Impaired Functional Connectivity Underlies Fragile X Syndrome
Abstract
1. Introduction
2. Results
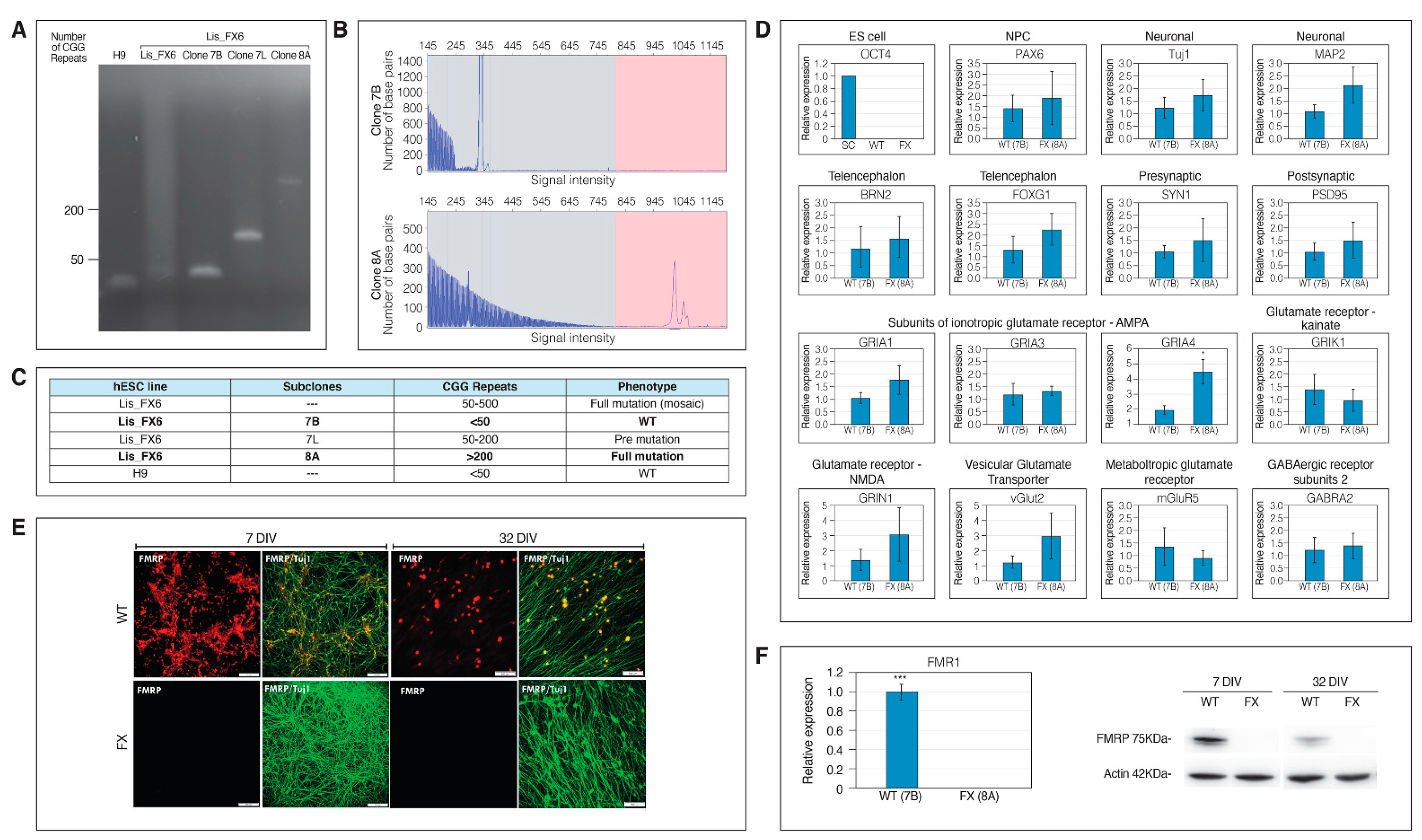
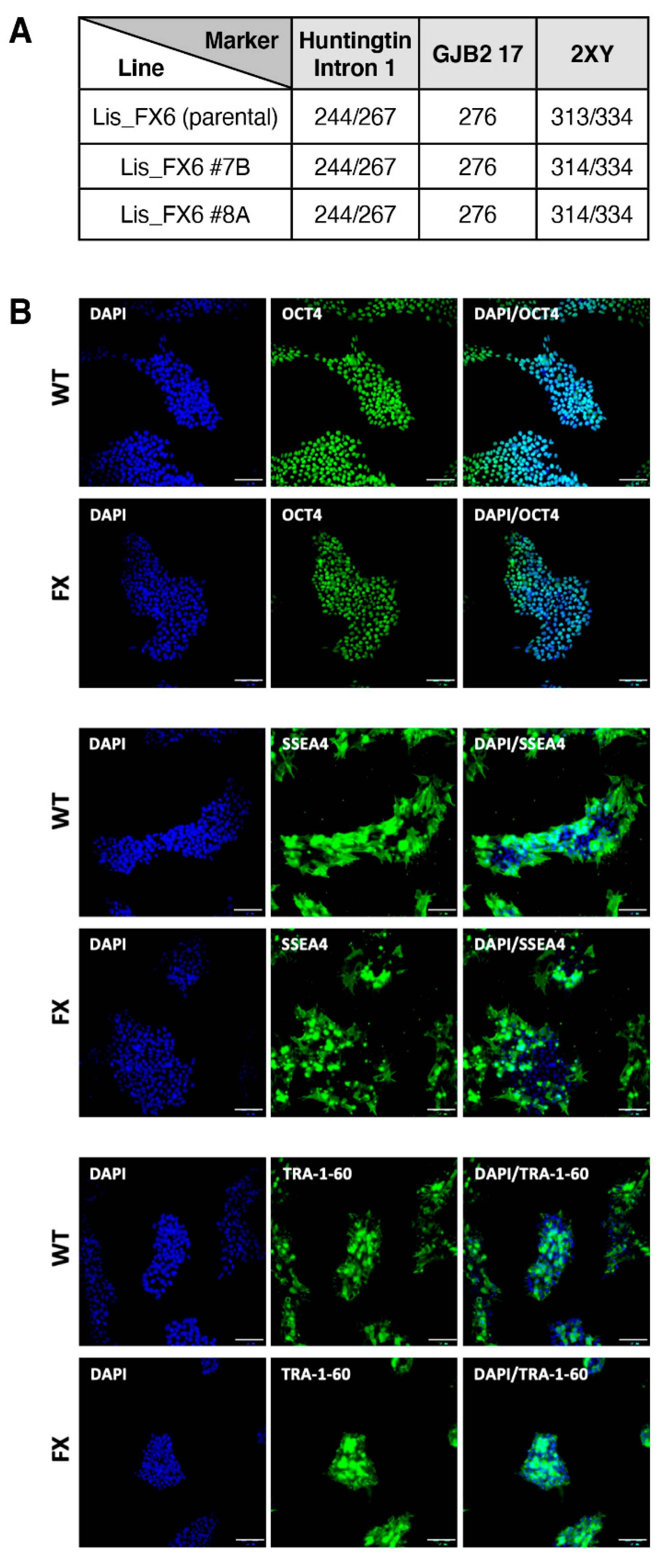
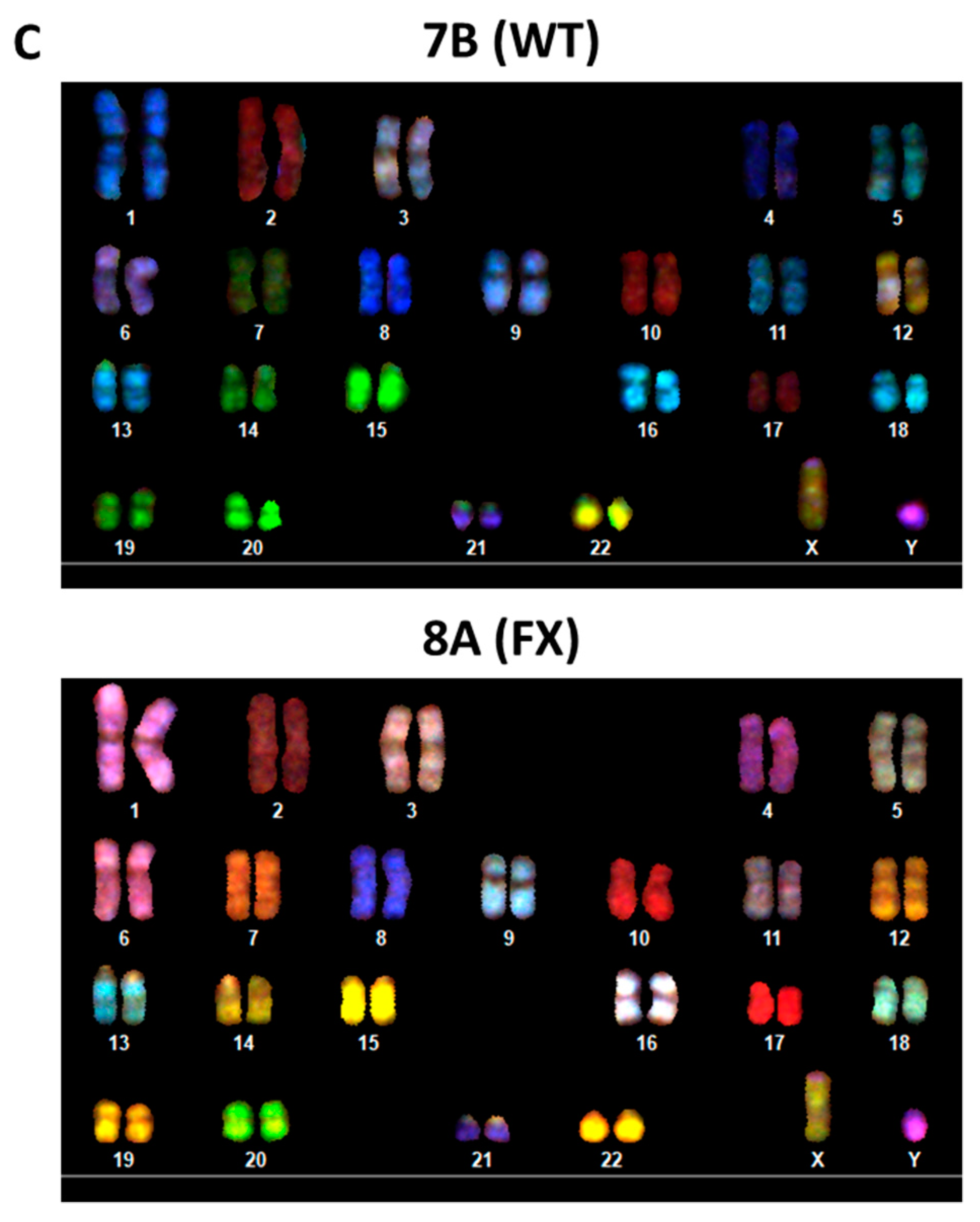
2.1. Generation of Isogenic FXS Full-Mutation hESC Subclones
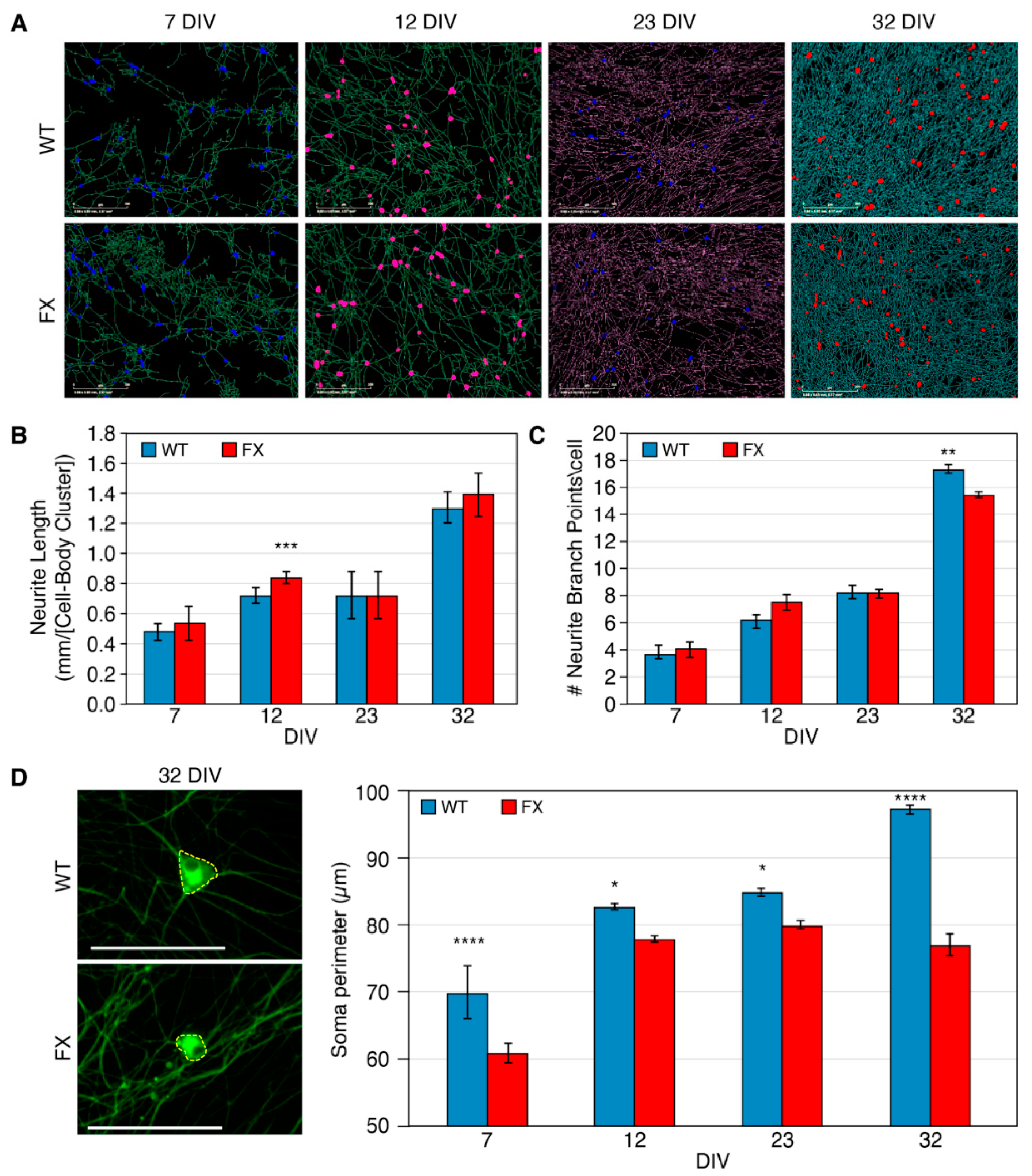
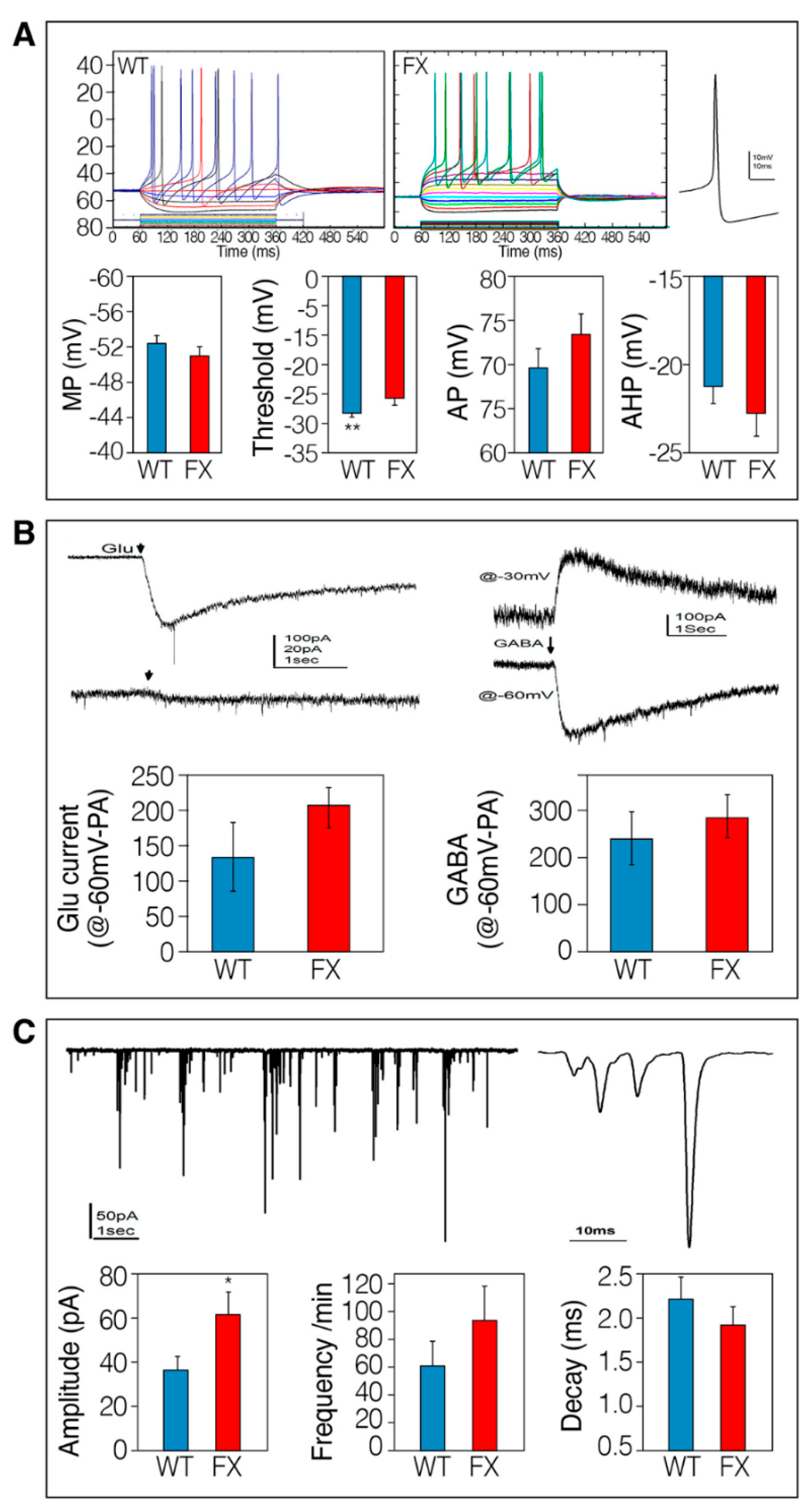
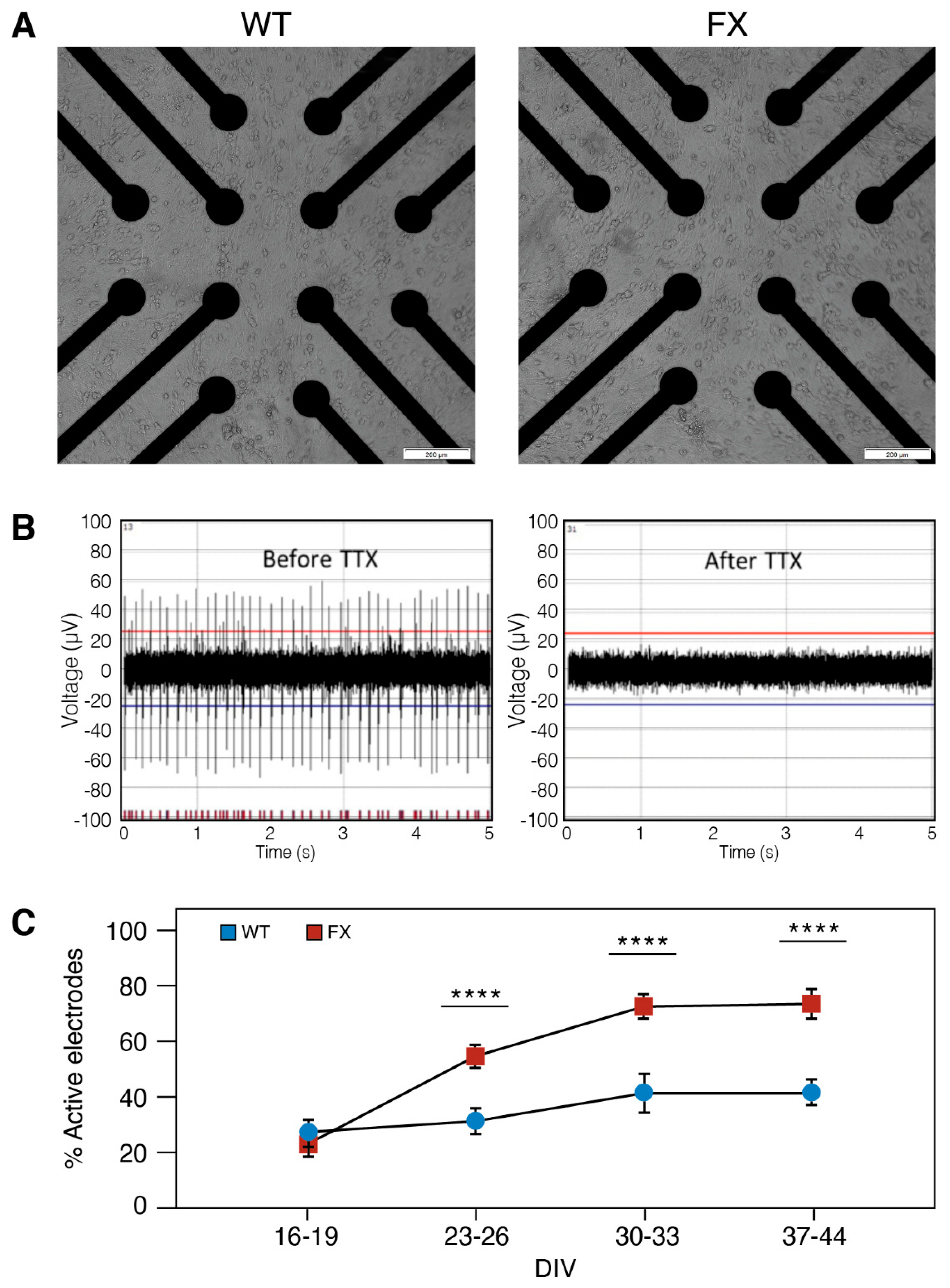
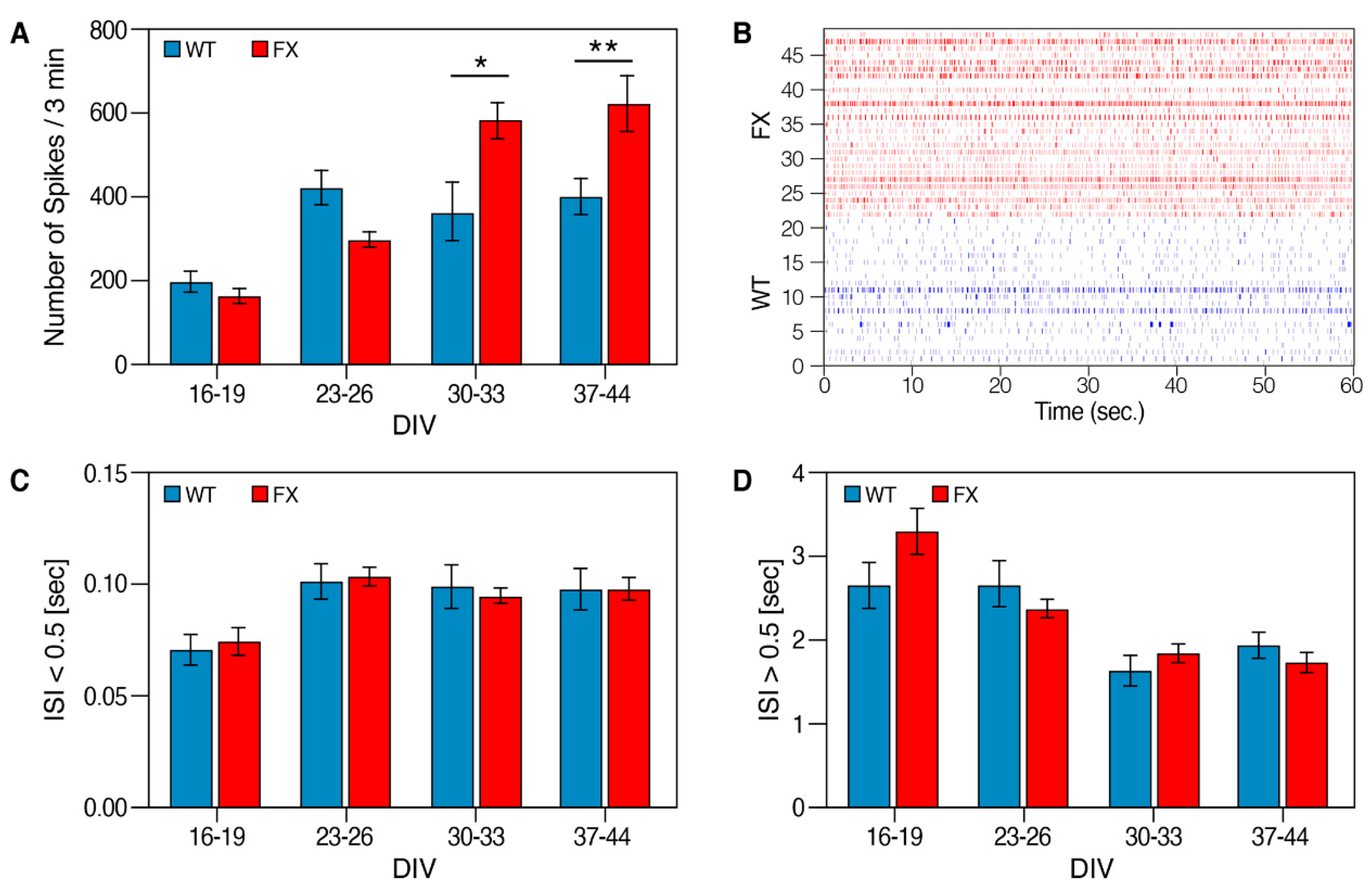
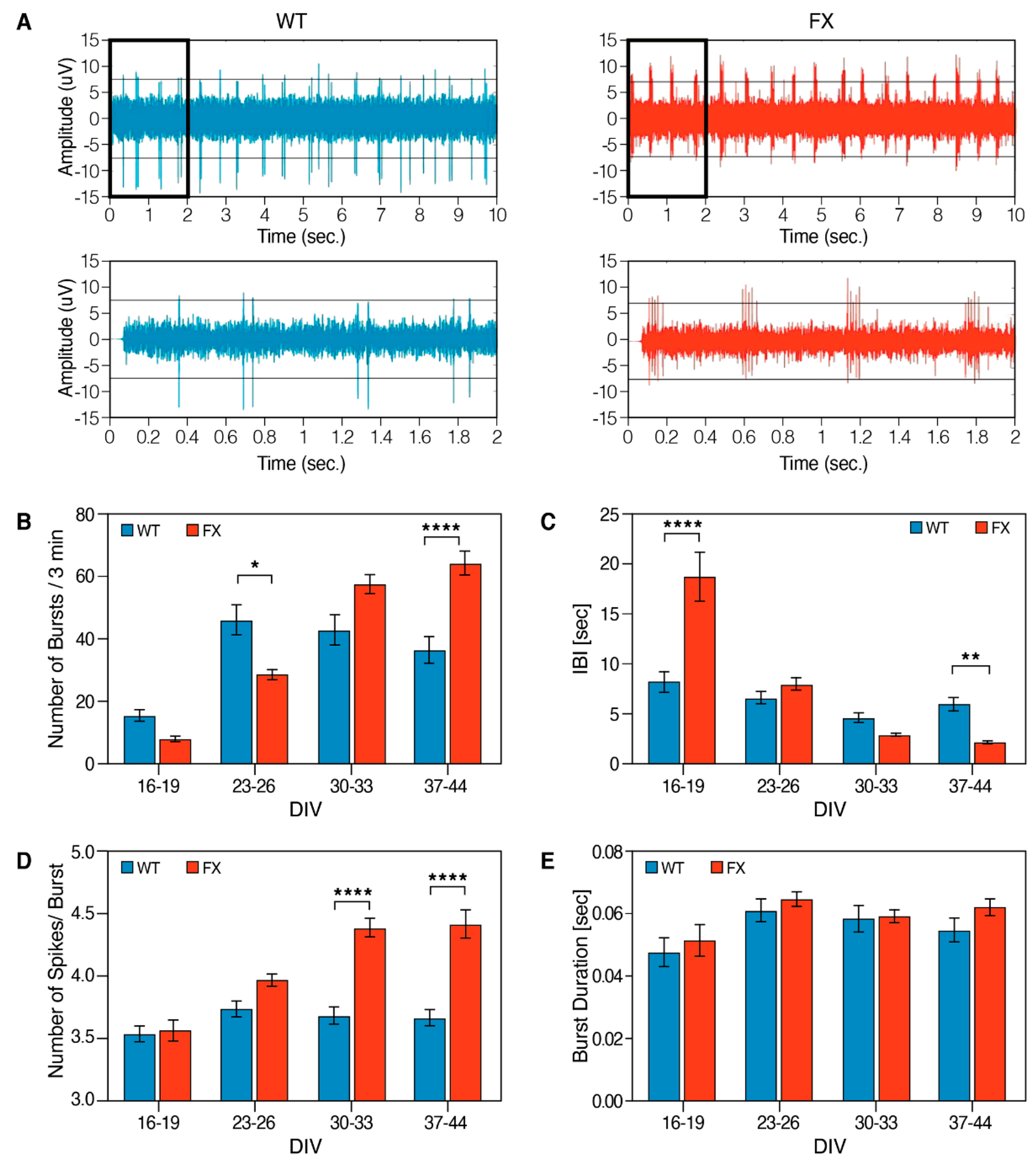
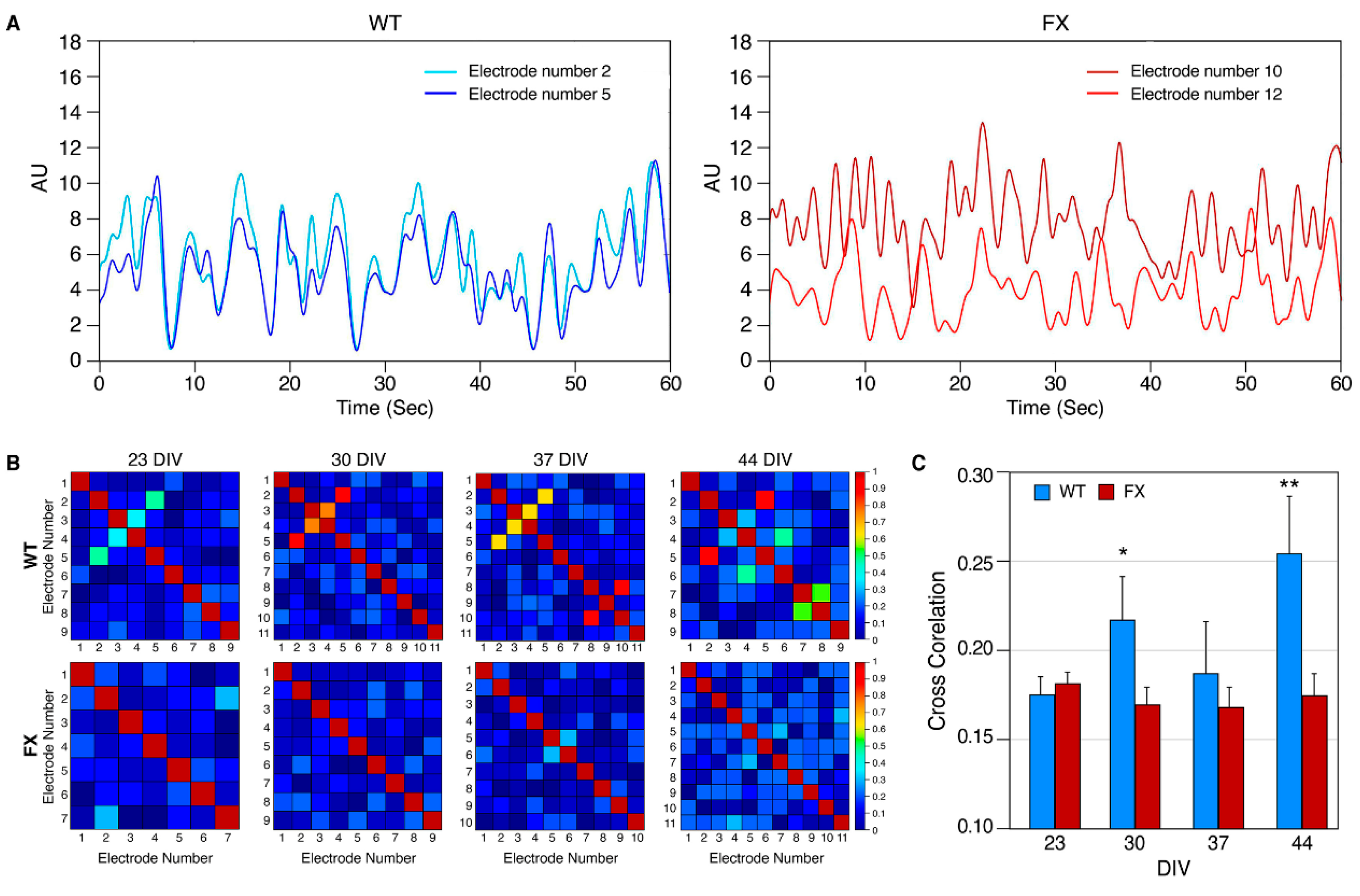
2.2. The Effect of FMRP Expression on Neuronal Function and Network Development
3. Discussion
4. Material and Methods
4.1. Human Embryonic Stem Cell Culture
4.2. Single-Cell Cloning
4.3. Repeat-Number Assay
4.4. Generation of Induced Neurons from hESC
4.5. Rat Astrocytes Feeder Cell Culture
4.6. Immunofluorescence
4.7. Western Blot Analysis
4.8. RNA Extraction and Quantitative Real-Time PCR
4.9. High-Throughput Cell Morphology Analysis
4.10. Electrophysiological Recordings
4.11. Multielectrode Array (MEAs) Recordings and Data Analysis
4.12. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Contractor, A.; Klyachko, V.A.; Portera-Cailliau, C. Altered Neuronal and Circuit Excitability in Fragile X Syndrome. Neuron 2015, 87, 699–715. [Google Scholar] [CrossRef]
- Richter, J.D.; Zhao, X. The molecular biology of FMRP: New insights into fragile X syndrome. Nat. Rev. Neurosci. 2021, 22, 209–222. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, A.; McMillan, E.; Wallace, K.; Tubon, J.T.C.; Capowski, E.E.; Svendsen, C.N. Normal Neurogenesis but Abnormal Gene Expression in Human Fragile X Cortical Progenitor Cells. Stem Cells Dev. 2008, 17, 107–118. [Google Scholar] [CrossRef]
- Winograd, C.; Ceman, S. Fragile X family members have important and non-overlapping functions. Biomol. Concepts 2011, 2, 343–352. [Google Scholar] [CrossRef]
- Kazdoba, T.M.; Leach, P.T.; Silverman, J.L.; Crawley, J.N. Modeling fragile X syndrome in the Fmr1 knockout mouse. Intractable Rare Dis. Res. 2014, 3, 118–133. [Google Scholar] [CrossRef]
- Drozd, M.; Bardoni, B.; Capovilla, M. Modeling Fragile X Syndrome in Drosophila. Front. Mol. Neurosci. 2018, 11, 124. [Google Scholar] [CrossRef]
- Shamay-Ramot, A.; Khermesh, K.; Porath, H.; Barak, M.; Pinto, Y.; Wachtel, C.; Zilberberg, A.; Lerer-Goldshtein, T.; Efroni, S.; Levanon, E.; et al. Fmrp Interacts with Adar and Regulates RNA Editing, Synaptic Density and Locomotor Activity in Zebrafish. PLoS Genet. 2015, 11, e1005702. [Google Scholar] [CrossRef] [PubMed]
- Gibson, J.R.; Bartley, A.; Hays, S.A.; Huber, K.M. Imbalance of Neocortical Excitation and Inhibition and Altered UP States Reflect Network Hyperexcitability in the Mouse Model of Fragile X Syndrome. J. Neurophysiol. 2008, 100, 2615–2626. [Google Scholar] [CrossRef]
- Zhang, L.; Liang, Z.; Zhu, P.; Li, M.; Yi, Y.-H.; Liao, W.-P.; Su, T. Altered intrinsic properties and bursting activities of neurons in layer IV of somatosensory cortex from Fmr-1 knockout mice. Exp. Neurol. 2016, 280, 60–69. [Google Scholar] [CrossRef]
- Booker, S.A.; Domanski, A.P.F.; Dando, O.R.; Jackson, A.D.; Isaac, J.T.R.; Hardingham, G.E.; Wyllie, D.J.A.; Kind, P.C. Altered dendritic spine function and integration in a mouse model of fragile X syndrome. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Kurosaki, T.; Imamachi, N.; Pröschel, C.; Mitsutomi, S.; Nagao, R.; Akimitsu, N.; Maquat, L.E. Loss of the fragile X syndrome protein FMRP results in misregulation of nonsense-mediated mRNA decay. Nat. Cell Biol. 2021, 23, 40–48. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Bhattacharyya, A. Human Models Are Needed for Studying Human Neurodevelopmental Disorders. Am. J. Hum. Genet. 2018, 103, 829–857. [Google Scholar] [CrossRef] [PubMed]
- van Karnebeek, C.D.; Bowden, K.; Berry-Kravis, E. Treatment of Neurogenetic Developmental Conditions: From 2016 into the Future. Pediatr. Neurol. 2016, 65, 1–13. [Google Scholar] [CrossRef]
- Lee, A.W.; Ventola, P.; Budimirovic, D.; Berry-Kravis, E.; Visootsak, J. Clinical Development of Targeted Fragile X Syndrome Treatments: An Industry Perspective. Brain Sci. 2018, 8, 214. [Google Scholar] [CrossRef] [PubMed]
- Hagerman, R.J.; Berry-Kravis, E.; Hazlett, H.C.; Bailey, D.B., Jr.; Moine, H.; Kooy, R.F.; Tassone, F.; Gantois, I.; Sonenberg, N.; Mandel, J.L.; et al. Fragile X syndrome. Nat. Rev. Dis. Primers 2017, 3, 17065. [Google Scholar] [CrossRef]
- Telias, M.; Kuznitsov-Yanovsky, L.; Segal, M.; Ben-Yosef, D. Functional Deficiencies in Fragile X Neurons Derived from Human Embryonic Stem Cells. J. Neurosci. 2015, 35, 15295–15306. [Google Scholar] [CrossRef] [PubMed]
- Utami, K.H.; Skotte, N.H.; Colaço, A.R.; Yusof, N.A.B.M.; Sim, B.; Yeo, X.Y.; Bae, H.-G.; Garcia-Miralles, M.; Radulescu, C.I.; Chen, Q.; et al. Integrative Analysis Identifies Key Molecular Signatures Underlying Neurodevelopmental Deficits in Fragile X Syndrome. Biol. Psychiatry 2020, 88, 500–511. [Google Scholar] [CrossRef]
- Das Sharma, S.; Pal, R.; Reddy, B.K.; Selvaraj, B.T.; Raj, N.; Samaga, K.K.; Srinivasan, D.J.; Ornelas, L.; Sareen, D.; Livesey, M.; et al. Cortical neurons derived from human pluripotent stem cells lacking FMRP display altered spontaneous firing patterns. Mol. Autism 2020, 11, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.S.; Wu, H.; Krzisch, M.; Wu, X.; Graef, J.; Muffat, J.; Hnisz, D.; Li, C.; Yuan, B.; Xu, C.; et al. Rescue of Fragile X Syndrome Neurons by DNA Methylation Editing of the FMR1 Gene. Cell 2018, 172, 979–992. [Google Scholar] [CrossRef]
- Graef, J.D.; Wu, H.; Ng, C.; Sun, C.; Villegas, V.; Qadir, D.; Jesseman, K.; Warren, S.T.; Jaenisch, R.; Cacace, A.; et al. Partial FMRP expression is sufficient to normalize neuronal hyperactivity in Fragile X neurons. Eur. J. Neurosci. 2019, 51, 2143–2157. [Google Scholar] [CrossRef]
- Shi, Y.; Kirwan, P.; Smith, J.; Robinson, H.P.C.; Livesey, F.J. Human cerebral cortex development from pluripotent stem cells to functional excitatory synapses. Nat. Neurosci. 2012, 15, 477–486. [Google Scholar] [CrossRef]
- Zhang, Y.; Pak, C.; Han, Y.; Ahlenius, H.; Zhang, Z.; Chanda, S.; Marro, S.; Patzke, C.; Acuna, C.; Covy, J.; et al. Rapid Single-Step Induction of Functional Neurons from Human Pluripotent Stem Cells. Neuron 2013, 78, 785–798. [Google Scholar] [CrossRef] [PubMed]
- Avitzour, M.; Mor-Shaked, H.; Yanovsky-Dagan, S.; Aharoni, S.; Altarescu, G.; Renbaum, P.; Eldar-Geva, T.; Schonberger, O.; Levy-Lahad, E.; Epsztejn-Litman, S.; et al. FMR1 Epigenetic Silencing Commonly Occurs in Undifferentiated Fragile X-Affected Embryonic Stem Cells. Stem Cell Rep. 2014, 3, 699–706. [Google Scholar] [CrossRef] [PubMed]
- Telias, M.; Mayshar, Y.; Amit, A.; Ben-Yosef, D. Molecular Mechanisms Regulating Impaired Neurogenesis of Fragile X Syndrome Human Embryonic Stem Cells. Stem Cells Dev. 2015, 24, 2353–2365. [Google Scholar] [CrossRef]
- Park, C.-Y.; Halevy, T.; Lee, D.R.; Sung, J.J.; Lee, J.S.; Yanuka, O.; Benvenisty, N.; Kim, D.-W. Reversion of FMR1 Methylation and Silencing by Editing the Triplet Repeats in Fragile X iPSC-Derived Neurons. Cell Rep. 2015, 13, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Telias, M.; Segal, M.; Ben-Yosef, D. Neural differentiation of fragile X human embryonic stem cells reveals abnormal patterns of development despite successful neurogenesis. Dev. Biol. 2013, 374, 32–45. [Google Scholar] [CrossRef] [PubMed]
- Ooi, J.; Langley, S.R.; Xu, X.; Utami, K.H.; Sim, B.; Huang, Y.; Harmston, N.; Tay, Y.L.; Ziaei, A.; Zeng, R.; et al. Unbiased Profiling of Isogenic Huntington Disease hPSC-Derived CNS and Peripheral Cells Reveals Strong Cell-Type Specificity of CAG Length Effects. Cell Rep. 2019, 26, 2494–2508. [Google Scholar] [CrossRef]
- Tandon, R.; Brändl, B.; Baryshnikova, N.; Landshammer, A.; Steenpaß, L.; Keminer, O.; Pless, O.; Müller, F.-J. Generation of two human isogenic iPSC lines from fetal dermal fibroblasts. Stem Cell Res. 2018, 33, 120–124. [Google Scholar] [CrossRef]
- Soldner, F.; Laganière, J.; Cheng, A.W.; Hockemeyer, D.; Gao, Q.; Alagappan, R.; Khurana, V.; Golbe, L.I.; Myers, R.H.; Lindquist, S.; et al. Generation of Isogenic Pluripotent Stem Cells Differing Exclusively at Two Early Onset Parkinson Point Mutations. Cell 2011, 146, 318–331. [Google Scholar] [CrossRef]
- Urbach, A.; Bar-Nur, O.; Daley, G.Q.; Benvenisty, N. Differential Modeling of Fragile X Syndrome by Human Embryonic Stem Cells and Induced Pluripotent Stem Cells. Cell Stem Cell 2010, 6, 407–411. [Google Scholar] [CrossRef]
- Shi, Y.; Kirwan, P.; Livesey, F.J. Directed differentiation of human pluripotent stem cells to cerebral cortex neurons and neural networks. Nat. Protoc. 2012, 7, 1836–1846. [Google Scholar] [CrossRef] [PubMed]
- Sagata, N.; Iwaki, A.; Aramaki, T.; Takao, K.; Kura, S.; Tsuzuki, T.; Kawakami, R.; Ito, I.; Kitamura, T.; Sugiyama, H.; et al. Comprehensive behavioural study of GluR4 knockout mice: Implication in cognitive function. Genes, Brain Behav. 2010, 9, 899–909. [Google Scholar] [CrossRef] [PubMed]
- Huupponen, J.T.; Atanasova, T.; Taira, T.; E Lauri, S. GluA4 subunit of AMPA receptors mediates the early synaptic response to altered network activity in the developing hippocampus. J. Neurophysiol. 2016, 115, 2989–2996. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.J.; Esteban, J.A.; Hayashi, Y.; Malinow, R. Postnatal synaptic potentiation: Delivery of GluR4-containing AMPA receptors by spontaneous activity. Nat. Neurosci. 2000, 3, 1098–1106. [Google Scholar] [CrossRef]
- Comery, T.A.; Harris, J.B.; Willems, P.J.; Oostra, B.A.; Irwin, S.A.; Weiler, I.J.; Greenough, W.T. Abnormal dendritic spines in fragile X knockout mice: Maturation and pruning deficits. Proc. Natl. Acad. Sci. 1997, 94, 5401–5404. [Google Scholar] [CrossRef]
- Thomas, C.C.; Combe, C.L.; Dyar, K.A.; Inglis, F.M. Modest alterations in patterns of motor neuron dendrite morphology in the Fmr1 knockout mouse model for fragile X. Int. J. Dev. Neurosci. 2008, 26, 805–811. [Google Scholar] [CrossRef]
- Doers, M.; Musser, M.T.; Nichol, R.; Berndt, E.R.; Baker, M.; Gomez, T.; Zhang, S.-C.; Abbeduto, L.; Bhattacharyya, A. iPSC-Derived Forebrain Neurons from FXS Individuals Show Defects in Initial Neurite Outgrowth. Stem Cells Dev. 2014, 23, 1777–1787. [Google Scholar] [CrossRef]
- Odawara, A.; Katoh, H.; Matsuda, N.; Suzuki, I. Physiological maturation and drug responses of human induced pluripotent stem cell-derived cortical neuronal networks in long-term culture. Sci. Rep. 2016, 6, 26181. [Google Scholar] [CrossRef]
- Kathuria, A.; Nowosiad, P.; Jagasia, R.; Aigner, S.; Taylor, R.D.; Andreae, L.; Gatford, N.; Lucchesi, W.; Srivastava, D.P.; Price, J. Stem cell-derived neurons from autistic individuals with SHANK3 mutation show morphogenetic abnormalities during early development. Mol. Psychiatry 2017, 23, 735–746. [Google Scholar] [CrossRef]
- Patel, A.B.; Loerwald, K.W.; Huber, K.M.; Gibson, J.R. Postsynaptic FMRP Promotes the Pruning of Cell-to-Cell Connections among Pyramidal Neurons in the L5A Neocortical Network. J. Neurosci. 2014, 34, 3413–3418. [Google Scholar] [CrossRef]
- Stephan, A.H.; Barres, B.A.; Stevens, B. The Complement System: An Unexpected Role in Synaptic Pruning During Development and Disease. Annu. Rev. Neurosci. 2012, 35, 369–389. [Google Scholar] [CrossRef] [PubMed]
- Gatto, C.L.; Broadie, K. Temporal requirements of the fragile X mental retardation protein in the regulation of synaptic structure. Development 2008, 135, 2637–2648. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, S.; Doering, L.C. Astrocytes Prevent Abnormal Neuronal Development in the Fragile X Mouse. J. Neurosci. 2010, 30, 4508–4514. [Google Scholar] [CrossRef]
- Jacobs, S.; Nathwani, M.; Doering, L.C. Fragile X astrocytes induce developmental delays in dendrite maturation and synaptic protein expression. BMC Neurosci. 2010, 11, 132. [Google Scholar] [CrossRef]
- Olmos-Serrano, J.L.; Paluszkiewicz, S.M.; Martin, B.S.; Kaufmann, W.E.; Corbin, J.G.; Huntsman, M.M. Defective GABAergic Neurotransmission and Pharmacological Rescue of Neuronal Hyperexcitability in the Amygdala in a Mouse Model of Fragile X Syndrome. J. Neurosci. 2010, 30, 9929–9938. [Google Scholar] [CrossRef] [PubMed]
- Testa-Silva, G.; Loebel, A.; Giugliano, M.; de Kock, C.P.; Mansvelder, H.D.; Meredith, R.M. Hyperconnectivity and Slow Synapses during Early Development of Medial Prefrontal Cortex in a Mouse Model for Mental Retardation and Autism. Cereb. Cortex 2012, 22, 1333–1342. [Google Scholar] [CrossRef]
- Kang, Y.; Zhou, Y.; Li, Y.; Han, Y.; Xu, J.; Niu, W.; Li, Z.; Liu, S.; Feng, H.; Huang, W.; et al. A human forebrain organoid model of fragile X syndrome exhibits altered neurogenesis and highlights new treatment strategies. Nat. Neurosci. 2021, 24, 1377–1391. [Google Scholar] [CrossRef]
- Rosato-Siri, M.D.; Zoccolan, D.; Furlan, F.; Ballerini, L. Interneurone bursts are spontaneously associated with muscle contractions only during early phases of mouse spinal network development: A study in organotypic cultures. Eur. J. Neurosci. 2004, 20, 2697–2710. [Google Scholar] [CrossRef]
- Suresh, J.; Radojicic, M.; Pesce, L.L.; Bhansali, A.; Wang, J.; Tryba, A.; Marks, J.; van Drongelen, W. Network burst activity in hippocampal neuronal cultures: The role of synaptic and intrinsic currents. J. Neurophysiol. 2016, 115, 3073–3089. [Google Scholar] [CrossRef]
- Gonçalves, J.T.; Anstey, J.E.; Golshani, P.; Portera-Cailliau, C. Circuit level defects in the developing neocortex of Fragile X mice. Nat. Neurosci. 2013, 16, 903–909. [Google Scholar] [CrossRef]
- Bertero, A.; Liska, A.; Pagani, M.; Parolisi, R.; Masferrer, M.E.; Gritti, M.; Pedrazzoli, M.; Galbusera, A.; Sarica, A.; Cerasa, A.; et al. Autism-associated 16p11.2 microdeletion impairs prefrontal functional connectivity in mouse and human. Brain 2018, 141, 2055–2065. [Google Scholar] [CrossRef] [PubMed]
- Mohns, E.J.; Blumberg, M.S. Synchronous Bursts of Neuronal Activity in the Developing Hippocampus: Modulation by Active Sleep and Association with Emerging Gamma and Theta Rhythms. J. Neurosci. 2008, 28, 10134–10144. [Google Scholar] [CrossRef]
- Liu, Z.; Geng, L.; Li, R.; He, X.; Zheng, J.Q.; Xie, Z. Frequency Modulation of Synchronized Ca2+Spikes in Cultured Hippocampal Networks through G-Protein-Coupled Receptors. J. Neurosci. 2003, 23, 4156–4163. [Google Scholar] [CrossRef]
- Olshausen, B.A.; Field, D.J. Sparse coding of sensory inputs. Curr. Opin. Neurobiol. 2004, 14, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Just, M.A.; Cherkassky, V.L.; Keller, T.A.; Minshew, N.J. Cortical activation and synchronization during sentence comprehension in high-functioning autism: Evidence of underconnectivity. Brain 2004, 127, 1811–1821. [Google Scholar] [CrossRef]
- Gilbert, C.D.; Sigman, M. Brain States: Top-Down Influences in Sensory Processing. Neuron 2007, 54, 677–696. [Google Scholar] [CrossRef] [PubMed]
- Miller, L.J.; Mcintosh, D.N.; Mcgrath, J.; Shyu, V.; Lampe, M.; Taylor, A.K.; Tassone, F.; Neitzel, K.; Stackhouse, T.; Hagerman, R.J. Electrodermal Responses to Sensory Stimuli in Individuals With Fragile X Syndrome: A Preliminary Report. Am. J. Med. Genet. 1999, 83, 268–279. [Google Scholar] [CrossRef]
- Qiu, L.-F.; Hao, Y.-H.; Li, Q.-Z.; Xiong, Z.-Q. Fragile X syndrome and epilepsy. Neurosci. Bull. 2008, 24, 338–344. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhou, Y.; Kumari, D.; Sciascia, N.; Usdin, K. CGG-repeat dynamics and FMR1 gene silencing in fragile X syndrome stem cells and stem cell-derived neurons. Mol. Autism 2016, 7, 42. [Google Scholar] [CrossRef]
- Busskamp, V.; Lewis, N.; Guye, P.; Ng, A.; Shipman, S.L.; Byrne, S.M.; Sanjana, N.E.; Murn, J.; Li, Y.; Li, S.; et al. Rapid neurogenesis through transcriptional activation in human stem cells. Mol. Syst. Biol. 2014, 10, 760. [Google Scholar] [CrossRef]
- Schreiber, S.; Fellous, J.; Whitmer, D.; Tiesinga, P.; Sejnowski, T. A new correlation-based measure of spike timing reliability. Neurocomputing 2003, 52-54, 925–931. [Google Scholar] [CrossRef]
| Antibody | Experiment (Dilution) | Host | Supplier | Catalog # |
|---|---|---|---|---|
| OCT4 | IF (1:60) | Mouse | Santa-Cruz | sc-5279 |
| FMRP | IF (1:100); WB (1:2000) | Mouse | Biolegend | BLG-834601 |
| Tuj1 | IF (1:2000) | Mouse | Biolegend | BLG-801201 |
| Synapsin-I | IF (1:500); WB (1:1000) | Rabbit | Merck | AB1543 |
| GluA2 | IF (1:200); WB (1:5000) | Rabbit | Abcam | ab133477 |
| Alexa Fluor 488 | IF (1:500) | Rabbit | Thermo Fisher | A11008 |
| Alexa Fluor 488 | IF (1:400) | Mouse | Thermo Fisher | A21202 |
| Alexa Fluor 594 | IF (1:500) | Rabbit | Thermo Fisher | A11012 |
| β-actin | WB (1:5000) | Mouse | Abcam | ab8226 |
| horseradish peroxidase | WB (1:5000) | Mouse | Cell Signaling Technology | 7076s |
| horseradish peroxidase | (WB 1:5000) | Rabbit | Cell Signaling Technology | 7074s |
| Gene | Forward | Reverse |
|---|---|---|
| GUSB | ATCGATGACATCACCGTCACCA | TCCAAAAGACGCACTTCCAACTTG |
| OCT4 | TTCAGCCAAACGACCATCTG | GGTTCGCTTTCTCTTTCGGG |
| TuJ1 | TTT GGA CAT CTC TTC AGG CC | TTT CAC ACT CCT TCC GCA C |
| MAP2 | CCTAAGCCATGTGACATCCA | CTGCCTGGGGACTGTGTAAT |
| PAX6 | GTGTCCAACGGATGTGTGAG | CTAGCCAGGTTGCGAAGAAC |
| FMR1 | TCAAGGGTTGGACCTAATGC | GCAGGAAGCTCTCCCTCTCT |
| FOXG1 | TCAGAACTCGCTGGGCAACA | CAGAGCAGGGCACCGACA |
| BRN2 | AAAGTAACTGTCAAATGCGCG | GCTGTAGTGGTTAGACGCTG |
| SYN1 | CCCCAATCACAAAGAAATGCTC | ATGTCCTGGAAGTCATGCTG |
| PSD95 | AGTCAGAAATACCGCTACCAAG | CCGTTCACCTGCAACTCATATC |
| GRIA1 | TGATGGAAAATACGGAGCCC | CTTCCCGGACCAAAGTGATAG |
| GRIA3 | GTCTTTGGTTTTCCTTGGGTG | CAGCGAGATTGGCAGTATAGG |
| GRIA4 | CAAGGAGAGGAAATGCTGGG | ACGTCCATAGTGGTCAAACTG |
| GRIK1 | CCTGATGCCTAACACCACATTAACCT | CTGTATGTGTGGAACTTCGAGAGCA |
| GRIN1 | GAGAAGGAGAACATCACCGAC | GTC CCCATCCTCATTGAACTC |
| mGluR5 | GGTCCAAGAAAAGCAACATCATCAGAT | ATCATCAGTGGGCCAAGACCC |
| vGlut2 | TGGTCGTTGGCTATTCTCATAC | ATACTGGCATATCTTGGAGCG |
| GABRA2 | CAGCTACTTGCTGAGTGATAGCAAC | CAGGGTCCCACACCAAGAAAAC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gildin, L.; Rauti, R.; Vardi, O.; Kuznitsov-Yanovsky, L.; Maoz, B.M.; Segal, M.; Ben-Yosef, D. Impaired Functional Connectivity Underlies Fragile X Syndrome. Int. J. Mol. Sci. 2022, 23, 2048. https://doi.org/10.3390/ijms23042048
Gildin L, Rauti R, Vardi O, Kuznitsov-Yanovsky L, Maoz BM, Segal M, Ben-Yosef D. Impaired Functional Connectivity Underlies Fragile X Syndrome. International Journal of Molecular Sciences. 2022; 23(4):2048. https://doi.org/10.3390/ijms23042048
Chicago/Turabian StyleGildin, Lital, Rossana Rauti, Ofir Vardi, Liron Kuznitsov-Yanovsky, Ben M. Maoz, Menahem Segal, and Dalit Ben-Yosef. 2022. "Impaired Functional Connectivity Underlies Fragile X Syndrome" International Journal of Molecular Sciences 23, no. 4: 2048. https://doi.org/10.3390/ijms23042048
APA StyleGildin, L., Rauti, R., Vardi, O., Kuznitsov-Yanovsky, L., Maoz, B. M., Segal, M., & Ben-Yosef, D. (2022). Impaired Functional Connectivity Underlies Fragile X Syndrome. International Journal of Molecular Sciences, 23(4), 2048. https://doi.org/10.3390/ijms23042048








