COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19
Abstract
:1. Introduction
2. Proteomics of COVID-19
2.1. Plasma Proteomics Studies
2.2. Serum Proteomics Studies
2.3. Infected-Cells Proteomics Studies
3. Metabolomics of COVID-19
3.1. Plasma Metabolomics Studies
3.2. Serum Metabolomics Studies
4. Multiomics Studies of COVID-19
5. COVIDomics Data Integration
6. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Costanzo, M.; De Giglio, M.A.R.; Roviello, G.N. Anti-Coronavirus Vaccines: Past Investigations on SARS-CoV-1 and MERS-CoV, the Approved Vaccines from BioNTech/Pfizer, Moderna, Oxford/AstraZeneca and others under Development Against SARS-CoV- 2 Infection. Curr. Med. Chem. 2022, 29, 4–18. [Google Scholar] [CrossRef] [PubMed]
- De Giglio, M.A.R.; Roviello, G.N. SARS-CoV-2: Recent Reports on Antiviral Therapies Based on Lopinavir/Ritonavir, Darunavir/Umifenovir, Hydroxychloroquine, Remdesivir, Favipiravir and other Drugs for the Treatment of the New Coronavirus. Curr. Med. Chem. 2020, 27, 4536–4541. [Google Scholar] [CrossRef]
- Rehman, S.U.; Rehman, S.U.; Yoo, H.H. COVID-19 challenges and its therapeutics. Biomed. Pharmacother. 2021, 142, 112015. [Google Scholar] [CrossRef] [PubMed]
- Singh, T.U.; Parida, S.; Lingaraju, M.C.; Kesavan, M.; Kumar, D.; Singh, R.K. Drug repurposing approach to fight COVID-19. Pharmacol. Rep. 2020, 72, 1479–1508. [Google Scholar] [CrossRef]
- Borbone, N.; Piccialli, G.; Roviello, G.N.; Oliviero, G. Nucleoside Analogs and Nucleoside Precursors as Drugs in the Fight against SARS-CoV-2 and Other Coronaviruses. Molecules 2021, 26, 986. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Nalbandian, A.; Sehgal, K.; Gupta, A.; Madhavan, M.V.; McGroder, C.; Stevens, J.S.; Cook, J.R.; Nordvig, A.S.; Shalev, D.; Sehrawat, T.S.; et al. Post-acute COVID-19 syndrome. Nat. Med. 2021, 27, 601–615. [Google Scholar] [CrossRef]
- Marshall, J.C.; Murthy, S.; Diaz, J.; Adhikari, N.K.; Angus, D.C.; Arabi, Y.M.; Baillie, K.; Bauer, M.; Berry, S.; Blackwood, B.; et al. A minimal common outcome measure set for COVID-19 clinical research. Lancet Infect. Dis. 2020, 20, e192–e197. [Google Scholar] [CrossRef]
- Zhang, Y.-Z.; Holmes, E.C. A Genomic Perspective on the Origin and Emergence of SARS-CoV-2. Cell 2020, 181, 223–227. [Google Scholar] [CrossRef]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef]
- Bell, L.C.; Meydan, C.; Kim, J.; Foox, J.; Butler, D.; Mason, C.E.; Shapira, S.D.; Noursadeghi, M.; Pollara, G. Transcriptional response modules characterize IL-1β and IL-6 activity in COVID-19. iScience 2021, 24, 101896. [Google Scholar] [CrossRef] [PubMed]
- Fagone, P.; Ciurleo, R.; Lombardo, S.D.; Iacobello, C.; Palermo, C.I.; Shoenfeld, Y.; Bendtzen, K.; Bramanti, P.; Nicoletti, F. Transcriptional landscape of SARS-CoV-2 infection dismantles pathogenic pathways activated by the virus, proposes unique sex-specific differences and predicts tailored therapeutic strategies. Autoimmun. Rev. 2020, 19, 102571. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.-C.; Clark, D.J.; Schnaubelt, M.; Teo, G.C.; Leprevost, F.D.V.; Bocik, W.; Boja, E.S.; Hiltke, T.; Nesvizhskii, A.I.; Zhang, H. Deep Proteomics Using Two Dimensional Data Independent Acquisition Mass Spectrometry. Anal. Chem. 2020, 92, 4217–4225. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, Y.; Watanabe, E.; Umeyama, T.; Nakajima, D.; Hattori, M.; Honda, K.; Ohara, O. Optimization of Data-Independent Acquisition Mass Spectrometry for Deep and Highly Sensitive Proteomic Analysis. Int. J. Mol. Sci. 2019, 20, 5932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakajima, D.; Ohara, O.; Kawashima, Y. Data-Independent Acquisition Mass Spectrometry-Based Deep Proteome Analysis for Hydrophobic Proteins from Dried Blood Spots Enriched by Sodium Carbonate Precipitation. In Clinical Proteomics; Humana: New Yor, NY, USA, 2022; pp. 39–52. ISBN 978-1-0716-1935-3. [Google Scholar]
- Costanzo, M.; Caterino, M.; Cevenini, A.; Kollipara, L.; Shevchuk, O.; Nguyen, C.D.L.; Sickmann, A.; Ruoppolo, M. Data Independent Acquisition Mass Spectrometry for Proteomic Advances into Isolated Methylmalonic Acidemia. In NATO Science for Peace and Security Series A: Chemistry and Biology; Springer: Dordrecht, The Netherlands, 2020; pp. 221–223. [Google Scholar]
- Barigazzi, E.; Santorelli, L.; Morello, W.; Raimondo, F.; Crapella, B.; Ghio, L.; Tamburello, C.; Montini, G.; Pitto, M. New Insight into Idiopathic Nephrotic Syndrome: Strategy Based on Urinary Exosomes. In NATO Science for Peace and Security Series A: Chemistry and Biology; Springer: Dordrecht, The Netherlands, 2020; pp. 217–218. [Google Scholar]
- Santorelli, L.; Stella, M.; Chinello, C.; Capitoli, G.; Piga, I.; Smith, A.; Grasso, A.; Grasso, M.; Bovo, G.; Magni, F. Does the Urinary Proteome Reflect ccRCC Stage and Grade Progression? Diagnostics 2021, 11, 2369. [Google Scholar] [CrossRef]
- Kong, S.W.; Hernandez-Ferrer, C. Assessment of coverage for endogenous metabolites and exogenous chemical compounds using an untargeted metabolomics platform. Pac. Symp. Biocomput. 2020, 25, 587–598. [Google Scholar]
- Manganelli, V.; Salvatori, I.; Costanzo, M.; Capozzi, A.; Caissutti, D.; Caterino, M.; Valle, C.; Ferri, A.; Sorice, M.; Ruoppolo, M.; et al. Overexpression of Neuroglobin Promotes Energy Metabolism and Autophagy Induction in Human Neuroblastoma SH-SY5Y Cells. Cells 2021, 10, 3394. [Google Scholar] [CrossRef]
- Costanzo, M.; Caterino, M.; Cevenini, A.; Jung, V.; Chhuon, C.; Lipecka, J.; Fedele, R.; Guerrera, I.C.; Ruoppolo, M. Dataset of a comparative proteomics experiment in a methylmalonyl-CoA mutase knockout HEK 293 cell model. Data Brief 2020, 33, 106453. [Google Scholar] [CrossRef]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef]
- Giacco, A.; Paoli, G.D.; Senese, R.; Cioffi, F.; Silvestri, E.; Moreno, M.; Ruoppolo, M.; Caterino, M.; Costanzo, M.; Lombardi, A.; et al. The saturation degree of fatty acids and their derived acylcarnitines determines the direct effect of metabolically active thyroid hormones on insulin sensitivity in skeletal muscle cells. FASEB J. 2018, 33, 1811–1823. [Google Scholar] [CrossRef] [Green Version]
- Ruoppolo, M.; Caterino, M.; Albano, L.; Pecce, R.; Di Girolamo, M.G.; Crisci, D.; Costanzo, M.; Milella, L.; Franconi, F.; Campesi, I. Targeted metabolomic profiling in rat tissues reveals sex differences. Sci. Rep. 2018, 8, 4663. [Google Scholar] [CrossRef] [PubMed]
- De Pasquale, V.; Caterino, M.; Costanzo, M.; Fedele, R.; Ruoppolo, M.; Pavone, L.M. Targeted Metabolomic Analysis of a Mucopolysaccharidosis IIIB Mouse Model Reveals an Imbalance of Branched-Chain Amino Acid and Fatty Acid Metabolism. Int. J. Mol. Sci. 2020, 21, 4211. [Google Scholar] [CrossRef] [PubMed]
- Heiles, S. Advanced tandem mass spectrometry in metabolomics and lipidomics—methods and applications. Anal. Bioanal. Chem. 2021, 413, 5927–5948. [Google Scholar] [CrossRef] [PubMed]
- Rampler, E.; El Abiead, Y.; Schoeny, H.; Rusz, M.; Hildebrand, F.; Fitz, V.; Koellensperger, G. Recurrent Topics in Mass Spectrometry-Based Metabolomics and Lipidomics—Standardization, Coverage, and Throughput. Anal. Chem. 2020, 93, 519–545. [Google Scholar] [CrossRef]
- Abu-Farha, M.; Thanaraj, T.A.; Qaddoumi, M.G.; Hashem, A.; Abubaker, J.; Al-Mulla, F. The Role of Lipid Metabolism in COVID-19 Virus Infection and as a Drug Target. Int. J. Mol. Sci. 2020, 21, 3544. [Google Scholar] [CrossRef]
- Galbraith, M.D.; Kinning, K.T.; Sullivan, K.D.; Baxter, R.; Araya, P.; Jordan, K.R.; Russell, S.; Smith, K.P.; Granrath, R.E.; Shaw, J.R.; et al. Seroconversion stages COVID19 into distinct pathophysiological states. eLife 2021, 10. [Google Scholar] [CrossRef]
- Li, C.-X.; Gao, J.; Zhang, Z.; Chen, L.; Li, X.; Zhou, M.; Wheelock, Å.M. Multiomics integration-based molecular characterizations of COVID-19. Brief. Bioinform. 2021, 23, bbab485. [Google Scholar] [CrossRef]
- Bhawal, R.; Oberg, A.L.; Zhang, S.; Kohli, M. Challenges and Opportunities in Clinical Applications of Blood-Based Proteomics in Cancer. Cancers 2020, 12, 2428. [Google Scholar] [CrossRef]
- Caterino, M.; Ruoppolo, M.; Costanzo, M.; Albano, L.; Crisci, D.; Sotgiu, G.; Saderi, L.; Montella, A.; Franconi, F.; Campesi, I. Sex Affects Human Premature Neonates’ Blood Metabolome According to Gestational Age, Parenteral Nutrition, and Caffeine Treatment. Metabolites 2021, 11, 158. [Google Scholar] [CrossRef]
- Assarsson, E.; Lundberg, M.; Holmquist, G.; Björkesten, J.; Thorsen, S.B.; Ekman, D.; Eriksson, A.; Dickens, E.R.; Ohlsson, S.; Edfeldt, G.; et al. Homogenous 96-Plex PEA Immunoassay Exhibiting High Sensitivity, Specificity, and Excellent Scalability. PLoS ONE 2014, 9, e95192. [Google Scholar] [CrossRef] [Green Version]
- Patel, H.; Ashton, N.J.; Dobson, R.J.B.; Andersson, L.-M.; Yilmaz, A.; Blennow, K.; Gisslen, M.; Zetterberg, H. Proteomic blood profiling in mild, severe and critical COVID-19 patients. Sci. Rep. 2021, 11, 6357. [Google Scholar] [CrossRef] [PubMed]
- Haljasmägi, L.; Salumets, A.; Rumm, A.P.; Jürgenson, M.; Krassohhina, E.; Remm, A.; Sein, H.; Kareinen, L.; Vapalahti, O.; Sironen, T.; et al. Longitudinal proteomic profiling reveals increased early inflammation and sustained apoptosis proteins in severe COVID-19. Sci. Rep. 2020, 10, 20533. [Google Scholar] [CrossRef] [PubMed]
- Zhong, W.; Altay, O.; Arif, M.; Edfors, F.; Doganay, L.; Mardinoglu, A.; Uhlen, M.; Fagerberg, L. Next generation plasma proteome profiling of COVID-19 patients with mild to moderate symptoms. eBioMedicine 2021, 74, 103723. [Google Scholar] [CrossRef] [PubMed]
- Filbin, M.R.; Mehta, A.; Schneider, A.M.; Kays, K.R.; Guess, J.R.; Gentili, M.; Fenyves, B.G.; Charland, N.C.; Gonye, A.L.; Gushterova, I.; et al. Longitudinal proteomic analysis of severe COVID-19 reveals survival-associated signatures, tissue-specific cell death, and cell-cell interactions. Cell Rep. Med. 2021, 2, 100287. [Google Scholar] [CrossRef] [PubMed]
- Bauer, W.; Weber, M.; Diehl-Wiesenecker, E.; Galtung, N.; Prpic, M.; Somasundaram, R.; Tauber, R.; Schwenk, J.M.; Micke, P.; Kappert, K. Plasma Proteome Fingerprints Reveal Distinctiveness and Clinical Outcome of SARS-CoV-2 Infection. Viruses 2021, 13, 2456. [Google Scholar] [CrossRef] [PubMed]
- Shu, T.; Ning, W.; Wu, D.; Xu, J.; Han, Q.; Huang, M.; Zou, X.; Yang, Q.; Yuan, Y.; Bie, Y.; et al. Plasma Proteomics Identify Biomarkers and Pathogenesis of COVID-19. Immunity 2020, 53, 1108–1122.e5. [Google Scholar] [CrossRef]
- Park, J.; Kim, H.; Kim, S.Y.; Kim, Y.; Lee, J.-S.; Dan, K.; Seong, M.-W.; Han, D. In-depth blood proteome profiling analysis revealed distinct functional characteristics of plasma proteins between severe and non-severe COVID-19 patients. Sci. Rep. 2020, 10, 22418. [Google Scholar] [CrossRef]
- Messner, C.B.; Demichev, V.; Wendisch, D.; Michalick, L.; White, M.; Freiwald, A.; Textoris-Taube, K.; Vernardis, S.I.; Egger, A.-S.; Kreidl, M.; et al. Ultra-High-Throughput Clinical Proteomics Reveals Classifiers of COVID-19 Infection. Cell Syst. 2020, 11, 11–24. [Google Scholar] [CrossRef]
- Demichev, V.; Tober-Lau, P.; Lemke, O.; Nazarenko, T.; Thibeault, C.; Whitwell, H.; Röhl, A.; Freiwald, A.; Szyrwiel, L.; Ludwig, D.; et al. A time-resolved proteomic and prognostic map of COVID-19. Cell Syst. 2021, 12, 780–794.e7. [Google Scholar] [CrossRef]
- Hou, X.; Zhang, X.; Wu, X.; Lu, M.; Wang, D.; Xu, M.; Wang, H.; Liang, T.; Dai, J.; Duan, H.; et al. Serum Protein Profiling Reveals a Landscape of Inflammation and Immune Signaling in Early-stage COVID-19 Infection. Mol. Cell. Proteom. 2020, 19, 1749–1759. [Google Scholar] [CrossRef]
- Chen, Y.; Yao, H.; Zhang, N.; Wu, J.; Gao, S.; Guo, J.; Lu, X.; Cheng, L.; Luo, R.; Liang, X.; et al. Proteomic Analysis Identifies Prolonged Disturbances in Pathways Related to Cholesterol Metabolism and Myocardium Function in the COVID-19 Recovery Stage. J. Proteome Res. 2021, 20, 3463–3474. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Nakai, Y.; Shin, J.; Hara, M.; Takeda, Y.; Kubo, S.; Jeremiah, S.S.; Ino, Y.; Akiyama, T.; Moriyama, K.; et al. Identification of serum prognostic biomarkers of severe COVID-19 using a quantitative proteomic approach. Sci. Rep. 2021, 11, 20638. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Han, D.; Kim, S.Y.; Hong, K.H.; Jang, M.; Kim, M.J.; Kim, Y.; Park, J.H.; Cho, S.I.; Park, W.B.; et al. Longitudinal proteomic profiling provides insights into host response and proteome dynamics in COVID-19 progression. Proteomics 2021, 21, 2000278. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, A.; Thomas, T.; Dzieciatkowska, M.; Hill, R.C.; Francis, R.O.; Hudson, K.E.; Zimring, J.C.; Hod, E.A.; Spitalnik, S.L.; Hansen, K.C. Serum Proteomics in COVID-19 Patients: Altered Coagulation and Complement Status as a Function of IL-6 Level. J. Proteome Res. 2020, 19, 4417–4427. [Google Scholar] [CrossRef]
- Bojkova, D.; Klann, K.; Koch, B.; Widera, M.; Krause, D.; Ciesek, S.; Cinatl, J.; Münch, C. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature 2020, 583, 469–472. [Google Scholar] [CrossRef]
- Grenga, L.; Gallais, F.; Pible, O.; Gaillard, J.-C.; Gouveia, D.; Batina, H.; Bazaline, N.; Ruat, S.; Culotta, K.; Miotello, G.; et al. Shotgun proteomics analysis of SARS-CoV-2-infected cells and how it can optimize whole viral particle antigen production for vaccines. Emerg. Microbes Infect. 2020, 9, 1712–1721. [Google Scholar] [CrossRef]
- Bock, J.-O.; Ortea, I. Re-analysis of SARS-CoV-2-infected host cell proteomics time-course data by impact pathway analysis and network analysis: A potential link with inflammatory response. Aging 2020, 12, 11277–11286. [Google Scholar] [CrossRef]
- Danlos, F.-X.; Grajeda-Iglesias, C.; Durand, S.; Sauvat, A.; Roumier, M.; Cantin, D.; Colomba, E.; Rohmer, J.; Pommeret, F.; Baciarello, G.; et al. Metabolomic analyses of COVID-19 patients unravel stage-dependent and prognostic biomarkers. Cell Death Dis. 2021, 12, 258. [Google Scholar] [CrossRef]
- Blasco, H.; Bessy, C.; Plantier, L.; Lefevre, A.; Piver, E.; Bernard, L.; Marlet, J.; Stefic, K.; Benz-de Bretagne, I.; Cannet, P.; et al. The specific metabolome profiling of patients infected by SARS-CoV-2 supports the key role of tryptophan-nicotinamide pathway and cytosine metabolism. Sci. Rep. 2020, 10, 16824. [Google Scholar] [CrossRef]
- Danchin, A.; Marlière, P. Cytosine drives evolution of SARS-CoV-2. Environ. Microbiol. 2020, 22, 1977–1985. [Google Scholar] [CrossRef] [Green Version]
- Heer, C.D.; Sanderson, D.J.; Voth, L.S.; Alhammad, Y.M.; Schmidt, M.S.; Trammell, S.A.; Perlman, S.; Cohen, M.S.; Fehr, A.R.; Brenner, C. Coronavirus infection and PARP expression dysregulate the NAD metabolome: An actionable component of innate immunity. J. Biol. Chem. 2020, 295, 17986–17996. [Google Scholar] [CrossRef] [PubMed]
- Fraser, D.D.; Slessarev, M.; Martin, C.M.; Daley, M.; Patel, M.A.; Miller, M.R.; Patterson, E.K.; O’Gorman, D.B.; Gill, S.E.; Wishart, D.S.; et al. Metabolomics Profiling of Critically Ill Coronavirus Disease 2019 Patients: Identification of Diagnostic and Prognostic Biomarkers. Crit. Care Explor. 2020, 2, e0272. [Google Scholar] [CrossRef] [PubMed]
- Sindelar, M.; Stancliffe, E.; Schwaiger-Haber, M.; Anbukumar, D.S.; Adkins-Travis, K.; Goss, C.W.; O’Halloran, J.A.; Mudd, P.A.; Liu, W.-C.; Albrecht, R.A.; et al. Longitudinal Metabolomics of Human Plasma Reveals Prognostic Markers of COVID-19 Disease Severity. Cell Rep. Med. 2021, 2, 100369. [Google Scholar] [CrossRef] [PubMed]
- Bizkarguenaga, M.; Bruzzone, C.; Gil-Redondo, R.; SanJuan, I.; Martin-Ruiz, I.; Barriales, D.; Palacios, A.; Pasco, S.T.; González-Valle, B.; Laín, A.; et al. Uneven metabolic and lipidomic profiles in recovered COVID-19 patients as investigated by plasma NMR metabolomics. NMR Biomed. 2021, 35, e4637. [Google Scholar] [CrossRef]
- Khodadoust, M.M. Inferring a causal relationship between ceramide levels and COVID-19 respiratory distress. Sci. Rep. 2021, 11, 20866. [Google Scholar] [CrossRef]
- Raines, N.H.; Cheung, M.D.; Wilson, L.S.; Edberg, J.C.; Erdmann, N.B.; Schmaier, A.A.; Berryhill, T.F.; Manickas-Hill, Z.; Li, J.Z.; Yu, X.G.; et al. Nicotinamide Adenine Dinucleotide Biosynthetic Impairment and Urinary Metabolomic Alterations Observed in Hospitalized Adults With COVID-19–Related Acute Kidney Injury. Kidney Int. Rep. 2021, 6, 3002–3013. [Google Scholar] [CrossRef]
- Roberts, I.; Muelas, M.W.; Taylor, J.M.; Davison, A.S.; Xu, Y.; Grixti, J.M.; Gotts, N.; Sorokin, A.; Goodacre, R.; Kell, D.B. Untargeted metabolomics of COVID-19 patient serum reveals potential prognostic markers of both severity and outcome. Metabolomics 2021, 18, 6. [Google Scholar] [CrossRef]
- Xiao, N.; Nie, M.; Pang, H.; Wang, B.; Hu, J.; Meng, X.; Li, K.; Ran, X.; Long, Q.; Deng, H.; et al. Integrated cytokine and metabolite analysis reveals immunometabolic reprogramming in COVID-19 patients with therapeutic implications. Nat. Commun. 2021, 12, 1618. [Google Scholar] [CrossRef]
- Li, T.; Ning, N.; Li, B.; Luo, D.; Qin, E.; Yu, W.; Wang, J.; Yang, G.; Nan, N.; He, Z.; et al. Longitudinal Metabolomics Reveals Ornithine Cycle Dysregulation Correlates with Inflammation and Coagulation in COVID-19 Severe Patients. Front. Microbiol. 2021, 12, 723818. [Google Scholar] [CrossRef]
- Páez-Franco, J.C.; Torres-Ruiz, J.; Sosa-Hernández, V.A.; Cervantes-Díaz, R.; Romero-Ramírez, S.; Pérez-Fragoso, A.; Meza-Sánchez, D.E.; Germán-Acacio, J.M.; Maravillas-Montero, J.L.; Mejía-Domínguez, N.R.; et al. Metabolomics analysis reveals a modified amino acid metabolism that correlates with altered oxygen homeostasis in COVID-19 patients. Sci. Rep. 2021, 11, 6350. [Google Scholar] [CrossRef]
- Shi, D.; Yan, R.; Lv, L.; Jiang, H.; Lu, Y.; Sheng, J.; Xie, J.; Wu, W.; Xia, J.; Xu, K.; et al. The serum metabolome of COVID-19 patients is distinctive and predictive. Metabolism 2021, 118, 154739. [Google Scholar] [CrossRef] [PubMed]
- Ansone, L.; Briviba, M.; Silamikelis, I.; Terentjeva, A.; Perkons, I.; Birzniece, L.; Rovite, V.; Rozentale, B.; Viksna, L.; Kolesova, O.; et al. Amino Acid Metabolism is Significantly Altered at the Time of Admission in Hospital for Severe COVID-19 Patients: Findings from Longitudinal Targeted Metabolomics Analysis. Microbiol. Spectr. 2021, 9, e0033821. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Liu, C.; Li, D.; Huang, Q.; Liu, D.; Zhang, Y.; Ye, C.; Zhou, D.; Wang, Y.; Tan, Y.; et al. Metabolomic analyses reveals new stage-specific features of the COVID-19. Eur. Respir. J. 2021, 59, 2100284. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.; Stefanoni, D.; Reisz, J.A.; Nemkov, T.; Bertolone, L.; Francis, R.O.; Hudson, K.E.; Zimring, J.C.; Hansen, K.C.; Hod, E.A.; et al. COVID-19 infection alters kynurenine and fatty acid metabolism, correlating with IL-6 levels and renal status. JCI Insight 2020, 5, e140327. [Google Scholar] [CrossRef]
- Caterino, M.; Costanzo, M.; Fedele, R.; Cevenini, A.; Gelzo, M.; Di Minno, A.; Andolfo, I.; Capasso, M.; Russo, R.; Annunziata, A.; et al. The Serum Metabolome of Moderate and Severe COVID-19 Patients Reflects Possible Liver Alterations Involving Carbon and Nitrogen Metabolism. Int. J. Mol. Sci. 2021, 22, 9548. [Google Scholar] [CrossRef] [PubMed]
- Caterino, M.; Gelzo, M.; Sol, S.; Fedele, R.; Annunziata, A.; Calabrese, C.; Fiorentino, G.; D’Abbraccio, M.; Dell’Isola, C.; Fusco, F.M.; et al. Dysregulation of lipid metabolism and pathological inflammation in patients with COVID-19. Sci. Rep. 2021, 11, 2941. [Google Scholar] [CrossRef] [PubMed]
- Cas, M.D.; Ottolenghi, S.; Morano, C.; Rinaldo, R.; Roda, G.; Chiumello, D.; Centanni, S.; Samaja, M.; Paroni, R. Link between serum lipid signature and prognostic factors in COVID-19 patients. Sci. Rep. 2021, 11, 21633. [Google Scholar] [CrossRef]
- Torretta, E.; Garziano, M.; Poliseno, M.; Capitanio, D.; Biasin, M.; Santantonio, T.A.; Clerici, M.; Caputo, S.L.; Trabattoni, D.; Gelfi, C. Severity of COVID-19 Patients Predicted by Serum Sphingolipids Signature. Int. J. Mol. Sci. 2021, 22, 10198. [Google Scholar] [CrossRef]
- Kaur, G.; Ji, X.; Rahman, I. SARS-CoV2 Infection Alters Tryptophan Catabolism and Phospholipid Metabolism. Metabolites 2021, 11, 659. [Google Scholar] [CrossRef]
- Su, Y.; Chen, D.; Yuan, D.; Lausted, C.; Choi, J.; Dai, C.L.; Voillet, V.; Duvvuri, V.R.; Scherler, K.; Troisch, P.; et al. Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19. Cell 2020, 183, 1479–1495.e20. [Google Scholar] [CrossRef]
- Chen, Y.; Zheng, Y.; Yu, Y.; Wang, Y.; Huang, Q.; Qian, F.; Sun, L.; Song, Z.; Chen, Z.; Feng, J.; et al. Blood molecular markers associated with COVID-19 immunopathology and multi-organ damage. EMBO J. 2020, 39, e105896. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, S.; Nordqvist, H.; Ambikan, A.T.; Gupta, S.; Sperk, M.; Svensson-Akusjärvi, S.; Mikaeloff, F.; Benfeitas, R.; Saccon, E.; Ponnan, S.M.; et al. Metabolic Perturbation Associated With COVID-19 Disease Severity and SARS-CoV-2 Replication. Mol. Cell. Proteom. 2021, 20, 100159. [Google Scholar] [CrossRef] [PubMed]
- Suvarna, K.; Salkar, A.; Palanivel, V.; Bankar, R.; Banerjee, N.; Pai, M.G.J.; Srivastava, A.; Singh, A.; Khatri, H.; Agrawal, S.; et al. A Multi-omics Longitudinal Study Reveals Alteration of the Leukocyte Activation Pathway in COVID-19 Patients. J. Proteome Res. 2021, 20, 4667–4680. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Chen, D.; Ding, W.; Wu, P.; Hou, H.; Bai, Y.; Zhou, Y.; Li, K.; Xiang, S.; Liu, P.; et al. The trans-omics landscape of COVID-19. Nat. Commun. 2021, 12, 4543. [Google Scholar] [CrossRef]
- Li, Y.; Hou, G.; Zhou, H.; Wang, Y.; Tun, H.M.; Zhu, A.; Zhao, J.; Xiao, F.; Lin, S.; Liu, D.; et al. Multi-platform omics analysis reveals molecular signature for COVID-19 pathogenesis, prognosis and drug target discovery. Signal Transduct. Target. Ther. 2021, 6, 155. [Google Scholar] [CrossRef]
- Wang, C.; Li, X.; Ning, W.; Gong, S.; Yang, F.; Fang, C.; Gong, Y.; Wu, D.; Huang, M.; Gou, Y.; et al. Multi-omic profiling of plasma reveals molecular alterations in children with COVID-19. Theranostics 2021, 11, 8008–8026. [Google Scholar] [CrossRef]
- Wang, Y.; Bai, C.; Ruan, Y.; Liu, M.; Chu, Q.; Qiu, L.; Yang, C.; Li, B. Coordinative metabolism of glutamine carbon and nitrogen in proliferating cancer cells under hypoxia. Nat. Commun. 2019, 10, 201. [Google Scholar] [CrossRef] [Green Version]
- Costanzo, M.; Caterino, M.; Cevenini, A.; Jung, V.; Chhuon, C.; Lipecka, J.; Fedele, R.; Guerrera, I.C.; Ruoppolo, M. Proteomics Reveals that Methylmalonyl-CoA Mutase Modulates Cell Architecture and Increases Susceptibility to Stress. Int. J. Mol. Sci. 2020, 21, 4998. [Google Scholar] [CrossRef]
- Costanzo, M.; Cevenini, A.; Marchese, E.; Imperlini, E.; Raia, M.; Del Vecchio, L.; Caterino, M.; Ruoppolo, M. Label-Free Quantitative Proteomics in a Methylmalonyl-CoA Mutase-Silenced Neuroblastoma Cell Line. Int. J. Mol. Sci. 2018, 19, 3580. [Google Scholar] [CrossRef] [Green Version]
- Shen, B.; Yi, X.; Sun, Y.; Bi, X.; Du, J.; Zhang, C.; Quan, S.; Zhang, F.; Sun, R.; Qian, L.; et al. Proteomic and Metabolomic Characterization of COVID-19 Patient Sera. Cell 2020, 182, 59–72.e15. [Google Scholar] [CrossRef]
- Cornillet, M.; Strunz, B.; Rooyackers, O.; Ponzetta, A.; Chen, P.; Muvva, J.R.; Akber, M.; Buggert, M.; Chambers, B.J.; Dzidic, M.; et al. COVID-19-specific metabolic imprint yields insights into multiorgan system perturbations. Eur. J. Immunol. 2021; Epub ahead of print. [Google Scholar] [CrossRef]
- Wilk, A.J.; Lee, M.J.; Wei, B.; Parks, B.; Pi, R.; Martínez-Colón, G.J.; Ranganath, T.; Zhao, N.Q.; Taylor, S.; Becker, W.; et al. Multi-omic profiling reveals widespread dysregulation of innate immunity and hematopoiesis in COVID-19. J. Exp. Med. 2021, 218, e20210582. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chen, C.; Chen, W.; Huang, L.; Fu, Z.; Ye, K.; Lv, L.; Nong, Z.; Zhou, X.; Lu, W.; et al. Proteomics and metabonomics analyses of COVID-19 complications in patients with pulmonary fibrosis. Sci. Rep. 2021, 11, 14601. [Google Scholar] [CrossRef]
- Caterino, M.; Ruoppolo, M.; Mandola, A.; Costanzo, M.; Orrù, S.; Imperlini, E. Protein–protein interaction networks as a new perspective to evaluate distinct functional roles of voltage-dependent anion channel isoforms. Mol. BioSyst. 2017, 13, 2466–2476. [Google Scholar] [CrossRef] [PubMed]
- De Pasquale, V.; Costanzo, M.; Siciliano, R.A.; Mazzeo, M.F.; Pistorio, V.; Bianchi, L.; Marchese, E.; Ruoppolo, M.; Pavone, L.M.; Caterino, M. Proteomic Analysis of Mucopolysaccharidosis IIIB Mouse Brain. Biomolecules 2020, 10, 355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Melo, M.G.; Fontana, A.O.; Viertl, D.; Allenbach, G.; Prior, J.O.; Rotman, S.; Feichtinger, R.G.; Mayr, J.A.; Costanzo, M.; Caterino, M.; et al. A knock-in rat model unravels acute and chronic renal toxicity in glutaric aciduria type I. Mol. Genet. Metab. 2021, 134, 287–300. [Google Scholar] [CrossRef]
- Asakura, H.; Ogawa, H. COVID-19-associated coagulopathy and disseminated intravascular coagulation. Int. J. Hematol. 2020, 113, 45–57. [Google Scholar] [CrossRef]
- Wool, G.D.; Miller, J.L. The Impact of COVID-19 Disease on Platelets and Coagulation. Pathobiology 2020, 88, 15–27. [Google Scholar] [CrossRef]
- Gąsecka, A.; Borovac, J.A.; Guerreiro, R.A.; Giustozzi, M.; Parker, W.; Caldeira, D.; Chiva-Blanch, G. Thrombotic Complications in Patients with COVID-19: Pathophysiological Mechanisms, Diagnosis, and Treatment. Cardiovasc. Drugs Ther. 2020, 35, 215–229. [Google Scholar] [CrossRef]
- Perico, L.; Benigni, A.; Casiraghi, F.; Ng, L.F.P.; Renia, L.; Remuzzi, G. Immunity, endothelial injury and complement-induced coagulopathy in COVID-19. Nat. Rev. Nephrol. 2020, 17, 46–64. [Google Scholar] [CrossRef]
- Chowdhury, M.A.; Hossain, N.; Kashem, M.A.; Shahid, M.A.; Alam, A. Immune response in COVID-19: A review. J. Infect. Public Health 2020, 13, 1619–1629. [Google Scholar] [CrossRef]
- Moriarty, P.M.; Gorby, L.K.; Stroes, E.S.; Kastelein, J.P.; Davidson, M.; Tsimikas, S. Lipoprotein(a) and Its Potential Association with Thrombosis and Inflammation in COVID-19: A Testable Hypothesis. Curr. Atheroscler. Rep. 2020, 22, 48. [Google Scholar] [CrossRef] [PubMed]
- Kočar, E.; Režen, T.; Rozman, D. Cholesterol, lipoproteins, and COVID-19: Basic concepts and clinical applications. Biochim. Biophys. Acta (BBA) Mol. Cell Biol. Lipids 2020, 1866, 158849. [Google Scholar] [CrossRef] [PubMed]
- Caterino, M.; Ruoppolo, M.; Villani, G.R.D.; Marchese, E.; Costanzo, M.; Sotgiu, G.; Dore, S.; Franconi, F.; Campesi, I. Influence of Sex on Urinary Organic Acids: A Cross-Sectional Study in Children. Int. J. Mol. Sci. 2020, 21, 582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Melo, M.G.; Remacle, N.; Cudré-Cung, H.-P.; Roux, C.; Poms, M.; Cudalbu, C.; Barroso, M.; Gersting, S.W.; Feichtinger, R.G.; Mayr, J.A.; et al. The first knock-in rat model for glutaric aciduria type I allows further insights into pathophysiology in brain and periphery. Mol. Genet. Metab. 2021, 133, 157–181. [Google Scholar] [CrossRef]
- Costanzo, M.; Fiocchetti, M.; Ascenzi, P.; Marino, M.; Caterino, M.; Ruoppolo, M. Proteomic and Bioinformatic Investigation of Altered Pathways in Neuroglobin-Deficient Breast Cancer Cells. Molecules 2021, 26, 2397. [Google Scholar] [CrossRef]
- Costanzo, M.; Caterino, M.; Salvatori, I.; Manganelli, V.; Ferri, A.; Misasi, R.; Ruoppolo, M. Proteome data of neuroblastoma cells overexpressing Neuroglobin. Data Brief 2022, 41, 107843. [Google Scholar] [CrossRef]
- Li, S.; Ma, F.; Yokota, T.; Garcia, G.; Palermo, A.; Wang, Y.; Farrell, C.; Wang, Y.-C.; Wu, R.; Zhou, Z.; et al. Metabolic reprogramming and epigenetic changes of vital organs in SARS-CoV-2–induced systemic toxicity. JCI Insight 2021, 6, e145027. [Google Scholar] [CrossRef]
- Casari, I.; Manfredi, M.; Metharom, P.; Falasca, M. Dissecting lipid metabolism alterations in SARS-CoV-2. Prog. Lipid Res. 2021, 82, 101092. [Google Scholar] [CrossRef]
- Tanner, J.E.; Alfieri, C. The Fatty Acid Lipid Metabolism Nexus in COVID-19. Viruses 2021, 13, 90. [Google Scholar] [CrossRef]

| Authors | Biologic Matrix | Patients | Technology | Pathway/Protein Dysregulation |
|---|---|---|---|---|
| Bauer et al. (2021) | Plasma | 44 non-hospitalized COVID-19 53 hospitalized COVID-19 44 non-COVID-19 | PEA | Inflammation |
| Chen Y. et al. (2021) | Serum | 10 moderate COVID-19 6 severe COVID-19 10 healthy controls | DIA-MS | Cholesterol metabolism, coagulation, cardiovascular system |
| D’Alessandro et al. (2020) | Serum | 33 COVID-19 16 non-COVID-19 | LC-MS/MS | IL-6 signaling, coagulation, complement, antimicrobial enzymes |
| Demichev et al. (2021) | Plasma | 139 (WHO grade 3–7) COVID-19 16 non-COVID-19 | DIA-MS | Coagulation, complement, immune system, inflammation |
| Haljasmägi et al. (2020) | Plasma | 25 moderate COVID-19 15 severe COVID-19 18 healthy controls | PEA | Apoptosis, inflammation, neuronal injury |
| Hou et al. (2020) | Serum | 15 COVID-19 13 influenza | antibody microarray | Immune system, inflammation |
| Kimura et al. (2021) | Serum | 10 severe COVID-19 | DIA-MS | cardiovascular system, inflammation |
| Lee et al. (2021) | Serum | 13 non-severe COVID-19 12 severe COVID-19 | DIA-MS | Coagulation, immune system, inflammation, lipid metabolism |
| Messner et al. (2021) | Plasma | 31 (mild + severe) COVID-19 + 17 COVID-19 15 healthy controls | DIA-MS | Coagulation, complement, inflammation |
| Park et al. (2020) | Plasma | 3 mild COVID-19 5 severe COVID-19 | LC-MS/MS | Coagulation, neutrophils activation |
| Patel et al. (2021) | Plasma | 26 mild COVID-19 9 severe COVID-19 24 critical COVID-19 28 healthy controls | PEA | Cytokine-cytokine receptor interaction |
| Shu et al. (2020) | Plasma | 10 mild COVID-19 7 severe COVID-19 5 fatal COVID-19 8 healthy controls | LC-MS/MS | Coagulation, complement, energy metabolism, immune system, inflammation |
| Zhong et al. (2021) | Plasma | 50 (mild + moderate) COVID-19 | PEA | Cytokine-related, immune-related |
| Authors | Biologic Matrix | Patients | Technology | Pathway/Metabolite Dysregulation |
|---|---|---|---|---|
| Ansone et al. (2021) | Serum | 32 hospitalized COVID-19 39 healthy controls | LC-MS/MS | Amino acid metabolism, tryptophan metabolism, urea cycle |
| Bizkarguenaga et al. (2021) | Plasma | 69 recovered COVID-19 71 healthy controls | NMR | TG, cholesterol, phospholipids |
| Blasco et al. (2020) | Plasma | 55 COVID-19 45 healthy controls | LC-MS/MS | NAD metabolism, pyrimidine metabolism, tryptophan metabolism |
| Caterino M. et al. (2021) | Serum | 20 mild COVID-19 16 moderate COVID-19 17 severe COVID-19 9 healthy controls | LC-MS/MS | Carbon and nitrogen metabolism, energy metabolism, purine and pyrimidine metabolism |
| Caterino M. et al. (2021) | Serum | 20 mild COVID-19 16 moderate COVID-19 16 severe COVID-19 9 healthy controls | LC-MS/MS | Cer, TG |
| Danlos et al. (2021) | Plasma | 23 mild COVID-19 21 moderate COVID-19 28 critical COVID-19 29 healthy controls | GC-MS LC-MS/MS | Tryptophan metabolism |
| Dei Cas et al. (2021) | Serum | 49 COVID-19 10 healthy controls | LC-MS/MS | Acylcarnitines, PC, PE, CE, DAG, lysoPE, SM |
| Fraser et al. (2020) | Plasma | 10 COVID19+ patients 10 COVID19– patients 10 healthy controls | LC-MS/MS NMR | Tryptophan metabolism, lysoPC |
| Jia et al. (2021) | Serum | 18 mild COVID-19 12 severe COVID-19 69 recovered COVID-19 13 healthy controls + 90 COVID-19 28 healthy controls | LC-MS/MS | Amino acid metabolism, TCA cycle, urea cycle |
| Kaur et al. (2021) | Serum | 6 COVID-19 6 recovered COVID-19 | LC-MS/MS | PC, SM, arachidonic acid, tryptophan metabolism |
| Khodadoust et al. (2021) | Plasma | 32 mild COVID-19 18 severe COVID-19 (n.f.) healthy controls | LC-MS/MS | Cer |
| Li T. et al. (2021) | Serum | 30 (mild + moderate) COVID-19 17 severe COVID-19 20 healthy controls | LC-MS/MS | Amino acid metabolism, carbohydrate metabolism, urea cycle |
| Páez-Franco et al. (2021) | Serum | 19 mild COVID-19 46 severe COVID-19 27 healthy controls | GC-MS | Amino acid metabolism, energy metabolism |
| Roberts et al. (2021) | Serum | 71 mild COVID-19 49 severe COVID-19 + 90 COVID-19 | LC-MS/MS | Acylcarnitines, energy metabolism, tryptophan metabolism |
| Shi et al. (2021) | Serum | 79 COVID-19 30 COVID-19-like 78 healthy controls | GC-MS | Amino acid metabolism, energy metabolism |
| Sindelar et al. (2021) | Plasma | 272 COVID-19 67 negative controls | LC-MS/MS | Cer, lysoPC, PC |
| Thomas et al. (2020) | Serum | 33 COVID-19 16 negative controls | LC-MS/MS | Carbon and nitrogen metabolism, tryptophan metabolism |
| Torretta et al. (2021) | Serum | 11 mild COVID-19 28 moderate COVID-19 12 severe COVID-19 8 critical COVID-19 27 healthy controls | LC-MS/MS | Cer, SM, sphingosine |
| Xiao et al. (2021) | Serum | 14 mild COVID-19 (+7 mild COVID-19) 23 severe COVID-19 17 healthy controls | LC-MS/MS | Arginine metabolism, purine metabolism, tryptophan metabolism |
| Authors | Biologic Matrix | Patients | Omics Used | Technology | Proteomic/Metabolomic Dysregulation |
|---|---|---|---|---|---|
| Chen Y.-M. et al. (2020) | Plasma | 50 mild COVID-19 16 severe COVID-19 17 healthy controls | Proteomics Metabolomics | DIA-MS NMR | TCA cycle, glycolytic pathway, platelet signaling pathway, TG, cholesterol, phospholipids |
| Cornillet et al. (2021) | Serum | 27 (moderate + severe) COVID-19 17 healthy controls | Proteomics Metabolomics | PEA LC-MS/MS | Immune system, neurological inflammation |
| Krishnan et al. (2021) | Plasma | 41 (mild + severe) COVID-19 31 healthy controls | Proteomics Metabolomics | PEA LC-MS/MS | Cytokine-cytokine receptor interaction, chemokine signaling, TNF signaling pathway, glycolysis, TCA cycle |
| Li Y. et al. (2021) | Plasma | 10 non-severe COVID-19 10 severe COVID-19 10 healthy controls + 5 non-severe COVID-19 5 severe COVID-19 | Proteomics Metabolomics | DIA-MS LC-MS/MS | Complement, inflammation, host-virus interaction, lipid metabolism, DAG, TG, PC, PG |
| Shen et al. (2020) | Serum | 25 non-severe COVID-19 28 severe COVID-19 25 non-COVID-19 25 healthy controls | Proteomics Metabolomics | LC-MS/MS LC-MS/MS | Coagulation, complement, immune system, inflammation, arginine metabolism, lipid metabolism, NAD and tryptophan metabolism |
| Su et al. (2020) | Plasma | 139 COVID-19 133 healthy controls | Proteomics Metabolomics | PEA LC-MS/MS | Amino acid metabolism, tryptophan metabolism, urea cycle |
| Suvarna et al. (2021) | Plasma | 13 COVID-19 | Proteomics Metabolomics | LC-MS/MS LC-MS/MS | Coagulation, complement, myeloid leukocyte activation, arginine amino acid metabolism |
| Wang et al. (2021) | Plasma | 18 mild COVID-19 12 healthy controls | Proteomics Metabolomics | LC-MS/MS LC-MS/MS | Coagulation, extra-cellular matrix organization, arginine metabolism, carbon metabolism, choline metabolism, pyrimidine and tryptophan metabolism |
| Wilk et al. (2021) | Blood (immune cells) | 64 (mild-to-fatal) COVID-19 8 healthy controls | Proteomics | CyTOF | Immune system, neutrophil and NK cell hyperactivation |
| Wu et al. (2021) | Plasma | 231 (asymptomatic, mild, severe, critical) COVID-19 | Proteomics Metabolomics | DIA-MS LC-MS/MS | Inflammation, macrophage migration, neutrophil degranulation, apoptosis, arginine metabolism, tryptophan metabolism, Cer, lysoPC, PE |
| Yang et al. (2021) | Serum | 85 COVID-19 41 non-pulmonary fibrosis 44 pulmonary fibrosis | Proteomics Metabolomics | DIA-MS LC-MS/MS | Immune system, cell adhesion, PPAR signaling, D-arginine and D-ornithine metabolism (urea cycle) |
| Protein Symbol | UniProt ID | Protein Name | STRING Cluster |
|---|---|---|---|
| A2M | P01023 | Alpha-2-macroglobulin | Cluster 1 |
| ACTB | P60709 | Actin, cytoplasmic 1 | |
| AHSG | P02765 | Alpha-2-HS-glycoprotein | |
| ALB | P02768 | Albumin | |
| C1R | P00736 | Complement C1r subcomponent | |
| C5 | P01031 | Complement C5 | |
| CFB | P00751 | Complement Factor B | |
| CFH | P08603 | Complement factor H | |
| CFI | P05156 | Complement factor I | |
| CRP | P02741 | C-reactive protein | |
| CST3 | P01034 | Cystatin-C | |
| CTSB | P07858 | Cathepsin B | |
| CTSL | P07711 | Procathepsin L | |
| F9 | P00740 | Coagulation factor IX | |
| F10 | P00742 | Coagulation factor X | |
| F12 | P00748 | Coagulation factor XII | |
| F13B | P05160 | Coagulation factor XIII B chain | |
| FGA | P02671 | Fibrinogen alpha chain | |
| FGG | P02679 | Fibrinogen gamma chain | |
| GSN | P06396 | Gelsolin | |
| HRG | P04196 | Histidine-rich glycoprotein | |
| HSPA8 | P11142 | Heat shock cognate 71 kDa protein | |
| ITIH4 | Q14624 | Inter-alpha-trypsin inhibitor heavy chain H4 | |
| KLKB1 | P03952 | Plasma kallikrein | |
| KNG1 | P01042 | Kininogen-1 | |
| LGALS3BP | Q08380 | Galectin-3-binding protein | |
| LRG1 | P02750 | Leucine-rich alpha-2-glycoprotein | |
| MPO | P05164 | Myeloperoxidase | |
| ORM1 | P02763 | Alpha-1-acid glycoprotein 1 | |
| PIGR | P01833 | Polymeric immunoglobulin receptor | |
| PLG | P00747 | Plasminogen | |
| PRG4 | Q92954 | Proteoglycan 4 | |
| PROS1 | P07225 | Vitamin K-dependent protein S | |
| SERPINA1 | P01009 | Alpha-1-antitrypsin | |
| SERPINA3 | P01011 | Alpha-1-antichymotrypsin | |
| SERPINA10 | Q9UK55 | Protein Z-dependent protease inhibitor | |
| SERPINC1 | P01008 | Antithrombin-III | |
| SERPINF2 | P08697 | Alpha-2-antiplasmin | |
| TF | P02787 | Transferrin | |
| TTR | P02766 | Transthyretin | |
| VIM | P08670 | Vimentin | |
| CCL2 | P13500 | C-C motif chemokine 2 | Cluster 2 |
| CCL7 | P80098 | C-C motif chemokine 7 | |
| CCL8 | P80075 | C-C motif chemokine 8 | |
| CD14 | P08571 | Monocyte differentiation antigen CD14 | |
| CCL23 | P55773 | C-C motif chemokine 23 | |
| CD274 | Q9NZQ7 | Programmed cell death 1 ligand 1 | |
| CHI3L1 | P36222 | Chitinase-3-like protein 1 | |
| CXCL10 | P02778 | C-X-C motif chemokine 10 | |
| CXCL11 | O14625 | C-X-C motif chemokine 11 | |
| DEFA1 | P59665 | Neutrophil defensin 1 | |
| HGF | P14210 | Hepatocyte growth factor | |
| IL-10 | P22301 | Interleukin-10 | |
| IL-18R1 | Q13478 | Interleukin-18 receptor 1 | |
| IL-6 | P08887 | Interleukin-6 receptor subunit alpha | |
| LBP | P18428 | Lipopolysaccharide-binding protein | |
| LCN2 | P80188 | Neutrophil gelatinase-associated lipocalin | |
| LGALS9 | O00182 | Galectin-9 | |
| S100A11 | P31949 | Protein S100-A11 | |
| S100A12 | P80511 | Protein S100-A12 | |
| S100A8 | P05109 | Protein S100-A8 | |
| S100A9 | P06702 | Protein S100-A9 | |
| SAA1 | P0DJI8 | Serum amyloid A-1 protein | |
| TGFB1 | P01137 | Transforming growth factor beta-1 proprotein | |
| TNF | P01375 | Tumor necrosis factor | |
| VEGFA | P15692 | Vascular endothelial growth factor A | |
| APOA1 | P02647 | Apolipoprotein A1 | Cluster 3 |
| APOA2 | P02652 | Apolipoprotein A2 | |
| APOC1 | P02654 | Apolipoprotein C1 | |
| APOC3 | P02656 | Apolipoprotein C3 | |
| APOD | P05090 | Apolipoprotein D | |
| APOL1 | O14791 | Apolipoprotein L1 | |
| APOM | O95445 | Apolipoprotein M | |
| C8A | P07357 | Complement component C8 alpha chain | |
| CETP | P11597 | Cholesteryl ester transfer protein | |
| CFHR5 | Q9BXR6 | Complement factor H-related protein 5 | |
| FGB | P02675 | Fibrinogen beta chain | |
| IGFALS | P35858 | Insulin-like growth factor-binding protein complex acid labile subunit | |
| ITIH3 | Q06033 | Inter-alpha-trypsin inhibitor heavy chain H3 | |
| PI16 | Q6UXB8 | Peptidase inhibitor 16 | |
| SAA2 | P0DJI9 | Serum amyloid A-2 protein | |
| SAA4 | P35542 | Serum amyloid A-4 protein | |
| SCARB2 | Q14108 | Lysosome membrane protein 2 |
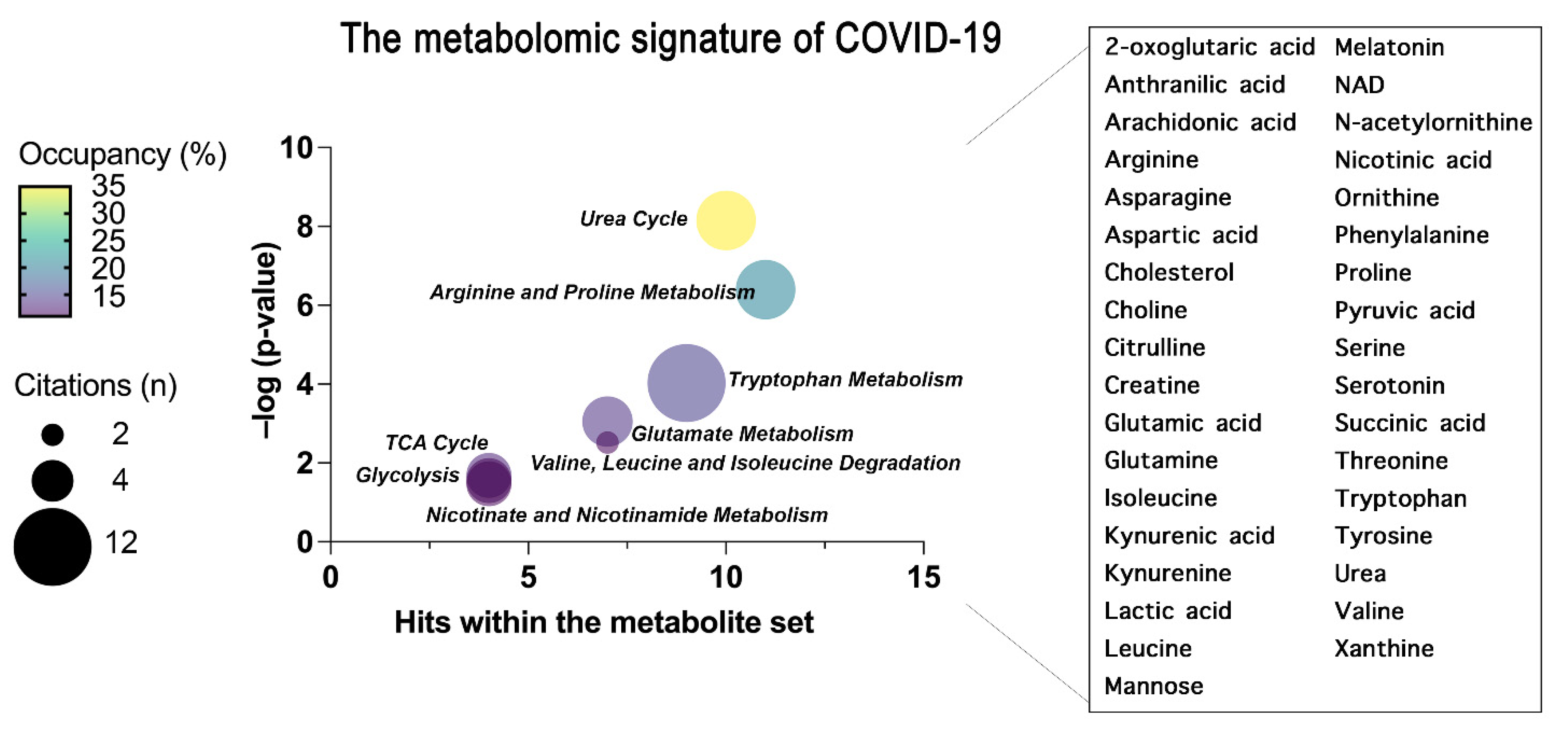
| KEGG Pathway | Metabolite Set |
|---|---|
| Urea Cycle | 2-oxoglutaric acid, Arginine, Aspartic acid, Citrulline, Glutamic acid, Glutamine, NAD, Ornithine, Pyruvic acid, Urea |
| Arginine and Proline Metabolism | 2-oxoglutaric acid, Arginine, Aspartic acid, Citrulline, Glutamic acid, NAD, Ornithine, Proline, Succinic acid, Urea |
| Tryptophan Metabolism | 2-oxoglutaric acid, Anthranilic acid, Glutamic acid, Kynurenic acid, Kynurenine, Melatonin, NAD, Serotonin, Tryptophan |
| Glutamate Metabolism | 2-oxoglutaric acid, Aspartic acid, Glutamic acid, Glutamine, NAD, Pyruvic acid, Succinic acid |
| Valine, Leucine and Isoleucine Degradation | 2-oxoglutaric acid, Glutamic acid, Isoleucine, Leucine, NAD, Succinic acid, Valine |
| TCA Cycle | 2-oxoglutaric acid, NAD, Pyruvic acid, Succinic acid |
| Glycolysis | 2-oxoglutaric acid, Lactic acid, NAD, Pyruvic acid |
| Nicotinate and Nicotinamide Metabolism | Glutamic acid, Glutamine, NAD, Nicotinic acid |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costanzo, M.; Caterino, M.; Fedele, R.; Cevenini, A.; Pontillo, M.; Barra, L.; Ruoppolo, M. COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19. Int. J. Mol. Sci. 2022, 23, 2414. https://doi.org/10.3390/ijms23052414
Costanzo M, Caterino M, Fedele R, Cevenini A, Pontillo M, Barra L, Ruoppolo M. COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19. International Journal of Molecular Sciences. 2022; 23(5):2414. https://doi.org/10.3390/ijms23052414
Chicago/Turabian StyleCostanzo, Michele, Marianna Caterino, Roberta Fedele, Armando Cevenini, Mariarca Pontillo, Lucia Barra, and Margherita Ruoppolo. 2022. "COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19" International Journal of Molecular Sciences 23, no. 5: 2414. https://doi.org/10.3390/ijms23052414
APA StyleCostanzo, M., Caterino, M., Fedele, R., Cevenini, A., Pontillo, M., Barra, L., & Ruoppolo, M. (2022). COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19. International Journal of Molecular Sciences, 23(5), 2414. https://doi.org/10.3390/ijms23052414










