Circular RNA Formation and Degradation Are Not Directed by Universal Pathways
Abstract
1. Introduction
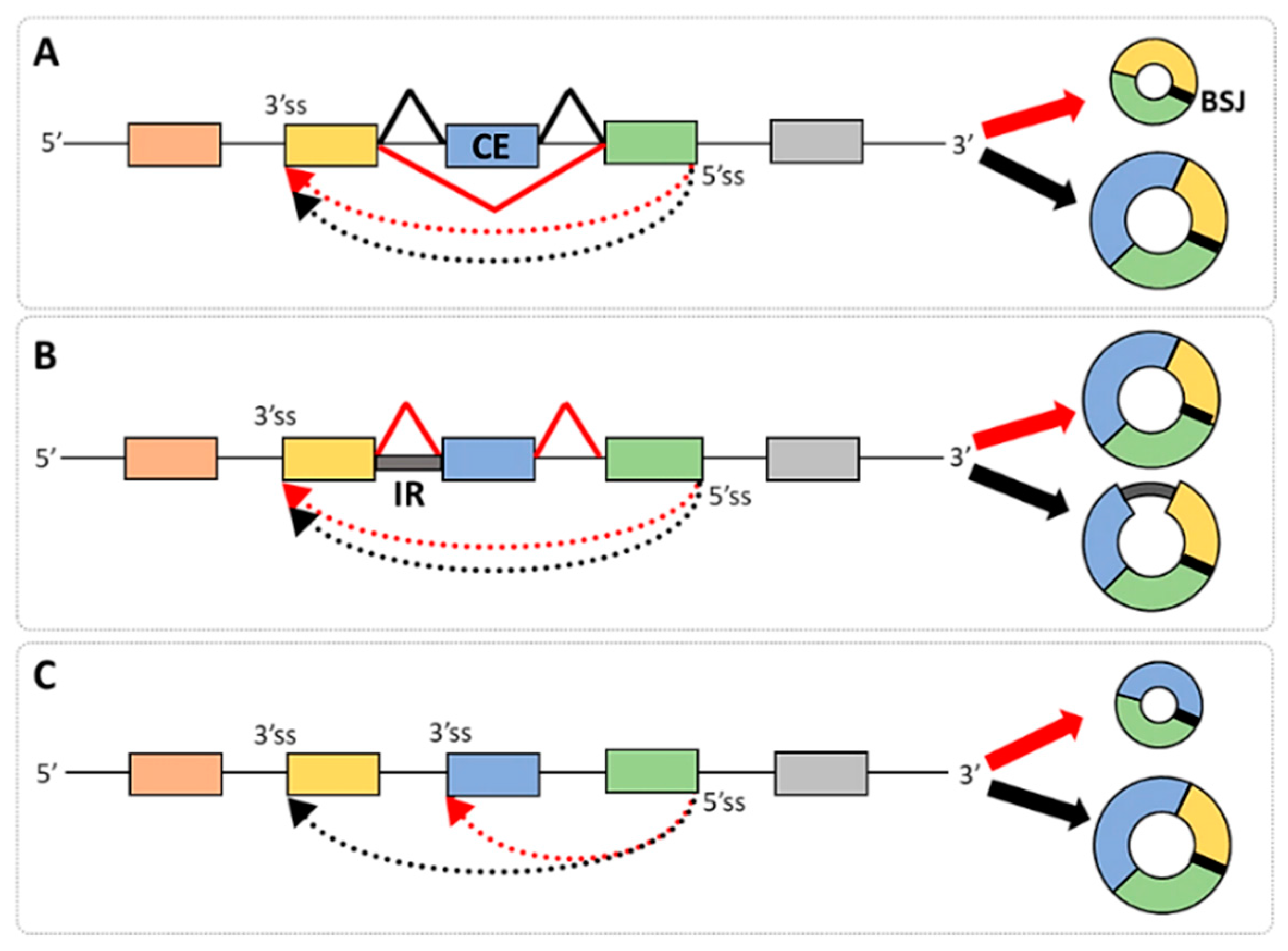
2. Alternative Splicing and circRNA Isoforms
3. Types of Circular Transcripts
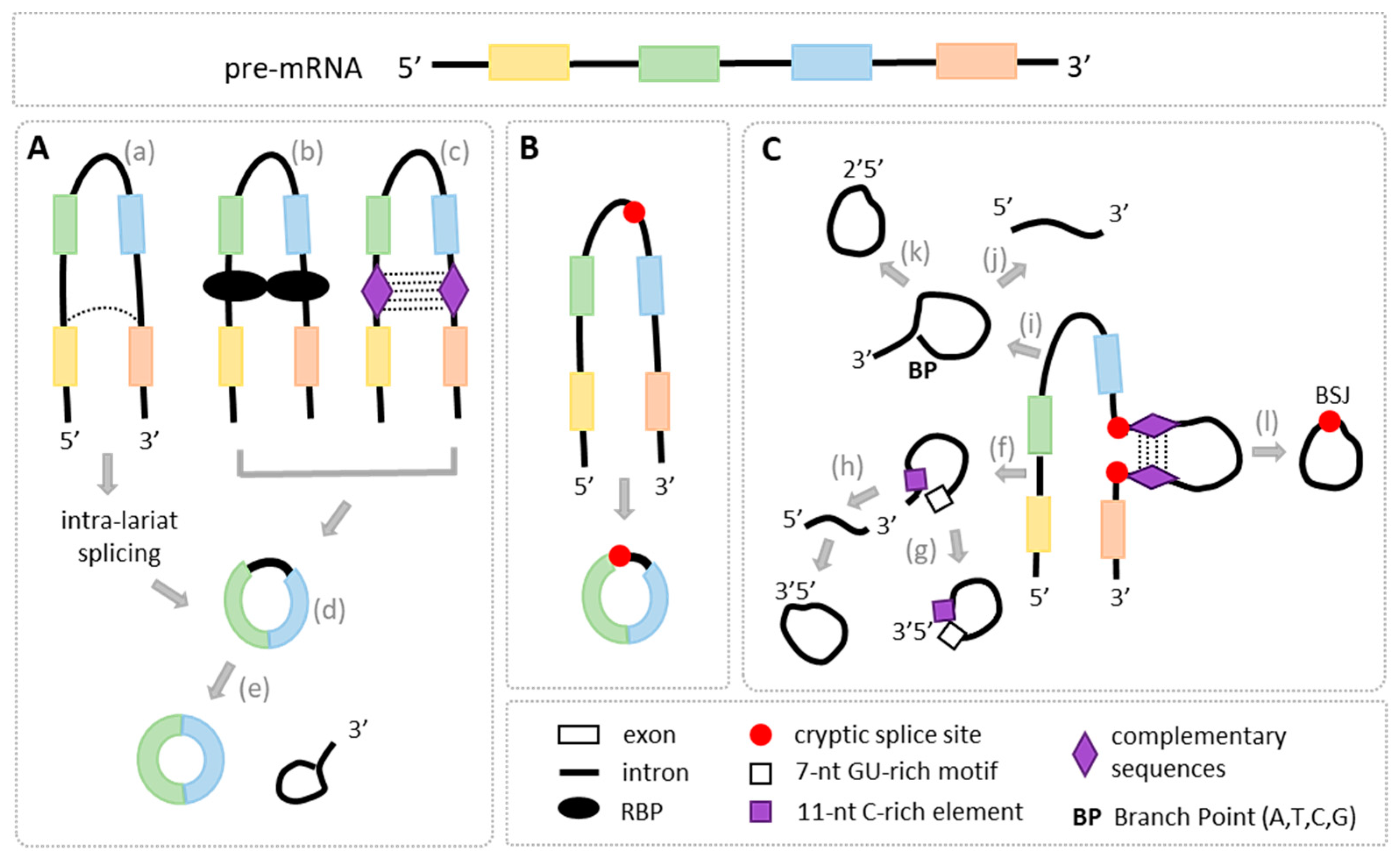
4. Regulators of circRNA Biogenesis
4.1. cis-Regulatory Elements
4.1.1. The Features of Introns and Exons Participating in circRNA Generation
4.1.2. The Features of Canonical and Cryptic Splice Sites Involved in BSJ Formation
4.2. trans-Regulatory Factors
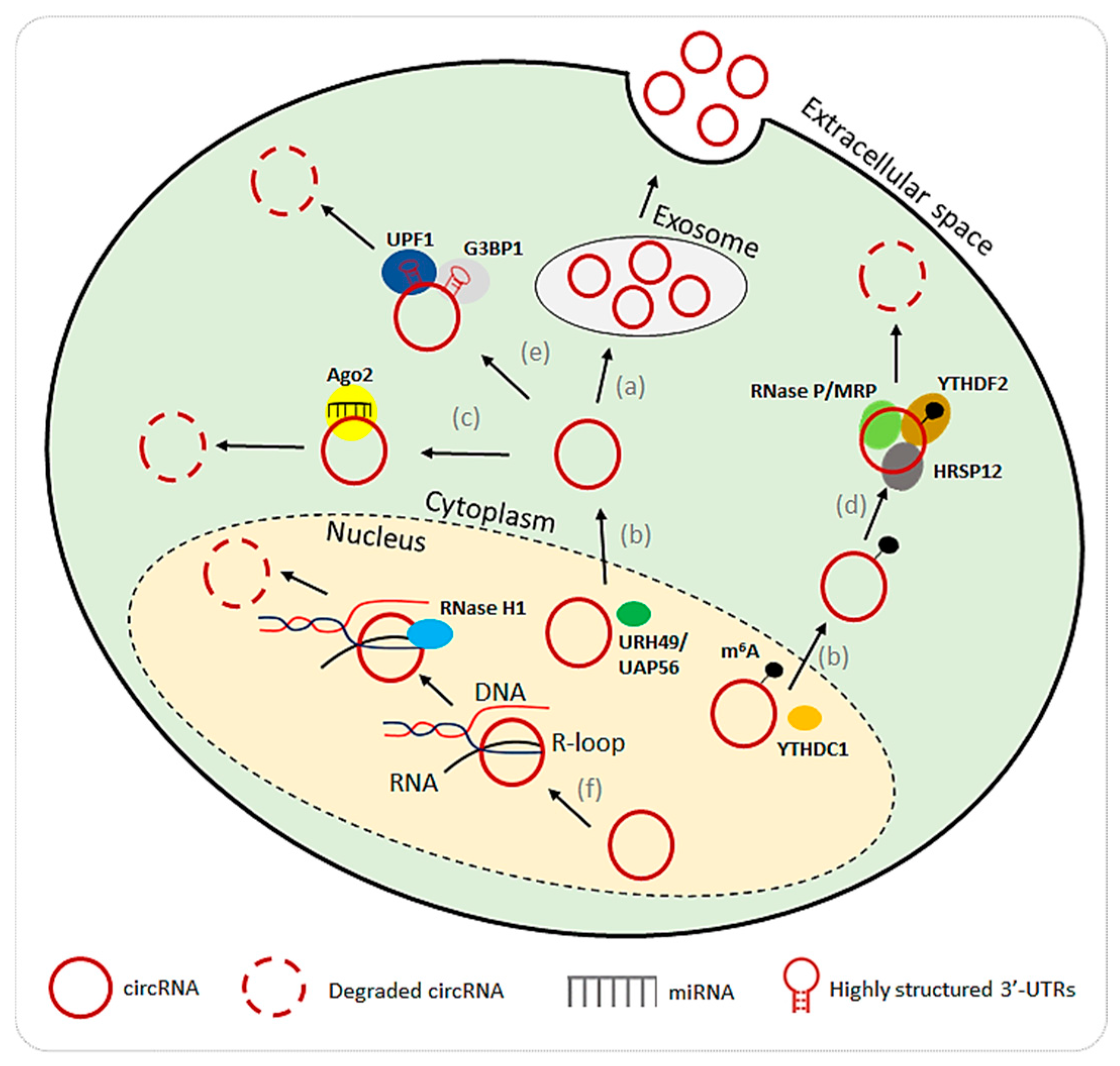
5. Degradation of Circular RNAs
6. Conclusions and Future Directions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Sanger, H.L.; Klotz, G.; Riesner, D.; Gross, H.J.; Kleinschmidt, A.K. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl. Acad. Sci. USA 1976, 73, 3852–3856. [Google Scholar] [CrossRef]
- Capel, B.; Swain, A.; Nicolis, S.; Hacker, A.; Walter, M.; Koopman, P.; Goodfellow, P.; Lovell-Badge, R. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 1993, 73, 1019–1030. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs Are the Predominant Transcript Isoform from Hundreds of Human Genes in Diverse Cell Types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Westholm, J.O.; Miura, P.; Olson, S.; Shenker, S.; Joseph, B.; Sanfilippo, P.; Celniker, S.E.; Graveley, B.R.; Lai, E.C. Genome-wide Analysis of Drosophila Circular RNAs Reveals Their Structural and Sequence Properties and Age-Dependent Neural Accumulation. Cell Rep. 2014, 9, 1966–1980. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Ivanov, A.; Memczak, S.; Wyler, E.; Torti, F.; Porath, H.T.; Orejuela, M.R.; Piechotta, M.; Levanon, E.Y.; Landthaler, M.; Dieterich, C.; et al. Analysis of Intron Sequences Reveals Hallmarks of Circular RNA Biogenesis in Animals. Cell Rep. 2014, 10, 170–177. [Google Scholar] [CrossRef]
- Lu, T.; Cui, L.; Zhou, Y.; Zhu, C.; Fan, D.; Gong, H.; Zhao, Q.; Zhou, C.; Zhao, Y.; Lu, D.; et al. Transcriptome-wide investigation of circular RNAs in rice. RNA 2015, 21, 2076–2087. [Google Scholar] [CrossRef] [PubMed]
- Szabo, L.; Salzman, J. Detecting circular RNAs: Bioinformatic and experimental challenges. Nat. Rev. Genet. 2016, 17, 679–692. [Google Scholar] [CrossRef]
- Abaza, T.; El-Aziz, M.K.A.; Daniel, K.A.; Karousi, P.; Papatsirou, M.; Fahmy, S.A.; Hamdy, N.M.; Kontos, C.K.; Youness, R.A. Emerging Role of Circular RNAs in Hepatocellular Carcinoma Immunotherapy. Int. J. Mol. Sci. 2023, 24, 16484. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Fu, J.; Zhou, Y. Circular RNAs and Their Emerging Roles in Immune Regulation. Front. Immunol. 2018, 9, 2977. [Google Scholar] [CrossRef]
- Liu, J.; Liu, T.; Wang, X.; He, A. Circles reshaping the RNA world: From waste to treasure. Mol. Cancer 2017, 16, 58. [Google Scholar] [CrossRef] [PubMed]
- Ragan, C.; Goodall, G.J.; Shirokikh, N.E.; Preiss, T. Insights into the biogenesis and potential functions of exonic circular RNA. Sci. Rep. 2019, 9, 2048. [Google Scholar] [CrossRef]
- Hu, D.G.; Mackenzie, P.I.; Hulin, J.-A.; McKinnon, R.A.; Meech, R. Circular RNAs of UDP-Glycosyltransferase (UGT) Genes Expand the Complexity and Diversity of the UGT Transcriptome. Mol. Pharmacol. 2021, 99, 488–503. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Sunagawa, Y.; Yamada, S.; Sonohara, F.; Kurimoto, K.; Tanaka, N.; Suzuki, Y.; Inokawa, Y.; Takami, H.; Hayashi, M.; Kanda, M.; et al. Genome-wide identification and characterization of circular RNA in resected hepatocellular carcinoma and background liver tissue. Sci. Rep. 2021, 11, 6016. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, K.; Nielsen, A.F.; Venø, M.T.; Kjems, J. Nanopore long-read sequencing of circRNAs. Methods 2021, 196, 23–29. [Google Scholar] [CrossRef]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef]
- Liang, D.; Wilusz, J.E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 2014, 28, 2233–2247. [Google Scholar] [CrossRef]
- Ren, L.; Jiang, Q.; Mo, L.; Tan, L.; Dong, Q.; Meng, L.; Yang, N.; Li, G. Mechanisms of circular RNA degradation. Commun. Biol. 2022, 5, 1355. [Google Scholar] [CrossRef] [PubMed]
- Huang, A.; Zheng, H.; Wu, Z.; Chen, M.; Huang, Y. Circular RNA-protein interactions: Functions, mechanisms, and identification. Theranostics 2020, 10, 3503–3517. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Joseph, S.; Xia, M.; Teng, F.; Chen, X.; Huang, R.; Zhai, L.; Deng, W. Circular RNAs Acting as miRNAs’ Sponges and Their Roles in Stem Cells. J. Clin. Med. 2022, 11, 2909. [Google Scholar] [CrossRef]
- Li, T.-R.; Jia, Y.-J.; Wang, Q.; Shao, X.-Q.; Lv, R.-J. Circular RNA: A new star in neurological diseases. Int. J. Neurosci. 2016, 127, 726–734. [Google Scholar] [CrossRef]
- Altesha, M.; Ni, T.; Khan, A.; Liu, K.; Zheng, X. Circular RNA in cardiovascular disease. J. Cell. Physiol. 2018, 234, 5588–5600. [Google Scholar] [CrossRef]
- Czubak, K.; Taylor, K.; Piasecka, A.; Sobczak, K.; Kozlowska, K.; Philips, A.; Sedehizadeh, S.; Brook, J.D.; Wojciechowska, M.; Kozlowski, P. Global Increase in Circular RNA Levels in Myotonic Dystrophy. Front. Genet. 2019, 10, 649. [Google Scholar] [CrossRef]
- Czubak, K.; Sedehizadeh, S.; Kozlowski, P.; Wojciechowska, M. An Overview of Circular RNAs and Their Implications in Myotonic Dystrophy. Int. J. Mol. Sci. 2019, 20, 4385. [Google Scholar] [CrossRef]
- Zaphiropoulos, P.G. Exon Skipping and Circular RNA Formation in Transcripts of the Human Cytochrome P-450 2C18 Gene in Epidermis and of the Rat Androgen Binding Protein Gene in Testis. Mol. Cell. Biol. 1997, 17, 2985–2993. [Google Scholar] [CrossRef]
- Surono, A.; Takeshima, Y.; Wibawa, T.; Ikezawa, M.; Nonaka, I.; Matsuo, M. Circular dystrophin RNAs consisting of exons that were skipped by alternative splicing. Hum. Mol. Genet. 1999, 8, 493–500. [Google Scholar] [CrossRef]
- Kelly, S.; Greenman, C.; Cook, P.R.; Papantonis, A. Exon Skipping Is Correlated with Exon Circularization. J. Mol. Biol. 2015, 427, 2414–2417. [Google Scholar] [CrossRef]
- Kumari, A.; Sedehizadeh, S.; Brook, J.D.; Kozlowski, P.; Wojciechowska, M. Differential fates of introns in gene expression due to global alternative splicing. Hum. Genet. 2021, 141, 31–47. [Google Scholar] [CrossRef]
- Gao, Y.; Wang, J.; Zheng, Y.; Zhang, J.; Chen, S.; Zhao, F. Comprehensive identification of internal structure and alternative splicing events in circular RNAs. Nat. Commun. 2016, 7, 12060. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-O.; Dong, R.; Zhang, Y.; Zhang, J.-L.; Luo, Z.; Zhang, J.; Chen, L.-L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [PubMed]
- Beltrán-García, J.; Osca-Verdegal, R.; Nacher-Sendra, E.; Pallardó, F.V.; García-Giménez, J.L. Circular RNAs in Sepsis: Biogenesis, Function, and Clinical Significance. Cells 2020, 9, 1544. [Google Scholar] [CrossRef]
- Ji, P.; Wu, W.; Chen, S.; Zheng, Y.; Zhou, L.; Zhang, J.; Cheng, H.; Yan, J.; Zhang, S.; Yang, P.; et al. Expanded Expression Landscape and Prioritization of Circular RNAs in Mammals. Cell Rep. 2019, 26, 3444–3460.e5. [Google Scholar] [CrossRef]
- Feng, J.; Chen, K.; Dong, X.; Xu, X.; Jin, Y.; Zhang, X.; Chen, W.; Han, Y.; Shao, L.; Gao, Y.; et al. Genome-wide identification of cancer-specific alternative splicing in circRNA. Mol. Cancer 2019, 18, 35. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sharpless, N.E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 2014, 32, 453–461. [Google Scholar] [CrossRef]
- Chen, I.; Chen, C.; Chuang, T. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip. Rev. RNA 2015, 6, 563–579. [Google Scholar] [CrossRef] [PubMed]
- Lyu, D.; Huang, S. The emerging role and clinical implication of human exonic circular RNA. RNA Biol. 2016, 14, 1000–1006. [Google Scholar] [CrossRef] [PubMed]
- Kameyama, T.; Suzuki, H.; Mayeda, A. Re-splicing of mature mRNA in cancer cells promotes activation of distant weak alternative splice sites. Nucleic Acids Res. 2012, 40, 7896–7906. [Google Scholar] [CrossRef] [PubMed]
- Ju, H.; Hu, Z.; Wei, D.; Huang, J.; Zhang, X.; Rui, M.; Li, Z.; Zhang, X.; Hu, J.; Guo, W.; et al. A novel intronic circular RNA, circGNG7, inhibits head and neck squamous cell carcinoma progression by blocking the phosphorylation of heat shock protein 27 at Ser78 and Ser82. Cancer Commun. 2021, 41, 1152–1172. [Google Scholar] [CrossRef]
- Taggart, A.J.; Lin, C.-L.; Shrestha, B.; Heintzelman, C.; Kim, S.; Fairbrother, W.G. Large-scale analysis of branchpoint usage across species and cell lines. Genome Res. 2017, 27, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular Intronic Long Noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Osman, I.; Tay, M.L.-I.; Pek, J.W. Stable intronic sequence RNAs (sisRNAs): A new layer of gene regulation. Cell. Mol. Life Sci. 2016, 73, 3507–3519. [Google Scholar] [CrossRef]
- Chan, S.N.; Pek, J.W. Stable Intronic Sequence RNAs (sisRNAs): An Expanding Universe. Trends Biochem. Sci. 2019, 44, 258–272. [Google Scholar] [CrossRef]
- Talhouarne, G.J.S.; Gall, J.G. Lariat intronic RNAs in the cytoplasm of vertebrate cells. Proc. Natl. Acad. Sci. USA 2018, 115, E7970–E7977. [Google Scholar] [CrossRef]
- Saini, H.; A Bicknell, A.; Eddy, S.R.; Moore, M.J.; States, U. Free circular introns with an unusual branchpoint in neuronal projections. eLife 2019, 8, e47809. [Google Scholar] [CrossRef]
- Jin, J.; He, X.; Silva, E. Stable intronic sequence RNAs (sisRNAs) are selected regions in introns with distinct properties. BMC Genom. 2020, 21, 287. [Google Scholar] [CrossRef]
- Zhong, Y.; Yang, Y.; Wang, X.; Ren, B.; Wang, X.; Shan, G.; Chen, L. Systematic identification and characterization of exon–intron circRNAs. Genome Res. 2024, 34, 376–393. [Google Scholar] [CrossRef]
- Aktas, T.; Avsar Ilik, I.; Maticzka, D.; Bhardwaj, V.; Pessoa Rodrigues, C.; Mittler, G.; Manke, T.; Backofen, R.; Akhtar, A. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 2017, 544, 115–119. [Google Scholar] [CrossRef]
- Kramer, M.C.; Liang, D.; Tatomer, D.C.; Gold, B.; March, Z.M.; Cherry, S.; Wilusz, J.E. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev. 2015, 29, 2168–2182. [Google Scholar] [CrossRef]
- Kokot, K.E.; Kneuer, J.M.; John, D.; Rebs, S.; Möbius-Winkler, M.N.; Erbe, S.; Müller, M.; Andritschke, M.; Gaul, S.; Sheikh, B.N.; et al. Reduction of A-to-I RNA editing in the failing human heart regulates formation of circular RNAs. Basic Res. Cardiol. 2022, 117, 32. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.O.; Wang, H.B.; Zhang, Y.; Lu, X.; Chen, L.L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef]
- Zhang, X.-O.; Pratt, H.; Weng, Z. Investigating the Potential Roles of SINEs in the Human Genome. Annu. Rev. Genom. Hum. Genet. 2021, 22, 199–218. [Google Scholar] [CrossRef]
- Chen, L.-L.; DeCerbo, J.N.; Carmichael, G.G. Alu element-mediated gene silencing. EMBO J. 2008, 27, 1694–1705. [Google Scholar] [CrossRef]
- Ottesen, E.W.; Luo, D.; Seo, J.; Singh, N.N.; Singh, R.N. Human Survival Motor Neuron genes generate a vast repertoire of circular RNAs. Nucleic Acids Res. 2019, 47, 2884–2905. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Bao, J.; Hu, J.; Liu, L.; Xu, D. Circular RNA in cardiovascular disease: Expression, mechanisms and clinical prospects. J. Cell. Mol. Med. 2020, 25, 1817–1824. [Google Scholar] [CrossRef] [PubMed]
- Szabo, L.; Morey, R.; Palpant, N.J.; Wang, P.L.; Afari, N.; Jiang, C.; Parast, M.M.; Murry, C.E.; Laurent, L.C.; Salzman, J. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol. 2015, 16, 126. [Google Scholar] [CrossRef] [PubMed]
- Aufiero, S.; Hoogenhof, M.M.G.v.D.; Reckman, Y.J.; Beqqali, A.; van der Made, I.; Kluin, J.; Khan, M.A.F.; Pinto, Y.M.; Creemers, E.E. Cardiac circRNAs arise mainly from constitutive exons rather than alternatively spliced exons. RNA 2018, 24, 815–827. [Google Scholar] [CrossRef]
- Chen, L.-L. The biogenesis and emerging roles of circular RNAs. Nat. Rev. Mol. Cell Biol. 2016, 17, 205–211. [Google Scholar] [CrossRef]
- Starke, S.; Jost, I.; Rossbach, O.; Schneider, T.; Schreiner, S.; Hung, L.-H.; Bindereif, A. Exon Circularization Requires Canonical Splice Signals. Cell Rep. 2015, 10, 103–111. [Google Scholar] [CrossRef]
- Gao, X.; Ma, X.-K.; Li, X.; Li, G.-W.; Liu, C.-X.; Zhang, J.; Wang, Y.; Wei, J.; Chen, J.; Chen, L.-L.; et al. Knockout of circRNAs by base editing back-splice sites of circularized exons. Genome Biol. 2022, 23, 16. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Hu, Z.; Zhou, J.; Tian, C.; Tian, G.; He, M.; Gao, L.; Chen, L.; Li, T.; Peng, H.; et al. Interior circular RNA. RNA Biol. 2019, 17, 87–97. [Google Scholar] [CrossRef]
- Di Liddo, A.; Machado, C.d.O.F.; Fischer, S.; Ebersberger, S.; Heumüller, A.W.; E Weigand, J.; Müller-McNicoll, M.; Zarnack, K. A combined computational pipeline to detect circular RNAs in human cancer cells under hypoxic stress. J. Mol. Cell Biol. 2019, 11, 829–844. [Google Scholar] [CrossRef]
- Wilusz, J. Circular RNA and Splicing: Skip Happens. J. Mol. Biol. 2015, 427, 2411–2413. [Google Scholar] [CrossRef] [PubMed]
- Conn, S.J.; Pillman, K.A.; Toubia, J.; Conn, V.M.; Salmanidis, M.; Phillips, C.A.; Roslan, S.; Schreiber, A.W.; Gregory, P.A.; Goodall, G.J. The RNA Binding Protein Quaking Regulates Formation of circRNAs. Cell 2015, 160, 1125–1134. [Google Scholar] [CrossRef]
- Yuan, Y.; Compton, S.A.; Sobczak, K.; Stenberg, M.G.; Thornton, C.A.; Griffith, J.D.; Swanson, M.S. Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res. 2007, 35, 5474–5486. [Google Scholar] [CrossRef] [PubMed]
- Teplova, M.; Hafner, M.; Teplov, D.; Essig, K.; Tuschl, T.; Patel, D.J. Structure–function studies of STAR family Quaking proteins bound to their in vivo RNA target sites. Genes Dev. 2013, 27, 928–940. [Google Scholar] [CrossRef] [PubMed]
- Errichelli, L.; Dini Modigliani, S.; Laneve, P.; Colantoni, A.; Legnini, I.; Capauto, D.; Rosa, A.; De Santis, R.; Scarfò, R.; Peruzzi, G.; et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat. Commun. 2017, 8, 14741. [Google Scholar] [CrossRef] [PubMed]
- Rybak-Wolf, A.; Stottmeister, C.; Glažar, P.; Jens, M.; Pino, N.; Giusti, S.; Hanan, M.; Behm, M.; Bartok, O.; Ashwal-Fluss, R.; et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell 2015, 58, 870–885. [Google Scholar] [CrossRef]
- Middleton, R.; Gao, D.; Thomas, A.; Singh, B.; Au, A.; Wong, J.J.-L.; Bomane, A.; Cosson, B.; Eyras, E.; Rasko, J.E.J.; et al. IRFinder: Assessing the impact of intron retention on mammalian gene expression. Genome Biol. 2017, 18, 51. [Google Scholar] [CrossRef]
- Monteuuis, G.; Wong, J.J.L.; Bailey, C.G.; Schmitz, U.; Rasko, J.E.J. The changing paradigm of intron retention: Regulation, ramifications and recipes. Nucleic Acids Res. 2019, 47, 11497–11513. [Google Scholar] [CrossRef] [PubMed]
- Lasda, E.; Parker, R. Circular RNAs Co-Precipitate with Extracellular Vesicles: A Possible Mechanism for circRNA Clearance. PLoS ONE 2016, 11, e0148407. [Google Scholar] [CrossRef]
- Huang, C.; Liang, D.; Tatomer, D.C.; Wilusz, J.E. A length-dependent evolutionarily conserved pathway controls nuclear export of circular RNAs. Genes Dev. 2018, 32, 639–644. [Google Scholar] [CrossRef]
- Chen, R.-X.; Chen, X.; Xia, L.-P.; Zhang, J.-X.; Pan, Z.-Z.; Ma, X.-D.; Han, K.; Chen, J.-W.; Judde, J.-G.; Deas, O.; et al. N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat. Commun. 2019, 10, 4695. [Google Scholar] [CrossRef] [PubMed]
- Piwecka, M.; Glažar, P.; Hernandez-Miranda, L.R.; Memczak, S.; Wolf, S.A.; Rybak-Wolf, A.; Filipchyk, A.; Klironomos, F.; Cerda Jara, C.A.; Fenske, P.; et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 2017, 357, eaam8526. [Google Scholar] [CrossRef]
- Pan, Z.; Li, G.F.; Sun, M.L.; Xie, L.; Liu, D.; Zhang, Q.; Yang, X.X.; Xia, S.; Liu, X.; Zhou, H.; et al. MicroRNA-1224 Splicing CircularRNA-Filip1l in an Ago2-Dependent Manner Regulates Chronic Inflammatory Pain via Targeting Ubr5. J. Neurosci. Off. J. Soc. Neurosci. 2019, 39, 2125–2143. [Google Scholar] [CrossRef]
- Xu, J.; Wan, Z.; Tang, M.; Lin, Z.; Jiang, S.; Ji, L.; Gorshkov, K.; Mao, Q.; Xia, S.; Cen, D.; et al. N6-methyladenosine-modified CircRNA-SORE sustains sorafenib resistance in hepatocellular carcinoma by regulating β-catenin signaling. Mol. Cancer 2020, 19, 163. [Google Scholar] [CrossRef]
- Guo, Y.; Guo, Y.; Chen, C.; Fan, D.; Wu, X.; Zhao, L.; Shao, B.; Sun, Z.; Ji, Z. Circ3823 contributes to growth, metastasis and angiogenesis of colorectal cancer: Involvement of miR-30c-5p/TCF7 axis. Mol. Cancer 2021, 20, 93. [Google Scholar] [CrossRef] [PubMed]
- Park, O.H.; Ha, H.; Lee, Y.; Boo, S.H.; Kwon, D.H.; Song, H.K.; Kim, Y.K. Endoribonucleolytic Cleavage of m6A-Containing RNAs by RNase P/MRP Complex. Mol. Cell 2019, 74, 494–507. [Google Scholar] [CrossRef]
- Zhang, M.; Wang, T.; Xiao, G.; Xie, Y. Large-Scale Profiling of RBP-circRNA Interactions from Public CLIP-Seq Datasets. Genes 2020, 11, 54. [Google Scholar] [CrossRef]
- Fischer, J.W.; Busa, V.F.; Shao, Y.; Leung, A.K. Structure-Mediated RNA Decay by UPF1 and G3BP1. Mol. Cell 2020, 78, 70–84.e6. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, J.-L.; Lei, Y.-N.; Liu, X.-Q.; Xue, W.; Zhang, Y.; Nan, F.; Gao, X.; Zhang, J.; Wei, J.; et al. Linking circular intronic RNA degradation and function in transcription by RNase H1. Sci. China Life Sci. 2021, 64, 1795–1809. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Srinivasan, A.; Mroczko-Młotek, E.; Wojciechowska, M. Circular RNA Formation and Degradation Are Not Directed by Universal Pathways. Int. J. Mol. Sci. 2025, 26, 726. https://doi.org/10.3390/ijms26020726
Srinivasan A, Mroczko-Młotek E, Wojciechowska M. Circular RNA Formation and Degradation Are Not Directed by Universal Pathways. International Journal of Molecular Sciences. 2025; 26(2):726. https://doi.org/10.3390/ijms26020726
Chicago/Turabian StyleSrinivasan, Arvind, Emilia Mroczko-Młotek, and Marzena Wojciechowska. 2025. "Circular RNA Formation and Degradation Are Not Directed by Universal Pathways" International Journal of Molecular Sciences 26, no. 2: 726. https://doi.org/10.3390/ijms26020726
APA StyleSrinivasan, A., Mroczko-Młotek, E., & Wojciechowska, M. (2025). Circular RNA Formation and Degradation Are Not Directed by Universal Pathways. International Journal of Molecular Sciences, 26(2), 726. https://doi.org/10.3390/ijms26020726




