Biological Detoxification of Mycotoxins: Current Status and Future Advances
Abstract
:1. Introduction
2. Chemical Structure and Toxic Effects of Mycotoxins
2.1. Aflatoxins (AFs)
2.2. Trichothecenes
2.3. Ochratoxins
2.4. Zearalenone
2.5. Fumonisins
2.6. Patulin
3. Mycotoxin Removal by Probiotics
3.1. Fast Glance to Probiotic Properties
3.2. Mycotoxins Detoxification by Probiotics and the Related Mechanism
3.2.1. Cell-Binding of Mycotoxins
3.2.2. Biodegradation
4. Biodegradation of Mycotoxins by Microbial Consortia
5. Mycotoxin Degradation by Recombinant Enzymes
6. Application, Prospect and Challenges
6.1. Unexplored Mechanisms of Mycotoxins Degradation
6.2. Biosafety Evaluation of Degradation Products
6.3. Limited Practical Application in Real Matrices
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Taheur, F.B.; Kouidhi, B.; al Qurashi, Y.M.A.; Salah-Abbes, J.B.; Chaieb, K. Review: Biotechnology of mycotoxins detoxification using microorganisms and enzymes. Toxicon 2019, 160, 12–22. [Google Scholar] [CrossRef]
- Savi, G.D.; Piacentini, K.C.; Scussel, V.M. Ozone treatment efficiency in Aspergillus and Penicillium growth inhibition and mycotoxin degradation of stored wheat grains (Triticum aestivum L.). J. Food Process. Preserv. 2015, 39, 940–948. [Google Scholar] [CrossRef]
- Temba, B.A.; Sultanbawa, Y.; Kriticos, D.J.; Fox, G.P.; Harvey, J.J.; Fletcher, M.T. Tools for defusing a major global food and feed safety risk: Nonbiological postharvest procedures to decontaminate mycotoxins in foods and feeds. J. Agric. Food Chem. 2016, 64, 8959–8972. [Google Scholar] [CrossRef]
- Klaric, M.S.; Cvetnic, Z.; Pepeljnjak, S.; Kosalec, I. Co-occurrence of aflatoxins, ochratoxin A, fumonisins, and zearalenone in cereals and feed, determined by competitive direct enzyme-linked immunosorbent assay and thin-layer chromatography. Arh. Hig. Rada Toksikol. 2009, 60, 427–434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schatzmayr, G.; Zehner, F.; Taubel, M.; Schatzmayr, D.; Klimitsch, A.; Loibner, A.P.; Binder, E.M. Microbiologicals for deactivating mycotoxins. Mol. Nutr. Food Res. 2006, 50, 543–551. [Google Scholar] [CrossRef]
- Bocarov-Stancic, A.; Adamovic, M.; Salma, N.; Bodroza-Solarov, M.; Vuckovic, J.; Pantic, V. In vitro efficacy of mycotoxins adsorption by natural mineral adsorbents. Biotechnol. Anim. Husb. 2011, 27, 1241–1251. [Google Scholar] [CrossRef]
- He, J.; Zhou, T.; Young, J.C.; Boland, G.J.; Scott, P.M. Chemical and biological transformations for detoxification of trichothecene mycotoxins in human and animal food chains: A review. Trends Food Sci. Technol. 2010, 21, 67–76. [Google Scholar] [CrossRef]
- Li, P.; Su, R.; Yin, R.; Lai, D.; Wang, M.; Liu, Y.; Zhou, L. Detoxification of mycotoxins through biotransformation. Toxins 2020, 12, 121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, W.; Chang, J.; Wan, P.; Liu, C.; Yin, Q.; Lu, F.; Gao, T. Effect of the combined compound probiotics with mycotoxin–degradation enzyme on detoxifying aflatoxin B1, and zearalenone. J. Toxicol. Sci. 2018, 43, 377–385. [Google Scholar] [CrossRef] [Green Version]
- Afsah-Hejri, L.; Hajeb, P.; Ehsani, R.J. Application of ozone for degradation of mycotoxins in food: A review. Compr. Rev. Food Sci. Food Saf. 2020, 19, 1777–1808. [Google Scholar] [CrossRef]
- Zhu, Y.; Hassan, Y.I.; Lepp, D.; Shao, S.; Zhou, T. Strategies and methodologies for developing microbial detoxification systems to mitigate mycotoxins. Toxins 2017, 9, 130. [Google Scholar] [CrossRef] [Green Version]
- Lyagin, I.; Efremenko, E. Enzymes for detoxification of various mycotoxins: Origins and mechanisms of catalytic action. Molecules 2019, 24, 2362. [Google Scholar] [CrossRef] [Green Version]
- Sato, I.; Ito, M.; Ishizaka, M.; Ikunaga, Y.; Sato, Y.; Yoshida, S.; Koitabashi, M.; Tsushima, S. Thirteen novel deoxynivalenol-degrading bacteria are classified within two genera with distinct degradation mechanisms. FEMS Microbiol. Lett. 2012, 327, 110–117. [Google Scholar] [CrossRef] [Green Version]
- Liuzzi, V.C.; Fanelli, F.; Tristezza, M.; Haidukowski, M.; Picardi, E.; Manzari, C.; Lionetti, C.; Grieco, F.; Logrieco, A.F.; Thon, M.R.; et al. Transcriptional analysis of Acinetobacter sp. neg1, capable of degrading ochratoxin a. Front. Microbiol. 2016, 7, 2162. [Google Scholar] [CrossRef] [Green Version]
- Monachese, M.; Burton, J.P.; Reid, G. Bioremediation and tolerance of humans to heavy metals through microbial processes: A potential role for probiotics? Appl. Environ. Microbiol. 2012, 78, 6397–6404. [Google Scholar] [CrossRef] [Green Version]
- Rajasekharan, S.K.; Lee, J.H.; Zhao, Y.; Lee, J. The mycotoxin zearalenone hinders Candida albicans biofilm formation and hyphal morphogenesis. Indian J. Microbiol. 2018, 58, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xie, Y. Review on microbial degradation of zearalenone and aflatoxins. Grain Oil Sci. Technol. 2020, 3, 117–125. [Google Scholar] [CrossRef]
- Liu, B.; Peng, X.; Meng, X. Effective biodegradation of mycotoxin patulin by porcine pancreatic lipase. Front. Microbiol. 2018, 9, 615. [Google Scholar] [CrossRef] [Green Version]
- Al-Nussairawi, M.; Risa, A.; Garai, E.; Varga, E.; Szabo, I.; Csenki-Bakos, Z.; Kriszt, B.; Cserhati, M. Mycotoxin biodegradation ability of the Cupriavidus Genus. Curr. Microbiol. 2020, 77, 2430–2440. [Google Scholar] [CrossRef] [PubMed]
- Silva, L.J.G.; Pereira, A.; Pena, A.; Lino, C.M. Citrinin in foods and supplements: A review of occurrence and analytical methodologies. Foods 2020, 10, 14. [Google Scholar] [CrossRef]
- Marin, S.; Ramos, A.J.; Cano-Sancho, G.; Sanchis, V. Mycotoxins: Occurrence, toxicology, and exposure assessment. Food Chem. Toxicol. 2013, 60, 218–237. [Google Scholar] [CrossRef] [PubMed]
- Khan, G.I. Eco-friendly bacterial biodegradation of mycotoxins. Pure Appl. Biol. 2020, 9, 1708–1717. [Google Scholar] [CrossRef]
- Loi, M.; Fanelli, F.; Liuzzi, V.C.; Logrieco, A.F.; Mule, G. Mycotoxin biotransformation by native and commercial enzymes: Present and future perspectives. Toxins 2017, 9, 111. [Google Scholar] [CrossRef] [PubMed]
- Gil-Serna, J.; Vázquez, C.; Patiño, B. Mycotoxins in functional beverages: A Review. Beverages 2020, 6, 52. [Google Scholar] [CrossRef]
- Vanhoutte, I.; Audenaert, K.; de Gelder, L. Biodegradation of mycotoxins: Tales from known and unexplored worlds. Front. Microbiol. 2016, 7, 561. [Google Scholar] [PubMed] [Green Version]
- Winter, G.; Pereg, L. A review on the relation between soil and mycotoxins: Effect of aflatoxin on field, food and finance. Eur. J. Soil Sci. 2019, 70, 882–897. [Google Scholar] [CrossRef]
- Adebo, O.A.; Njobeh, P.B.; Sidu, S.; Adebiyi, J.A.; Mavumengwana, V. Aflatoxin B1, degradation by culture and lysate of a Pontibacter specie. Food Control 2017, 80, 99–103. [Google Scholar] [CrossRef]
- Afshar, P.; Shokrzadeh, M.; Raeisi, S.N.; Ghorbani-HasanSaraei, A.; Nasiraii, L.R. Aflatoxins biodetoxification strategies based on probiotic bacteria. Toxicon 2020, 178, 50–58. [Google Scholar] [CrossRef]
- Nazhand, A.; Durazzo, A.; Lucarini, M.; Souto, E.B.; Santini, A. Characteristics, occurrence, detection and detoxification of aflatoxins in foods and feeds. Foods 2020, 9, 644. [Google Scholar] [CrossRef]
- Polak-Sliwinska, M.; Paszczyk, B. Trichothecenes in food and feed, relevance to human and animal health and methods of detection: A systematic review. Molecules 2021, 26, 454. [Google Scholar] [PubMed]
- Bezuidenhout, C.C.; Prinsloo, M.; van der Walt, A.M. Multiplex PCR-based detection of potential fumonisin-producing Fusarium in traditional African vegetables. Environ. Toxicol. 2006, 21, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.; Cen, Y.; Ye, W.; Li, S.; Zhang, W. Recent advances on macrocyclic trichothecenes, their bioactivities and biosynthetic pathway. Toxins 2020, 12, 417. [Google Scholar] [CrossRef]
- Khaneghah, A.M.; Farhadi, A.; Nematollahi, A.; Vasseghian, Y.; Fakhri, Y. A systematic review and meta-analysis to investigate the concentration and prevalence of trichothecenes in the cereal-based food. Trends Food Sci. Technol. 2020, 102, 193–202. [Google Scholar] [CrossRef]
- Van der Fels-Klerx, H.; Stratakou, I. T-2, toxin and HT-2, toxin in grain and grain-based commodities in Europe: Occurrence, factors affecting occurrence, co-occurrence and toxicological effects. World Mycotoxin J. 2010, 3, 349–367. [Google Scholar] [CrossRef]
- McCormick, S.P.; Stanley, A.M.; Stover, N.A.; Alexander, N.J. Trichothecenes: From simple to complex mycotoxins. Toxins 2011, 3, 802–814. [Google Scholar] [CrossRef] [PubMed]
- Rocha, O.; Ansari, K.; Doohan, F.M. Effects of trichothecene mycotoxins on eukaryotic cells: A review. Food Addit. Contam. 2005, 22, 369–378. [Google Scholar] [CrossRef]
- Zhou, T.; He, J.; Gong, J. Microbial transformation of trichothecene mycotoxins. World Mycotoxin J. 2008, 1, 23–30. [Google Scholar] [CrossRef]
- Stanciu, O.; Juan, C.; Miere, D.; Berrada, H.; Loghin, F.; Manes, J. First study on trichothecene and zearalenone exposure of the Romanian population through wheat-based products consumption. Food Chem. Toxicol. 2018, 121, 336–342. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Mahato, D.K.; Sharma, B.; Borah, R.; Haque, S.; Mahmud, M.M.C.; Shah, A.K.; Rawal, D.; Bora, H.; Bui, S. Ochratoxins in food and feed: Occurrence and its impact on human health and management strategies. Toxicon 2020, 187, 151–162. [Google Scholar]
- Bayman, P.; Baker, J.L. Ochratoxins: A global perspective. Mycopathologia 2006, 162, 215–223. [Google Scholar] [CrossRef]
- Reddy, L.; Bhoola, K. Ochratoxins-food contaminants: Impact on human health. Toxins 2010, 2, 771–779. [Google Scholar] [CrossRef] [Green Version]
- Rai, A.; Dixit, S.; Singh, S.P.; Gautam, N.K.; Das, M.; Tripathi, A. Presence of zearalenone in cereal grains and its exposure risk assessment in indian population. J. Food Sci. 2018, 83, 3126–3133. [Google Scholar] [CrossRef]
- Al-Jaal, B.A.; Jaganjac, M.; Barcaru, A.; Horvatovich, P.; Latiff, A. Aflatoxin, fumonisin, ochratoxin, zearalenone and deoxynivalenol biomarkers in human biological fluids: A systematic literature review, 2001–2018. Food Chem. Toxicol. 2019, 129, 211–228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kowalska, K.; Habrowska-Gorczynska, D.E.; Piastowska-Ciesielska, A.W. Zearalenone as an endocrine disruptor in humans. Environ. Toxicol. Pharmacol 2016, 48, 141–149. [Google Scholar] [CrossRef]
- Popiel, D.; Koczyk, G.; Dawidziuk, A.; Gromadzka, K.; Blaszczyk, L.; Chelkowski, J. Zearalenone lactonohydrolase activity in Hypocreales and its evolutionary relationships within the epoxide hydrolase subset of a/b-hydrolases. BMC Microbiol. 2014, 14, 82. [Google Scholar] [CrossRef]
- Li, Y.-S.; Zhou, Y.; Lu, S.-Y.; Guo, D.-J.; Ren, H.-L.; Meng, X.-M.; Zhi, B.-H.; Lin, C.; Wang, Z.; Li, X.-B.; et al. Development of a one-step test strip for rapid screening of fumonisins B1, B2, and B3, in maize. Food Control 2012, 24, 72–77. [Google Scholar] [CrossRef]
- Mirasoli, M.; Buragina, A.; Dolci, L.S.; Simoni, P.; Anfossi, L.; Giraudi, G.; Roda, A. Chemiluminescence-based biosensor for fumonisins quantitative detection in maize samples. Biosens. Bioelectron. 2012, 32, 283–287. [Google Scholar] [CrossRef] [PubMed]
- Silva, L.J.; Pena, A.; Lino, C.M.; Fernandez, M.F.; Manes, J. Fumonisins determination in urine by LC-MS-MS. Anal. Bioanal. Chem. 2010, 396, 809–816. [Google Scholar] [CrossRef] [PubMed]
- Voss, K.A.; Smith, G.W.; Haschek, W.M. Fumonisins: Toxicokinetics, mechanism of action and toxicity. Animal Feed Sci. Technol. 2007, 137, 299–325. [Google Scholar] [CrossRef]
- Morgavi, D.P.; Riley, R.T. Fusarium and their toxins: Mycology, occurrence, toxicity, control and economic impact. Animal Feed Sci. Technol. 2007, 137, 199–200. [Google Scholar] [CrossRef]
- Diao, E.; Hou, H.; Hu, W.; Dong, H.; Li, X. Removing and detoxifying methods of patulin: A review. Trends Food Sci. Technol. 2018, 81, 139–145. [Google Scholar] [CrossRef]
- Rodriguez, A.; Luque, M.I.; Andrade, M.J.; Rodriguez, M.; Asensio, M.A.; Cordoba, J.J. Development of real-time PCR methods to quantify patulin-producing molds in food products. Food Microbiol. 2011, 28, 1190–1199. [Google Scholar] [CrossRef] [PubMed]
- Tannous, J.; el Khoury, R.; Snini, S.P.; Lippi, Y.; el Khoury, A.; Atoui, A.; Lteif, R.; Oswald, I.P.; Puel, O. Sequencing, physical organization and kinetic expression of the patulin biosynthetic gene cluster from Penicillium expansum. Int. J. Food Microbiol. 2014, 189, 51–60. [Google Scholar] [PubMed]
- Zhang, H.; Ahima, J.; Yang, Q.; Zhao, L.; Zhang, X.; Zheng, X. A review on citrinin: Its occurrence, risk implications, analytical techniques, biosynthesis, physiochemical properties and control. Food Res. Int. 2021, 141, 110075. [Google Scholar] [CrossRef] [PubMed]
- Wan, M.L.Y.; Forsythe, S.J.; El-Nezami, H. Probiotics interaction with foodborne pathogens: A potential alternative to antibiotics and future challenges. Crit. Rev. Food Sci. Nutr. 2019, 59, 3320–3333. [Google Scholar] [CrossRef] [PubMed]
- Parvez, S.; Malik, K.A.; Kang, S.A.; Kim, H.Y. Probiotics and their fermented food products are beneficial for health. J. Appl. Microbiol. 2006, 100, 1171–1185. [Google Scholar] [CrossRef] [PubMed]
- Dalié, D.K.D.; Deschamps, A.M.; Richard-Forget, F. Lactic acid bacteria—Potential for control of mould growth and mycotoxins: A review. Food Control 2010, 21, 370–380. [Google Scholar] [CrossRef]
- Gaggia, F.; Mattarelli, P.; Biavati, B. Probiotics and prebiotics in animal feeding for safe food production. Int. J. Food Microbiol. 2010, 141, S15–S28. [Google Scholar] [CrossRef]
- Kesarcodi-Watson, A.; Kaspar, H.; Lategan, M.J.; Gibson, L. Probiotics in aquaculture: The need, principles and mechanisms of action and screening processes. Aquaculture 2008, 274, 1–14. [Google Scholar] [CrossRef]
- Zoghia, A.; Khosravi-Darani, K.; Sohrabvandib, S. Surface binding of toxins and heavy metals by probiotics. Mini Rev. Med. Chem. 2014, 14, 84–98. [Google Scholar]
- Shetty, P.H.; Jespersen, L. Saccharomyces cerevisiae and lactic acid bacteria as potential mycotoxin decontaminating agents. Trends Food Sci. Technol. 2006, 17, 48–55. [Google Scholar] [CrossRef]
- Chiocchetti, G.M.; Jadan-Piedra, C.; Monedero, V.; Zuniga, M.; Velez, D.; Devesa, V. Use of lactic acid bacteria and yeasts to reduce exposure to chemical food contaminants and toxicity. Crit. Rev. Food Sci. Nutr. 2019, 59, 1534–1545. [Google Scholar] [CrossRef] [PubMed]
- Hatab, S.; Yue, T.; Mohamad, O. Removal of patulin from apple juice using inactivated lactic acid bacteria. J. Appl. Microbiol. 2012, 112, 892–899. [Google Scholar] [CrossRef] [PubMed]
- Peltonen, K.; El-Nezami, H.; Haskard, C.; Ahokas, J.; Salminen, S. Aflatoxin B1, binding by dairy strains of lactic acid bacteria and bifidobacteria. J. Dairy Sci. 2001, 84, 2152–2156. [Google Scholar] [CrossRef]
- Prete, V.D.; Rodriguez, H.; Carrascosa, A.V. In vitro removal of ochratoxin a by wine lactic acid bacteria. J. Food Prot. 2007, 70, 2155–2160. [Google Scholar] [CrossRef] [PubMed]
- Sangsila, A.; Faucet-Marquis, V.; Pfohl-Leszkowicz, A.; Itsaranuwat, P. Detoxification of zearalenone by Lactobacillus pentosus strains. Food Control 2016, 62, 187–192. [Google Scholar] [CrossRef]
- Vinderola, G.; Ritieni, A. Role of probiotics against mycotoxins and their deleterious effects. J. Food Res. 2014, 4, 10–21. [Google Scholar] [CrossRef]
- Chapot-Chartier, M.P.; Kulakauskas, S. Cell wall structure and function in lactic acid bacteria. Microb. Cell Factories 2014, 13, S9. [Google Scholar] [CrossRef] [Green Version]
- Niderkorn, V.; Morgavi, D.P.; Aboab, B.; Lemaire, M.; Boudra, H. Cell wall component and mycotoxin moieties involved in the binding of fumonisin B1, and B2, by lactic acid bacteria. J. Appl. Microbiol. 2009, 106, 977–985. [Google Scholar] [CrossRef] [PubMed]
- Haskard, C.; Binnion, C.; Ahokas, J. Factors affecting the sequestration of aflatoxin by Lactobacillus rhamnosus strain GG. Chem. Biol. Interact. 2000, 128, 39–49. [Google Scholar] [CrossRef]
- Hernandez-Mendoza, A.; Guzman-de-Pena, D.; Garcia, H.S. Key role of teichoic acids on aflatoxin B binding by probiotic bacteria. J. Appl. Microbiol. 2009, 107, 395–403. [Google Scholar] [CrossRef] [PubMed]
- Jouany, J.P.; Yiannikouris, A.; Bertin, G. The chemical bonds between mycotoxins and cell wall components of Saccharomyces cerevisiae have been identified. Arch. Zootech. 2005, 8, 26–50. [Google Scholar]
- Yiannikouris, A.; Poughon, L.; Cameleyre, X.; Dussap, C.G.; François, J.; Bertin, G.; Jouany, J.P. A novel technique to evaluate interactions between Saccharomyces cerevisiae cell wall and mycotoxins: Application to zearalenone. Biotechnol. Lett. 2003, 25, 783–789. [Google Scholar] [CrossRef] [PubMed]
- Harkai, P.; Szabó, I.; Cserháti, M.; Krifaton, C.; Risa, A.; Radó, J.; Balázs, A.; Berta, K.; Kriszt, B. Biodegradation of aflatoxin-B1, and zearalenone by Streptomyces sp. collection. Int. Biodeterior. Biodegrad. 2016, 108, 48–56. [Google Scholar] [CrossRef] [Green Version]
- Rao, K.R.; Vipin, A.V.; Hariprasad, P.; Appaiah, K.A.A.; Venkateswaran, G. Biological detoxification of Aflatoxin B1, by Bacillus licheniformis CFR1. Food Control 2017, 71, 234–241. [Google Scholar]
- Juodeikiene, G.; Bartkiene, E.; Cernauskas, D.; Cizeikiene, D.; Zadeike, D.; Lele, V.; Bartkevics, V. Antifungal activity of lactic acid bacteria and their application for Fusarium mycotoxin reduction in malting wheat grains. LWT Food Sci. Technol. 2018, 89, 307–314. [Google Scholar] [CrossRef]
- Nathanail, A.V.; Gibson, B.; Han, L.; Peltonen, K.; Ollilainen, V.; Jestoi, M.; Laitila, A. The lager yeast Saccharomyces pastorianus removes and transforms Fusarium trichothecene mycotoxins during fermentation of brewer’s wort. Food Chem. 2016, 203, 448–455. [Google Scholar] [CrossRef] [PubMed]
- Festa, S.; Coppotelli, B.M.; Morelli, I.S. Bacterial diversity and functional interactions between bacterial strains from a phenanthrene-degrading consortium obtained from a chronically contaminated-soil. Int. Biodeterior. Biodegrad. 2013, 85, 42–51. [Google Scholar] [CrossRef]
- Zafra, G.; Absalon, A.E.; Anducho-Reyes, M.A.; Fernandez, F.J.; Cortes-Espinosa, D.V. Construction of PAH-degrading mixed microbial consortia by induced selection in soil. Chemosphere 2017, 172, 120–126. [Google Scholar] [CrossRef]
- Beeton, S.; Bull, A.T. Biotransformation and detoxification of T-2, toxin by soil and freshwater bacteria. Appl. Environ. Microbiol. 1989, 55, 190–197. [Google Scholar] [CrossRef] [Green Version]
- He, W.J.; Yuan, Q.S.; Zhang, Y.B.; Guo, M.W.; Gong, A.D.; Zhang, J.B.; Wu, A.B.; Huang, T.; Qu, B.; Li, H.P.; et al. Aerobic de-epoxydation of trichothecene mycotoxins by a soil bacterial consortium isolated using in situ soil enrichment. Toxins 2016, 8, 277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Zhao, C.; Zhang, D.; Zhao, M.; Peng, M.; Guo, P.; Cui, Z. Microbial degradation of zearalenone by a novel microbial consortium, NZDC-6, and its application on contaminated corncob by semisolid fermentation. J. Agric. Food Chem. 2020, 68, 1634–1644. [Google Scholar] [CrossRef]
- Zhao, Z.; Zhang, Y.; Gong, A.; Liu, N.; Chen, S.; Zhao, X.; Li, X.; Chen, L.; Zhou, C.; Wang, J. Biodegradation of mycotoxin fumonisin B1, by a novel bacterial consortium SAAS79. Appl. Microbiol. Biotechnol. 2019, 103, 7129–7140. [Google Scholar] [CrossRef]
- Wang, Y.; Zhao, C.; Zhang, D.; Zhao, M.; Zheng, D.; Lyu, Y.; Cheng, W.; Guo, P.; Cui, Z. Effective degradation of aflatoxin B1, using a novel thermophilic microbial consortium TADC7. Bioresour. Technol. 2017, 224, 166–173. [Google Scholar] [CrossRef]
- Kosicki, R.; Błajet-Kosicka, A.; Grajewski, J.; Twarużek, M. Multiannual mycotoxin survey in feed materials and feedingstuffs. Animal Feed Sci. Technol. 2016, 215, 165–180. [Google Scholar] [CrossRef]
- Sun, L.H.; Lei, M.Y.; Zhang, N.Y.; Gao, X.; Li, C.; Krumm, C.S.; Qi, D.S. Individual and combined cytotoxic effects of aflatoxin B1, zearalenone, deoxynivalenol and fumonisin B1, on BRL 3A rat liver cells. Toxicon 2015, 95, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, C.; Zhang, D.; Zhao, M.; Zheng, D.; Peng, M.; Cheng, W.; Guo, P.; Cui, Z. Simultaneous degradation of aflatoxin B1, and zearalenone by a microbial consortium. Toxicon 2018, 146, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Wang, T.; Wang, P.; Yin, Q.; Liu, C.; Zhu, Q.; Lu, F.; Gao, T. Compound probiotics alleviating aflatoxin B1, and zearalenone toxic effects on broiler production performance and gut microbiota. Ecotoxicol. Environ. Saf. 2020, 194, 110420. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Chang, J.; Wang, P.; Liu, C.; Yin, Q.; Song, A.; Gao, T.; Dang, X.; Lu, F. Effect of compound probiotics and mycotoxin degradation enzymes on alleviating cytotoxicity of swine jejunal epithelial cells induced by aflatoxin B(1) and zearalenone. Toxins 2019, 11, 12. [Google Scholar] [CrossRef] [Green Version]
- Hartinger, D.; Moll, W. Fumonisin elimination and prospects for detoxification by enzymatic transformation. World Mycotoxin J. 2011, 4, 271–283. [Google Scholar] [CrossRef]
- Alberts, J.F.; Gelderblom, W.C.; Botha, A.; van Zyl, W.H. Degradation of aflatoxin B(1) by fungal laccase enzymes. Int. J. Food Microbiol. 2009, 135, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Ece, S.; Lambertz, C.; Fischer, R.; Commandeur, U. Heterologous expression of a Streptomyces cyaneus laccase for biomass modification applications. AMB Express 2017, 7, 86. [Google Scholar] [CrossRef]
- Yu, Y.; Wu, H.; Tang, Y.; Qiu, L. Cloning, expression of a peroxiredoxin gene from Acinetobacter sp. SM04, and characterization of its recombinant protein for zearalenone detoxification. Microbiol. Res. 2012, 167, 121–126. [Google Scholar] [CrossRef]
- Tang, Y.; Xiao, J.; Chen, Y.; Yu, Y.; Xiao, X.; Yu, Y.; Wu, H. Secretory expression and characterization of a novel peroxiredoxin for zearalenone detoxification in Saccharomyces cerevisiae. Microbiol. Res. 2013, 168, 6–11. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.; Wang, Q.; Zhou, Y.; Yin, L.; Zhang, G.; Ma, Y. High-level expression of a ZEN-detoxifying gene by codon optimization and biobrick in Pichia pastoris. Microbiol. Res. 2016, 193, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Bi, K.; Zhang, W.; Xiao, Z.; Zhang, D. Characterization, expression and application of a zearalenone degrading enzyme from Neurospora crassa. AMB Express 2018, 8, 194. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Yin, L.; Hu, H.; Selvaraj, J.N.; Zhou, Y.; Zhang, G. Expression, functional analysis and mutation of a novel neutral zearalenone-degrading enzyme. Int. J. Biol. Macromol. 2018, 118, 1284–1292. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.C.; Hsu, T.C.; Cheng, K.C.; Liu, J.R. Expression of the Clonostachys rosea lactonohydrolase gene by Lactobacillus reuteri to increase its zearalenone-removing ability. Microb Cell Fact. 2017, 16, 69. [Google Scholar] [CrossRef] [PubMed]
- Heinl, S.; Hartinger, D.; Thamhesl, M.; Vekiru, E.; Krska, R.; Schatzmayr, G.; Moll, W.D.; Grabherr, R. Degradation of fumonisin B1, by the consecutive action of two bacterial enzymes. J. Biotechnol. 2010, 145, 120–129. [Google Scholar] [CrossRef]
- Hartinger, D.; Schwartz, H.; Hametner, C.; Schatzmayr, G.; Haltrich, D.; Moll, W.D. Enzyme characteristics of aminotransferase FumI of Sphingopyxis sp. MTA144, for deamination of hydrolyzed fumonisin B(1). Appl. Microbiol. Biotechnol. 2011, 91, 757–768. [Google Scholar] [CrossRef] [Green Version]
- Dobritzsch, D.; Wang, H.; Schneider, G.; Yu, S. Structural and functional characterization of ochratoxinase, a novel mycotoxin-degrading enzyme. Biochem. J. 2014, 462, 441–452. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Zhang, Y.; Yin, T.; Wang, J.; Zhang, X. Heterologous expression and characterization of a novel ochratoxin a degrading enzyme, N-acyl-L-amino acid amidohydrolase, from Alcaligenes faecalis. Toxins 2019, 11, 518. [Google Scholar] [CrossRef] [Green Version]
- Carere, J.; Hassan, Y.I.; Lepp, D.; Zhou, T. The enzymatic detoxification of the mycotoxin deoxynivalenol: Identification of DepA from the DON epimerization pathway. Microb. Biotechnol. 2018, 11, 1106–1111. [Google Scholar] [CrossRef] [PubMed]
- Rawal, S.; Yip, S.S.M.; Coulombe, R.A. Cloning, expression and functional characterization of cytochrome P450, 3A37, from turkey liver with high aflatoxin B1, epoxidation activity. Chem. Res. Toxicol. 2010, 23, 1322–1329. [Google Scholar] [CrossRef]
- Ito, M.; Sato, I.; Ishizaka, M.; Yoshida, S.; Koitabashi, M.; Yoshida, S.; Tsushima, S. Bacterial cytochrome P450, system catabolizing the Fusarium toxin deoxynivalenol. Appl. Environ. Microbiol. 2013, 79, 1619–1628. [Google Scholar] [CrossRef] [Green Version]
- Azam, M.S.; Yu, D.; Liu, N.; Wu, A. Degrading ochratoxin A and zearalenone mycotoxins using a multifunctional recombinant enzyme. Toxins 2019, 11, 301. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Qin, X.; Hao, Z.; Luo, H.; Yao, B.; Su, X. Degradation of four major mycotoxins by eight manganese peroxidases in presence of a dicarboxylic acid. Toxins 2019, 11, 566. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, G.; Wang, Y.; Man, H.; Lee, Y.W.; Shi, J.; Xu, J. Metabolomics-guided analysis reveals a two-step epimerization of deoxynivalenol catalyzed by the bacterial consortium IFSN-C1. Appl. Microbiol. Biotechnol. 2020, 104, 6045–6056. [Google Scholar] [CrossRef] [PubMed]
- Pinedo, C.; Wright, S.A.I.; Collado, I.G.; Goss, R.J.M.; Castoria, R.; Hrelia, P.; Maffei, F.; Duran-Patron, R. Isotopic labeling studies reveal the patulin detoxification pathway by the biocontrol yeast Rhodotorula kratochvilovae LS11. J. Nat. Prod. 2018, 81, 2692–2699. [Google Scholar] [CrossRef] [PubMed]
- Yu, K.; Yi, S.; Li, B.; Guo, F.; Peng, X.; Wang, Z.; Wu, Y.; Alvarez-Cohen, L.; Zhang, T. An integrated meta-omics approach reveals substrates involved in synergistic interactions in a bisphenol A (BPA)-degrading microbial community. Microbiome 2019, 7, 16. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Yang, H.; Zheng, X.; Zhao, L.; Gu, X.; Wang, K.; Apaliya, M.T.; Ahima, J.; Zhang, H. Protein and transcript profiling analysis of the response of Yarrowia lipolytica Y-2, in the degradation of ochratoxin A. Ann. Appl. Biol. 2019, 175, 98–110. [Google Scholar] [CrossRef]
- Tomin, M.; Tomic, S. Oxidase or peptidase? A computational insight into a putative aflatoxin oxidase from Armillariella tabescens. Proteins 2019, 87, 390–400. [Google Scholar] [CrossRef]
- Ianiri, G.; Idnurm, A.; Wright, S.A.; Duran-Patron, R.; Mannina, L.; Ferracane, R.; Ritieni, A.; Castoria, R. Searching for genes responsible for patulin degradation in a biocontrol yeast provides insight into the basis for resistance to this mycotoxin. Appl. Environ. Microbiol. 2013, 79, 3101–3115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berry, J.P.; Gantar, M.; Gibbs, P.D.; Schmale, M.C. The zebrafish (Danio rerio) embryo as a model system for identification and characterization of developmental toxins from marine and freshwater microalgae. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2007, 145, 61–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haq, M.; Gonzalez, N.; Mintz, K.; Jaja-Chimedza, A.; de Jesus, C.L.; Lydon, C.; Welch, A.; Berry, J.P. Teratogenicity of ochratoxin A and the degradation product, ochratoxin alpha, in the zebrafish (Danio rerio) embryo model of vertebrate development. Toxins 2016, 8, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adebiyi, J.A.; Kayitesi, E.; Adebo, O.A.; Changwa, R.; Njobeh, P.B. Food fermentation and mycotoxin detoxification: An African perspective. Food Control 2019, 106, 106731. [Google Scholar] [CrossRef]
- Oliveira, P.; Brosnan, B.; Jacob, F.; Furey, A.; Coffey, A.; Zannini, E.; Arendt, E.K. Lactic acid bacteria bioprotection applied to the malting process. Part II: Substrate impact and mycotoxin reduction. Food Control 2015, 51, 444–452. [Google Scholar] [CrossRef]

| Mycotoxins | Chemical Structure | Main Toxic Groups | Main Degradation Products | Organ/System Affected | Main Clinical Signs | Producing Fungi |
|---|---|---|---|---|---|---|
| Aflatoxins (B1, B2, G1, G2) | 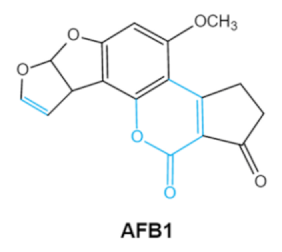 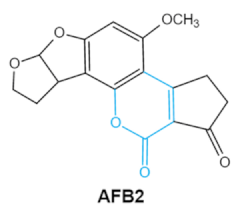  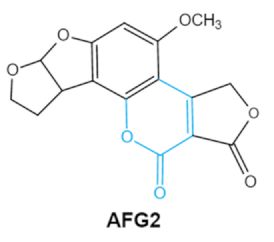 | Lactone ring Double bond in difuran ring moiety | AFB1-8,9dihydrodiol, AFB1-8,9-epoxide, dihydrohydroxyaflatoxin B1, | Liver, kidney, immune system | Hepatitis, carcinogenic, abdominal pain, vomiting, increased susceptibility to disease, immunosuppressive and carcinogenic effects | Aspergillus. Flavus A. Parasiticus A. nomius |
| Zearalenones (ZEA) |  | Lactone ring C-4 hydroxyl group | α-/β-zearalenol, α-/β-zearalanol,zearalenone-4-sulfate, 1-(3,5-dihydroxyphenyl)-6′-hydroxy-l’-undecen-l0′-one, (5S)-5-({2,4-dihydroxy-6-[(1E)-5-hydroxypent-1-en-1-yl]benzoyl}oxy)hexanoic acid | Reproductive tract, mainly female | Hyperestrogenism, Reproductive disorders | Fusarium graminearum(F. roseum) F. culmorum F. equiseti F. cerealis F. verticillioides F. incarnatum |
| Ochratoxins (A,B,C) (OTs) | 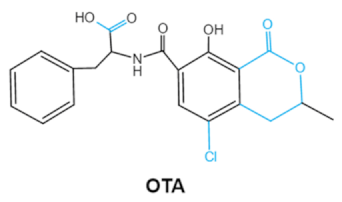 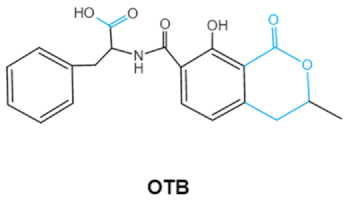 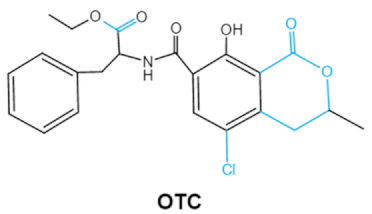 | Isocoumarin moiety Carboxyl group of the phenylalanine moietyCl group | L-βphenylalanine, OTα | Liver, kidney, immune system, inhibit RNA, DNA and protein synthesis in kidney | Nephritis, enlargement of kidney and hepatitis | A. ochraceus A. carbonarius A. niger P. verrucosum P. nordicum |
| Fumonisins FBs (B1, B2) | 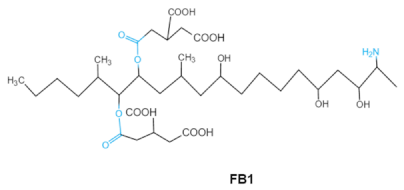  | Two tricarballylic acid side chains Free amino group | 2-oxo-12,16-dimethyl-3,5,10,14,15-icosanepentol hemiketal, NacetylAP1, | Lungs and heart (pig), central nervous system (horse), liver, immune system | Porcine pulmonary edema (PPE), equine leukoencephalomalacia | Fusarium section Liseola |
| Trichothecenes (DON, T-2, HT-2) TCNs | 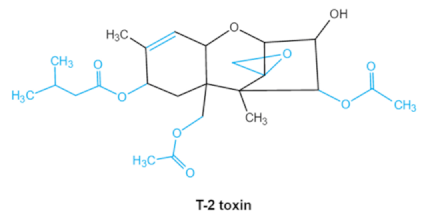 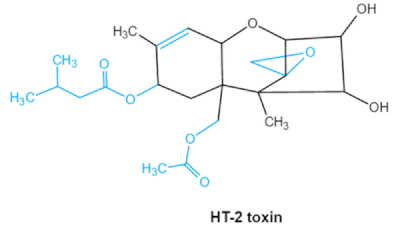  | Epoxide group Acylated side groups C9-10 double bond | HT-2 toxin, T-2 triol, T-2 tetraol, de-epoxy T-2 tetraol, 3α,7α,15α-triacetoxy-deoxynivalenol, de-epoxy deoxynivalenol, 3-acetyldeoxynivalenol, diaacetoxydeoxynivalenol. Epoxymonoacetoxyscirpenol, de-epoxyscirpentrio | Central nervous system, gastrointestinal tract, liver, immune system | Anorexia, vomiting, abdominal pains, cardiovascular dysfunction | F. acuminatum F. sporotrichioides F. langsethiae Fusariumgraminearum, F. culmorum F. cerealis F. culmorum F. graminearum F. sporotrichioides F. poae |
| Patulin PAT | 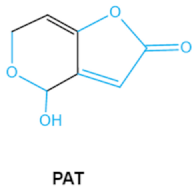 | Furan, pyran or lactone ring Hemiacetal | Ascladiol, hydroascladiol, desoxypatulinic acid, 3-keto-5-hydroxypentanal, glyoxylic acid | Gut epithelium, liver, kidney, immune system | Oral and epithelial lesion, loss of appetite Nausea, vomitin, gastric ulcers | Penicillim expansum Bysochlamis nívea Aspergillus clavatus P. griseofulvum |
| Mycotoxins | Microorganism | Reduction Rate (%) | Toxin Level | Degradation Condition | Reference |
|---|---|---|---|---|---|
| AFB1 | Streptomyces cacaoi subsp. Asoensis K234 | 88.34 ± 15.62 | 1 μg mL−1 | 5 days, 28 °C,170 rpm liquid LB medium | Harkai et al. (2016) |
| Streptomyces luteogriseus K144 | 79.93 | 1 μg mL−1 | 5 days, 28 °C,170 rpm liquid LB medium | ||
| Bacillus licheniformis CFR1 | 94.73 ± 1.09 | 500 ppb | liquid nutrient broth (NB) at 37 °C, 72 h | Rao et al. (2017) | |
| T-2 | P. pentosaceus KTU05-10 | 78.0 | 12.8–19.5 μg kg−1 | malting wheat grains with bacterial suspension at 18 °C for 30 min | Juodeikiene et al. (2018) |
| Saccharomyces pastorianus A15 | 31.0 | 5000 µg L−1 | 15 °C, 120 rpm, four days 11.5° Plato wort | Nathanail et al. (2016) | |
| HT-2 | P. pentosaceus KTU05-10 | 79.0 | 258–819 μg L−1 | malting wheat grains with bacterial suspension at 18 °C for 30 min | Juodeikiene et al. (2018) |
| ZEA | P. acidilactici | 38.0 | 19.5–873.7 μg L−1 | malting wheat grains with bacterial suspension at 18 °C for 30 min | Juodeikiene et al. (2018) |
| Streptomyces rimosus (K145, K189) | 100.0 | 1 μg mL−1 | 5 days, 28 °C,170 rpm liquid LB medium | Harkai et al. (2016) | |
| DON | P. pentosaceus KTU05-10 | 47.0 | 3370–6930 μg kg−1 | malting wheat grains with bacterial suspension at 18 °C for 30 min | Juodeikiene et al. (2018) |
| Saccharomyces pastorianus A15 | 15.0 | 400 µg L−1 | 11.5° Plato wort, 15 °C, 120 rpm for 4 days | Nathanail et al. (2016) |
| Enzyme | Gene Source | Target Toxin | Degrading Products | Expression System | Degrading Properties | Degradation Mechanism | Reference |
|---|---|---|---|---|---|---|---|
| Cytochrome P450 3A37 | Turkey liver | AFB1 | exo-AFBO aflatoxin Q1 | E. coli | exo-AFBO, Km: 287 ±21 µmol L−1 Vmax: 1.45 ± 0.07 nmol min−1 P450 aflatoxin Q1, Km: 302 ±51 µmol L−1 Vmax: 7.86 ±0.75 nmol min−1 | Epoxidation | Rawal et al. (2010) |
| Peroxiredoxin (Prx) | Acinetobacter sp. SM04 | ZEA | NM | E. coli | The optimum degradation pH and temperature was 9.0 and 70 °C in presence of H2O2 | Oxidation | Yu et al. (2012) |
| Lactonohydrolase Zhd518 | Clonostachys rosea | ZEA | NM | E. coli | Activity of 207.0 U mg−1 with the optimal temperature and pH at 40 °C and 8.0 | NM | Wang et al. (2018) |
| Peroxiredoxin | Acinetobacter sp. SM04 | ZEA | NM | Saccharomyces cerevisiae | Optimal activity at pH 9.0, 80 °C and H2O2 concentration of 20 mmol L−1 Thermal stable, alkali resistance | Oxidation | Tang et al. (2013) |
| Lactone hydrolase ZHD | Gliocladium roseum | ZEA | α-zearalenol and β-zearalenol | Pichia pastoris | Enzyme activity in shake flask fermentation was 22.5 U mL−1 with the specific activity of 4976.5 U mg−1 The maximum enzyme activity of the supernatant was 150.1 U mL−1 in 5-L fermenter | Cleavage of lactone ring | Xiang et al. (2016) |
| Lactonohydrolase | Clonostachys rosea | ZEA | 1-(3,5-dihydroxy-phenyl)-10-hydroxy-1-undecen6-one | Lactobacillus reuteri Pg4 | Did not affect cell growth, acid and bile salt tolerance | Cleavage of lactone ring | Yang et al. (2017) |
| Lactonase | Neurospora crassa | ZEA | NM | Pichia pastoris | Optimal activity at pH 8.0 and 45 °C, highly stable at pH 6.0–8.0 for 1 h at 37 °C, the maximal enzyme activity reached 290.6 U mL−1 in 30-L fermenter | NM | Bi et al. (2018) |
| Carboxylesterases, type B | Sphingopyxis sp. MTA144 | FB1 | NM | E. coli | NM | Deesterification | Heinl et al. (2010) |
| Aminotransferases, class III | HFB1 | Oxygen independence, temperature range 6–50 °C with an optimum at 35 °C, and pH adaptation 6–10 with an optimum at pH 8.5 | Deamination | ||||
| Aminotransferase FumI | Sphingopyxis sp. MTA144 | HFB1 | NM | E. coli | Optimal activity at pH 8.5 and 35 °C, low salt concentration, the kinetic parameters Km = 1.1 μmol L−1 and kcat = 104 min−1 | Eamination | Hartinger et al. (2011) |
| Putative amidase | Aspergillus niger | OTA | NM | NM | Thermostable, optimal activity at pH 6.0 and 66 °C | Hydrolysis | Dobritzsch et al. (2014) |
| N-acyl-L-amino acid amidohydrolase | Alcaligenes faecalis | OTA | β-phenylalanine | E. coli | Optimal activity at pH 6.5 and 50 °C | OTA amide bond hydrolysis | Zhang et al. (2019) |
| Dehydrogenase DepA | Devosia mutans 17-2-E-8 | DON | 3-keto-DON | E. coli | NM | Oxidation of C3 position | Carere et al. (2017) |
| Cytochrome P450 | Sphingomonas sp. strain KSM1 | DON | 16-hydroxy-deoxynivalenol | E. coli | kcat/Km of 6.4 mmol L−1 s−1 | Hydroxylation | Ito et al. (2013) |
| Fusion ZHDCP enzyme | Clonostachy rosea B. amyloliquefaciens ASAG | ZEA OTA | HZEA, DZEA, OTAα | E. coli | 100% degradation rate at pH 7 and 30 °C in 2 h | Hydrolysis Removal of an amino acid from the end of a peptide chain | Azam et al. (2019) |
| Manganese peroxidase | Irpex lacteus, Phanerochaete chrysosporium, Ceriporiopsis subvermispora, Nematoloma frowardii | AFB1 ZEA DON FB1 | AFB1-8,9-epoxide | E. coli | In the presence of dicarboxylic acid malonate | Oxidoreduction | Wang et al. (2019) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, L.; Xie, M.; Wei, D. Biological Detoxification of Mycotoxins: Current Status and Future Advances. Int. J. Mol. Sci. 2022, 23, 1064. https://doi.org/10.3390/ijms23031064
Liu L, Xie M, Wei D. Biological Detoxification of Mycotoxins: Current Status and Future Advances. International Journal of Molecular Sciences. 2022; 23(3):1064. https://doi.org/10.3390/ijms23031064
Chicago/Turabian StyleLiu, Lu, Mei Xie, and Dong Wei. 2022. "Biological Detoxification of Mycotoxins: Current Status and Future Advances" International Journal of Molecular Sciences 23, no. 3: 1064. https://doi.org/10.3390/ijms23031064
APA StyleLiu, L., Xie, M., & Wei, D. (2022). Biological Detoxification of Mycotoxins: Current Status and Future Advances. International Journal of Molecular Sciences, 23(3), 1064. https://doi.org/10.3390/ijms23031064






