MicroRNA: an Emerging Therapeutic Target and Intervention Tool
Abstract
1. Introduction
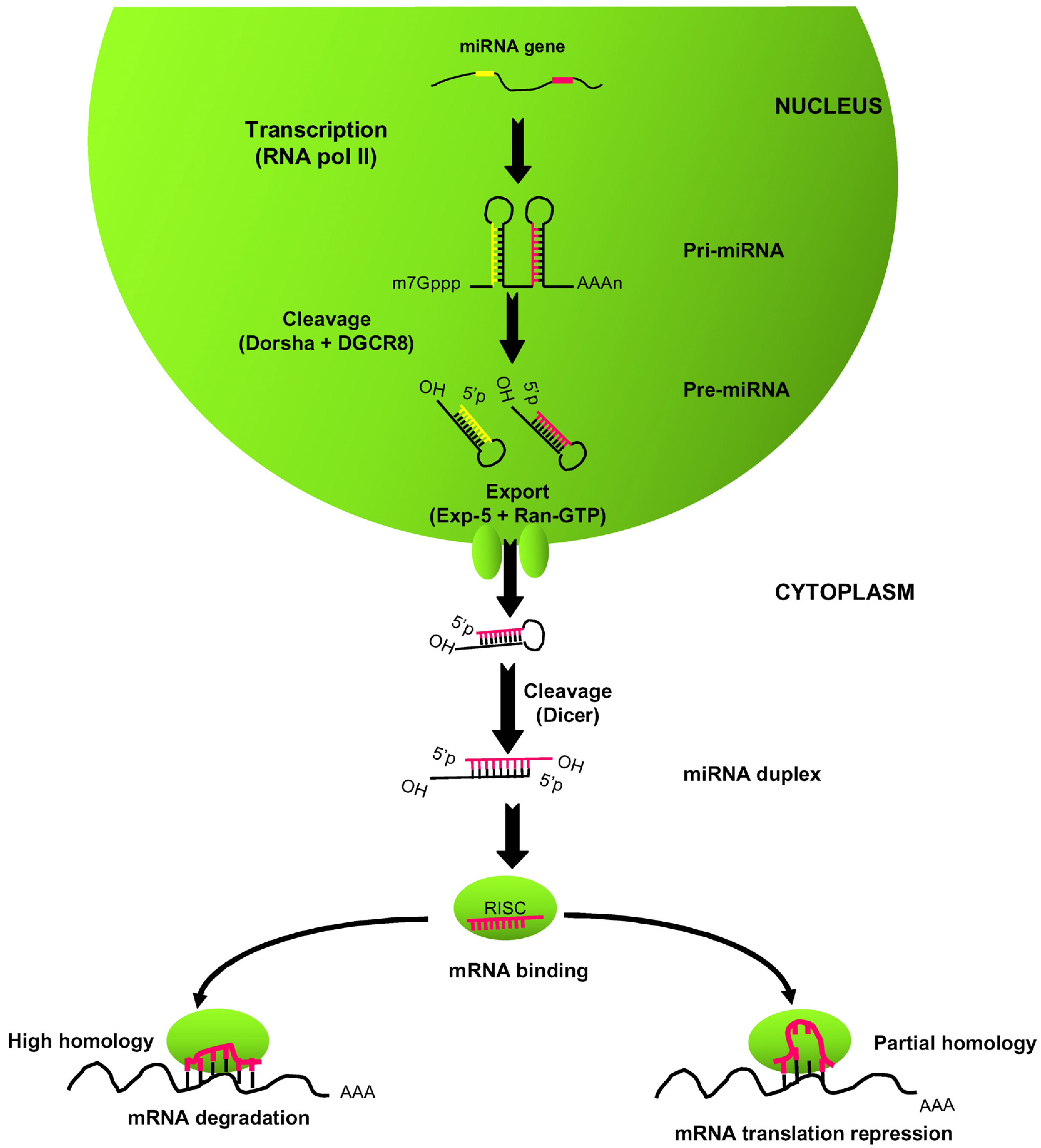
2. miRNA Biogenesis and Mechanisms of Action
3. Association of miRNA with Diseases
4. miRNA as Therapeutic Target and Tool
4.1 Antisense inhibition of mature miRNA
4.2 Other strategies for miRNA inhibition
4.3 Replacement of miRNAs
4.4 miRNA scaffolds
4.5 Artificial miRNAs
5. Future Prospect
Acknowledgment
References
- Rodriguez, A; Griffiths-Jones, S; Ashurst, JL; Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 2004, 14(10A), 1902–1910. [Google Scholar]
- Pfeffer, S; Sewer, A; Lagos-Quintana, M; Sheridan, R; Sander, C; Grasser, FA; van Dyk, LF; Ho, CK; Shuman, S; Chien, M; Russo, JJ; Ju, J; Randall, G; Lindenbach, BD; Rice, CM; Simon, V; Ho, DD; Zavolan, M; Tuschl, T. Identification of microRNAs of the herpesvirus family. Nat. Methods 2005, 2(4), 269–276. [Google Scholar]
- Pillai, RS; Bhattacharyya, SN; Filipowicz, W. Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell. Biol. 2007, 17(3), 118–126. [Google Scholar]
- Petersen, CP; Bordeleau, ME; Pelletier, J; Sharp, PA. Short RNAs repress translation after initiation in mammalian cells. Mol. Cell. 2006, 21(4), 533–542. [Google Scholar]
- Farh, KK; Grimson, A; Jan, C; Lewis, BP; Johnston, WK; Lim, LP; Burge, CB; Bartel, DP. The widespread impact of mammalian MicroRNAs on mRNA repression and evolution. Science 2005, 310(5755), 1817–1821. [Google Scholar]
- Borchert, GM; Lanier, W; Davidson, BL. RNA polymerase III transcribes human microRNAs. Nat. Struct. Mol. Biol. 2006, 13(12), 1097–1101. [Google Scholar]
- Andersson, MG; Haasnoot, PC; Xu, N; Berenjian, S; Berkhout, B; Akusjarvi, G. Suppression of RNA interference by adenovirus virus-associated RNA. J. Virol. 2005, 79(15), 9556–9565. [Google Scholar]
- Samols, MA; Hu, J; Skalsky, RL; Renne, R. Cloning and identification of a microRNA cluster within the latency-associated region of Kaposi’s sarcoma-associated herpesvirus. J. Virol. 2005, 79(14), 9301–9305. [Google Scholar]
- Cullen, BR. Transcription and processing of human microRNA precursors. Mol. Cell 2004, 16(6), 861–865. [Google Scholar]
- Cai, X; Hagedorn, CH; Cullen, BR. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004, 10(12), 1957–1966. [Google Scholar]
- Lee, Y; Ahn, C; Han, J; Choi, H; Kim, J; Yim, J; Lee, J; Provost, P; Radmark, O; Kim, S; Kim, VN. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425(6956), 415–419. [Google Scholar]
- Kim, VN. MicroRNA biogenesis: coordinated cropping and dicing. Nat. Rev. Mol. Cell. Biol. 2005, 6(5), 376–385. [Google Scholar]
- Yi, R; Qin, Y; Macara, IG; Cullen, BR. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes. Dev. 2003, 17(24), 3011–3016. [Google Scholar]
- Bohnsack, MT; Czaplinski, K; Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 2004, 10(2), 185–191. [Google Scholar]
- Basyuk, E; Suavet, F; Doglio, A; Bordonne, R; Bertrand, E. Human let-7 stem-loop precursors harbor features of RNase III cleavage products. Nucl. Acids Res. 2003, 31(22), 6593–6597. [Google Scholar]
- Zamore, PD. Thirty-three years later, a glimpse at the ribonuclease III active site. Mol. Cell. 2001, 8(6), 1158–1160. [Google Scholar]
- Chendrimada, TP; Gregory, RI; Kumaraswamy, E; Norman, J; Cooch, N; Nishikura, K; Shiekhattar, R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005, 436(7051), 740–744. [Google Scholar]
- Lee, Y; Hur, I; Park, SY; Kim, YK; Suh, MR; Kim, VN. The role of PACT in the RNA silencing pathway. EMBO J. 2006, 25(3), 522–532. [Google Scholar]
- Maniataki, E; Mourelatos, Z. A human, ATP-independent, RISC assembly machine fueled by pre-miRNA. Genes. Dev. 2005, 19(24), 2979–2990. [Google Scholar]
- Hammond, SM; Bernstein, E; Beach, D; Hannon, GJ. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 2000, 404(6775), 293–296. [Google Scholar]
- Meister, G; Landthaler, M; Patkaniowska, A; Dorsett, Y; Teng, G; Tuschl, T. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell. 2004, 15(2), 185–197. [Google Scholar]
- Zeng, Y; Yi, R; Cullen, BR. MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc. Natl. Acad. Sci. USA 2003, 100(17), 9779–9784. [Google Scholar]
- Hutvagner, G; Zamore, PD. A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002, 297(5589), 2056–2060. [Google Scholar]
- Yekta, S; Shih, IH; Bartel, DP. MicroRNA-directed cleavage of HOXB8 mRNA. Science 2004, 304(5670), 594–596. [Google Scholar]
- Olsen, PH; Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 1999, 216(2), 671–680. [Google Scholar]
- Lewis, BP; Burge, CB; Bartel, DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005, 120(1), 15–20. [Google Scholar]
- Brennecke, J; Stark, A; Russell, RB; Cohen, SM. Principles of microRNA-target recognition. PLoS Biol. 2005, 3(3), e85. [Google Scholar]
- Lytle, JR; Yario, TA; Steitz, JA. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl. Acad. Sci. USA 2007, 104(23), 9667–9672. [Google Scholar]
- Care, A; Catalucci, D; Felicetti, F; Bonci, D; Addario, A; Gallo, P; Bang, ML; Segnalini, P; Gu, Y; Dalton, ND; Elia, L; Latronico, MV; Hoydal, M; Autore, C; Russo, MA; Dorn, GW, 2nd; Ellingsen, O; Ruiz-Lozano, P; Peterson, KL; Croce, CM; Peschle, C; Condorelli, G. MicroRNA-133 controls cardiac hypertrophy. Nat. Med. 2007, 13(5), 613–618. [Google Scholar]
- Wiemer, EA. The role of microRNAs in cancer: no small matter. Eur. J. Cancer. 2007, 43(10), 1529–1544. [Google Scholar]
- Sullivan, CS; Grundhoff, AT; Tevethia, S; Pipas, JM; Ganem, D. SV40-encoded microRNAs regulate viral gene expression and reduce susceptibility to cytotoxic T cells. Nature 2005, 435(7042), 682–686. [Google Scholar]
- Krutzfeldt, J; Stoffel, M. MicroRNAs: a new class of regulatory genes affecting metabolism. Cell. Metab. 2006, 4(1), 9–12. [Google Scholar]
- Nelson, PT; Keller, JN. RNA in brain disease: no longer just “the messenger in the middle”. J. Neuropathol. Exp. Neurol. 2007, 66(6), 461–468. [Google Scholar]
- Calin, GA; Dumitru, CD; Shimizu, M; Bichi, R; Zupo, S; Noch, E; Aldler, H; Rattan, S; Keating, M; Rai, K; Rassenti, L; Kipps, T; Negrini, M; Bullrich, F; Croce, CM. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99(24), 15524–15529. [Google Scholar]
- Calin, GA; Liu, CG; Sevignani, C; Ferracin, M; Felli, N; Dumitru, CD; Shimizu, M; Cimmino, A; Zupo, S; Dono, M; Dell’Aquila, ML; Alder, H; Rassenti, L; Kipps, TJ; Bullrich, F; Negrini, M; Croce, CM. MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc. Natl. Acad. Sci. USA 2004, 101(32), 11755–11760. [Google Scholar]
- He, H; Jazdzewski, K; Li, W; Liyanarachchi, S; Nagy, R; Volinia, S; Calin, GA; Liu, CG; Franssila, K; Suster, S; Kloos, RT; Croce, CM; de la Chapelle, A. The role of microRNA genes in papillary thyroid carcinoma. Proc. Natl. Acad. Sci. USA 2005, 102(52), 19075–19080. [Google Scholar]
- Zhang, L; Huang, J; Yang, N; Greshock, J; Megraw, MS; Giannakakis, A; Liang, S; Naylor, TL; Barchetti, A; Ward, MR; Yao, G; Medina, A; O’Brien-Jenkins, A; Katsaros, D; Hatzigeorgiou, A; Gimotty, PA; Weber, BL; Coukos, G. microRNAs exhibit high frequency genomic alterations in human cancer. Proc. Natl. Acad. Sci. USA 2006, 103(24), 9136–9141. [Google Scholar]
- He, L; Thomson, JM; Hemann, MT; Hernando-Monge, E; Mu, D; Goodson, S; Powers, S; Cordon-Cardo, C; Lowe, SW; Hannon, GJ; Hammond, SM. A microRNA polycistron as a potential human oncogene. Nature 2005, 435(7043), 828–833. [Google Scholar]
- Venturini, L; Battmer, K; Castoldi, M; Schultheis, B; Hochhaus, A; Muckenthaler, MU; Ganser, A; Eder, M; Scherr, M. Expression of the miR-17-92 polycistron in chronic myeloid leukemia (CML) CD34+ cells. Blood 2007, 109(10), 4399–4405. [Google Scholar]
- O’Brien, SJ; Nash, WG; Goodwin, JL; Lowy, DR; Chang, EH. Dispersion of the ras family of transforming genes to four different chromosomes in man. Nature 1983, 302(5911), 839–842. [Google Scholar]
- Brown, BD; Venneri, MA; Zingale, A; Sergi Sergi, L; Naldini, L. Endogenous microRNA regulation suppresses transgene expression in hematopoietic lineages and enables stable gene transfer. Nat. Med. 2006, 12(5), 585–591. [Google Scholar]
- Hayashita, Y; Osada, H; Tatematsu, Y; Yamada, H; Yanagisawa, K; Tomida, S; Yatabe, Y; Kawahara, K; Sekido, Y; Takahashi, T. A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res. 2005, 65(21), 9628–9632. [Google Scholar]
- Cimmino, A; Calin, GA; Fabbri, M; Iorio, MV; Ferracin, M; Shimizu, M; Wojcik, SE; Aqeilan, RI; Zupo, S; Dono, M; Rassenti, L; Alder, H; Volinia, S; Liu, CG; Kipps, TJ; Negrini, M; Croce, CM. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 2005, 102(39), 13944–13949. [Google Scholar]
- Eis, PS; Tam, W; Sun, L; Chadburn, A; Li, Z; Gomez, MF; Lund, E; Dahlberg, JE. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc. Natl. Acad. Sci. USA 2005, 102(10), 3627–3632. [Google Scholar]
- Calin, GA; Ferracin, M; Cimmino, A; Di Leva, G; Shimizu, M; Wojcik, SE; Iorio, MV; Visone, R; Sever, NI; Fabbri, M; Iuliano, R; Palumbo, T; Pichiorri, F; Roldo, C; Garzon, R; Sevignani, C; Rassenti, L; Alder, H; Volinia, S; Liu, CG; Kipps, TJ; Negrini, M; Croce, CM. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 2005, 353(17), 1793–1801. [Google Scholar]
- Roldo, C; Missiaglia, E; Hagan, JP; Falconi, M; Capelli, P; Bersani, S; Calin, GA; Volinia, S; Liu, CG; Scarpa, A; Croce, CM. MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J. Clin. Oncol. 2006, 24(29), 4677–4684. [Google Scholar]
- Yanaihara, N; Caplen, N; Bowman, E; Seike, M; Kumamoto, K; Yi, M; Stephens, RM; Okamoto, A; Yokota, J; Tanaka, T; Calin, GA; Liu, CG; Croce, CM; Harris, CC. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006, 9(3), 189–198. [Google Scholar]
- Cummins, JM; Velculescu, VE. Implications of micro-RNA profiling for cancer diagnosis. Oncogene. 2006, 25(46), 6220–6227. [Google Scholar]
- Iorio, MV; Ferracin, M; Liu, CG; Veronese, A; Spizzo, R; Sabbioni, S; Magri, E; Pedriali, M; Fabbri, M; Campiglio, M; Menard, S; Palazzo, JP; Rosenberg, A; Musiani, P; Volinia, S; Nenci, I; Calin, GA; Querzoli, P; Negrini, M; Croce, CM. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005, 65(16), 7065–7070. [Google Scholar]
- Murakami, Y; Yasuda, T; Saigo, K; Urashima, T; Toyoda, H; Okanoue, T; Shimotohno, K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006, 25(17), 2537–2545. [Google Scholar]
- Kumar, MS; Lu, J; Mercer, KL; Golub, TR; Jacks, T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat. Genet. 2007, 39(5), 673–677. [Google Scholar]
- Zhang, B; Farwell, MA. microRNAs: a new emerging class of players for disease diagnostics and gene therapy. J. Cell. Mol. Med. 2008, 12(1), 3–21. [Google Scholar]
- Zhang, B; Pan, X; Cobb, GP; Anderson, TA. microRNAs as oncogenes and tumor suppressors. Dev. Biol. 2007, 302(1), 1–12. [Google Scholar]
- Ikeda, S; Kong, SW; Lu, J; Bisping, E; Zhang, H; Allen, PD; Golub, TR; Pieske, B; Pu, WT. Altered microRNA expression in human heart disease. Physiol. Genomics 2007, 31(3), 367–373. [Google Scholar]
- Thum, T; Galuppo, P; Wolf, C; Fiedler, J; Kneitz, S; van Laake, LW; Doevendans, PA; Mummery, CL; Borlak, J; Haverich, A; Gross, C; Engelhardt, S; Ertl, G; Bauersachs, J. MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure. Circulation 2007, 116(3), 258–267. [Google Scholar]
- Sayed, D; Hong, C; Chen, IY; Lypowy, J; Abdellatif, M. MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ. Res. 2007, 100(3), 416–424. [Google Scholar]
- McCarthy, JJ; Esser, KA. MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J. Appl. Physiol. 2007, 102(1), 306–313. [Google Scholar]
- Lecellier, CH; Dunoyer, P; Arar, K; Lehmann-Che, J; Eyquem, S; Himber, C; Saib, A; Voinnet, O. A cellular microRNA mediates antiviral defense in human cells. Science 2005, 308(5721), 557–560. [Google Scholar]
- Jopling, CL; Yi, M; Lancaster, AM; Lemon, SM; Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science 2005, 309(5740), 1577–81. [Google Scholar]
- Vidal, L; Blagden, S; Attard, G; de Bono, J. Making sense of antisense. Eur. J. Cancer. 2005, 41(18), 2812–2818. [Google Scholar]
- Jackson, AL; Burchard, J; Schelter, J; Chau, BN; Cleary, M; Lim, L; Linsley, PS. Widespread siRNA “off-target” transcript silencing mediated by seed region sequence complementarity. RNA 2006, 12(7), 1179–1187. [Google Scholar]
- Birmingham, A; Anderson, EM; Reynolds, A; Ilsley-Tyree, D; Leake, D; Fedorov, Y; Baskerville, S; Maksimova, E; Robinson, K; Karpilow, J; Marshall, WS; Khvorova, A. 3′ UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nat. Methods 2006, 3(3), 199–204. [Google Scholar]
- Fedorov, Y; Anderson, EM; Birmingham, A; Reynolds, A; Karpilow, J; Robinson, K; Leake, D; Marshall, WS; Khvorova, A. Off-target effects by siRNA can induce toxic phenotype. RNA 2006, 12(7), 1188–1196. [Google Scholar]
- Grimm, D; Streetz, KL; Jopling, CL; Storm, TA; Pandey, K; Davis, CR; Marion, P; Salazar, F; Kay, MA. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 2006, 441(7092), 537–541. [Google Scholar]
- Soifer, HS; Rossi, JJ; Saetrom, P. MicroRNAs in disease and potential therapeutic applications. Mol. Ther. 2007, 15(12), 2070–1079. [Google Scholar]
- Wurdinger, T; Costa, FF. Molecular therapy in the microRNA era. Pharmacogenomics J. 2007, 7(5), 297–304. [Google Scholar]
- Davis, S; Lollo, B; Freier, S; Esau, C. Improved targeting of miRNA with antisense oligonucleotides. Nucl. Acids Res. 2006, 34(8), 2294–2304. [Google Scholar]
- Hutvagner, G; Simard, MJ; Mello, CC; Zamore, PD. Sequence-specific inhibition of small RNA function. PLoS Biol. 2004, 2(4), E98. [Google Scholar]
- Meister, G; Landthaler, M; Dorsett, Y; Tuschl, T. Sequence-specific inhibition of microRNA- and siRNA-induced RNA silencing. RNA 2004, 10(3), 544–550. [Google Scholar]
- Boutla, A; Delidakis, C; Tabler, M. Developmental defects by antisense-mediated inactivation of micro-RNAs 2 and 13 in Drosophila and the identification of putative target genes. Nucl. Acids Res. 2003, 31(17), 4973–4980. [Google Scholar]
- Orom, UA; Kauppinen, S; Lund, AH. LNA-modified oligonucleotides mediate specific inhibition of microRNA function. Gene. 2006, 372, 137–141. [Google Scholar]
- Leaman, D; Chen, PY; Fak, J; Yalcin, A; Pearce, M; Unnerstall, U; Marks, DS; Sander, C; Tuschl, T; Gaul, U. Antisense-mediated depletion reveals essential and specific functions of microRNAs in Drosophila development. Cell 2005, 121(7), 1097–1108. [Google Scholar]
- Esau, C; Davis, S; Murray, SF; Yu, XX; Pandey, SK; Pear, M; Watts, L; Booten, SL; Graham, M; McKay, R; Subramaniam, A; Propp, S; Lollo, BA; Freier, S; Bennett, CF; Bhanot, S; Monia, BP. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell. Metab. 2006, 3(2), 87–98. [Google Scholar]
- Krutzfeldt, J; Kuwajima, S; Braich, R; Rajeev, KG; Pena, J; Tuschl, T; Manoharan, M; Stoffel, M. Specificity, duplex degradation and subcellular localization of antagomirs. Nucl. Acids Res. 2007, 35(9), 2885–2892. [Google Scholar]
- Yang, M; Mattes, J. Discovery, biology and therapeutic potential of RNA interference, microRNA and antagomirs. Pharmacol. Ther. 2008, 117(1), 94–104. [Google Scholar]
- Bijsterbosch, MK; Rump, ET; De Vrueh, RL; Dorland, R; van Veghel, R; Tivel, KL; Biessen, EA; van Berkel, TJ; Manoharan, M. Modulation of plasma protein binding and in vivo liver cell uptake of phosphorothioate oligodeoxynucleotides by cholesterol conjugation. Nucl. Acids Res. 2000, 28(14), 2717–2725. [Google Scholar]
- Metzler, M; Wilda, M; Busch, K; Viehmann, S; Borkhardt, A. High expression of precursor microRNA-155/BIC RNA in children with Burkitt lymphoma. Genes Chromosomes Cancer 2004, 39(2), 167–169. [Google Scholar]
- Prakash, TP; Allerson, CR; Dande, P; Vickers, TA; Sioufi, N; Jarres, R; Baker, BF; Swayze, EE; Griffey, RH; Bhat, B. Positional effect of chemical modifications on short interference RNA activity in mammalian cells. J. Med. Chem. 2005, 48(13), 4247–4253. [Google Scholar]
- Chang, J; Nicolas, E; Marks, D; Sander, C; Lerro, A; Buendia, MA; Xu, C; Mason, WS; Moloshok, T; Bort, R; Zaret, KS; Taylor, JM. miR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol. 2004, 1(2), 106–113. [Google Scholar]
- Esau, CC; Monia, BP. Therapeutic potential for microRNAs. Adv. Drug Deliv. Rev. 2007, 59(2–3), 101–114. [Google Scholar]
- Krutzfeldt, J; Rajewsky, N; Braich, R; Rajeev, KG; Tuschl, T; Manoharan, M; Stoffel, M. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005, 438(7068), 685–689. [Google Scholar]
- Elmen, J; Lindow, M; Silahtaroglu, A; Bak, M; Christensen, M; Lind-Thomsen, A; Hedtjarn, M; Hansen, JB; Hansen, HF; Straarup, EM; McCullagh, K; Kearney, P; Kauppinen, S. Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nuc. Acids Res. 2008, 36(4), 1153–1162. [Google Scholar]
- Vermeulen, A; Robertson, B; Dalby, AB; Marshall, WS; Karpilow, J; Leake, D; Khvorova, A; Baskerville, S. Double-stranded regions are essential design components of potent inhibitors of RISC function. RNA 2007, 13(5), 723–730. [Google Scholar]
- Zellweger, T; Miyake, H; Cooper, S; Chi, K; Conklin, BS; Monia, BP; Gleave, ME. Antitumor activity of antisense clusterin oligonucleotides is improved in vitro and in vivo by incorporation of 2′-O-(2-methoxy)ethyl chemistry. J. Pharmacol. Exp. Ther. 2001, 298(3), 934–940. [Google Scholar]
- McKay, RA; Miraglia, LJ; Cummins, LL; Owens, SR; Sasmor, H; Dean, NM. Characterization of a potent and specific class of antisense oligonucleotide inhibitor of human protein kinase C-alpha expression. J. Biol. Chem. 1999, 274(3), 1715–1722. [Google Scholar]
- Grunweller, A; Hartmann, RK. Locked nucleic acid oligonucleotides: the next generation of antisense agents? BioDrugs 2007, 21(4), 235–243. [Google Scholar]
- Dykxhoorn, DM; Lieberman, J. Running interference: prospects and obstacles to using small interfering RNAs as small molecule drugs. Annu. Rev. Biomed. Eng. 2006, 8, 377–402. [Google Scholar]
- Soutschek, J; Akinc, A; Bramlage, B; Charisse, K; Constien, R; Donoghue, M; Elbashir, S; Geick, A; Hadwiger, P; Harborth, J; John, M; Kesavan, V; Lavine, G; Pandey, RK; Racie, T; Rajeev, KG; Rohl, I; Toudjarska, I; Wang, G; Wuschko, S; Bumcrot, D; Koteliansky, V; Limmer, S; Manoharan, M; Vornlocher, HP. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 2004, 432(7014), 173–178. [Google Scholar]
- Zimmermann, TS; Lee, AC; Akinc, A; Bramlage, B; Bumcrot, D; Fedoruk, MN; Harborth, J; Heyes, JA; Jeffs, LB; John, M; Judge, AD; Lam, K; McClintock, K; Nechev, LV; Palmer, LR; Racie, T; Rohl, I; Seiffert, S; Shanmugam, S; Sood, V; Soutschek, J; Toudjarska, I; Wheat, AJ; Yaworski, E; Zedalis, W; Koteliansky, V; Manoharan, M; Vornlocher, HP; MacLachlan, I. RNAi-mediated gene silencing in non-human primates. Nature 2006, 441(7089), 111–114. [Google Scholar]
- Guo, S; Tschammer, N; Mohammed, S; Guo, P. Specific delivery of therapeutic RNAs to cancer cells via the dimerization mechanism of phi29 motor pRNA. Hum. Gene. Ther. 2005, 16(9), 1097–1109. [Google Scholar]
- Muratovska, A; Eccles, MR. Conjugate for efficient delivery of short interfering RNA (siRNA) into mammalian cells. FEBS Lett. 2004, 558(1–3), 63–68. [Google Scholar]
- Duan, W; Chan, JH; McKay, K; Crosby, JR; Choo, HH; Leung, BP; Karras, JG; Wong, WS. Inhaled p38alpha mitogen-activated protein kinase antisense oligonucleotide attenuates asthma in mice. Am. J. Respir. Crit. Care Med. 2005, 171(6), 571–578. [Google Scholar]
- Butler, M; Hayes, CS; Chappell, A; Murray, SF; Yaksh, TL; Hua, XY. Spinal distribution and metabolism of 2′-O-(2-methoxyethyl)-modified oligonucleotides after intrathecal administration in rats. Neuroscience 2005, 131(3), 705–715. [Google Scholar]
- Danis, R; Criswell, M; Orge, F; Wancewicz, E; Stecker, K; Henry, S; Monia, B. Intravitreous anti-raf-1 kinase antisense oligonucleotide as an angioinhibitory agent in porcine preretinal neovascularization. Curr. Eye Res. 2003, 26(1), 45–54. [Google Scholar]
- Lund, E; Guttinger, S; Calado, A; Dahlberg, JE; Kutay, U. Nuclear export of microRNA precursors. Science 2004, 303(5654), 95–98. [Google Scholar]
- Aagaard, L; Amarzguioui, M; Sun, G; Santos, LC; Ehsani, A; Prydz, H; Rossi, JJ. A facile lentiviral vector system for expression of doxycycline-inducible shRNAs: knockdown of the pre-miRNA processing enzyme Drosha. Mol. Ther. 2007, 15(5), 938–945. [Google Scholar]
- Lee, YS; Kim, HK; Chung, S; Kim, KS; Dutta, A. Depletion of human micro-RNA miR-125b reveals that it is critical for the proliferation of differentiated cells but not for the down-regulation of putative targets during differentiation. J. Biol. Chem. 2005, 280(17), 16635–16641. [Google Scholar]
- Robb, GB; Brown, KM; Khurana, J; Rana, TM. Specific and potent RNAi in the nucleus of human cells. Nat. Struct. Mol. Biol. 2005, 12(2), 133–137. [Google Scholar]
- Morris, KV; Rossi, JJ. Lentivirus-mediated RNA interference therapy for human immunodeficiency virus type 1 infection. Hum. Gene. Ther. 2006, 17(5), 479–486. [Google Scholar]
- Morris, KV; Chan, SW; Jacobsen, SE; Looney, DJ. Small interfering RNA-induced transcriptional gene silencing in human cells. Science 2004, 305(5688), 1289–1292. [Google Scholar]
- Ebert, MS; Neilson, JR; Sharp, PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 2007, 4(9), 721–726. [Google Scholar]
- Thomas, CE; Ehrhardt, A; Kay, MA. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 2003, 4(5), 346–358. [Google Scholar]
- De Geest, BR; Van Linthout, SA; Collen, D. Humoral immune response in mice against a circulating antigen induced by adenoviral transfer is strictly dependent on expression in antigen-presenting cells. Blood 2003, 101(7), 2551–2556. [Google Scholar]
- Brown, BD; Cantore, A; Annoni, A; Sergi, LS; Lombardo, A; Della Valle, P; D’Angelo, A; Naldini, L. A microRNA-regulated lentiviral vector mediates stable correction of hemophilia B mice. Blood 2007, 110(13), 4144–4152. [Google Scholar]
- Xiao, J; Yang, B; Lin, H; Lu, Y; Luo, X; Wang, Z. Novel approaches for gene-specific interference via manipulating actions of microRNAs: examination on the pacemaker channel genes HCN2 and HCN4. J. Cell. Physiol. 2007, 212(2), 285–292. [Google Scholar]
- Doench, JG; Petersen, CP; Sharp, PA. siRNAs can function as miRNAs. Genes Dev. 2003, 17(4), 438–442. [Google Scholar]
- Chiu, YL; Rana, TM. siRNA function in RNAi: a chemical modification analysis. RNA 2003, 9(9), 1034–1048. [Google Scholar]
- Czauderna, F; Fechtner, M; Dames, S; Aygun, H; Klippel, A; Pronk, GJ; Giese, K; Kaufmann, J. Structural variations and stabilising modifications of synthetic siRNAs in mammalian cells. Nucl. Acids Res. 2003, 31(11), 2705–2716. [Google Scholar]
- McCaffrey, AP; Meuse, L; Pham, TT; Conklin, DS; Hannon, GJ; Kay, MA. RNA interference in adult mice. Nature 2002, 418(6893), 38–39. [Google Scholar]
- Brummelkamp, TR; Bernards, R; Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 2002, 296(5567), 550–553. [Google Scholar]
- Miyagishi, M; Taira, K. U6 promoter-driven siRNAs with four uridine 3′ overhangs efficiently suppress targeted gene expression in mammalian cells. Nat. Biotechnol. 2002, 20(5), 497–500. [Google Scholar]
- Costinean, S; Zanesi, N; Pekarsky, Y; Tili, E; Volinia, S; Heerema, N; Croce, CM. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc. Natl. Acad. Sci. USA 2006, 103(18), 7024–7029. [Google Scholar]
- Boden, D; Pusch, O; Silbermann, R; Lee, F; Tucker, L; Ramratnam, B. Enhanced gene silencing of HIV-1 specific siRNA using microRNA designed hairpins. Nucl. Acids Res. 2004, 32(3), 1154–1158. [Google Scholar]
- Chung, KH; Hart, CC; Al-Bassam, S; Avery, A; Taylor, J; Patel, PD; Vojtek, AB; Turner, DL. Polycistronic RNA polymerase II expression vectors for RNA interference based on BIC/miR-155. Nucl. Acids Res. 2006, 34(7), e53. [Google Scholar]
- Zeng, Y; Cai, X; Cullen, BR. Use of RNA polymerase II to transcribe artificial microRNAs. Methods Enzymol. 2005, 392, 371–380. [Google Scholar]
- Zeng, Y; Wagner, EJ; Cullen, BR. Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells. Mol. Cell 2002, 9(6), 1327–33. [Google Scholar]
- Zhou, H; Xia, XG; Xu, Z. An RNA polymerase II construct synthesizes short-hairpin RNA with a quantitative indicator and mediates highly efficient RNAi. Nucl. Acids Res. 2005, 33(6), e62. [Google Scholar]
- Stegmeier, F; Hu, G; Rickles, RJ; Hannon, GJ; Elledge, SJ. A lentiviral microRNA-based system for single-copy polymerase II-regulated RNA interference in mammalian cells. Proc. Natl. Acad. Sci. USA 2005, 102(37), 13212–13217. [Google Scholar]
- Chen, S; Ni, M; Yu, B; Lv, T; Lu, M; Gong, F. Construction and identification of a human liver specific microRNA eukaryotic expression vector. Cell. Mol. Immunol. 2007, 4(6), 473–477. [Google Scholar]
- Wang, J; Theunissen, TW; Orkin, SH. Site-directed, virus-free, and inducible RNAi in embryonic stem cells. Proc. Natl. Acad. Sci. USA 2007, 104(52), 20850–20855. [Google Scholar]
- Bridge, AJ; Pebernard, S; Ducraux, A; Nicoulaz, AL; Iggo, R. Induction of an interferon response by RNAi vectors in mammalian cells. Nat. Genet. 2003, 34(3), 263–264. [Google Scholar]
- Sledz, CA; Holko, M; de Veer, MJ; Silverman, RH; Williams, BR. Activation of the interferon system by short-interfering RNAs. Nat. Cell Biol. 2003, 5(9), 834–839. [Google Scholar]
- McManus, MT; Petersen, CP; Haines, BB; Chen, J; Sharp, PA. Gene silencing using micro-RNA designed hairpins. RNA 2002, 8(6), 842–850. [Google Scholar]
- Castanotto, D; Sakurai, K; Lingeman, R; Li, H; Shively, L; Aagaard, L; Soifer, H; Gatignol, A; Riggs, A; Rossi, JJ. Combinatorial delivery of small interfering RNAs reduces RNAi efficacy by selective incorporation into RISC. Nucl. Acids Res. 2007, 35(15), 5154–5164. [Google Scholar]
- Dickins, RA; McJunkin, K; Hernando, E; Premsrirut, PK; Krizhanovsky, V; Burgess, DJ; Kim, SY; Cordon-Cardo, C; Zender, L; Hannon, GJ; Lowe, SW. Tissue-specific and reversible RNA interference in transgenic mice. Nat. Genet. 2007, 39(7), 914–921. [Google Scholar]
- McBride, JL; Boudreau, RL; Harper, SQ; Staber, PD; Monteys, AM; Martins, I; Gilmore, BL; Burstein, H; Peluso, RW; Polisky, B; Carter, BJ; Davidson, BL. Artificial miRNAs mitigate shRNA-mediated toxicity in the brain: implications for the therapeutic development of RNAi. Proc. Natl. Acad. Sci. USA 2008, 105(15), 5868–5873. [Google Scholar]
- Han, J; Lee, Y; Yeom, KH; Nam, JW; Heo, I; Rhee, JK; Sohn, SY; Cho, Y; Zhang, BT; Kim, VN. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 2006, 125(5), 887–901. [Google Scholar]
- Saetrom, P; Snove, O; Nedland, M; Grunfeld, TB; Lin, Y; Bass, MB; Canon, JR. Conserved microRNA characteristics in mammals. Oligonucleotides 2006, 16(2), 115–144. [Google Scholar]
- Kawasaki, H; Taira, K. Short hairpin type of dsRNAs that are controlled by tRNA(Val) promoter significantly induce RNAi-mediated gene silencing in the cytoplasm of human cells. Nucl. Acids Res. 2003, 31(2), 700–707. [Google Scholar]
- Marquez, RT; McCaffrey, AP. Advances in microRNAs: implications for gene therapists. Hum. Gene. Ther. 2008, 19(1), 27–38. [Google Scholar]
- Lo, HL; Chang, T; Yam, P; Marcovecchio, PM; Li, S; Zaia, JA; Yee, JK. Inhibition of HIV-1 replication with designed miRNAs expressed from RNA polymerase II promoters. Gene. Ther. 2007, 14(21), 1503–1512. [Google Scholar]
- Silva, JM; Li, MZ; Chang, K; Ge, W; Golding, MC; Rickles, RJ; Siolas, D; Hu, G; Paddison, PJ; Schlabach, MR; Sheth, N; Bradshaw, J; Burchard, J; Kulkarni, A; Cavet, G; Sachidanandam, R; McCombie, WR; Cleary, MA; Elledge, SJ; Hannon, GJ. Second-generation shRNA libraries covering the mouse and human genomes. Nat. Genet. 2005, 37(11), 1281–1288. [Google Scholar]
- Mineno, J; Okamoto, S; Ando, T; Sato, M; Chono, H; Izu, H; Takayama, M; Asada, K; Mirochnitchenko, O; Inouye, M; Kato, I. The expression profile of microRNAs in mouse embryos. Nucl. Acids Res. 2006, 34(6), 1765–1771. [Google Scholar]
- Yang, W; Paschen, W. Conditional gene silencing in mammalian cells mediated by a stress-inducible promoter. Biochem. Biophys. Res. Commun. 2008, 365(3), 521–527. [Google Scholar]
- Zhu, X; Santat, LA; Chang, MS; Liu, J; Zavzavadjian, JR; Wall, EA; Kivork, C; Simon, MI; Fraser, ID. A versatile approach to multiple gene RNA interference using microRNA-based short hairpin RNAs. BMC Mol. Biol. 2007, 8, 98. [Google Scholar]
- Zhang, B; Pan, X; Cobb, GP; Anderson, TA. Plant microRNA: a small regulatory molecule with big impact. Dev. Biol. 2006, 289(1), 3–16. [Google Scholar]
- Warthmann, N; Chen, H; Ossowski, S; Weigel, D; Herve, P. Highly specific gene silencing by artificial miRNAs in rice. PLoS ONE 2008, 3(3), e1829. [Google Scholar]
- Ossowski, S; Schwab, R; Weigel, D. Gene silencing in plants using artificial microRNAs and other small RNAs. Plant J. 2008, 53(4), 674–690. [Google Scholar]
- Qu, J; Ye, J; Fang, R. Artificial microRNA-mediated virus resistance in plants. J. Virol. 2007, 81(12), 6690–6699. [Google Scholar]
- Liang, Z; Wu, H; Reddy, S; Zhu, A; Wang, S; Blevins, D; Yoon, Y; Zhang, Y; Shim, H. Blockade of invasion and metastasis of breast cancer cells via targeting CXCR4 with an artificial microRNA. Biochem. Biophys. Res. Commun. 2007, 363(3), 542–546. [Google Scholar]
- Scaria, V; Hariharan, M; Brahmachari, SK; Maiti, S; Pillai, B. microRNA: an Emerging Therapeutic. ChemMedChem. 2007, 2(6), 789–792. [Google Scholar]
Share and Cite
Liu, Z.; Sall, A.; Yang, D. MicroRNA: an Emerging Therapeutic Target and Intervention Tool. Int. J. Mol. Sci. 2008, 9, 978-999. https://doi.org/10.3390/ijms9060978
Liu Z, Sall A, Yang D. MicroRNA: an Emerging Therapeutic Target and Intervention Tool. International Journal of Molecular Sciences. 2008; 9(6):978-999. https://doi.org/10.3390/ijms9060978
Chicago/Turabian StyleLiu, Zhen, Alhousseynou Sall, and Decheng Yang. 2008. "MicroRNA: an Emerging Therapeutic Target and Intervention Tool" International Journal of Molecular Sciences 9, no. 6: 978-999. https://doi.org/10.3390/ijms9060978
APA StyleLiu, Z., Sall, A., & Yang, D. (2008). MicroRNA: an Emerging Therapeutic Target and Intervention Tool. International Journal of Molecular Sciences, 9(6), 978-999. https://doi.org/10.3390/ijms9060978



