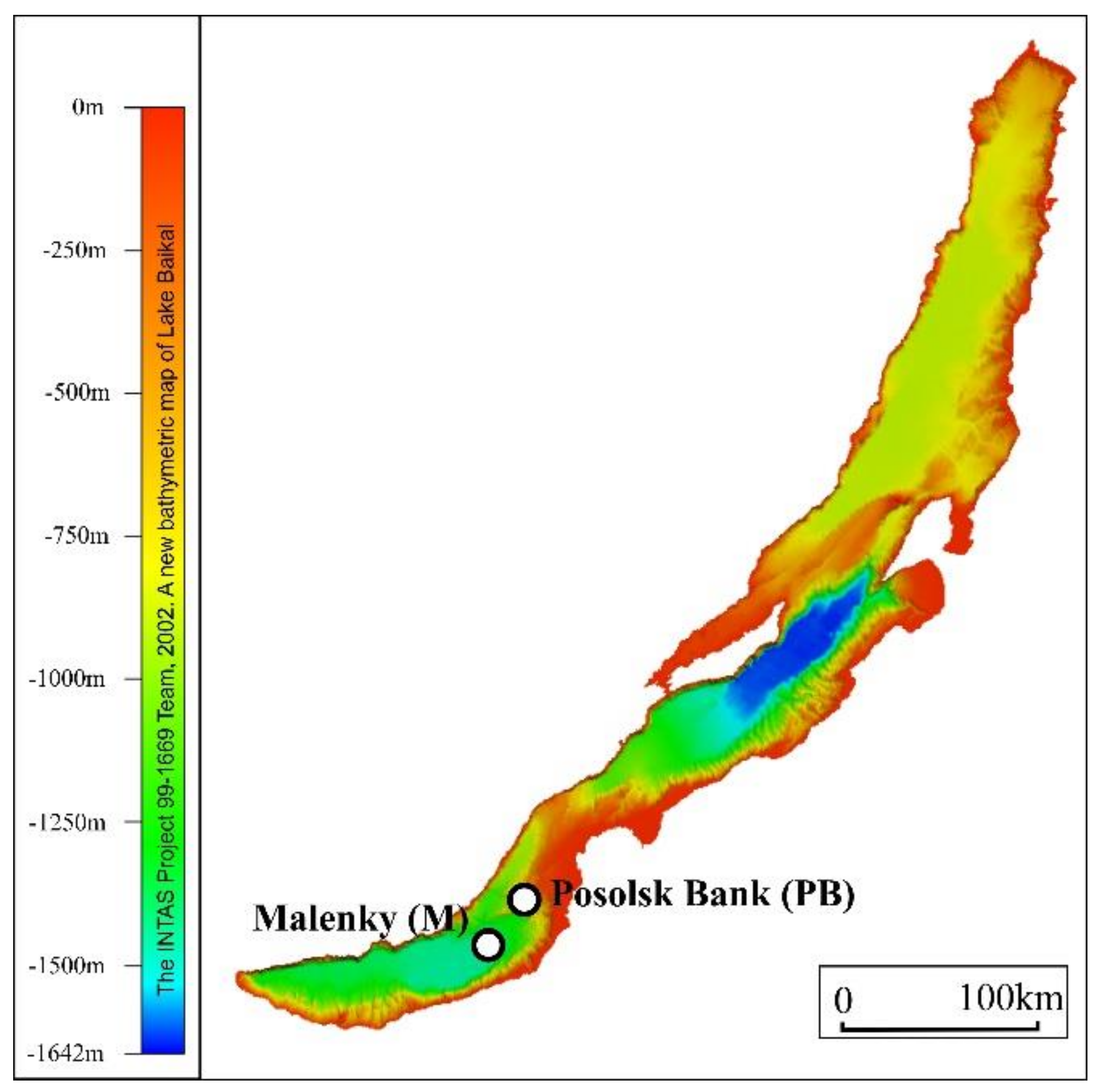
2.1. Description of Sampling Sites, Sample Collection, and Pretreatment
2.2. Measurements of Methane Concentrations and Analysis of the Chemical Composition of Pore Waters
2.3. Molecular Analyses
2.4. 16S rRNA Gene High-Throughput Sequencing of Archaeal Communities
3. Results
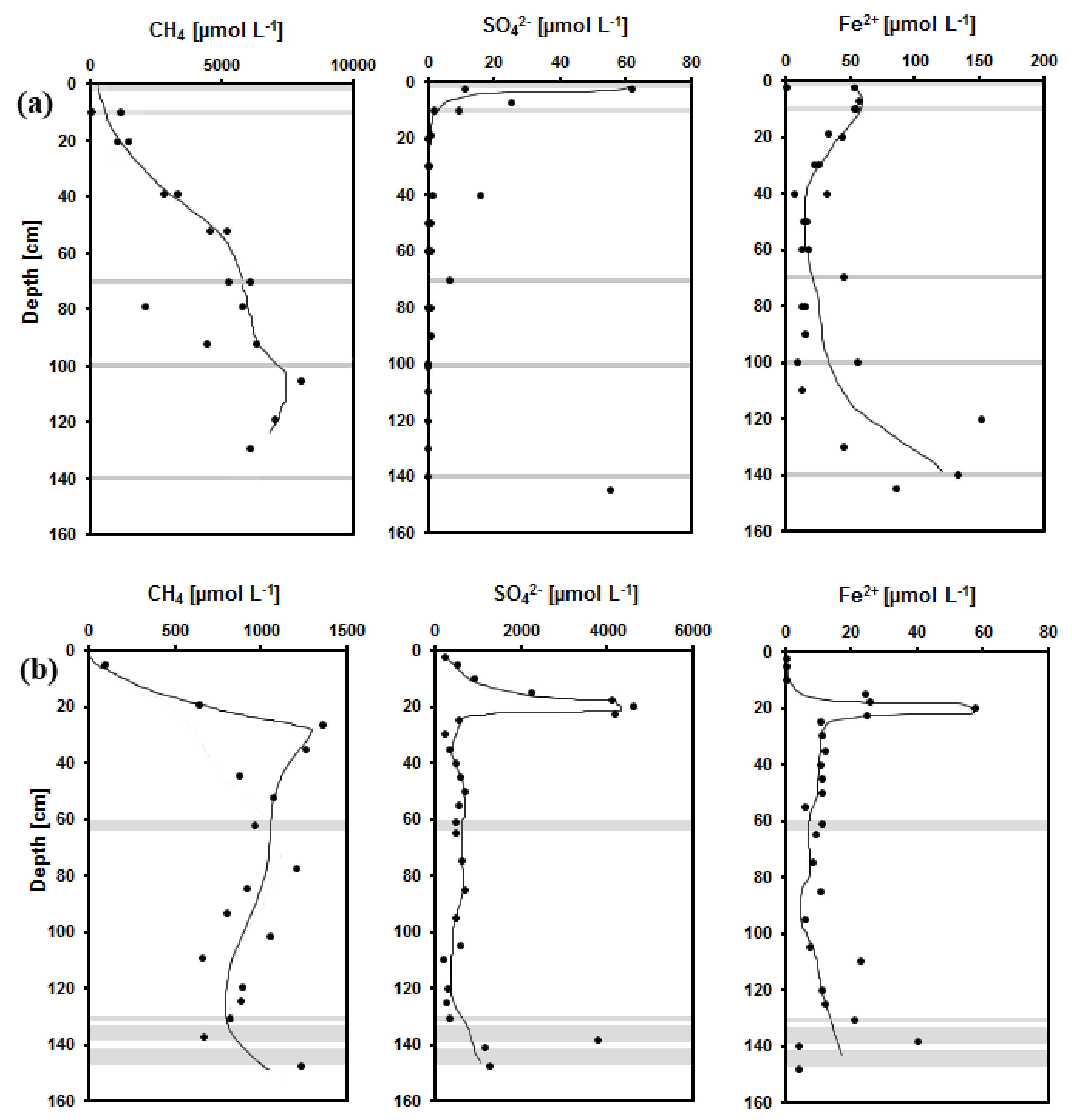
3.1. Chemical Composition of Pore Waters
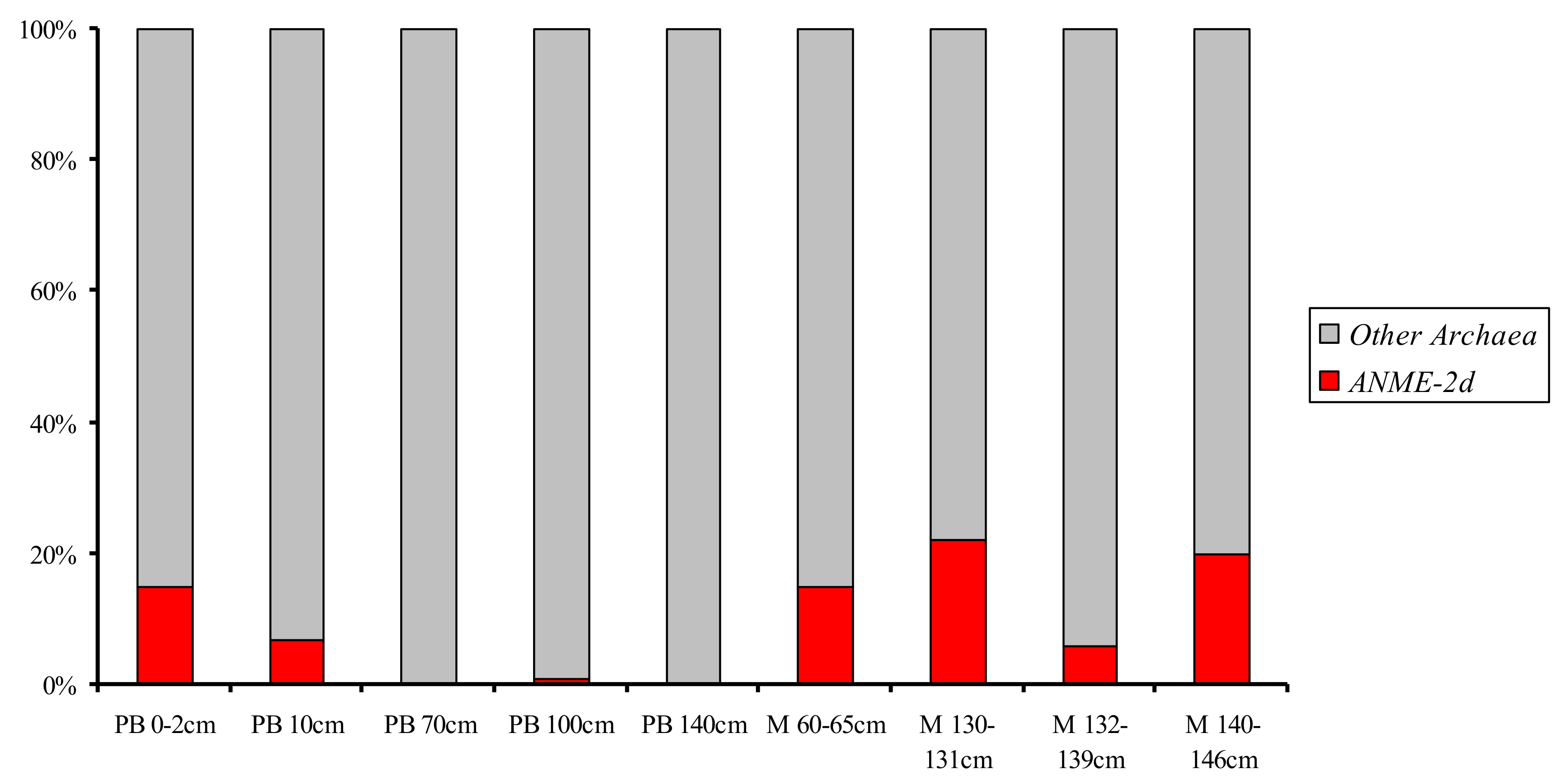
3.2. Diversity of the ANME-2d 16S rRNA Genes in Libraries
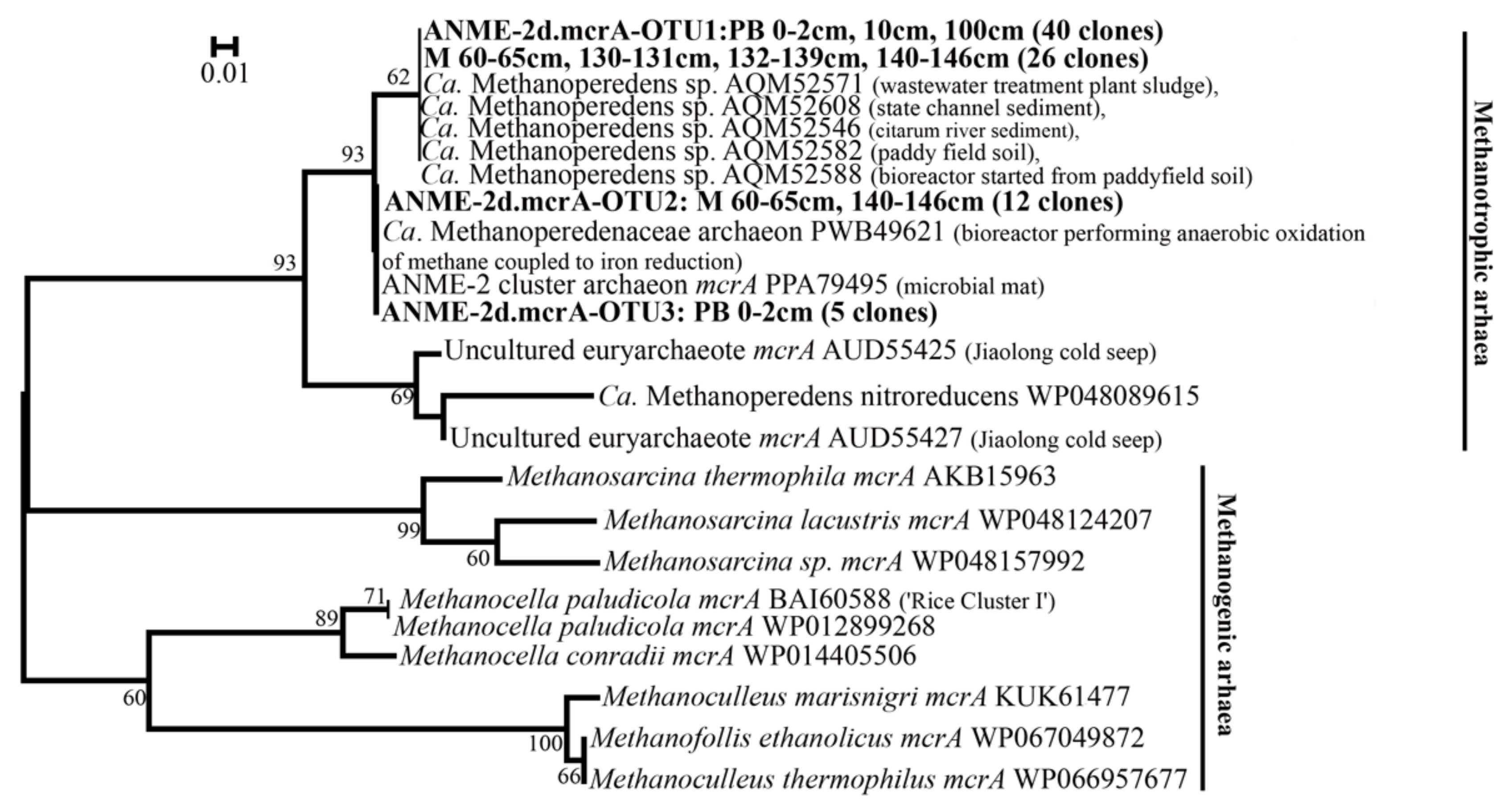
3.3. Diversity of mcrA Genes of ANME-2d Sequences in Clone Libraries
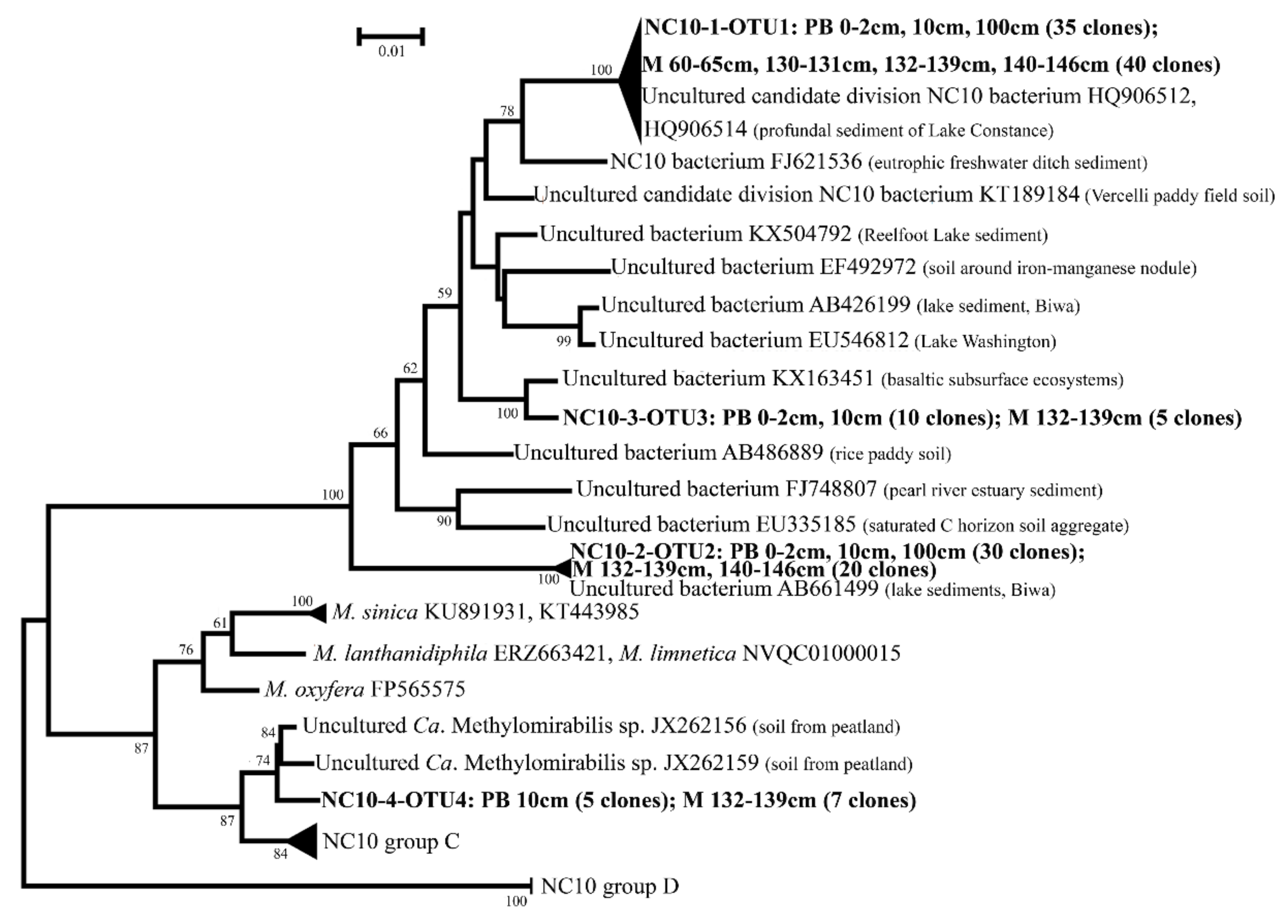
3.4. Diversity of NC10 Bacteria in 16S rRNA Gene Clone Libraries
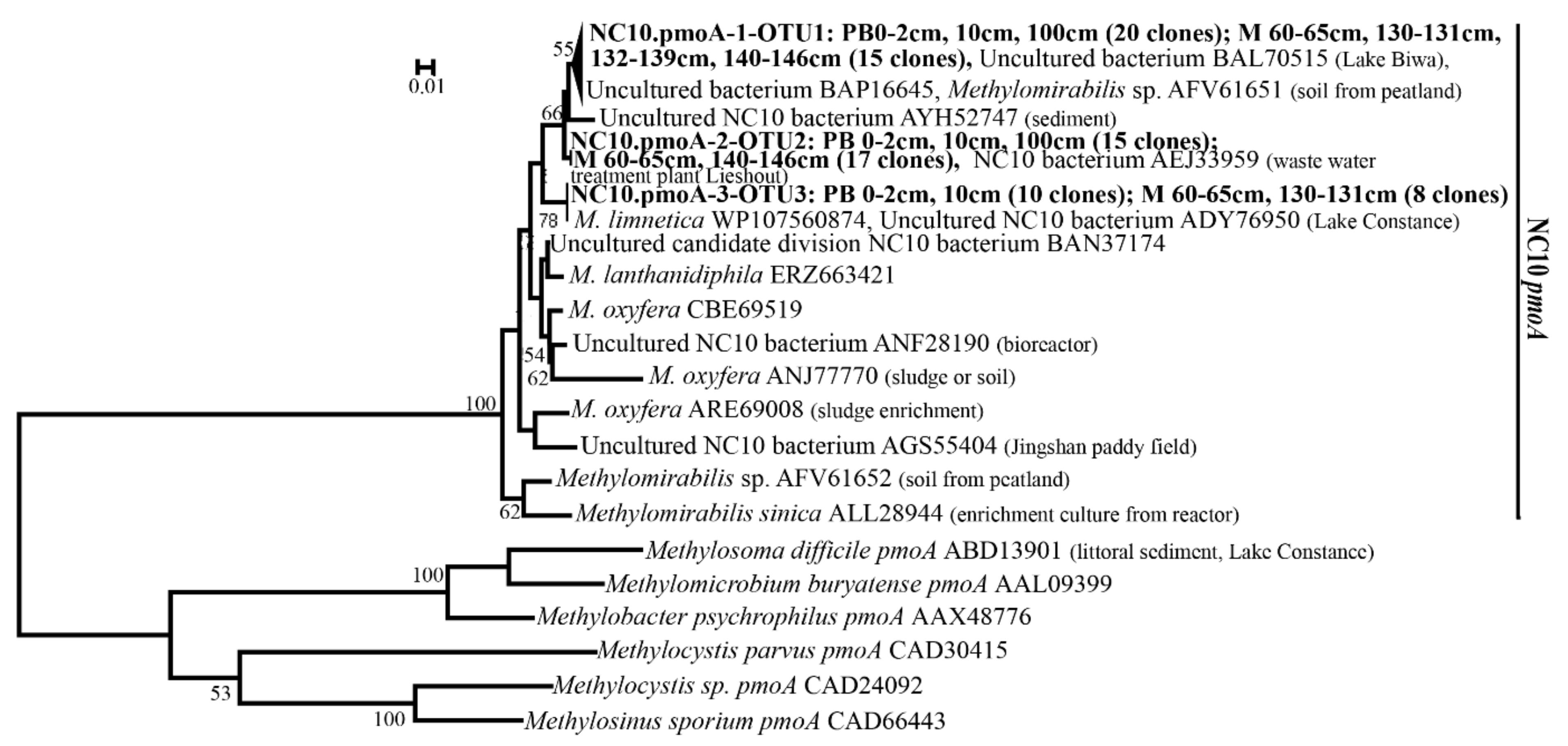
3.5. Diversity of pmoA Genes of the NC10 Bacteria in Clone Libraries
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Kortelainen, P.; Pajunen, H.; Rantakari, M.; Saarnisto, M. A Large carbon pool and small sink in boreal holocene lake sediments. Glob. Chang. Biol. 2004, 10, 1648–1653. [Google Scholar] [CrossRef]
- Molot, L.A.; Dillon, P.J. Storage of terrestrial carbon in boreal lake sediments and evasion to the stmosphere. Glob. Biogeochem. Cycles 1996, 10, 483–492. [Google Scholar] [CrossRef]
- Bastviken, D.; Cole, J.; Pace, M.; Tranvik, L. Methane Emissions from Lakes: Dependence of Lake characteristics, two regional assessments, and a global estimate. Glob. Biogeochem. Cycles 2004, 18, GB4009. [Google Scholar] [CrossRef]
- Chistoserdova, L. Methylotrophs in natural habitats: Current insights through metagenomics. Appl. Microbiol. Biotechnol. 2015, 99, 5763–5779. [Google Scholar] [CrossRef]
- Dedysh, S.N.; Derakshani, M.; Liesack, W. Detection and enumeration of methanotrophs in acidic Sphagnum peat by 16S rRNA fuorescence in situ hybridization, including the use of newly developed oligonucleotide probes for Methylocella palustris. Appl. Environ. Microbiol. 2001, 67, 4850–4857. [Google Scholar] [CrossRef]
- Gal’chemko, V.F. Methanotrophic Bacteria; GEOS: Moscow, Russia, 2001; p. 500. [Google Scholar]
- Hanson, R.S.; Hanson, T.E. Methanotrophic bacteria. Microbiol. Rev. 1996, 60, 439–471. [Google Scholar]
- Kallistova, A.Y.; Goel, G.; Nozhevnikova, A.N. Microbial diversity of methanogenic communities in the systems for anaerobic treatment of organic waste. Microbiology 2014, 83, 462–483. (In Russian) [Google Scholar] [CrossRef]
- Pimenov, N.V.; Kallistova, A.Y.; Rusanov, I.I.; Yusupov, S.K.; Montonen, L.; Jurgens, G.; Münster, U.; Nozhevnikova, A.N.; Ivanov, M.V. Methane formation and oxidation in the meromictic oligotrophic Lake Gek-Gel (Azerbaijan). Microbiology 2010, 79, 247–252. (In Russian) [Google Scholar] [CrossRef]
- Op den Camp, H.J.M.; Islam, T.; Stott, M.B.; Harhangi, H.R.; Hynes, A.; Schouten, S.; Jetten, M.S.M.; Birkeland, N.K.; Pol, A.; Dunfield, P.F. Environmental, genomic and taxonomic perspectives on methanotrophic Verrucomicrobia. Environ. Microbiol. Rep. 2009, 1, 293–306. [Google Scholar] [CrossRef]
- Boetius, A.; Ravenschlag, K.; Schubert, C.; Rickert, D.; Widdel, F.; Gieseke, A.; Amann, R.; Jørgensen, B.B.; Witte, U.; Pfannkuche, O. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 2000, 407, 623–626. [Google Scholar] [CrossRef]
- Hoehler, T.M.; Alperin, M.J.; Albert, D.B.; Martens, C.S. Field and laboratory studies of methane oxidation in an anoxic marine sediment: Evidence for a methanogen-sulfate reducer consortium. Glob. Biogeochem. Cycles 1994, 8, 451–463. [Google Scholar] [CrossRef]
- Reeburgh, W.S. Methane consumption in Cariaco trench waters and sediments. Earth Planet. Sci. Lett. 1976, 28, 337–344. [Google Scholar] [CrossRef]
- Valentine, D.L.; Reeburgh, W.S. New perspectives on anaerobic methane oxidation. Environ. Microbiol. 2000, 2, 477–484. [Google Scholar] [CrossRef] [PubMed]
- Knittel, K.; Losekann, T.; Boetius, A.; Kort, R.; Amann, R. Diversity and distribution of methanotrophic archaea at cold seeps. Appl. Environ. Microbiol. 2005, 71, 467–479. [Google Scholar] [CrossRef]
- Knittel, K.; Boetius, A. Anaerobic oxidation of methane: Progress with an unknown process. Annu. Rev. Microbiol. 2009, 63, 311–334. [Google Scholar] [CrossRef]
- Nauhaus, K.; Boetius, A.; Kruger, M.; Widdel, F. In vitro demonstration of anaerobic oxidation of methane coupled to sulphate reduction in sediment from a marine gas hydrate area. Environ. Microbiol. 2002, 4, 296–305. [Google Scholar] [CrossRef]
- Hinrichs, K.; Boetius, A. The anaerobic oxidation of methane: New insights in microbial ecology and biogeochemistry. In Ocean margin systems; Springer: Berlin, Germany, 2002; pp. 457–477. [Google Scholar]
- Ding, J.; Ding, Z.W.; Fu, L.; Lu, Y.Z.; Cheng, S.H.; Zeng, R.J. New primers for detecting and quantifying denitrifying anaerobic methane oxidation archaea in different ecological niches. Appl. Microbiol. Biotechnol. 2015, 99, 9805–9812. [Google Scholar] [CrossRef]
- Egger, M.; Rasigraf, O.; Sapart, C.J.; Jilbert, T.; Jetten, M.S.; Rockmann, T.; van der Veen, C.; Banda, N.; Kartal, B.; Ettwig, K.; et al. Iron-mediated anaerobic oxidation of methane in brackish coastal sediments. Environ. Sci. Technol. 2015, 49, 277–283. [Google Scholar] [CrossRef]
- Ettwig, K.F.; Butler, M.K.; Le Paslier, D.; Pelletier, E.; Mangenot, S.; Kuypers, M.M.; Schreiber, F.; Dutilh, B.E.; Zedelius, J.; de Beer, D.; et al. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 2010, 464, 543–548. [Google Scholar] [CrossRef]
- Haroon, M.F.; Hu, S.; Shi, Y.; Imelfort, M.; Keller, J.; Hugenholtz, P.; Yuan, Z.; Tyson, G.W. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 2013, 500, 567–570. [Google Scholar] [CrossRef]
- Raghoebarsing, A.A.; Pol, A.; van de Pas-Schoonen, K.T.; Smolders, A.J.; Ettwig, K.F.; Rijpstra, W.I.; Schouten, S.; Damste, J.S.; Op den Camp, H.J.; Jetten, M.S.; et al. A Microbial consortium couples anaerobic methane oxidation to denitrification. Nature 2006, 440, 918–921. [Google Scholar] [CrossRef] [PubMed]
- Weber, H.S.; Habicht, K.S.; Thamdrup, B. Anaerobic methanotrophic archaea of the ANME-2d cluster are active in a low-sulfate, iron-rich freshwater sediment. Front. Microbiol. 2017, 8, 619. [Google Scholar] [CrossRef] [PubMed]
- Luesken, F.A.; Zhu, B.; van Alen, T.A.; Butler, M.K.; Diaz, M.R.; Song, B.; Op den Camp, H.J.; Jetten, M.S.; Ettwig, K.F. pmoA primers for detection of anaerobic methanotrophs. Appl. Environ. Microbiol. 2011, 77, 3877–3880. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Wang, J.; Bradford, L.M.; Ettwig, K.; Hu, B.; Lueders, T. Nitric oxide dismutase (nod) genes as a functional marker for the diversity and phylogeny of methane-driven oxygenic denitrifiers. Front. Microbiol. 2019, 10, 1577. [Google Scholar] [CrossRef] [PubMed]
- Deutzmann, J.S.; Schink, B. Anaerobic Oxidation of Methane in Sediments of Lake Constance, an Oligotrophic Freshwater Lake. Appl. Environ. Microbiol. 2011, 77, 4429–4436. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.L.; Shen, L.D.; Lian, X.; Zhu, Q.; Liu, S.; Huang, Q.; He, Z.F.; Geng, S.; Cheng, D.Q.; Lou, L.P.; et al. Evidence for nitrite-dependent anaerobic methane oxidation as a previously overlooked microbial methane sink in wetlands. Proc. Natl. Acad. Sci. USA 2014, 111, 4495–4500. [Google Scholar] [CrossRef]
- Zhou, L.; Wang, Y.; Long, X.E.; Guo, J.; Zhu, G. High abundance and diversity of nitrite-dependent anaerobic methane-oxidizing bacteria in a paddy field profile. FEMS Microbiol. Lett. 2014, 360, 33–41. [Google Scholar] [CrossRef]
- Zhu, B.; van Dijk, G.; Fritz, C.; Smolders, A.J.; Pol, A.; Jetten, M.S.; Ettwig, K.F. Anaerobic oxidization of methane in a minerotrophic peatland: Enrichment of nitrite-dependent methane-oxidizing bacteria. Appl. Environ. Microbiol. 2012, 78, 8657–8665. [Google Scholar] [CrossRef]
- Zhu, B.; Bradford, L.; Huang, S.; Szalay, A.; Leix, C.; Weissbach, M.; Táncsics, A.; Drewes, J.E.; Lueders, T. Unexpected diversity and high abundance of putative nitric oxide dismutase (nod) genes in contaminated aquifers and wastewater treatment systems. Appl. Environ. Microbiol. 2017, 4, 02750–16. [Google Scholar] [CrossRef]
- Lu, Y.-Z.; Liang, F.; Li, N.; Ding, J.; Bai, Y.-N.; Samaras, P.; Zeng, R.J. The content of trace element iron is a key factor for competition between anaerobic ammonium oxidation and methane-dependent denitrification processes. Chemosphere 2018, 198, 370–376. [Google Scholar] [CrossRef]
- Fu, L.; Li, S.W.; Ding, Z.W.; Ding, J.; Lu, Y.Z.; Zeng, R.J. Iron reduction in the DAMO/Shewanella oneidensis MR-1 coculture system and the fate of Fe(II). Water Res. 2016, 88, 808–815. [Google Scholar] [CrossRef] [PubMed]
- Timmers, P.H.; Welte, C.U.; Koehorst, J.J.; Plugge, C.M.; Jetten, M.S.; Stams, A.J. Reverse methanogenesis and respiration in methanotrophic archaea. Archaea 2017, 2017, 1654237. [Google Scholar] [CrossRef] [PubMed]
- Welte, C.U.; Rasigraf, O.; Vaksmaa, A.; Versantvoort, W.; Arshad, A.; Op den Camp, H.J.; Jetten, M.S.; Lüke, C.; Reimann, J. Nitrate- and nitrite-dependent anaerobic oxidation of methane. Environ. Microbiol. Rep. 2016, 8, 941. [Google Scholar] [CrossRef] [PubMed]
- Arshad, A.; Speth, D.R.; De Graaf, R.M.; Op den Camp, H.J.M.; Jetten, M.S.; Welte, C.U. A metagenomics-based metabolic model of nitrate-dependent anaerobic oxidation of methane by Methanoperedens-like archaea. Front. Microbiol. 2015, 6, 1423. [Google Scholar] [CrossRef]
- Hales, B.A.; Edwards, C.; Ritchie, D.A.; Hall, G.; Pickup, R.W.; Saunders, J.R. Isolation and identification of methanogen-specific DNA from blanket bog feat by PCR amplification and sequence analysis. Appl. Environ. Microbiol. 1996, 62, 668–675. [Google Scholar]
- Juottonen, H.; Galand, P.E.; Yrjala, K. Detection of Methanogenic Archaea in Peat: Comparison of PCR primers targeting the mcrA gene. Res. Microbiol. 2006, 157, 914–921. [Google Scholar] [CrossRef]
- Luton, P.E.; Wayne, J.M.; Sharp, R.J.; Riley, P.W. The mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill. Microbiology 2002, 148, 3521–3530. [Google Scholar] [CrossRef]
- Nunoura, T.; Oida, H.; Miyazaki, J.; Miyashita, A.; Imachi, H.; Takai, K. Quantification of mcrA by fluorescent PCR in methanogenic and methanotrophic microbial communities. FEMS Microbiol. Ecol. 2008, 64, 240–247. [Google Scholar] [CrossRef]
- Vaksmaa, A.; Guerrero-Cruz, S.; van Alen, T.A.; Cremers, G.; Ettwig, K.F.; Luke, C.; Jetten, M.S.M. Enrichment of anaerobic nitrate-dependent methanotrophic “Candidatus Methanoperedens nitroreducens” archaea from an italian paddy field soil. Appl. Microbiol. Biotechnol. 2017, 101, 7075–7084. [Google Scholar] [CrossRef]
- Evans, P.N.; Parks, D.H.; Chadwick, G.L.; Robbins, S.J.; Orphan, V.J.; Golding, S.D.; Tyson, G.V. Methane metabolism in the archaeal phylum Bathyarchaeota revealed by genome-centric metagenomics. Science 2015, 350, 434–438. [Google Scholar] [CrossRef]
- Vanwonterghem, I.; Evans, P.N.; Parks, D.H.; Jensen, P.D.; Woodcroft, B.J.; Hugenholtz, P.; Hugenholtz, G.V. Methylotrophic methanogenesis discovered in the archaeal phylum Verstraetearchaeota. Nat. Microbiol. 2016, 1, 16170. [Google Scholar] [CrossRef] [PubMed]
- Ettwig, K.F.; Zhu, B.; Speth, D.; Keltjens, J.T.; Jetten, M.S.M.; Kartal, B. Archaea Catalyze iron-dependent anaerobic oxidation of methane. Proc. Natl. Acad. Sci. USA 2016, 113, 12792–12796. [Google Scholar] [CrossRef] [PubMed]
- Beal, E.J.; House, C.H.; Orphan, V.J. Manganese- and iron-dependent marine methane oxidation. Science 2009, 325, 184–187. [Google Scholar] [CrossRef] [PubMed]
- Sivan, O.; Adler, M.; Pearson, A.; Gelman, F.; Bar-Or, I.; John, S.G.; Eckert, W. Geochemical evidence for iron-mediated anaerobic oxidation of methane. Limnol. Oceanogr. 2011, 56, 1536–1544. [Google Scholar] [CrossRef]
- Norði, K.A.; Thamdrup, B.; Schubert, C.J. Anaerobic oxidation of methane in an iron-rich danish freshwater lake sediment. Limnol. Oceanogr. 2013, 58, 546–554. [Google Scholar] [CrossRef]
- Savvichev, A.S.; Kokryatskaya, N.M.; Zabelina, S.A.; Rusanov, I.I.; Zakharova, E.E.; Veslopolova, E.F.; Lunina, O.N.; Patutina, E.O.; Bumazhkin, B.K.; Gruzdev, D.S.; et al. Microbial Processes of the carbon and sulfur cycles in an ice-covered, iron-rich neromictic Lake Svetloe (Arkhangelsk Region, Russia). Environ. Microbiol. 2017, 19, 659–672. [Google Scholar] [CrossRef]
- Cai, C.; Leu, A.O.; Xie, G.J.; Guo, J.; Feng, Y.; Zhao, J.X.; Tyson, G.W.; Yuan, Z.; Hu, S. A methanotrophic archaeon couples anaerobic oxidation of methane to Fe(III) reduction. ISME J. 2018, 12, 1929–1939. [Google Scholar] [CrossRef]
- Weber, H.S.; Thamdrup, B.; Habicht, K.S. High sulfur isotope fractionation associated with anaerobic oxidation of methane in a low sulfate, iron-rich environment. Front. Earth Sci. 2016, 4, 61. [Google Scholar] [CrossRef]
- Khlystov, O.M.; Khabuev, A.V.; Minami, H.; Hachikubo, A.; Krylov, A.A. Gas hydrates in Lake Baikal. LFWB 2018, 1, 66–70. (In Russian) [Google Scholar] [CrossRef]
- Khlystov, O.M.; Poort, J.; Mazzini, A.; Akhmanov, G.G.; Minami, H.; Hachikubo, A.; Khabuev, A.V.; Kazakov, A.V.; De Batist, M.; Naudts, L.; et al. Shallow-rooted mud volcanism in Lake Baikal. Mar. Pet. Geol. 2019, 102, 580–589. [Google Scholar] [CrossRef]
- Dagurova, O.P.; Namsaraev, B.B.; Kozyreva, L.P.; Zemskaya, T.I.; Dulov, L.E. Bacterial processes of the methane cycle in bottom sediments of Lake Baikal. Microbiology 2004, 73, 202–210. (In Russian) [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2010, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Pimenov, N.V.; Zakharova, E.E.; Bryukhanov, A.L.; Korneeva, V.A.; Kuznetsov, B.B.; Tourova, T.P.; Pogodaeva, T.V.; Kalmychkov, G.V.; Zemskaya, T.I. Activity and structure of the sulfate-reducing bacterial community in the sediments of the Southern Part of Lake Baikal. Microbiology 2014, 83, 47–55. (In Russian) [Google Scholar] [CrossRef]
- Zemskaya, T.I.; Khlystov, O.M.; Egorov, A.V.; Pogodaeva, T.V.; Kalmychkov, G.V.; Shubenkova, O.V.; Chernitsyna, S.M.; Vorob’eva, S.S.; Grachev, M.A. Integrated Studies of Gas Hydrate Manifestations in the Sediments of Lake Baikal Mud Volcanoes. In Processes in the Boisphere: Changes in Soil and Plant Cover and RF Territorial Waters, Matter Turnover Caused by Global Climatic Changes and Catastrophic Processes; Zavarzin, G.A., Kudeyarov, V.N., Eds.; Pushchino: Moscow, Russia, 2008; Chapter 4; pp. 125–152. [Google Scholar]
- Zemskaya, T.I.; Pogodaeva, T.V.; Shubenkova, O.V.; Chernitsina, S.M.; Dagurova, O.P.; Buryukhaev, S.P.; Namsaraev, B.B.; Khlystov, O.M.; Egorov, A.V.; Krylov, A.A.; et al. Geochemical and microbiological characteristics of sediments near the Malenky Mud Volcano (Lake Baikal, Russia), with evidence of archaea intermediate between the marine anaerobic methanotrophs ANME-2 and ANME-3. Geo Mar. Lett. 2010, 30, 411–425. [Google Scholar]
- Schubert, C.J.; Vazquez, F.; Lösekann-Behrens, T.; Knittel, K.; Tonolla, M.; Boetius, A. Evidence for anaerobic oxidation of methane in sediments of a freshwater system (Lago di Cadagno). FEMS Microbiol. Ecol. 2011, 76, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Chernitsyna, S.M.; Mamaeva, E.V.; Lomakina, A.V.; Pogodaeva, T.V.; Galach’yants, Y.P.; Bukin, S.V.; Pimenov, N.V.; Khlystov, O.M.; Zemskaya, T.I. Phylogenetic diversity of microbial communities of the Posolsk Bank bottom sediments, Lake Baikal. Microbiology 2016, 85, 672–680. (In Russian) [Google Scholar] [CrossRef]
- Lomakina, A.V.; Mamaeva, E.V.; Galachyants, Y.P.; Petrova, D.P.; Pogodaeva, T.V.; Shubenkova, O.V.; Khabuev, A.V.; Morozov, I.V.; Zemskaya, T.I. Diversity of archaea in bottom sediments of the discharge area. Geomicrobiol. J. 2018, 35, 50–63. [Google Scholar] [CrossRef]
- Hu, S.; Zeng, R.J.; Burow, L.C.; Lant, P.; Keller, J.; Yuan, Z. Enrichment of denitrifying a anaerobic methane oxidizing microorganisms. Environ. Microbiol. Rep. 2009, 1, 377–384. [Google Scholar] [CrossRef]
- Lomakina, A.V.; Mamaeva, E.V.; Pogodaeva, T.V.; Kalmychkov, G.V.; Khalzov, I.A.; Zemskaya, T.I. Anaerobic methane oxidation in enrichment cultures from deep sediments of a mud volcano Peschanka (South Baikal). Microbiology 2018, 87, 317–325. (In Russian) [Google Scholar] [CrossRef]
- Zemskaya, T.I.; Lomakina, A.V.; Mamaeva, E.V.; Zakharenko, A.S.; Pogodaeva, T.V.; Petrova, D.P.; Galachyants, Y.P. Bacterial communities in sediments of Lake Baikal from areas with oil and gas discharge. Aquat. Microb. Ecol. 2015, 76, 95–109. [Google Scholar] [CrossRef]
- Bol’shakov, A.M.; Egorov, A.V. On the application of phase equilibrium degassing for gasometric research in water areas. Okeanologiya 1987, 37, 861–862. (In Russian) [Google Scholar]
- Pogodaeva, T.V.; Zemskaya, T.I.; Golobokova, L.P.; Khlystov, O.M.; Minami, H.; Sakagami, H. Chemical composition of pore waters of bottom sediments in different Baikal basins. Russ. Geol. Geophys. 2007, 48, 886–900. [Google Scholar] [CrossRef]
- Pogodaeva, T.V.; Lopatina, I.N.; Khlystov, O.M.; Egorov, A.V.; Zemskaya, T.I. Background composition of pore waters in Lake Baikal bottom sediments. J. Great Lakes Res. 2017, 43, 1030–1043. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning. A Laboratory Manual, 2nd ed.; Cold Spring Harbor: New York, NY, USA, 1989. [Google Scholar]
- Shubenkova, O.V.; Zemskaya, T.I.; Chernitsyna, S.M.; Khlystov, O.M.; Triboi, T.I. The first results of an investigation into the phylogenetic diversity of microorganisms in Southern Baikal sediments in the region of subsurface discharge of methane hydrates. Microbiology 2005, 74, 314–320. (In Russian) [Google Scholar] [CrossRef]
- Holmes, A.; Costello, A.; Lidstrom, M.; Murrell, J.C. Evidence that particulate methane monooxygenase and ammonia monooxygenase may be evolutionarily. FEMS Microbiol. Lett. 1995, 132, 203–208. [Google Scholar] [CrossRef]
- Sørensen, K.B.; Teske, A. Stratified communities of active archaea in deep marine subsurface sediments. Appl. Environ. Microbiol. 2006, 72, 4596–4603. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Quince, C.; Lanzen, A.; Davenport, R.J.; Turnbaugh, P.J. Removing noise from pyrosequenced amplicons. BMC Bioinform. 2011, 12, 38. [Google Scholar] [CrossRef]
- Lomakina, A.V.; Pogodaeva, T.V.; Morozov, I.V.; Zemskaya, T.I. Microbial communities of the discharge zone of oil- and gas-bearing fluids in low-mineral Lake Baikal. Microbiology 2014, 83, 278–287. (In Russian) [Google Scholar] [CrossRef]
- Kojima, H.; Tsutsumi, M.; Ishikawa, K.; Iwata, T.; Mußmann, M.; Fukui, M. Distribution of putative denitrifying methane oxidizing bacteria in sediment of a freshwater lake, Lake Biwa. Syst. Appl. Microbiol. 2012, 35, 233–238. [Google Scholar] [CrossRef]
- Wang, J.; Cai, C.; Li, Y.; Hua, M.; Wang, J.; Yang, H.; Zheng, P.; Hu, B. Denitrifying anaerobic methane oxidation: A previously overlooked methane sink in intertidal zone. Environ. Sci. Technol. 2019, 53, 203–212. [Google Scholar] [CrossRef]
- Krylov, A.A.; Minami, H.; Hachikubo, A.; Shoji, H.; Khlystov, O.M.; Zemskaya, T.I.; Pogodaeva, T.V.; Kida, M.; Naudts, L.; Poort, J. Crystallization of authigenic carbonates in mud volcanoes at Lake Baikal. Geochem. Int. 2008, 46, 985–995. (In Russian) [Google Scholar] [CrossRef]
- Holmer, M.; Storkholm, P. Sulphate reduction and sulphur cycling in Lake sediments: A review. Freshw. Biol. 2001, 46, 431–451. [Google Scholar] [CrossRef]
- Klerkx, J.; Zemskaya, T.I.; Matveeva, T.V.; Khlystov, O.M.; Namsaraev, B.B.; Dagurova, O.P.; Golobokova, L.P.; Vorob’eva, S.S.; Pogodaeva, T.P.; Granin, N.G.; et al. Methane hydrates in surface layer of deep-water sediments in Lake Baikal. Dokl. Earth Sci. 2003, 393, 822–826. (In Russian) [Google Scholar]
- Hu, S.; Zeng, R.J.; Keller, J.; Lant, P.A.; Yuan, Z. Effect of nitrate and nitrite on the selection of microorganisms in the denitrifying anaerobic methane oxidation process. Environ. Microbiol. 2011, 3, 315–319. [Google Scholar] [CrossRef]
- Zhu, G.; Jetten, M.; Kuschk, P.; Ettwig, K.; Yin, C. Potential roles of anaerobic ammonium and methane oxidation in the nitrogen cycle of wetland ecosystems. Appl. Microbiol. Biotechnol. 2010, 86, 1043–1055. [Google Scholar] [CrossRef]
- Klerkx, J.; De Batist, M.; Poort, J.; Hus, R.; Van Rensbergen, P.; Khlystov, O.M.; Granin, N. Tectonically Controlled Methane Escape in Lake Baikal. In The Geological Storage of Carbon Dioxide; Lombardi, S., Ed.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 203–219. [Google Scholar]
- Naudts, L.; Khlystov, O.; Granin, N.; Chensky, A.; Poort, J.; De Batist, M. Stratigraphic and structural control on the distribution of gas hydrates and active gas seeps on the Posolsky Bank, Lake Baikal. Geo Mar. Lett. 2012, 32, 395–406. [Google Scholar] [CrossRef]
- Ding, Z.; Ding, J.; Fu, L.; Zhang, F.; Zeng, R.J. Simultaneous enrichment of denitrifying methanotrophs and Anammox bacteria. Appl. Microbiol. Biotechnol. 2014, 98, 10211–10221. [Google Scholar] [CrossRef]
- Wang, Y.; Zhu, G.; Harhangi, H.R.; Zhu, B.; Jetten, M.S.M.; Yin, C.; Op den Camp, H.J.M. Co-occurrence and distribution of nitrite-dependent anaerobic ammonium and methane-oxidizing bacteria in a paddy soil. FEMS Microbiol. Lett. 2012, 336, 79–88. [Google Scholar] [CrossRef]
- Yang, J.; Jiang, H.; Wu, G.; Hou, W.; Sun, Y.; Lai, Z.; Dong, H. Co-Occurrence of nitrite-dependent anaerobic methane oxidizing and anaerobic ammonia oxidizing bacteria in two qinghai-tibetan saline lakes. Front. Earth Sci. 2012, 6, 383–391. [Google Scholar] [CrossRef]
- Ishii, S.; Yamamoto, M.; Kikuchi, M.; Oshima, K.; Hattori, M.; Otsuka, S.; Senoo, K. Microbial populations responsive to denitrification-inducing conditions in rice paddy soil, as revealed by comparative 16S rRNA gene analysis. Appl. Environ. Microbiol. 2009, 75, 7070–7078. [Google Scholar] [CrossRef]
- Ettwig, K.F.; van Alen, T.; van de Pas-Schoonen, K.T.; Jetten, M.S.M.; Strous, M. Enrichment and molecular detection of denitrifying methanotrophic bacteria of the NC10 phylum. Appl. Environ. Microbiol. 2009, 75, 3656–3662. [Google Scholar] [CrossRef] [PubMed]
- Riedinger, N.; Formolo, M.J.; Lyons, T.W.; Henkel, S.; Beck, A.; Kasten, S. An inorganic geochemical argument for coupled anaerobic oxidation of methane and iron reduction in marine sediments. Geobiology 2014, 12, 172–181. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Ouyang, L.; Zhu, Y.; Trimmer, M. Active pathways of anaerobic methane oxidation across contrasting riverbeds. ISME J. 2019, 13, 752–766. [Google Scholar] [CrossRef] [PubMed]

| Primer | Detected Microbial Groups | Sequence (5′–3′) | References |
|---|---|---|---|
| 159F | ANME-2d mcrA gene | AAAGTGCGGAGCAGCAATCACC | [41] |
| 345R | TCGTCCCATTCCTGCTGCATTGC | ||
| NC10-202Fdeg | NC10 16S rRNA gene | RACCAAAGGRGGCGAGCG | [27] |
| NC10-1043Rdeg | TCTCCRCGYTCCCTTGCG | ||
| A189F | NC10 pmoA gene | GGNGACTGGGACTTCTGG | [27,69] |
| NA720R | TCCCCATCCACACCCACCAG | ||
| pUC/M13F | Universal group | GTTTTCCCAGTCACGAC | Promega |
| pUC/M13R | CAGGAAACAGCTATGAC | Promega | |
| A2Fa | Archaea 16S rRNA gene | TTCCGGTTGATCCYGCCRGA | [70] |
| A519R | GGTDTTACCGCGGCKGCKGCTG |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lomakina, A.; Pogodaeva, T.; Kalmychkov, G.; Chernitsyna, S.; Zemskaya, T. Diversity of NC10 Bacteria and ANME-2d Archaea in Sediments of Fault Zones at Lake Baikal. Diversity 2020, 12, 10. https://doi.org/10.3390/d12010010
Lomakina A, Pogodaeva T, Kalmychkov G, Chernitsyna S, Zemskaya T. Diversity of NC10 Bacteria and ANME-2d Archaea in Sediments of Fault Zones at Lake Baikal. Diversity. 2020; 12(1):10. https://doi.org/10.3390/d12010010
Chicago/Turabian StyleLomakina, Anna, Tatyana Pogodaeva, Gennady Kalmychkov, Svetlana Chernitsyna, and Tamara Zemskaya. 2020. "Diversity of NC10 Bacteria and ANME-2d Archaea in Sediments of Fault Zones at Lake Baikal" Diversity 12, no. 1: 10. https://doi.org/10.3390/d12010010
APA StyleLomakina, A., Pogodaeva, T., Kalmychkov, G., Chernitsyna, S., & Zemskaya, T. (2020). Diversity of NC10 Bacteria and ANME-2d Archaea in Sediments of Fault Zones at Lake Baikal. Diversity, 12(1), 10. https://doi.org/10.3390/d12010010





