Genetic and Neurological Deficiencies in the Visual System of mct8 Mutant Zebrafish
Abstract
1. Introduction
2. Results
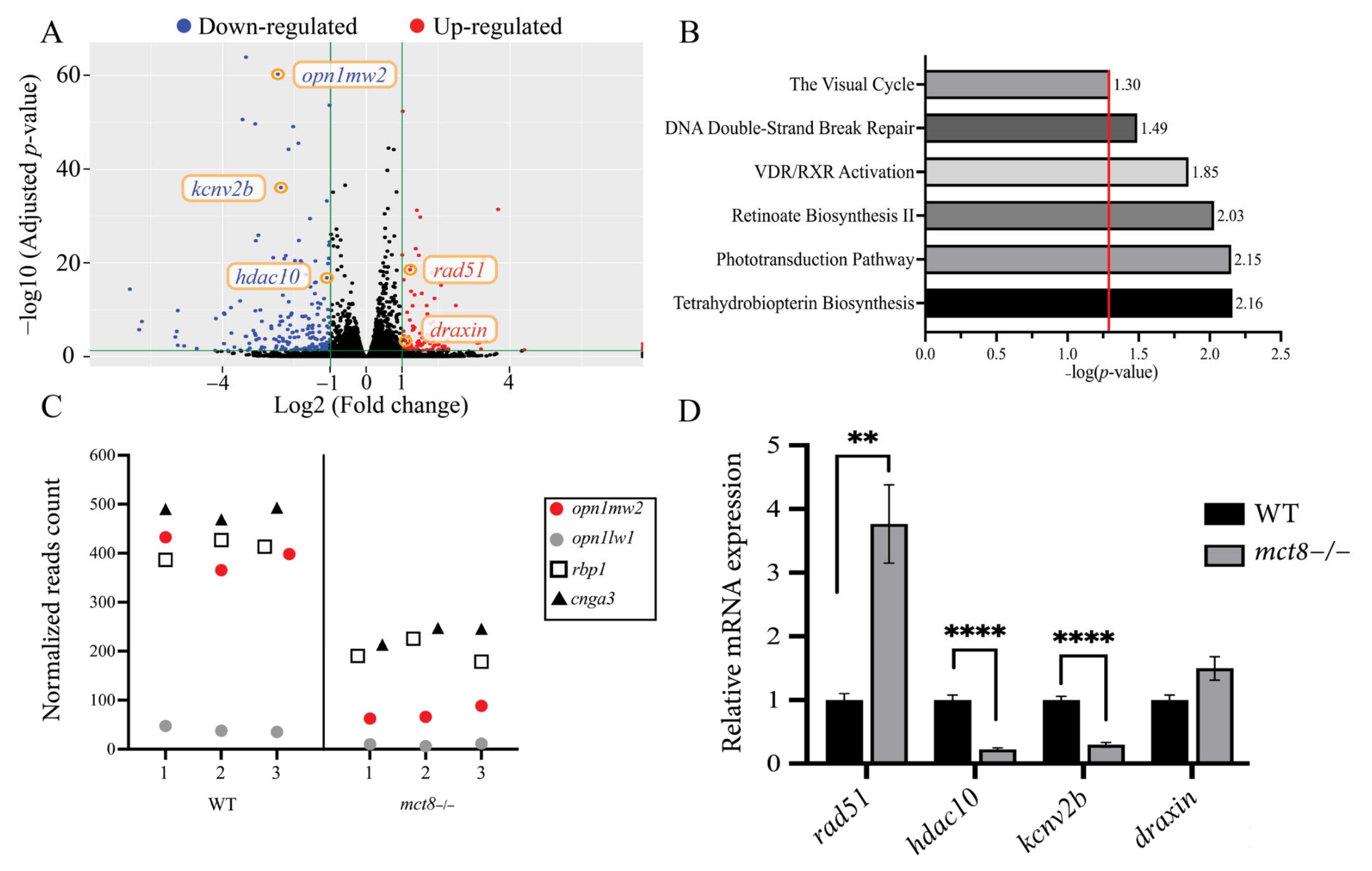
2.1. Profiling the Transcriptome of mct8−/− Larvae Revealed Altered Expression of Visual System Genes
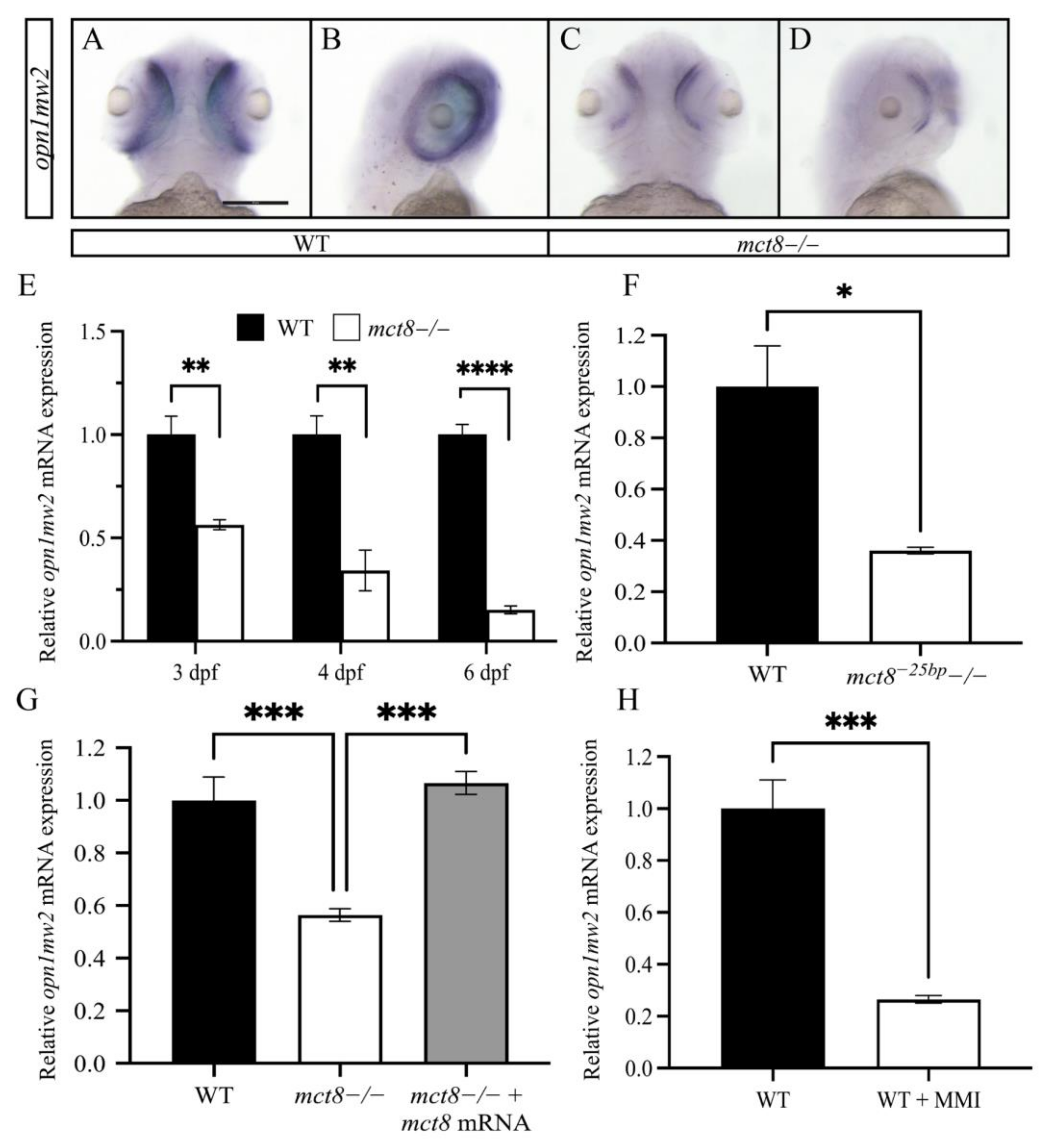
2.2. The Expression of opn1mw2 Is Reduced in mct8−/− Larvae and Is Regulated by THs
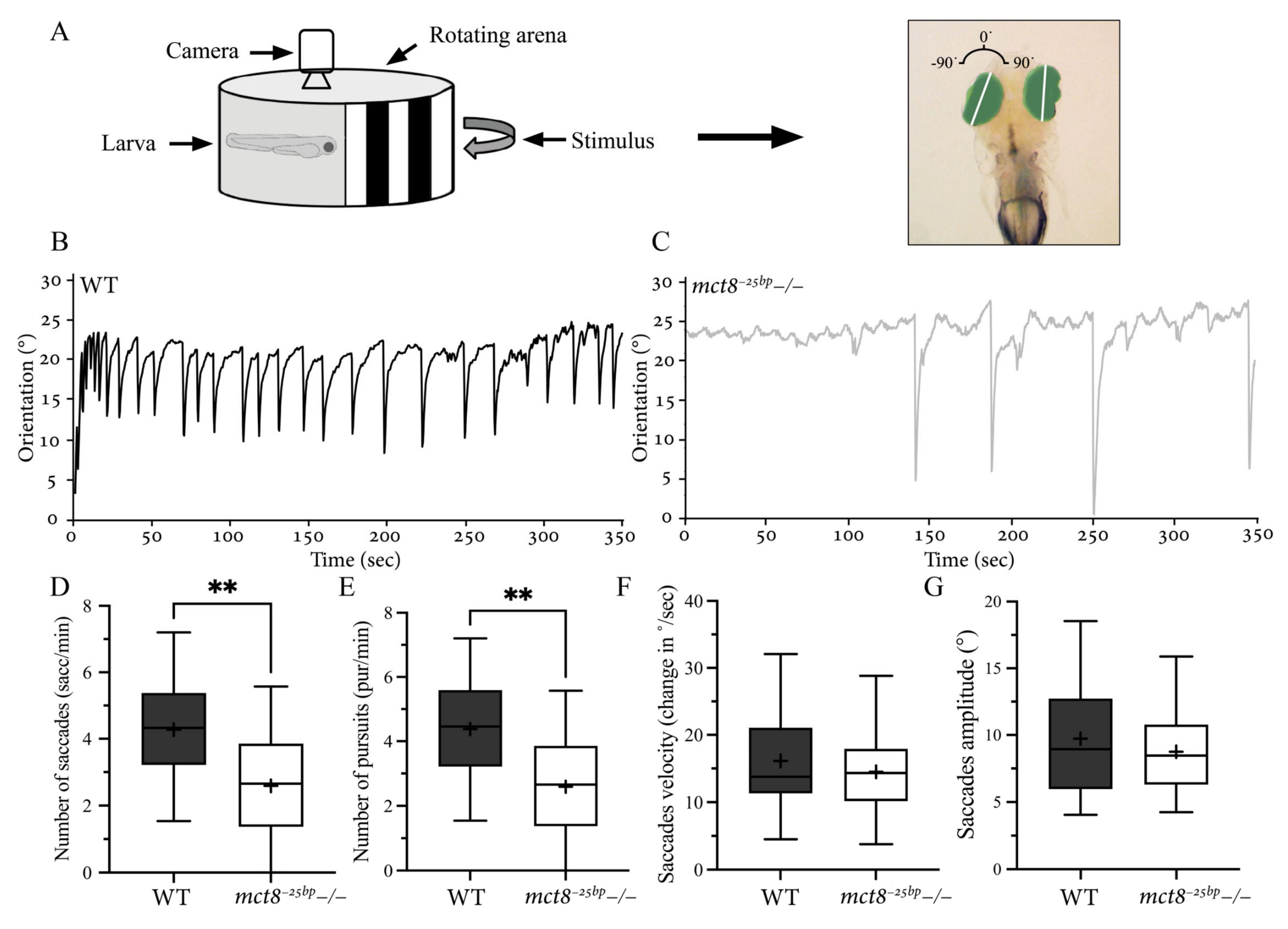
2.3. The Number of Conjugated Eye Movements Is Reduced in mct8−25bp−/− Larvae
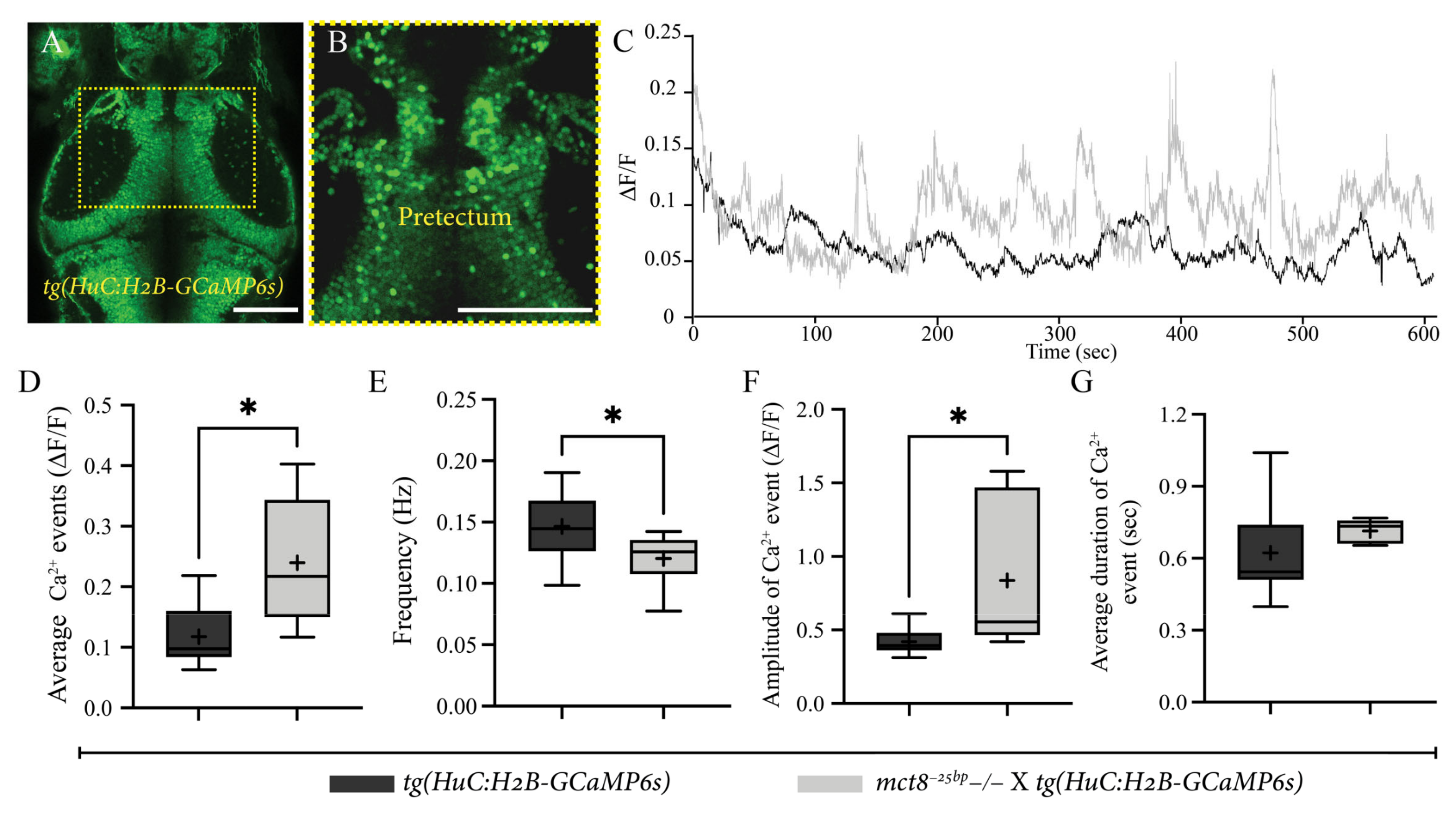
2.4. Altered Spontaneous Neuronal Activity in the Pretectum of mct8−25bp−/− Larvae
3. Discussion
4. Materials & Methods
4.1. Zebrafish Husbandry, Mutant, and Transgenic Lines
4.2. Whole Transcriptome RNA-seq and Pathway Analysis
4.3. Quantitative Real-Time PCR
4.4. Whole-Mount In Situ Hybridization (ISH) and Probe Preparation
4.5. MMI Pharmacological and Transient Expression Assays
4.6. Imaging and Image Analysis
4.7. OKR Assay
4.8. Genotyping
4.9. Data and Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Bernal, J. Thyroid Hormones in Brain Development and Function. In Endotext; Feingold, K.R., Anawalt, B., Boyce, A., Chrousos, G., de Herder, W.W., Dhatariya, K., Dungan, K., Grossman, A., Hershman, J.M., Hofland, J., et al., Eds.; MDText.com, Inc.: South Dartmouth, MA, USA, 2000. [Google Scholar]
- Weiss, R.E.; Refetoff, S. Effect of Thyroid Hormone On Growth: Lessons from the Syndrome of Resistance to Thyroid Hormone. Endocrinol. Metab. Clin. N. Am. 1996, 25, 719–730. [Google Scholar] [CrossRef]
- Gruters, A.; Krude, H.; Biebermann, H. Molecular Genetic Defects in Congenital Hypothyroidism. Eur. J. Endocrinol. 2004, 151, U39–U44. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ausó, E.; Lavado-Autric, R.; Cuevas, E.; del Rey, F.E.; de Escobar, G.M.; Berbel, P. A Moderate and Transient Deficiency of Maternal Thyroid Function at the Beginning of Fetal Neocorticogenesis Alters Neuronal Migration. Endocrinology 2004, 145, 4037–4047. [Google Scholar] [CrossRef]
- Friesema, E.C.H.; Ganguly, S.; Abdalla, A.; Fox, J.E.M.; Halestrap, A.P.; Visser, T.J. Identification of Monocarboxylate Transporter 8 as a Specific Thyroid Hormone Transporter. J. Biol. Chem. 2003, 278, 40128–40135. [Google Scholar] [CrossRef] [PubMed]
- Dumitrescu, A.M.; Liao, X.-H.; Best, T.B.; Brockmann, K.; Refetoff, S. A Novel Syndrome Combining Thyroid and Neurological Abnormalities Is Associated with Mutations in a Monocarboxylate Transporter Gene. Am. J. Hum. Genet. 2004, 74, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Heuer, H.; Visser, T.J. Minireview: Pathophysiological Importance of Thyroid Hormone Transporters. Endocrinology 2009, 150, 1078–1083. [Google Scholar] [CrossRef] [PubMed]
- Brockmann, K.; Dumitrescu, A.M.; Best, T.T.; Hanefeld, F.; Refetoff, S. X-Linked Paroxysmal Dyskinesia and Severe Global Retardation Caused by Defective MCT8 Gene. J. Neurol. 2005, 252, 663–666. [Google Scholar] [CrossRef] [PubMed]
- Biebermann, H.; Ambrugger, P.; Tarnow, P.; von Moers, A.; Schweizer, U.; Grueters, A. Extended Clinical Phenotype, Endocrine Investigations and Functional Studies of a Loss-of-Function Mutation A150V in the Thyroid Hormone Specific Transporter MCT8. Eur. J. Endocrinol. 2005, 153, 359–366. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Findlay, J.B.; Pappin, D.J. The Opsin Family of Proteins. Biochem. J. 1986, 238, 625–642. [Google Scholar] [CrossRef]
- Koyanagi, M.; Takano, K.; Tsukamoto, H.; Ohtsu, K.; Tokunaga, F.; Terakita, A. Jellyfish Vision Starts with CAMP Signaling Mediated by Opsin-Gs Cascade. Proc. Natl. Acad. Sci. USA 2008, 105, 15576–15580. [Google Scholar] [CrossRef]
- Roberts, M.R.; Srinivas, M.; Forrest, D.; de Escobar, G.M.; Reh, T.A. Making the Gradient: Thyroid Hormone Regulates Cone Opsin Expression in the Developing Mouse Retina. Proc. Natl. Acad. Sci. USA 2006, 103, 6218–6223. [Google Scholar] [CrossRef]
- Pessôa, C.N.; Santiago, L.A.; Santiago, D.A.; Machado, D.S.; Rocha, F.A.F.; Ventura, D.F.; Hokoç, J.N.; Pazos-Moura, C.C.; Wondisford, F.E.; Gardino, P.F.; et al. Thyroid Hormone Action Is Required for Normal Cone Opsin Expression during Mouse Retinal Development. Investig. Ophthalmol. Vis. Sci. 2008, 49, 2039–2045. [Google Scholar] [CrossRef] [PubMed]
- Eldred, K.C.; Hadyniak, S.E.; Hussey, K.A.; Brenerman, B.; Zhang, P.-W.; Chamling, X.; Sluch, V.M.; Welsbie, D.S.; Hattar, S.; Taylor, J.; et al. Thyroid Hormone Signaling Specifies Cone Subtypes in Human Retinal Organoids. Science 2018, 362, eaau6348. [Google Scholar] [CrossRef] [PubMed]
- Mayerl, S.; Müller, J.; Bauer, R.; Richert, S.; Kassmann, C.M.; Darras, V.M.; Buder, K.; Boelen, A.; Visser, T.J.; Heuer, H. Transporters MCT8 and OATP1C1 Maintain Murine Brain Thyroid Hormone Homeostasis. J. Clin. Investig. 2014, 124, 1987–1999. [Google Scholar] [CrossRef]
- Liao, X.-H.; Di Cosmo, C.; Dumitrescu, A.M.; Hernandez, A.; Van Sande, J.; St. Germain, D.L.; Weiss, R.E.; Galton, V.A.; Refetoff, S. Distinct Roles of Deiodinases on the Phenotype of Mct8 Defect: A Comparison of Eight Different Mouse Genotypes. Endocrinology 2011, 152, 1180–1191. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bárez-López, S.; Grijota-Martínez, C.; Ausó, E.; Fernández-de Frutos, M.; Montero-Pedrazuela, A.; Guadaño-Ferraz, A. Adult Mice Lacking Mct8 and Dio2 Proteins Present Alterations in Peripheral Thyroid Hormone Levels and Severe Brain and Motor Skill Impairments. Thyroid 2019, 29, 1669–1682. [Google Scholar] [CrossRef] [PubMed]
- Wirth, E.K.; Roth, S.; Blechschmidt, C.; Hölter, S.M.; Becker, L.; Racz, I.; Zimmer, A.; Klopstock, T.; Gailus-Durner, V.; Fuchs, H.; et al. Neuronal 3′,3,5-Triiodothyronine (T3) Uptake and Behavioral Phenotype of Mice Deficient in Mct8, the Neuronal T3 Transporter Mutated in Allan-Herndon-Dudley Syndrome. J. Neurosci. 2009, 29, 9439–9449. [Google Scholar] [CrossRef] [PubMed]
- Dumitrescu, A.M.; Liao, X.-H.; Weiss, R.E.; Millen, K.; Refetoff, S. Tissue-Specific Thyroid Hormone Deprivation and Excess in Monocarboxylate Transporter (Mct) 8-Deficient Mice. Endocrinology 2006, 147, 4036–4043. [Google Scholar] [CrossRef]
- Friesema, E.C.H.; Jansen, J.; Heuer, H.; Trajkovic, M.; Bauer, K.; Visser, T.J. Mechanisms of Disease: Psychomotor Retardation and High T3 Levels Caused by Mutations in Monocarboxylate Transporter 8. Nat. Clin. Pract. Endocrinol. Metab. 2006, 2, 512–523. [Google Scholar] [CrossRef]
- Porazzi, P.; Calebiro, D.; Benato, F.; Tiso, N.; Persani, L. Thyroid Gland Development and Function in the Zebrafish Model. Mol. Cell. Endocrinol. 2009, 312, 14–23. [Google Scholar] [CrossRef]
- Vancamp, P.; Houbrechts, A.M.; Darras, V.M. Insights from Zebrafish Deficiency Models to Understand the Impact of Local Thyroid Hormone Regulator Action on Early Development. Gen. Comp. Endocrinol. 2019, 279, 45–52. [Google Scholar] [CrossRef]
- Campinho, M.A.; Saraiva, J.; Florindo, C.; Power, D.M. Maternal Thyroid Hormones Are Essential for Neural Development in Zebrafish. Mol. Endocrinol. 2014, 28, 1136–1149. [Google Scholar] [CrossRef]
- Zada, D.; Tovin, A.; Lerer-Goldshtein, T.; Vatine, G.D.; Appelbaum, L. Altered Behavioral Performance and Live Imaging of Circuit-Specific Neural Deficiencies in a Zebrafish Model for Psychomotor Retardation. PLoS Genet. 2014, 10, e1004615. [Google Scholar] [CrossRef]
- Zada, D.; Tovin, A.; Lerer-Goldshtein, T.; Appelbaum, L. Pharmacological Treatment and BBB-Targeted Genetic Therapy for MCT8-Dependent Hypomyelination in Zebrafish. Dis. Models Mech. 2016, 9, 1339–1348. [Google Scholar] [CrossRef]
- Zada, D.; Blitz, E.; Appelbaum, L. Zebrafish—An Emerging Model to Explore Thyroid Hormone Transporters and Psychomotor Retardation. Mol. Cell. Endocrinol. 2017, 459, 53–58. [Google Scholar] [CrossRef]
- Visser, W.E.; Swagemakers, S.M.A.; Őzgűr, Z.; Schot, R.; Verheijen, F.W.; van Ijcken, W.F.J.; van der Spek, P.J.; Visser, T.J. Transcriptional Profiling of Fibroblasts from Patients with Mutations in MCT8 and Comparative Analysis with the Human Brain Transcriptome. Hum. Mol. Genet. 2010, 19, 4189–4200. [Google Scholar] [CrossRef]
- Silva, N.; Louro, B.; Trindade, M.; Power, D.M.; Campinho, M.A. Transcriptomics Reveal an Integrative Role for Maternal Thyroid Hormones during Zebrafish Embryogenesis. Sci. Rep. 2017, 7, 16657. [Google Scholar] [CrossRef]
- Morte, B.; Gil-Ibañez, P.; Heuer, H.; Bernal, J. Brain Gene Expression in Systemic Hypothyroidism and Mouse Models of MCT8 Deficiency: The Mct8-Oatp1c1-Dio2 Triad. Thyroid 2021, 31, 985–993. [Google Scholar] [CrossRef]
- Ramos-Lopez, E.; Kurylowicz, A.; Bednarczuk, T.; Paunkovic, J.; Seidl, C.; Badenhoop, K. Vitamin D Receptor Polymorphisms Are Associated with Graves’ Disease in German and Polish But Not in Serbian Patients. Thyroid 2005, 15, 1125–1130. [Google Scholar] [CrossRef]
- Cook, I.; Wang, T.; Leyh, T.S. Tetrahydrobiopterin Regulates Monoamine Neurotransmitter Sulfonation. Proc. Natl. Acad. Sci. USA 2017, 114, E5317–E5324. [Google Scholar] [CrossRef]
- Davidson, B.; Soodak, M.; Neary, J.T.; Strout, H.V.; Kieffer, J.D.; Mover, H.; Maloof, F. The Irreversible Inactivation of Thyroid Peroxidase by Methylmercaptoimidazole, Thiouracil, and Propylthiouracil in Vitro and Its Relationship to in Vivo Findings. Endocrinology 1978, 103, 871–882. [Google Scholar] [CrossRef]
- Brockerhoff, S.E.; Hurley, J.B.; Janssen-Bienhold, U.; Neuhauss, S.C.; Driever, W.; Dowling, J.E. A Behavioral Screen for Isolating Zebrafish Mutants with Visual System Defects. Proc. Natl. Acad. Sci. USA 1995, 92, 10545–10549. [Google Scholar] [CrossRef]
- Neuhauss, S.C.F.; Biehlmaier, O.; Seeliger, M.W.; Das, T.; Kohler, K.; Harris, W.A.; Baier, H. Genetic Disorders of Vision Revealed by a Behavioral Screen of 400 Essential Loci in Zebrafish. J. Neurosci. 1999, 19, 8603–8615. [Google Scholar] [CrossRef]
- Huang, M.Y.-Y.; Neuhauss, S. The Optokinetic Response in Zebrafish and Its Applications. Front. Biosci. J. Virtual Libr. 2008, 13, 1899–1916. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Hinz, J.; Zhang, Y.; Thiele, T.R.; Arrenberg, A.B. Parallel Channels for Motion Feature Extraction in the Pretectum and Tectum of Larval Zebrafish. Cell Rep. 2020, 30, 442–453.e6. [Google Scholar] [CrossRef] [PubMed]
- Reber, A.; Lannou, J.; Hess, B.J. Development of Optokinetic Neuronal Responses in the Pretectum and Horizontal Optokinetic Nystagmus in Unilaterally Enucleated Rats. Arch. Ital. Biol. 1989, 127, 225–242. [Google Scholar] [PubMed]
- Tsujimura, T.; Masuda, R.; Ashino, R.; Kawamura, S. Spatially Differentiated Expression of Quadruplicated Green-Sensitive RH2 Opsin Genes in Zebrafish Is Determined by Proximal Regulatory Regions and Gene Order to the Locus Control Region. BMC Genet. 2015, 16, 130. [Google Scholar] [CrossRef]
- Glaschke, A.; Weiland, J.; Turco, D.D.; Steiner, M.; Peichl, L.; Glösmann, M. Thyroid Hormone Controls Cone Opsin Expression in the Retina of Adult Rodents. J. Neurosci. 2011, 31, 4844–4851. [Google Scholar] [CrossRef]
- Sjoberg, M.; Vennstrom, B.; Forrest, D. Thyroid Hormone Receptors in Chick Retinal Development: Differential Expression of MRNAs for Alpha and N-Terminal Variant Beta Receptors. Development 1992, 114, 39–47. [Google Scholar] [CrossRef]
- Sevilla-Romero, E.; Muñoz, A.; Pinazo-Durán, M.D. Low Thyroid Hormone Levels Impair the Perinatal Development of the Rat Retina. ORE 2002, 34, 181–191. [Google Scholar] [CrossRef]
- Vancamp, P.; Bourgeois, N.M.A.; Houbrechts, A.M.; Darras, V.M. Knockdown of the Thyroid Hormone Transporter MCT8 in Chicken Retinal Precursor Cells Hampers Early Retinal Development and Results in a Shift towards More UV/Blue Cones at the Expense of Green/Red Cones. Exp. Eye Res. 2019, 178, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Land, M.F. Eye Movements of Vertebrates and Their Relation to Eye Form and Function. J. Comp. Physiol. A 2015, 201, 195–214. [Google Scholar] [CrossRef] [PubMed]
- Collery, R.F.; Volberding, P.J.; Bostrom, J.R.; Link, B.A.; Besharse, J.C. Loss of Zebrafish Mfrp Causes Nanophthalmia, Hyperopia, and Accumulation of Subretinal Macrophages. Investig. Ophthalmol. Vis. Sci. 2016, 57, 6805–6814. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Hinz, J.; Haikala, V.; Reiff, D.F.; Arrenberg, A.B. Selective Processing of All Rotational and Translational Optic Flow Directions in the Zebrafish Pretectum and Tectum. BMC Biol. 2019, 17, 29. [Google Scholar] [CrossRef]
- Missal, M.; Coimbra, A.; Lefèvre, P.; Olivier, E. A Quantitative Analysis of the Correlations between Eye Movements and Neural Activity in the Pretectum. Exp. Brain Res. 2002, 143, 373–382. [Google Scholar] [CrossRef]
- Kubo, F.; Hablitzel, B.; Dal Maschio, M.; Driever, W.; Baier, H.; Arrenberg, A.B. Functional Architecture of an Optic Flow-Responsive Area That Drives Horizontal Eye Movements in Zebrafish. Neuron 2014, 81, 1344–1359. [Google Scholar] [CrossRef]
- Naumann, E.A.; Fitzgerald, J.E.; Dunn, T.W.; Rihel, J.; Sompolinsky, H.; Engert, F. From Whole-Brain Data to Functional Circuit Models: The Zebrafish Optomotor Response. Cell 2016, 167, 947–960.e20. [Google Scholar] [CrossRef]
- Wu, Y.; dal Maschio, M.; Kubo, F.; Baier, H. An Optical Illusion Pinpoints an Essential Circuit Node for Global Motion Processing. Neuron 2020, 108, 722–734.e5. [Google Scholar] [CrossRef]
- Levitas-Djerbi, T.; Appelbaum, L. Modeling Sleep and Neuropsychiatric Disorders in Zebrafish. Curr. Opin. Neurobiol. 2017, 44, 89–93. [Google Scholar] [CrossRef]
- Pérez-Schuster, V.; Kulkarni, A.; Nouvian, M.; Romano, S.A.; Lygdas, K.; Jouary, A.; Dipoppa, M.; Pietri, T.; Haudrechy, M.; Candat, V.; et al. Sustained Rhythmic Brain Activity Underlies Visual Motion Perception in Zebrafish. Cell Rep. 2016, 17, 1098–1112. [Google Scholar] [CrossRef]
- Fernandes, A.M.; Mearns, D.S.; Donovan, J.C.; Larsch, J.; Helmbrecht, T.O.; Kölsch, Y.; Laurell, E.; Kawakami, K.; dal Maschio, M.; Baier, H. Neural Circuitry for Stimulus Selection in the Zebrafish Visual System. Neuron 2021, 109, 805–822.e6. [Google Scholar] [CrossRef]
- Förster, D.; Helmbrecht, T.O.; Mearns, D.S.; Jordan, L.; Mokayes, N.; Baier, H. Retinotectal Circuitry of Larval Zebrafish Is Adapted to Detection and Pursuit of Prey. eLife 2020, 9, e58596. [Google Scholar] [CrossRef]
- Admati, I.; Wasserman-Bartov, T.; Tovin, A.; Rozenblat, R.; Blitz, E.; Zada, D.; Lerer-Goldshtein, T.; Appelbaum, L. Neural Alterations and Hyperactivity of the Hypothalamic-Pituitary-Thyroid Axis in Oatp1c1 Deficiency. Thyroid 2020, 30, 161–174. [Google Scholar] [CrossRef]
- Grijota-Martínez, C.; Bárez-López, S.; Gómez-Andrés, D.; Guadaño-Ferraz, A. MCT8 Deficiency: The Road to Therapies for a Rare Disease. Front. Neurosci. 2020, 14, 380. [Google Scholar] [CrossRef]
- Van Geest, F.S.; Gunhanlar, N.; Groeneweg, S.; Visser, W.E. Monocarboxylate Transporter 8 Deficiency: From Pathophysiological Understanding to Therapy Development. Front. Endocrinol. 2021, 12, 723750. [Google Scholar] [CrossRef]
- Iwayama, H.; Liao, X.-H.; Braun, L.; Bárez-López, S.; Kaspar, B.; Weiss, R.E.; Dumitrescu, A.M.; Guadaño-Ferraz, A.; Refetoff, S. Adeno Associated Virus 9-Based Gene Therapy Delivers a Functional Monocarboxylate Transporter 8, Improving Thyroid Hormone Availability to the Brain of Mct8-Deficient Mice. Thyroid 2016, 26, 1311–1319. [Google Scholar] [CrossRef]
- Vatine, G.D.; Al-Ahmad, A.; Barriga, B.K.; Svendsen, S.; Salim, A.; Garcia, L.; Garcia, V.J.; Ho, R.; Yucer, N.; Qian, T.; et al. Modeling Psychomotor Retardation Using IPSCs from MCT8-Deficient Patients Indicates a Prominent Role for the Blood-Brain Barrier. Cell Stem Cell 2017, 20, 831–843.e5. [Google Scholar] [CrossRef]
- Elbaz, I.; Yelin-Bekerman, L.; Nicenboim, J.; Vatine, G.; Appelbaum, L. Genetic Ablation of Hypocretin Neurons Alters Behavioral State Transitions in Zebrafish. J. Neurosci. 2012, 32, 12961–12972. [Google Scholar] [CrossRef]
- Freeman, J.; Vladimirov, N.; Kawashima, T.; Mu, Y.; Sofroniew, N.J.; Bennett, D.V.; Rosen, J.; Yang, C.-T.; Looger, L.L.; Ahrens, M.B. Mapping Brain Activity at Scale with Cluster Computing. Nat. Methods 2014, 11, 941–950. [Google Scholar] [CrossRef]
- Hwang, W.Y.; Fu, Y.; Reyon, D.; Maeder, M.L.; Tsai, S.Q.; Sander, J.D.; Peterson, R.T.; Yeh, J.-R.J.; Joung, J.K. Efficient Genome Editing in Zebrafish Using a CRISPR-Cas System. Nat. Biotechnol. 2013, 31, 227–229. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python Framework to Work with High-Throughput Sequencing Data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential Expression Analysis for Sequence Count Data. Nat. Prec. 2010, 1. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing Real-Time PCR Data by the Comparative CT Method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Yelin-Bekerman, L.; Elbaz, I.; Diber, A.; Dahary, D.; Gibbs-Bar, L.; Alon, S.; Lerer-Goldshtein, T.; Appelbaum, L. Hypocretin Neuron-Specific Transcriptome Profiling Identifies the Sleep Modulator Kcnh4a. eLife 2015, 4, e08638. [Google Scholar] [CrossRef]
- Scheetz, S.D.; Shao, E.; Zhou, Y.; Cario, C.L.; Bai, Q.; Burton, E.A. An Open-Source Method to Analyze Optokinetic Reflex Responses in Larval Zebrafish. J. Neurosci. Methods 2018, 293, 329–337. [Google Scholar] [CrossRef]
- Kossack, M.E.; Draper, B.W. Genetic Regulation of Sex Determination and Maintenance in Zebrafish (Danio Rerio). Curr. Top. Dev. Biol. 2019, 134, 119–149. [Google Scholar] [CrossRef]
- Romano, S.A.; Pérez-Schuster, V.; Jouary, A.; Boulanger-Weill, J.; Candeo, A.; Pietri, T.; Sumbre, G. An Integrated Calcium Imaging Processing Toolbox for the Analysis of Neuronal Population Dynamics. PLoS Comput. Biol. 2017, 13, e1005526. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rozenblat, R.; Tovin, A.; Zada, D.; Lebenthal-Loinger, I.; Lerer-Goldshtein, T.; Appelbaum, L. Genetic and Neurological Deficiencies in the Visual System of mct8 Mutant Zebrafish. Int. J. Mol. Sci. 2022, 23, 2464. https://doi.org/10.3390/ijms23052464
Rozenblat R, Tovin A, Zada D, Lebenthal-Loinger I, Lerer-Goldshtein T, Appelbaum L. Genetic and Neurological Deficiencies in the Visual System of mct8 Mutant Zebrafish. International Journal of Molecular Sciences. 2022; 23(5):2464. https://doi.org/10.3390/ijms23052464
Chicago/Turabian StyleRozenblat, Rotem, Adi Tovin, David Zada, Ilana Lebenthal-Loinger, Tali Lerer-Goldshtein, and Lior Appelbaum. 2022. "Genetic and Neurological Deficiencies in the Visual System of mct8 Mutant Zebrafish" International Journal of Molecular Sciences 23, no. 5: 2464. https://doi.org/10.3390/ijms23052464
APA StyleRozenblat, R., Tovin, A., Zada, D., Lebenthal-Loinger, I., Lerer-Goldshtein, T., & Appelbaum, L. (2022). Genetic and Neurological Deficiencies in the Visual System of mct8 Mutant Zebrafish. International Journal of Molecular Sciences, 23(5), 2464. https://doi.org/10.3390/ijms23052464





