Bacteria with Different Assemblages in the Soil Profile Drive the Diverse Nutrient Cycles in the Sugarcane Straw Retention Ecosystem
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Site, Experimental Design, and Soil Sampling
2.2. Measurements of Soil Physiochemical Properties
2.3. Measurements of Soil Enzyme Activities
2.4. Soil DNA Extraction
2.5. PCR Amplification and Illumina MiSeq Sequencing
2.6. Processing and Analyzing of Sequencing Data
3. Results
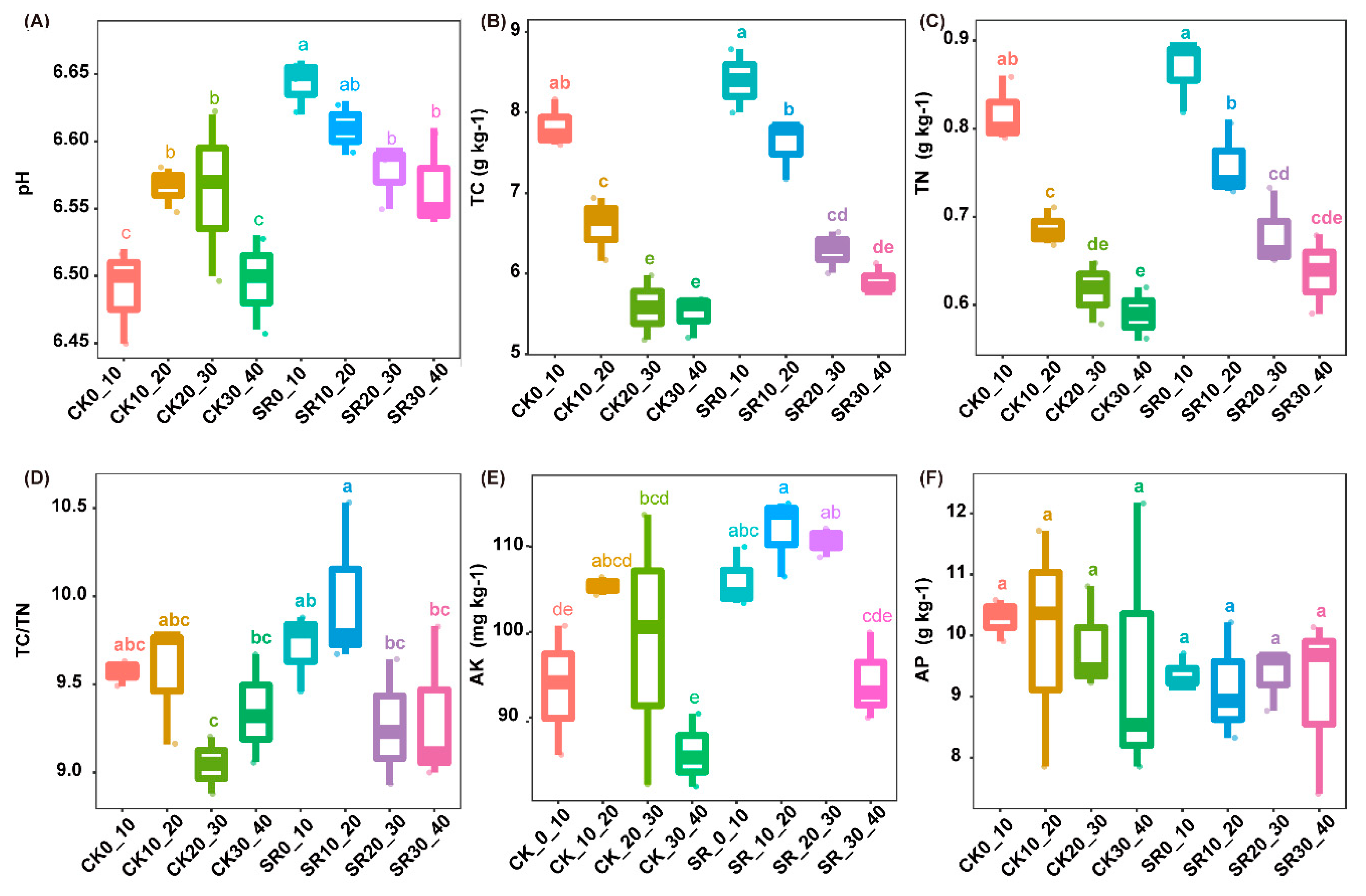
3.1. Soil Physiochemical Properties
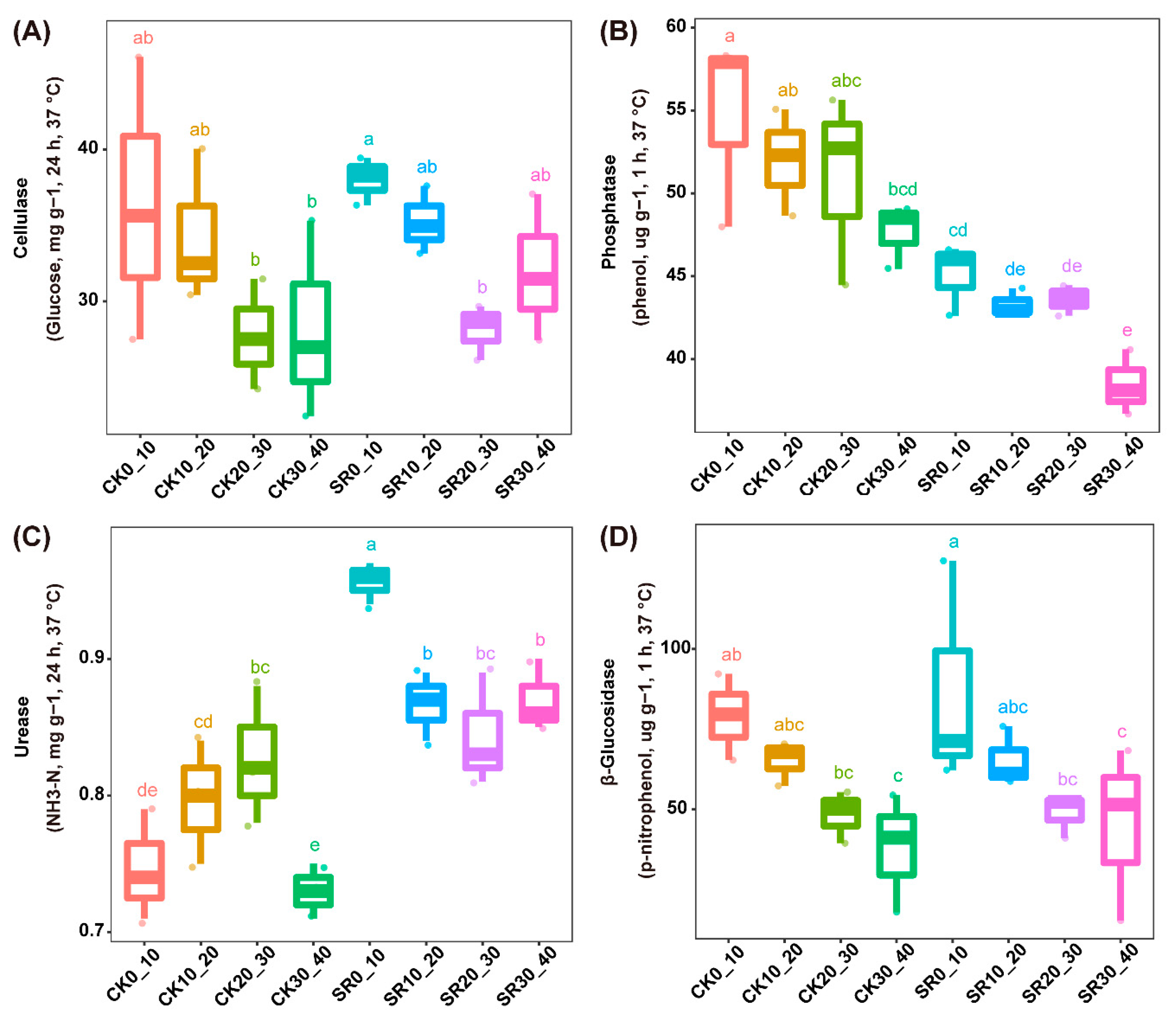
3.2. Soil Enzyme Activities
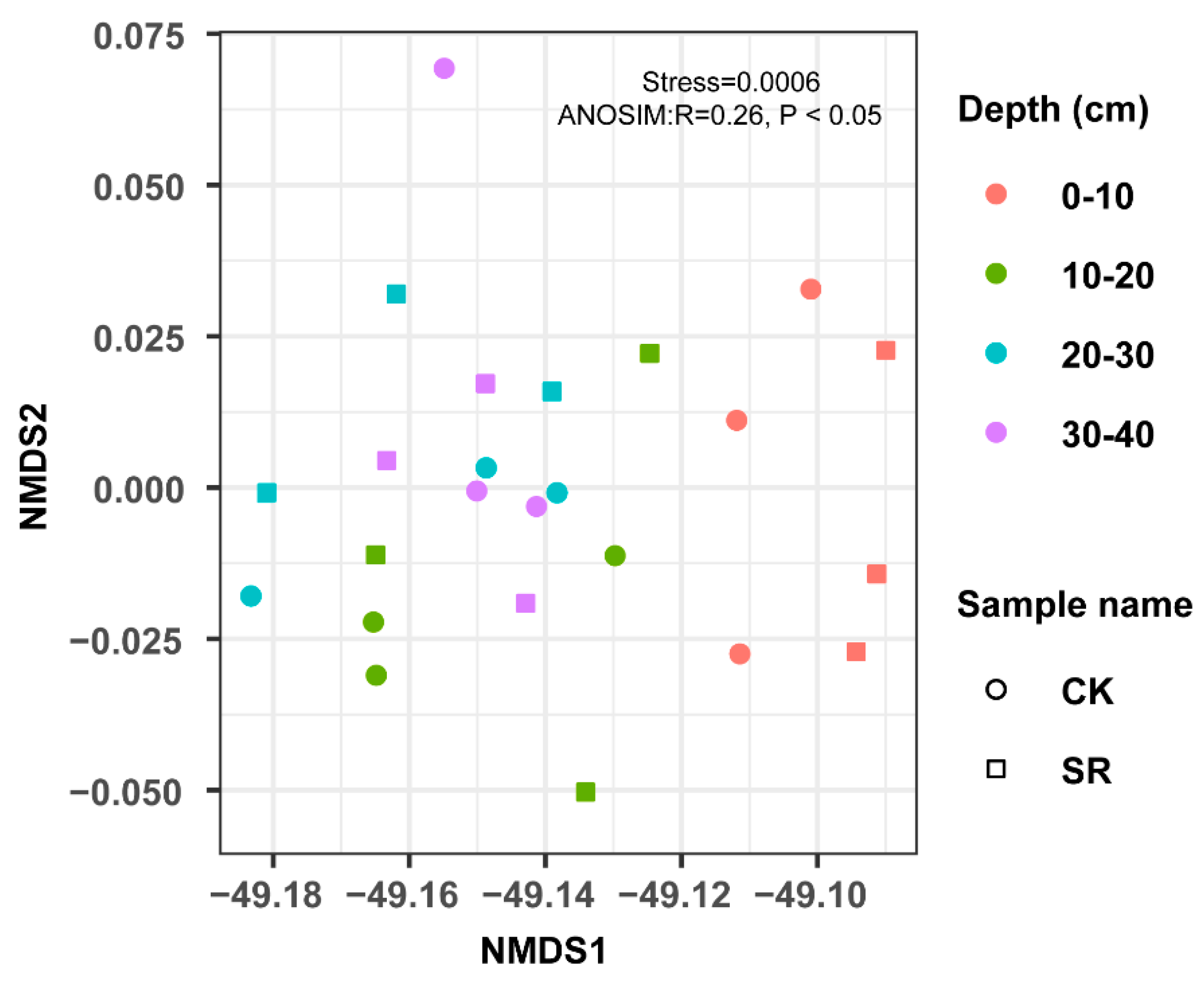
3.3. Bacterial Community Structural Diversity and Similarity
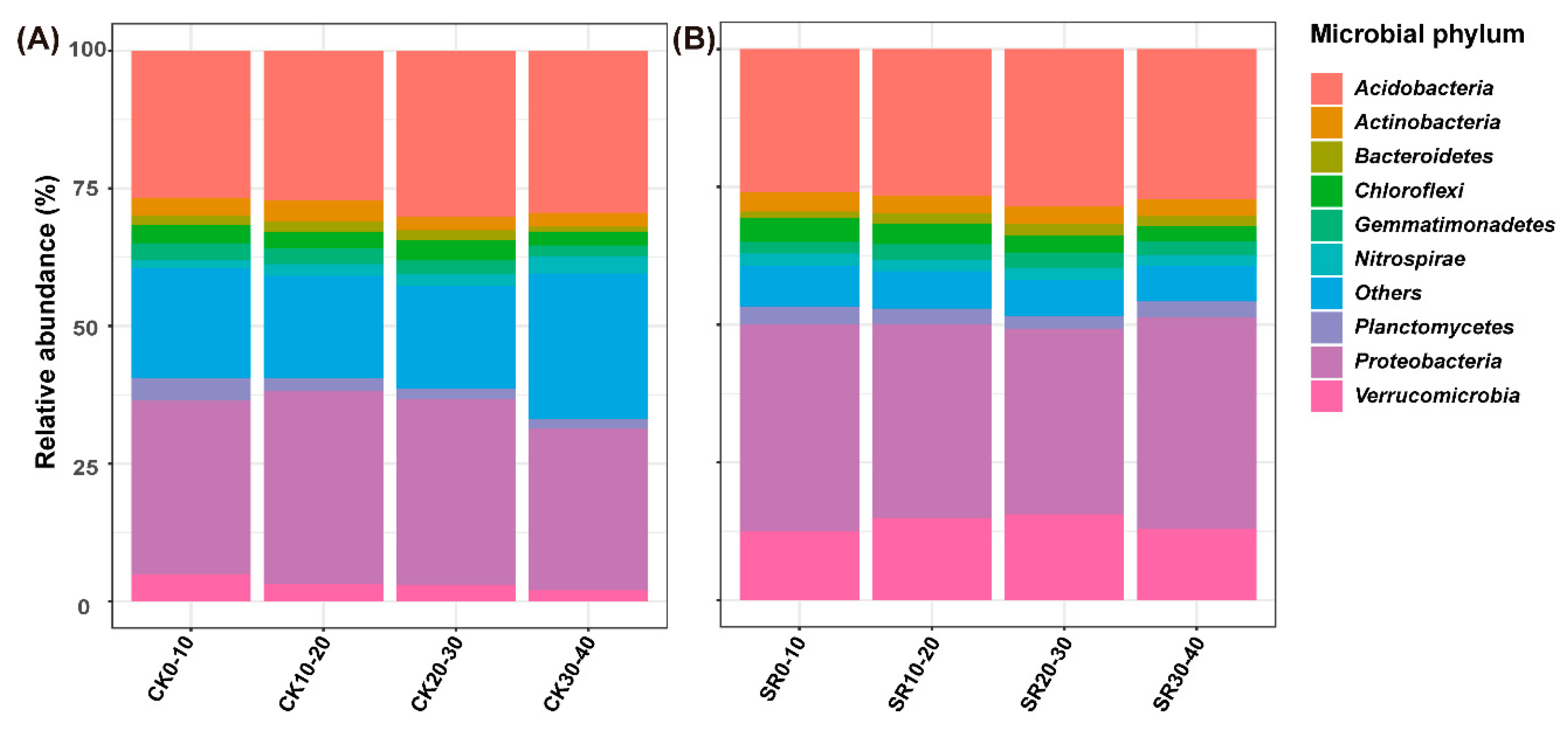
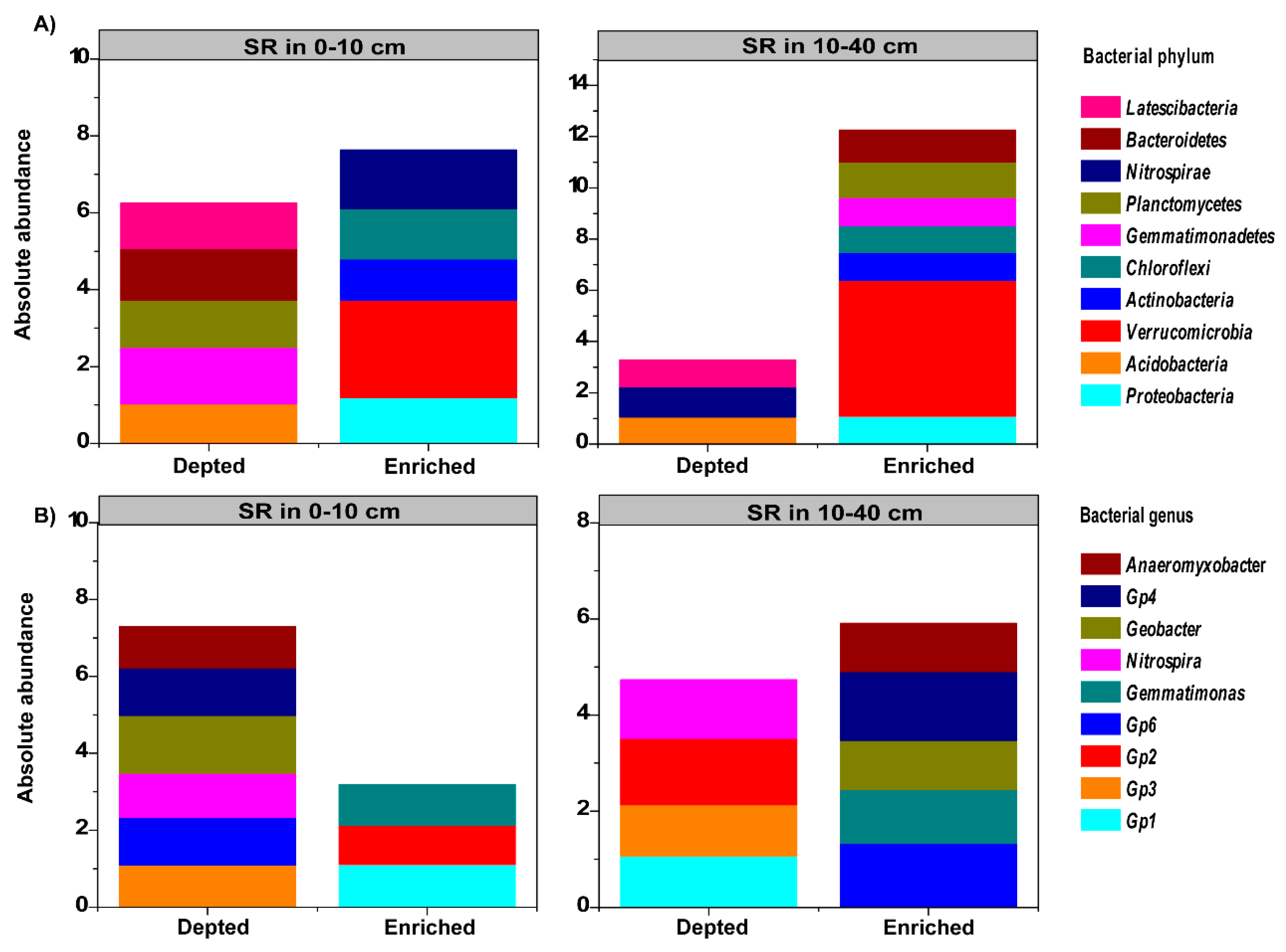
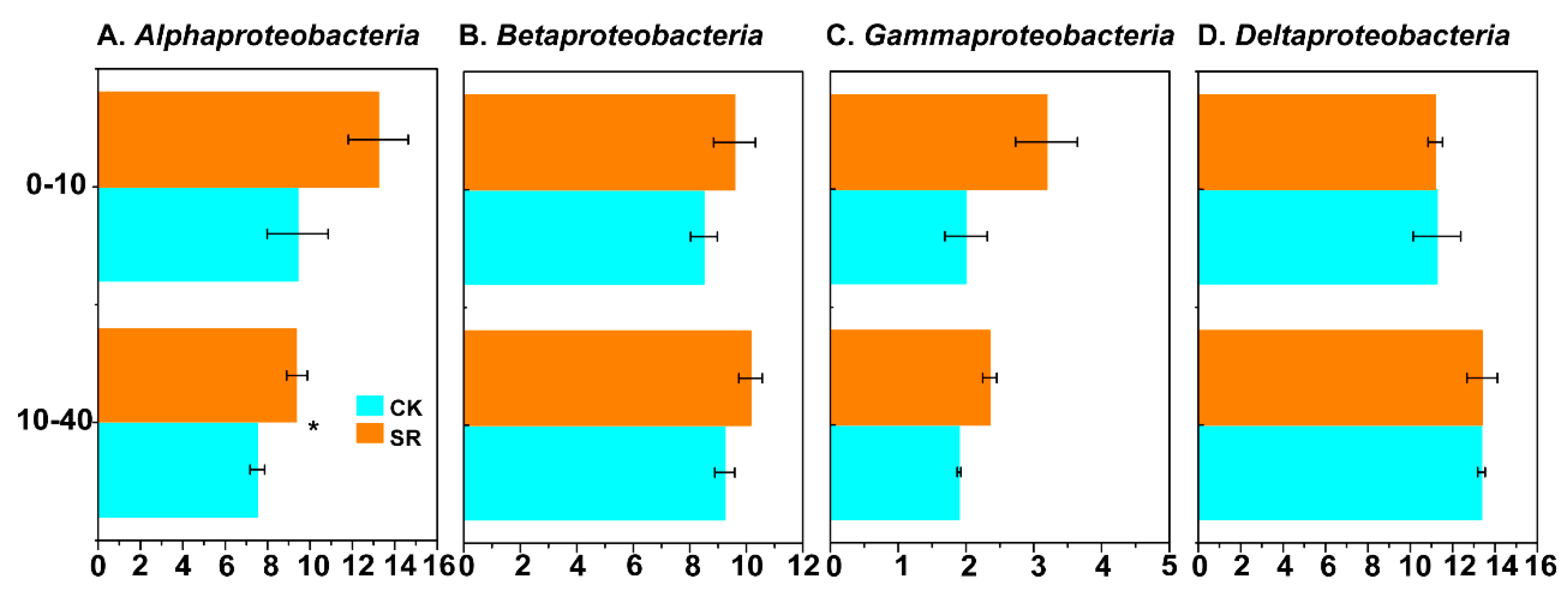
3.4. Relative Abundance of Dominant Phyla and Genera
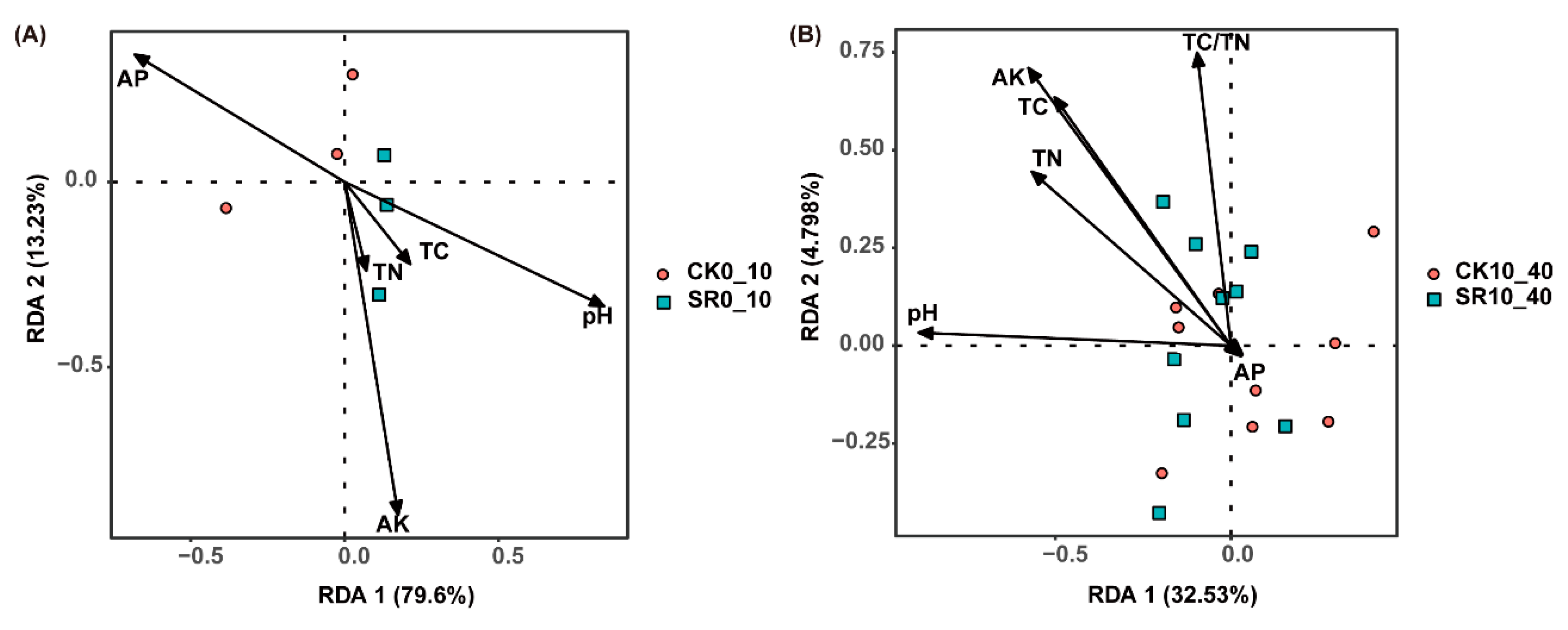
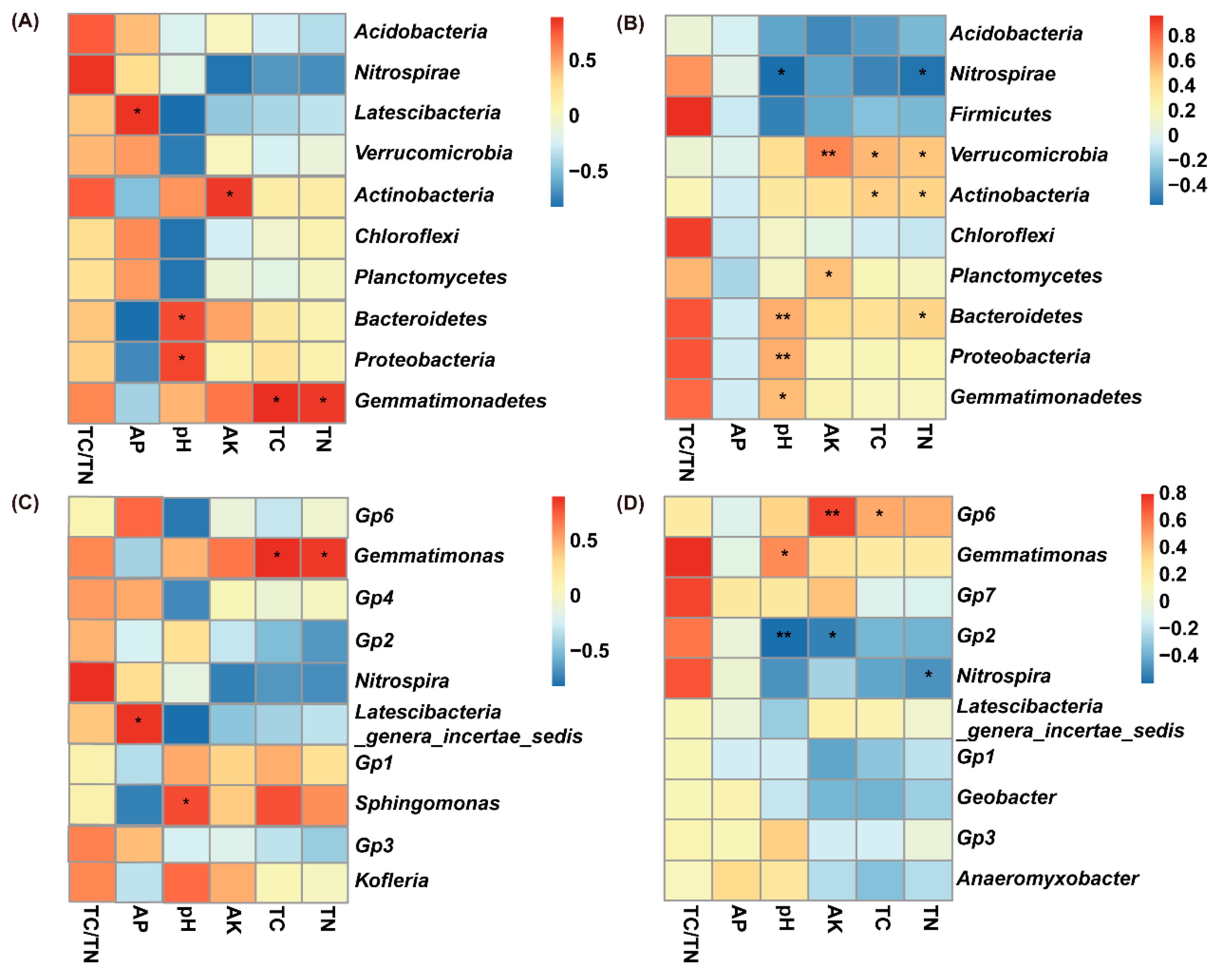
3.5. Pearson’s Correlation between Soil Attributes and Relative Abundance of Soil Bacterial Taxa
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lourenço, K.S.; Suleiman, A.K.A.; Pijl, A.; van Veen, J.A.; Cantarella, H.; Kuramae, E.E. Resilience of the resident soil microbiome to organic and inorganic amendment disturbances and to temporary bacterial invasion. Microbiome 2018, 6, 142. [Google Scholar] [CrossRef] [PubMed]
- Altieri, M.A. The ecological role of biodiversity in agroecosystems. Agric. Ecosyst. Environ. 1999. [Google Scholar] [CrossRef]
- Canfora, L.; Vendramin, E.; Felici, B.; Tarricone, L.; Florio, A.; Benedetti, A. Vineyard microbiome variations during different fertilisation practices revealed by 16s rRNA gene sequencing. Appl. Soil Ecol. 2018. [Google Scholar] [CrossRef]
- Will, C.; Thürmer, A.; Wollherr, A.; Nacke, H.; Herold, N.; Schrumpf, M.; Gutknecht, J.; Wubet, T.; Buscot, F.; Daniell, R. Horizon-specific bacterial community composition of german grassland soils, as revealed by pyrosequencing-based analysis of 16S rRNA genes. Appl. Environ. Microbiol. 2010, 76, 6751–6759. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Yan, K.; Tang, L.; Jia, Z.; Li, Y. Change in deep soil microbial communities due to long-term fertilization. Soil Biol. Biochem. 2014, 75, 264–272. [Google Scholar] [CrossRef]
- Gu, Y.; Wang, Y.; Lu, S.; Xiang, Q.; Yu, X.; Zhao, K.; Zou, L.; Chen, Q.; Tu, S.; Zhang, X. Long-term fertilization structures bacterial and archaeal communities along soil depth gradient in a paddy soil. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Tilman, D.; Balzer, C.; Hill, J.; Befort, B.L. Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. USA 2011. [Google Scholar] [CrossRef]
- Dari, B.; Sihi, D.; Bal, S.K.; Kunwar, S. Performance of direct-seeded rice under various dates of sowing and irrigation regimes in semi-arid region of India. Paddy Water Environ. 2017, 15, 395–401. [Google Scholar] [CrossRef]
- Chen, X.P.; Ju, X.T.; Yin, B.; Zhang, L.J.; Zhang, F.S.; Zhu, Z.L.; Xing, G.X.; Christie, P.; Cui, Z.L.; Zhang, S.L.; et al. Reducing environmental risk by improving N management in intensive Chinese agricultural systems. Proc. Natl. Acad. Sci. USA 2009, 106, 3041–3046. [Google Scholar] [CrossRef]
- Arafat, Y.; Tayyab, M.; Khan, M.U.; Chen, T.; Amjad, H.; Awais, S.; Lin, X.; Lin, W.; Lin, S. Long-Term Monoculture Negatively Regulates Fungal Community Composition and Abundance of Tea Orchards. Agronomy 2019, 9, 466. [Google Scholar] [CrossRef]
- Marschner, P.; Kandeler, E.; Marschner, B. Structure and function of the soil microbial community in a long-term fertilizer experiment. Soil Biol. Biochem. 2003. [Google Scholar] [CrossRef]
- Saha, S.; Gopinath, K.A.; Mina, B.L.; Gupta, H.S. Influence of continuous application of inorganic nutrients to a Maize-Wheat rotation on soil enzyme activity and grain quality in a rainfed Indian soil. Eur. J. Soil Biol. 2008, 44, 521–531. [Google Scholar] [CrossRef]
- Sihi, D.; Dari, B.; Sharma, D.K.; Pathak, H.; Nain, L.; Sharma, O.P. Evaluation of soil health in organic vs. conventional farming of basmati rice in North India. J. Plant Nutr. Soil Sci. 2017, 180, 389–406. [Google Scholar] [CrossRef]
- Seufert, V.; Ramankutty, N.; Foley, J.A. Comparing the yields of organic and conventional agriculture. Nature 2012, 485, 229–232. [Google Scholar] [CrossRef]
- Barak, P.; Jobe, B.O.; Krueger, A.R.; Peterson, L.A.; Laird, D.A. Effects of long-term soil acidification due to nitrogen fertilizer inputs in Wisconsin. Plant Soil 1997, 197, 61–69. [Google Scholar] [CrossRef]
- Nowinski, N.S.; Trumbore, S.E.; Schuur, E.A.G.; MacK, M.C.; Shaver, G.R. Nutrient addition prompts rapid destabilization of organic matter in an arctic tundra ecosystem. Ecosystems 2008, 11, 16–25. [Google Scholar] [CrossRef]
- Ramirez, K.S.; Lauber, C.L.; Knight, R.; Bradford, M.A.; Fierer, N. Consistent effects of nitrogen fertilization on soil bacterial communities in contrasting systems. Ecology 2010, 91, 3463–3470. [Google Scholar] [CrossRef]
- Kravchenko, A.N.; Snapp, S.S.; Robertson, G.P. Field-scale experiments reveal persistent yield gaps in low-input and organic cropping systems. Proc. Natl. Acad. Sci. USA 2017, 114, 926–931. [Google Scholar] [CrossRef]
- Dai, H.; Chen, Y.; Yang, X.; Cui, J.; Sui, P. The effect of different organic materials amendment on soil bacteria communities in barren sandy loam soil. Environ. Sci. Pollut. Res. 2017, 24, 24019–24028. [Google Scholar] [CrossRef]
- Das, S.S.; Jeong, S.T.; Das, S.S.; Kim, P.J. Composted cattle manure increases microbial activity and soil fertility more than composted swine manure in a submerged rice paddy. Front. Microbiol. 2017, 8, 1702. [Google Scholar] [CrossRef]
- Nair, V.D.; Nair, P.K.R.; Dari, B.; Freitas, A.M.; Chatterjee, N.; Pinheiro, F.M. Biochar in the agroecosystem-climate-change-sustainability Nexus. Front. Plant Sci. 2017. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Arafat, Y.; Letuma, P.; Ali, L.; Tayyab, M.; Waqas, M.; Li, Y.; Lin, W.; Lin, S.; Lin, W. Restoration of long-term monoculture degraded tea orchard by green and goat manures applications system. Sustainability 2019, 11, 1011. [Google Scholar] [CrossRef]
- Tayyab, M.; Caifang, Z.; Islam, W.; Khalil, F.; Ziqin, P.; Caifang, Z.; Arafat, Y.; Hui, L.; Rizwan, M.; Ahmad, K.; et al. Biochar: An efficient way to manage low water availability in plants. Appl. Ecol. Environ. Res. 2018, 16, 2565–2583. [Google Scholar] [CrossRef]
- Sarfraz, R.; Shakoor, A.; Abdullah, M.; Arooj, A.; Hussain, A.; Xing, S. Impact of integrated application of biochar and nitrogen fertilizers on maize growth and nitrogen recovery in alkaline calcareous soil. Soil Sci. Plant Nutr. 2017, 63, 488–498. [Google Scholar] [CrossRef]
- Liu, E.; Yan, C.; Mei, X.; He, W.; Bing, S.H.; Ding, L.; Liu, Q.; Liu, S.; Fan, T. Long-term effect of chemical fertilizer, straw, and manure on soil chemical and biological properties in northwest China. Geoderma 2010, 158, 173–180. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, S.; He, T.; Liu, L.; Wu, J. Response of organic carbon mineralization and microbial community to leaf litter and nutrient additions in subtropical forest soils. Soil Biol. Biochem. 2014, 71, 13–20. [Google Scholar] [CrossRef]
- Fanin, N.; Bertrand, I. Aboveground litter quality is a better predictor than belowground microbial communities when estimating carbon mineralization along a land-use gradient. Soil Biol. Biochem. 2016, 94, 48–60. [Google Scholar] [CrossRef]
- John, K.; Degtyarev, M.; Gorbunova, A.; Korobushkin, D.; Knöss, H.; Wolters, V.; Zaitsev, A.S. Enchytraeids simultaneously stimulate rice straw degradation and mitigate CO2 release in a paddy soil. Soil Biol. Biochem. 2019. [Google Scholar] [CrossRef]
- Tayyab, M.; Islam, W.; Lee, C.; Pang, Z.; Khalil, F.; Lin, S.; Lin, W.; Zhang, H. Short-Term Effects of Different Organic Amendments on Soil Fungal Composition. Sustainability 2019, 11, 198. [Google Scholar] [CrossRef]
- Guo, L.; Zheng, S.; Cao, C.; Li, C. Tillage practices and straw-returning methods affect topsoil bacterial community and organic C under a rice-wheat cropping system in central China. Sci. Rep. 2016, 6, 33155. [Google Scholar] [CrossRef]
- Tayyab, M.; Islam, W.; Arafat, Y.; Pang, Z.; Zhang, C.; Lin, Y.; Waqas, M.; Lin, S.; Lin, W.; Zhang, H. Effect of sugarcane straw and goat manure on soil nutrient transformation and bacterial communities. Sustainability 2018, 10, 2361. [Google Scholar] [CrossRef]
- Chen, Y.; Xin, L.; Liu, J.; Yuan, M.; Liu, S.; Jiang, W.; Chen, J.; Chen, Y.; Xin, L.; Liu, J.; et al. Changes in bacterial community of soil induced by long-term straw returning. Sci. Agric. 2017, 74, 349–356. [Google Scholar] [CrossRef]
- Yu, D.; Wen, Z.; Li, X.; Song, X.; Wu, H.; Yang, P. Effects of straw return on bacterial communities in a wheat-maize rotation system in the North China Plain. PLoS ONE 2018, 13, e0198087. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Butterly, C.R.; Chen, Q.; Mu, Z.; Wang, X.; Xi, Y.; Zhang, J.; Xiao, X. Surface Amendments Can Ameliorate Subsoil Acidity in Tea Garden Soils of High-Rainfall Environments. Pedosphere 2016. [Google Scholar] [CrossRef]
- Philippot, L.; Andersson, S.G.E.; Battin, T.J.; Prosser, J.I.; Schimel, J.P.; Whitman, W.B.; Hallin, S. The ecological coherence of high bacterial taxonomic ranks. Nat. Rev. Microbiol. 2010, 8, 523–529. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.; Lei, X.; Zhao, L.; Brookes, P.C.; Wang, F.; Chen, C.; Yang, W.; Xing, S. Fungal communities and functions response to long-term fertilization in paddy soils. Appl. Soil Ecol. 2018, 130, 251–258. [Google Scholar] [CrossRef]
- Hartman, W.H.; Richardson, C.J.; Vilgalys, R.; Bruland, G.L. Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc. Natl. Acad. Sci. USA 2008. [Google Scholar] [CrossRef]
- Faust, K.; Raes, J. Microbial interactions: From networks to models. Nat. Rev. Microbiol. 2012, 10, 538–550. [Google Scholar] [CrossRef]
- Nannipieri, P.; Giagnoni, L.; Renella, G.; Puglisi, E.; Ceccanti, B.; Masciandaro, G.; Fornasier, F.; Moscatelli, M.C.; Marinari, S. Soil enzymology: Classical and molecular approaches. Biol. Fertil. Soils 2012, 48, 743–762. [Google Scholar] [CrossRef]
- Akhtar, K.; Wang, W.; Ren, G.; Khan, A.; Feng, Y.; Yang, G. Changes in soil enzymes, soil properties, and maize crop productivity under wheat straw mulching in Guanzhong, China. Soil Tillage Res. 2018. [Google Scholar] [CrossRef]
- Zhao, S.; Li, K.; Zhou, W.; Qiu, S.; Huang, S.; He, P. Changes in soil microbial community, enzyme activities and organic matter fractions under long-term straw return in north-central China. Agric. Ecosyst. Environ. 2016, 216, 82–88. [Google Scholar] [CrossRef]
- Zhu, L.; Hu, N.; Yang, M.; Zhan, X.; Zhang, Z. Effects of different tillage and straw return on soil organic carbon in a rice-wheat rotation system. PLoS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; Yang, J.; Xue, Y.; Lv, W.; Zheng, X.; Pan, J. Influence of tillage practices and straw incorporation on soil aggregates, organic carbon, and crop yields in a rice-wheat rotation system. Sci. Rep. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bi, X.; Ren, L.; Gong, M.; He, Y.; Wang, L.; Ma, Z. Transfer of cadmium and lead from soil to mangoes in an uncontaminated area, Hainan Island, China. Geoderma 2010. [Google Scholar] [CrossRef]
- Watanabe, F.S.; Olsen, S.R. Test of an ascorbic acid method for determining phosphorus in water and NaHCO3 extracts from soil. Soil Sci. Soc. Am. J. 1965, 29, 677–678. [Google Scholar] [CrossRef]
- Pansu, M.; Gautheyrou, J. Handbook of Soil Analysis: Mineralogical, Organic and Inorganic Methods; Springer Science & Business Media: Berlin, Germany, 2007; ISBN 3540312110. [Google Scholar]
- Sun, J.; Zhang, Q.; Zhou, J.; Wei, Q. Pyrosequencing technology reveals the impact of different manure doses on the bacterial community in apple rhizosphere soil. Appl. Soil Ecol. 2014, 78, 28–36. [Google Scholar] [CrossRef]
- Jiang, J.; Song, Z.; Yang, X.; Mao, Z.; Nie, X.; Guo, H.; Peng, X. Microbial community analysis of apple rhizosphere around Bohai Gulf. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA\nsequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Chao, A. Non-parametric estimation of the number of classes in a population. Scand. J. Stat. 1984, 20, 265–270. [Google Scholar]
- Chao, A.; Lee, S.M. Estimating the number of classes via sample coverage. J. Am. Stat. Assoc. 1992, 87, 210–217. [Google Scholar] [CrossRef]
- Keylock, C.J. Simpson diversity and the Shannon-Wiener index as special cases of a generalized entropy. Oikos 2005, 109, 203–207. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Legendre, P.; Gallagher, E.D. Ecologically meaningful transformations for ordination of species data. Oecologia 2001. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; p. 2014. [Google Scholar]
- Ghimire, R.; Machado, S.; Bista, P. Soil pH, soil organic matter, and crop yields in winter wheat–summer fallow systems. Agron. J. 2017, 109, 706–717. [Google Scholar] [CrossRef]
- Cai, Z.; Xu, M.; Wang, B.; Zhang, L.; Wen, S.; Gao, S. Effectiveness of crop straws, and swine manure in ameliorating acidic red soils: A laboratory study. J. Soils Sediments 2018. [Google Scholar] [CrossRef]
- El-Baruni, B.; Olsen, S.R. Effect of manure on solubility of phosphorus in calcareous soils. Soil Sci. 1979, 128, 219–225. [Google Scholar] [CrossRef]
- Díaz-Zorita, M.; Grove, J.H. Duration of tillage management affects carbon and phosphorus stratification in phosphatic Paleudalfs. Soil Tillage Res. 2002. [Google Scholar] [CrossRef]
- Nannipieri, P.; Kandeler, E.; Ruggiero, P. Enzyme activities and microbiological and biochemical processes in soil. In Enzymes in the Environment: Activity, Ecology, and Applications; Marcel Dekker: New York, NY, USA, 2002; pp. 1–33. ISBN 9780824706142. [Google Scholar]
- Štursová, M.; Baldrian, P. Effects of soil properties and management on the activity of soil organic matter transforming enzymes and the quantification of soil-bound and free activity. Plant Soil 2011. [Google Scholar] [CrossRef]
- Li, F.; Chen, L.; Zhang, J.; Yin, J.; Huang, S. Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Fierer, N.; Bradford, M.A.; Jackson, R.B. Toward an ecological classification of soil bacteria. Ecology 2007, 88, 1354–1364. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Adhya, T.K. Effect of combine application of organic manure and inorganic fertilizer on methane and nitrous oxide emissions from a tropical flooded soil planted to rice. Geoderma 2014, 213, 185–192. [Google Scholar] [CrossRef]
- Lopes, A.R.; Manaia, C.M.; Nunes, O.C. Bacterial community variations in an alfalfa-rice rotation system revealed by 16S rRNA gene 454-pyrosequencing. FEMS Microbiol. Ecol. 2014, 87, 650–663. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Kloepper, J.W.; Huang, P.; McInroy, J.A.; Hu, C.H. Isolation and characterization of N2-fixing bacteria from giant reed and switchgrass for plant growth promotion and nutrient uptake. J. Basic Microbiol. 2018, 58, 459–471. [Google Scholar] [CrossRef] [PubMed]
- Mahadevan, R.; Palsson, B.; Lovley, D.R. In situ to in silico and back: Elucidating the physiology and ecology of Geobacter spp. using genome-scale modelling. Nat. Rev. Microbiol. 2011, 9, 39. [Google Scholar] [CrossRef] [PubMed]
- Sykes, G.; Skinner, F.A. Actinomycetales: Characteristics and Practical Importance; Academic Press: Cambridge, MA, USA, 1973; ISBN 0126799504. [Google Scholar]
- Hamamura, N.; Olson, S.H.; Ward, D.M.; Inskeep, W.P. Microbial population dynamics associated with crude-oil biodegradation in diverse soils. Appl. Environ. Microbiol. 2006. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, L.; França, L.; Rainey, F.A.; Schumann, P.; Nobre, M.F.; Da Costa, M.S. Gaiella occulta gen. nov., sp. nov., a novel representative of a deep branching phylogenetic lineage within the class Actinobacteria and proposal of Gaiellaceae fam. nov. and Gaiellales ord. nov. Syst. Appl. Microbiol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Freitag, T.E.; Chang, L.; Clegg, C.D.; Prosser, J.I. Influence of inorganic nitrogen management regime on the diversity of nitrite-oxidizing bacteria in agricultural grassland soils. Appl. Environ. Microbiol. 2005. [Google Scholar] [CrossRef]
- Xia, S.; Li, J.; Wang, R.; Li, J.; Zhang, Z. Tracking composition and dynamics of nitrification and denitrification microbial community in a biofilm reactor by PCR-DGGE and combining FISH with flow cytometry. Biochem. Eng. J. 2010. [Google Scholar] [CrossRef]
- Lucker, S.; Wagner, M.; Maixner, F.; Pelletier, E.; Koch, H.; Vacherie, B.; Rattei, T.; Damste, J.S.S.; Spieck, E.; Le Paslier, D.; et al. A Nitrospira metagenome illuminates the physiology and evolution of globally important nitrite-oxidizing bacteria. Proc. Natl. Acad. Sci. USA 2010. [Google Scholar] [CrossRef]
- Le Roux, X.; Bouskill, N.J.; Niboyet, A.; Barthes, L.; Dijkstra, P.; Field, C.B.; Hungate, B.A.; Lerondelle, C.; Pommier, T.; Tang, J.; et al. Predicting the responses of soil nitrite-oxidizers to multi-factorial global change: A trait-based approach. Front. Microbiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Nemergut, D.R.; Cleveland, C.C.; Wieder, W.R.; Washenberger, C.L.; Townsend, A.R. Plot-scale manipulations of organic matter inputs to soils correlate with shifts in microbial community composition in a lowland tropical rain forest. Soil Biol. Biochem. 2010. [Google Scholar] [CrossRef]
- Wei, C.; He, W.; Wei, L.; Ma, J.; Li, C. The performance and microbial communities of biodegradation- electron transfer with sulfur metabolism integrated process for flue gas desulfurization wastewater treatment. Bioprocess Biosyst. Eng. 2017. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sui, Y.; Yu, Z.; Shi, Y.; Chu, H.; Jin, J.; Liu, X.; Wang, G. High throughput sequencing analysis of biogeographical distribution of bacterial communities in the black soils of northeast China. Soil Biol. Biochem. 2014, 70, 113–122. [Google Scholar] [CrossRef]
- Wei, Z.; Hu, X.; Li, X.; Zhang, Y.; Jiang, L.; Li, J.; Guan, Z.; Cai, Y.; Liao, X. The rhizospheric microbial community structure and diversity of deciduous and evergreen forests in Taihu Lake area, China. PLoS ONE 2017, 12, e0174411. [Google Scholar] [CrossRef]
- Shen, Z.; Ruan, Y.; Xue, C.; Zhong, S.; Li, R.; Shen, Q. Soils naturally suppressive to banana Fusarium wilt disease harbor unique bacterial communities. Plant Soil 2015, 393, 21–33. [Google Scholar] [CrossRef]
- Hou, S.; Makarova, K.S.; Saw, J.H.W.; Senin, P.; Ly, B.V.; Zhou, Z.; Ren, Y.; Wang, J.; Galperin, M.Y.; Omelchenko, M.V.; et al. Complete genome sequence of the extremely acidophilic methanotroph isolate V4, Methylacidiphilum infernorum, a representative of the bacterial phylum Verrucomicrobia. Biol. Direct 2008. [Google Scholar] [CrossRef]
- Schlesner, H.; Jenkins, C.; Staley, J.T. The Phylum Verrucomicrobia: A Phylogenetically Heterogeneous Bacterial Group. In The Prokaryotes; Springer: New York, NY, USA, 2006. [Google Scholar]
- Chin, K.J.; Liesack, W.; Janssen, P.H. Opitutus terrae gen. nov., sp. nov., to accommodate novel strains of the division “Verrucomicrobia” isolated from rice paddy soil. Int. J. Syst. Evol. Microbiol. 2001, 51, 1965–1968. [Google Scholar] [CrossRef]
- Hengstmann, U.; Chin, K.J.; Janssen, P.H.; Liesack, W. Comparative phylogenetic assignment of environmental sequences of genes encoding 16S rRNA and numerically abundant culturable bacteria from an anoxic rice paddy soil. Appl. Environ. Microbiol. 1999, 65, 5050–5058. [Google Scholar]
- Carbonetto, B.; Rascovan, N.; Álvarez, R.; Mentaberry, A.; Vázquez, M.P. Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS ONE 2014. [Google Scholar] [CrossRef]
- Xu, H.J.; Wang, X.H.; Li, H.; Yao, H.Y.; Su, J.Q.; Zhu, Y.G. Biochar impacts soil microbial community composition and nitrogen cycling in an acidic soil planted with rape. Environ. Sci. Technol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Su, J.Q.; Ding, L.J.; Xue, K.; Yao, H.Y.; Quensen, J.; Bai, S.J.; Wei, W.X.; Wu, J.S.; Zhou, J.; Tiedje, J.M.; et al. Long-term balanced fertilization increases the soil microbial functional diversity in a phosphorus-limited paddy soil. Mol. Ecol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Bastian, F.; Bouziri, L.; Nicolardot, B.; Ranjard, L. Impact of wheat straw decomposition on successional patterns of soil microbial community structure. Soil Biol. Biochem. 2009. [Google Scholar] [CrossRef]
- Arafat, Y.; Wei, X.; Jiang, Y.; Chen, T.; Saqib, H.S.A.; Lin, S.; Lin, W. Spatial distribution patterns of root-associated bacterial communities mediated by root exudates in different aged ratooning tea monoculture systems. Int. J. Mol. Sci. 2017, 18, 1727. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, C.; Tayyab, M.; Abubakar, A.Y.; Yang, Z.; Pang, Z.; Islam, W.; Lin, Z.; Li, S.; Luo, J.; Fan, X.; et al. Bacteria with Different Assemblages in the Soil Profile Drive the Diverse Nutrient Cycles in the Sugarcane Straw Retention Ecosystem. Diversity 2019, 11, 194. https://doi.org/10.3390/d11100194
Zhang C, Tayyab M, Abubakar AY, Yang Z, Pang Z, Islam W, Lin Z, Li S, Luo J, Fan X, et al. Bacteria with Different Assemblages in the Soil Profile Drive the Diverse Nutrient Cycles in the Sugarcane Straw Retention Ecosystem. Diversity. 2019; 11(10):194. https://doi.org/10.3390/d11100194
Chicago/Turabian StyleZhang, Caifang, Muhammad Tayyab, Ahmad Yusuf Abubakar, Ziqi Yang, Ziqin Pang, Waqar Islam, Zhaoli Lin, Shiyan Li, Jun Luo, Xiaoliang Fan, and et al. 2019. "Bacteria with Different Assemblages in the Soil Profile Drive the Diverse Nutrient Cycles in the Sugarcane Straw Retention Ecosystem" Diversity 11, no. 10: 194. https://doi.org/10.3390/d11100194
APA StyleZhang, C., Tayyab, M., Abubakar, A. Y., Yang, Z., Pang, Z., Islam, W., Lin, Z., Li, S., Luo, J., Fan, X., Fallah, N., & Zhang, H. (2019). Bacteria with Different Assemblages in the Soil Profile Drive the Diverse Nutrient Cycles in the Sugarcane Straw Retention Ecosystem. Diversity, 11(10), 194. https://doi.org/10.3390/d11100194




