Two Single Nucleotide Polymorphisms in the Purinergic Receptor P2X7 Gene Are Associated with Disease Severity in Multiple Sclerosis
Abstract
1. Introduction
2. Results
2.1. Clinical Assessment
2.2. P2X7R Polymorphisms
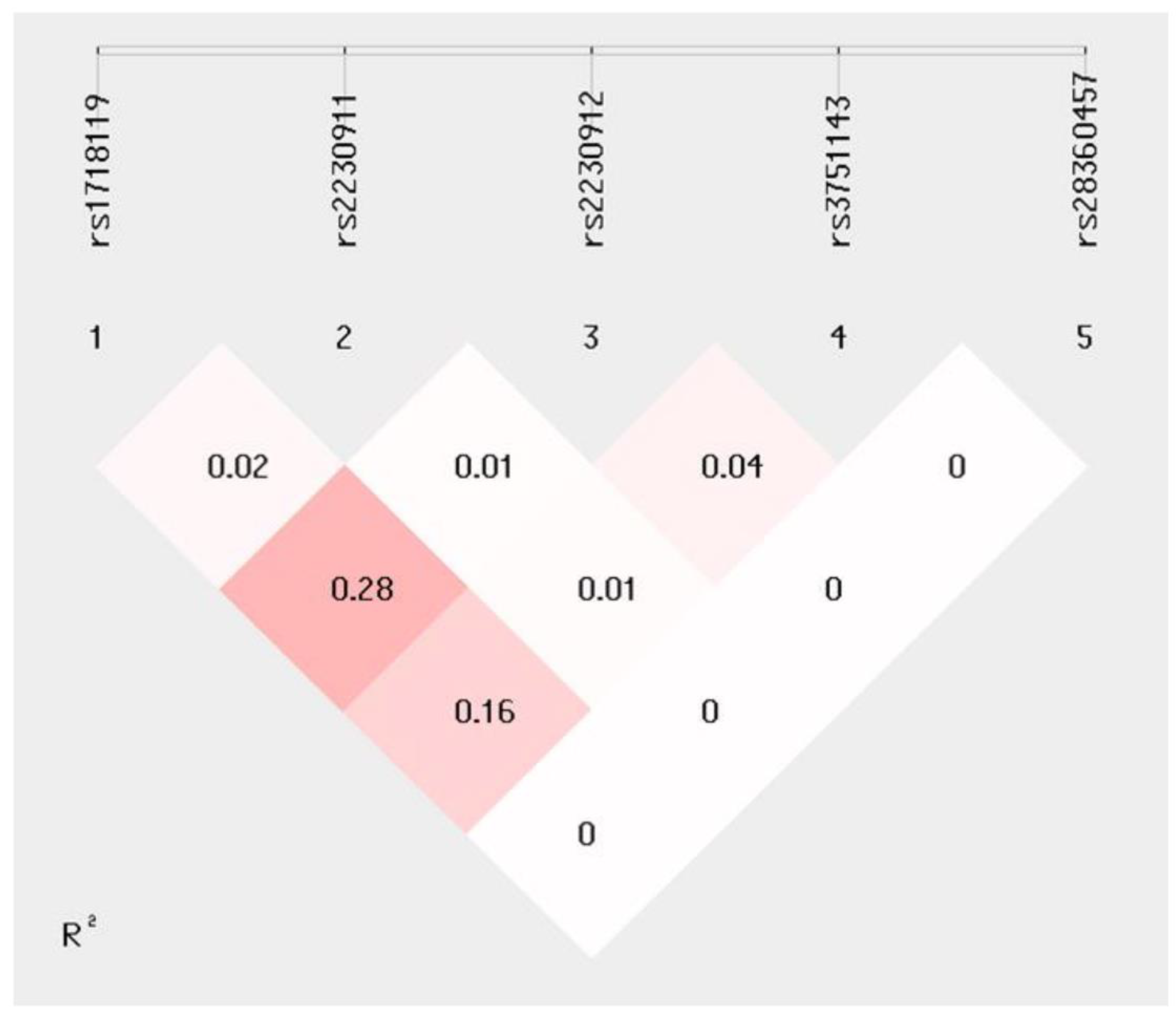
2.3. Haplotype Analyses
3. Discussion
4. Materials and Methods
4.1. Patients and Controls
4.1.1. Samples Collection and DNA Extraction
4.1.2. P2X7R SNPs Genotyping
4.1.3. HLA-DRB1*15 Characterization
4.1.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Walton, C.; King, R.; Rechtman, L.; Kaye, W.; Leray, E.; Marrie, R.A.; Robertson, N.; La Rocca, N.; Uitdehaag, B.; van der Mei, I.; et al. Rising prevalence of multiple sclerosis worldwide: Insights from the Atlas of MS, third edition. Mult. Scler. 2020, 26, 1816–1821. [Google Scholar] [CrossRef] [PubMed]
- Tarlinton, R.E.; Martynova, E.; Rizvanov, A.A.; Khaiboullina, S.; Verma, S. Role of Viruses in the Pathogenesis of Multiple Sclerosis. Viruses 2020, 12, 643. [Google Scholar] [CrossRef] [PubMed]
- Lopes Pinheiro, M.A.; Kooij, G.; Mizee, M.R.; Kamermans, A.; Enzmann, G.; Lyck, R.; Schwaninger, M.; Engelhardt, B.; de Vries, H.E. Immune cell trafficking across the barriers of the central nervous system in multiple sclerosis and stroke. Biochim. Biophys. Acta. 2016, 1862, 461–471. [Google Scholar] [CrossRef] [PubMed]
- Lassmann, H. Multiple Sclerosis Pathology. Cold Spring Harb. Perspect. Med. 2018, 8, a028936. [Google Scholar] [CrossRef] [PubMed]
- Barizzone, N.; Leone, M.; Pizzino, A.; Kockum, I.; Multiple MS Consortium; Martinelli-Boneschi, F.; D’Alfonso, S. A Scoping Review on Body Fluid Biomarkers for Prognosis and Disease Activity in Patients with Multiple Sclerosis. J. Pers. Med. 2022, 12, 1430. [Google Scholar] [CrossRef] [PubMed]
- Burnstock, G. Purinergic Signalling and Neurological Diseases: An Update. CNS Neurol. Disord. Drug Targets 2017, 16, 257–265. [Google Scholar] [CrossRef]
- Domercq, M.; Matute, C. Targeting P2X4 and P2X7 receptors in multiple sclerosis. Curr. Opin. Pharmacol. 2019, 47, 119–125. [Google Scholar] [CrossRef]
- Qian, Y.; Qian, C.; Xie, K.; Fan, Q.; Yan, Y.; Lu, R.; Wang, L.; Zhang, M.; Wang, Q.; Mou, S.; et al. P2X7 receptor signaling promotes inflammation in renal parenchymal cells suffering from ischemia-reperfusion injury. Cell Death Dis. 2021, 12, 132. [Google Scholar] [CrossRef]
- Rotondo, J.C.; Mazziotta, C.; Lanzillotti, C.; Stefani, C.; Badiale, G.; Campione, G.; Martini, F.; Tognon, M. The Role of Purinergic P2X7 Receptor in Inflammation and Cancer: Novel Molecular Insights and Clinical Applications. Cancers 2022, 14, 1116. [Google Scholar] [CrossRef]
- Di Virgilio, F.; Dal Ben, D.; Sarti, A.C.; Giuliani, A.L.; Falzoni, S. The P2X7 Receptor in Infection and Inflammation. Immunity. 2017, 47, 15–31. [Google Scholar] [CrossRef]
- Rivas-Yanez, E.; Barrera-Avalos, C.; Parra-Tello, B.; Briceno, P.; Rosemblatt, M.V.; Saavedra-Almarza, J.; Rosemblatt, M.; AcunaCastillo, C.; Bono, M.R.; Sauma, D. P2X7 Receptor at the Crossroads of T Cell Fate. Int. J. Mol. Sci. 2020, 21, 4937. [Google Scholar] [CrossRef] [PubMed]
- Karmakar, M.; Katsnelson, M.A.; Dubyak, G.R.; Pearlman, E. Neutrophil P2X7 receptors mediate NLRP3 inflammasome dependent IL-1beta secretion in response to ATP. Nat. Commun. 2016, 7, 10555. [Google Scholar] [CrossRef] [PubMed]
- Surprenant, A.; Rassendren, F.; Kawashima, E.; North, R.A.; Buell, G.; Muragaki, Y.; Mundlos, S.; Upton, J.; Olsen, B.R. The Cytolytic P2Z Receptor for Extracellular ATP Identified as a P2X Receptor (P2X7). Science 1996, 272, 735–738. [Google Scholar] [CrossRef]
- Di Virgilio, F.; Chiozzi, P.; Falzoni, S.; Ferrari, D.; Sanz, J.M.; Venketaraman, V.; Baricordi, O.R. Cytolytic P2X purinoceptors. Cell Death Differ. 1998, 5, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Savio, L.E.B.; de Andrade, M.P.; da Silva, C.G.; Coutinho-Silva, R. The P2X7 Receptor in Inflammatory Diseases: Angel or Demon? Front. Pharmacol. 2018, 9, 52. [Google Scholar] [CrossRef]
- Kopp, R.; Krautloher, A.; Ramírez-Fernández, A.; Nicke, A. P2X7 Interactions and Signaling—Making Head or Tail of It. Front. Mol. Neurosci. 2019, 12, 183. [Google Scholar] [CrossRef]
- Stokes, L.; Fuller, S.J.; Sluyter, R.; Skarratt, K.K.; Gu, B.J.; Wiley, J.S. Two haplotypes of the P2X(7) receptor containing the Ala-348 to Thr polymorphism exhibit a gain-of-function effect and enhanced interleukin-1beta secretion. FASEB J. 2010, 24, 2916–2927. [Google Scholar] [CrossRef]
- Church, L.D.; Cook, G.P.; McDermott, M.F. Primer: Inflammasomes and interleukin 1beta in inflammatory disorders. Nat. Clin. Pract. Rheumatol. 2008, 4, 34–42. [Google Scholar] [CrossRef]
- Shemon, A.N.; Sluyter, R.; Fernando, S.L.; Clarke, A.L.; Dao-Ung, L.P.; Skarratt, K.K.; Saunders, B.M.; Tan, K.S.; Gu, B.J.; Fuller, S.J.; et al. A Thr357 to Ser polymorphism in homozygous and compound heterozygous subjects causes absent or reduced P2X7 function and impairs ATP-induced mycobacterial killing by macrophages. J. Biol. Chem. 2006, 281, 2079–2086. [Google Scholar] [CrossRef]
- Gu, B.J.; Zhang, W.; Worthington, R.A.; Sluyter, R.; Dao-Ung, P.; Petrou, S.; Barden, J.A.; Wiley, J.S. A Glu-496 to Ala polymorphism leads to loss of function of the human P2X7 receptor. J. Biol. Chem. 2001, 276, 11135–11142. [Google Scholar] [CrossRef]
- Sidoryk-Węgrzynowicz, M.; Strużyńska, L. Astroglial and Microglial Purinergic P2X7 Receptor as a Major Contributor to Neuroinflammation during the Course of Multiple Sclerosis. Int. J. Mol. Sci. 2021, 22, 8404. [Google Scholar] [CrossRef] [PubMed]
- Gu, B.J.; Field, J.; Dutertre, S.; Ou, A.; Kilpatrick, T.J.; Lechner-Scott, J.; Scott, R.; Lea, R.A.; Taylor, B.V.; Stankovich, J.; et al. A rare P2X7 variant Arg307Gln with absent pore formation function protects against neuroinflammation in multiple sclerosis. Hum. Mol. Genet. 2015, 24, 5644–5654. [Google Scholar] [CrossRef]
- Miller, C.M.; Boulter, N.R.; Fuller, S.J.; Zakrzewski, A.M.; Lees, M.P.; Saunders, B.M.; Wiley, J.S.; Smith, N.C. The Role of the P2X7 Receptor in Infectious Diseases. PLoS Pathog. 2011, 7, e1002212. [Google Scholar] [CrossRef] [PubMed]
- Revesz, T.; Kidd, D.; Thompson, A.J. A comparison of the pathology of primary and secondary progressive multiple sclerosis. Brain 1994, 117, 759–765. [Google Scholar] [CrossRef] [PubMed]
- Amadio, S.; Parisi, C.; Piras, E.; Fabbrizio, P.; Apolloni, S.; Montilli, C.; Luchetti, S.; Ruggieri, S.; Gasperini, C.; Laghi-Pasini, F.; et al. Modulation of P2X7 Receptor during Inflammation in Multiple Sclerosis. Front. Immunol. 2017, 8, 1529. [Google Scholar] [CrossRef]
- Illes, P. P2X7 Receptors Amplify CNS Damage in Neurodegenerative Diseases. Int. J. Mol. Sci. 2020, 21, 5996. [Google Scholar] [CrossRef]
- Thompson, A.J.; Banwell, B.L.; Barkhof, F.; Carroll, W.M.; Coetzee, T.; Comi, G.; Correale, J.; Fazekas, F.; Filippi, M.; Freedman, M.S.; et al. Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol. 2018, 17, 162–173. [Google Scholar] [CrossRef]
- Kalincik, T.; Kister, I.; Bacon, T.E.; Malpas, C.B.; Sharmin, S.; Horakova, D.; Kubala-Havrdova, E.; Patti, F.; Izquierdo, G.; Eichau, S.; et al. Multiple Sclerosis Severity Score (MSSS) improves the accuracy of individualized prediction in MS. Mult. Scler. 2022, 28, 1752–1761. [Google Scholar] [CrossRef]
- Jersild, C.; Fog, T.; Hansen, G.S.; Thomsen, M.; Svejgaard, A.; Dupont, B. Histocompatibility determinants in multiple sclerosis, with special reference to clinical course. Lancet 1973, 2, 1221–1225. [Google Scholar] [CrossRef]
- Maniatis, T.; Fritsch, E.F.; Sambrook, J. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Long Island, NY, USA, 1989. [Google Scholar]
- de Bakker, P.I.; McVean, G.; Sabeti, P.C.; Miretti, M.M.; Green, T.; Marchini, J.; Ke, X.; Monsuur, A.J.; Whittaker, P.; Delgado, M.; et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 2006, 38, 1166–1172. [Google Scholar] [CrossRef]
- Shi, Y.Y.; He, L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 2005, 15, 97–98. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, Z.; He, Z.; Tang, W.; Li, T.; Zeng, Z.; He, L.; Shi, Y. A partition-ligation-combination-subdivision EM algorithm for haplotype inference with multiallelic markers: Update of the SHEsis (http://analysis.bio-x.cn). Cell Res. 2009, 19, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
| Patients | ||||||
|---|---|---|---|---|---|---|
| SPMS n = 34 | RRMS n = 94 | Total MS n = 128 | HCs n = 189 | p Value | ||
| GENDER M/F: N | 15/19 | 31/63 | 46/82 | 54/135 | ns | |
| AGE AT ONSET § years: mean ± SD | * 32.3 ±10.8 | ° 30.4 ±10.1 | ^ 30.9 ± 10.3 | § 54.3 ±13.2 | ^*° vs. § < 0.001 | |
| AGE AT MSSS years: mean ± SD | * 53.1 ±9.9 | ° 47.3 ±9.5 | 48.8 ± 9.9 | * vs. ° < 0.01 | ||
| Disease duration years: mean ± SD | * 22.7 ±7.7 | ° 17.9 ±9.5 | 19.2 ± 9.3 | * vs. ° < 0.01 | ||
| MSSS: mean ± SD | * 7.0 ±2.2 | ° 3.6 ±2.7 | 4.5 ± 2.9 | * vs. ° < 0.001 | ||
| DRB1*15 pos: N (%) | 7 (20.6) | ° 28 (29.8) | ^ 35(27.3) | § 24 (12.7) | ^° vs. § < 0.001 | |
| MSSS | MSSS | MSSS | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SPMS (n = 34) | RRMS (n = 94) | Total (n = 128) | |||||||||||
| P2x7R SNPs | N (%) | Mean | SD | p Value | N (%) | Mean | SD | p Value | N (%) | Mean | SD | p Value | |
| rs1718119 Ala348Thr | GG | 12 (35.3) | 7.2 | 2.5 | 35 (37.2) | 2.3 | 1.7 | 47 (36.7) | 3.5 | 2.9 | |||
| AG | 16 (47.1) | 6.7 | 2.1 | 44 (46.8) | 4.5 | 2.8 | 60 (46.9) | 5.1 | 2.8 | ||||
| AA | 6 (17.6) | 7.5 | 1.6 | 0.56 | 15 (16.0) | 3.9 | 2.8 | <0.001 | 21 (16.4) | 5 | 3.0 | 0.02 | |
| MAF | AA + AG | 6.8 | 1.9 | 0.59 | 4.4 | 2.8 | <0.001 | 5.0 | 2.8 | 0.005 | |||
| rs2230911 Thr357Ser | CC | 32 (94.1) | 7.1 | 2.1 | 84 (89.3) | 3.7 | 2.7 | 116 (90.6) | 4.6 | 2.9 | |||
| CG | 2(5.9) | 4.9 | 2.5 | 0.16 | 10 (10.6) | 3.0 | 2.2 | 0.47 | 12 (9.4) | 3.3 | 2.3 | 0.15 | |
| rs2230912 Gln464Arg | AA | 25 (73.5) | 6.9 | 2.1 | 71 (74.7) | 3.2 | 2.4 | 95 (74.2) | 4.2 | 2.8 | |||
| AG | 9 (26.5) | 6.9 | 2.5 | 20 (21.1) | 5.1 | 3 | 30 (23.4) | 5.7 | 2.9 | ||||
| GG | 0 | 0.99 | 4 (4.2) | 3.1 | 3.8 | 0.01 | 3 (2.4) | 3.1 | 3.8 | 0.03 | |||
| MAF | GG + AG | 6.9 | 2.1 | 0.99 | 4.8 | 3.1 | 0.01 | 5.4 | 3.1 | 0.03 | |||
| rs3751143 Glu496Ala | AA | 24 (70.6) | 7.0 | 2.3 | 55 (58.5) | 3.2 | 2.5 | 79 (61.7) | 4.3 | 3.0 | |||
| AC | 8 (23.5) | 6.3 | 1.9 | 36 (38.3) | 4.2 | 2.8 | 44 (34.4) | 4.6 | 2.7 | ||||
| CC | 2 (5.9) | 8.8 | 0.3 | 0.33 | 3 (3.2) | 3.6 | 2.2 | 0.16 | 5 (3.9) | 5.7 | 3.3 | 0.59 | |
| MAF | CC + AC | 6.8 | 2.0 | 0.74 | 4.2 | 2.7 | 0.06 | 4.7 | 2.8 | 0.48 | |||
| rs2836045 Arg307Gln | GG | 33(97.1) | 6.9 | 2.2 | 92 (97.9) | 3.6 | 2.7 | 125 (97.7) | 4.4 | 2.9 | |||
| AG | 1 (2.9) | 8.2 | / | 0.59 | 2 (2.1) | 3.7 | 0.6 | 0.94 | 3 (2.3) | 5.2 | 2.6 | 0.67 | |
| HAPLOTYPE ASSOCIATION WITH MSSS | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SPMS n = 34 | RRMS n = 94 | Total MS n = 128 | HCs n = 189 | Haplotype | rs1718119 | rs2230911 | rs2230912 | rs3751143 | rs2836045 | SPMS | RRMS | Total MS |
| freq | freq | freq | freq | Beta value | Beta value | Beta value | ||||||
| 0.41 | 0.32 | 0.34 | 0.35 | P2X7R-1 | G | C | A | A | G | −0.05 | °°−1.54 | §−0.94 |
| 0.24 | 0.25 | 0.25 | 0.21 | P2X7R-2 | A | C | A | A | G | 0.29 | 0.53 | 0.51 |
| 0.18 | 0.22 | 0.21 | 0.21 | P2X7R-3 | G | C | A | C | G | 0.08 | 0.87 | 0.45 |
| 0.13 | 0.14 | 0.14 | 0.16 | P2X7R-4 | A | C | G | A | G | −0.07 | * 1.11 | 0.81 |
| 0 | 0.05 | 0.04 | 0.05 | P2X7R-5 | G | G | A | A | G | / | −0.68 | −1.9 |
| 0 | 0.01 | 0.01 | 0.01 | P2X7R-6 | G | C | A | A | A | / | 0.4 | 1.02 |
| 0.01 | 0 | 0.004 | 0.01 | P2X7R-7 | A | C | A | A | A | 1.5 | / | / |
| 0.03 | 0 | 0.01 | 0.01 | P2X7R-8 | A | G | G | A | G | −2.12 | / | / |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guerini, F.R.; Agliardi, C.; Bolognesi, E.; Zanzottera, M.; Caputo, D.; Pasanisi, M.B.; Rovaris, M.; Clerici, M. Two Single Nucleotide Polymorphisms in the Purinergic Receptor P2X7 Gene Are Associated with Disease Severity in Multiple Sclerosis. Int. J. Mol. Sci. 2022, 23, 15381. https://doi.org/10.3390/ijms232315381
Guerini FR, Agliardi C, Bolognesi E, Zanzottera M, Caputo D, Pasanisi MB, Rovaris M, Clerici M. Two Single Nucleotide Polymorphisms in the Purinergic Receptor P2X7 Gene Are Associated with Disease Severity in Multiple Sclerosis. International Journal of Molecular Sciences. 2022; 23(23):15381. https://doi.org/10.3390/ijms232315381
Chicago/Turabian StyleGuerini, Franca Rosa, Cristina Agliardi, Elisabetta Bolognesi, Milena Zanzottera, Domenico Caputo, Maria Barbara Pasanisi, Marco Rovaris, and Mario Clerici. 2022. "Two Single Nucleotide Polymorphisms in the Purinergic Receptor P2X7 Gene Are Associated with Disease Severity in Multiple Sclerosis" International Journal of Molecular Sciences 23, no. 23: 15381. https://doi.org/10.3390/ijms232315381
APA StyleGuerini, F. R., Agliardi, C., Bolognesi, E., Zanzottera, M., Caputo, D., Pasanisi, M. B., Rovaris, M., & Clerici, M. (2022). Two Single Nucleotide Polymorphisms in the Purinergic Receptor P2X7 Gene Are Associated with Disease Severity in Multiple Sclerosis. International Journal of Molecular Sciences, 23(23), 15381. https://doi.org/10.3390/ijms232315381








