Age-Related Decline in Nrf2/ARE Signaling Is Associated with the Mitochondrial DNA Damage and Cognitive Impairments
Abstract
:1. Introduction
2. Results
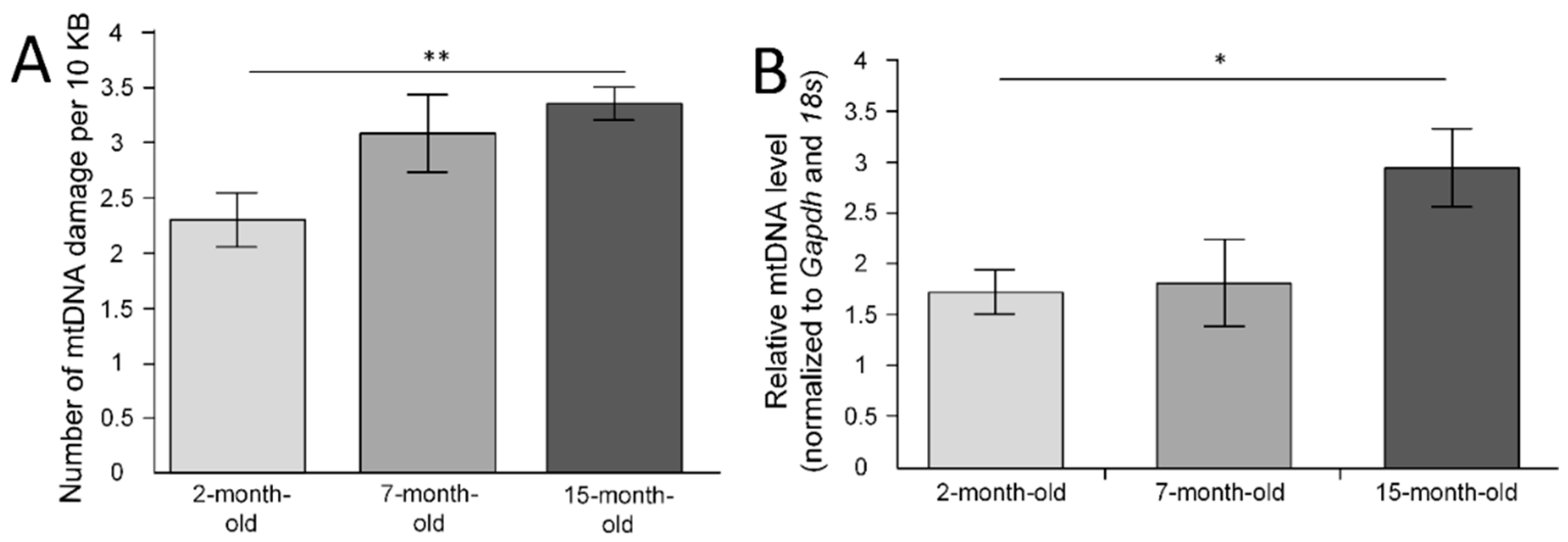
2.1. mtDNA Lesions in the Cerebral Cortex
2.2. mtDNA Copy Number in the Cerebral Cortex
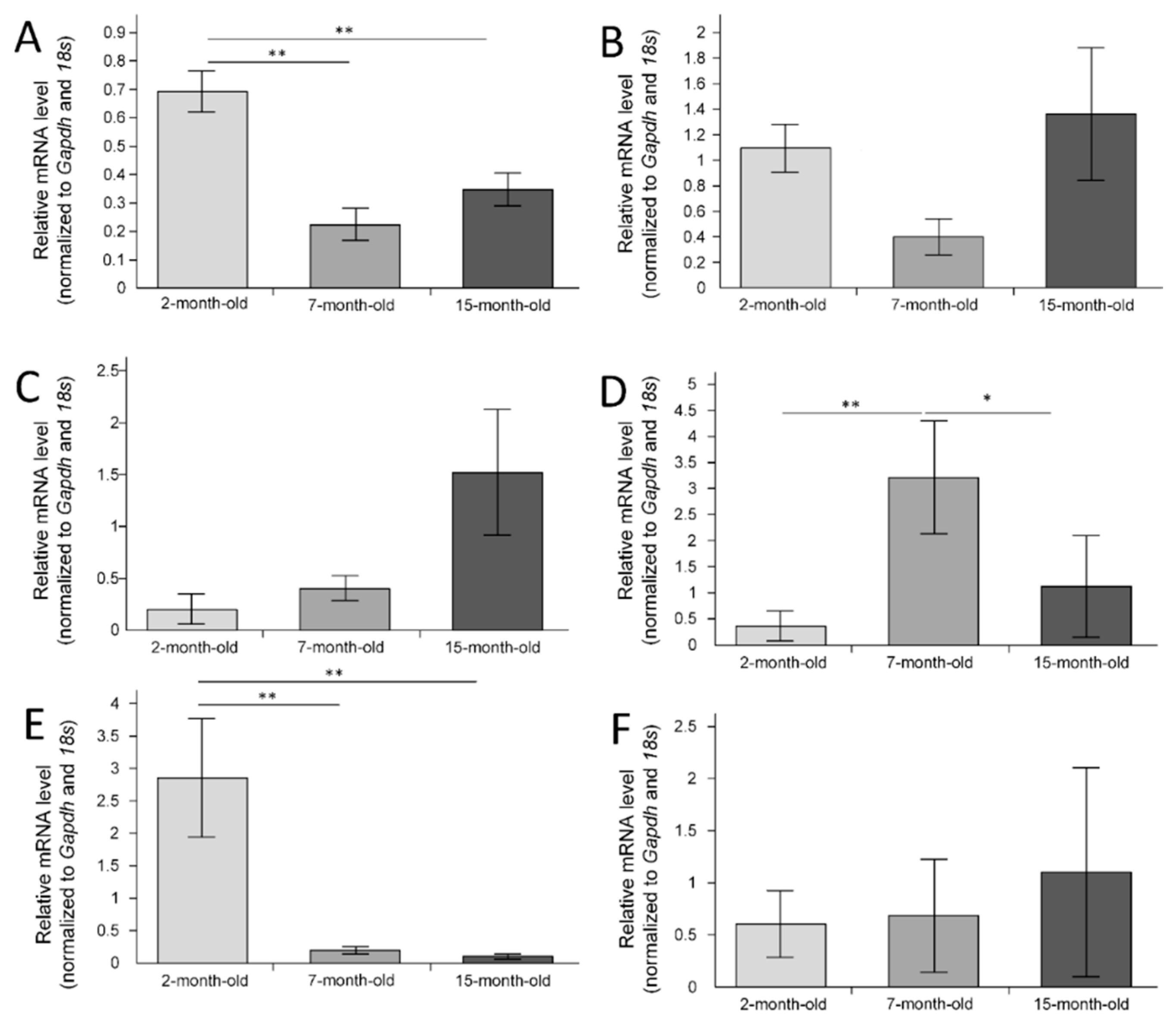
2.3. Gene Expression in the Cerebral Cortex
2.4. Gene Expression in the Hippocampus
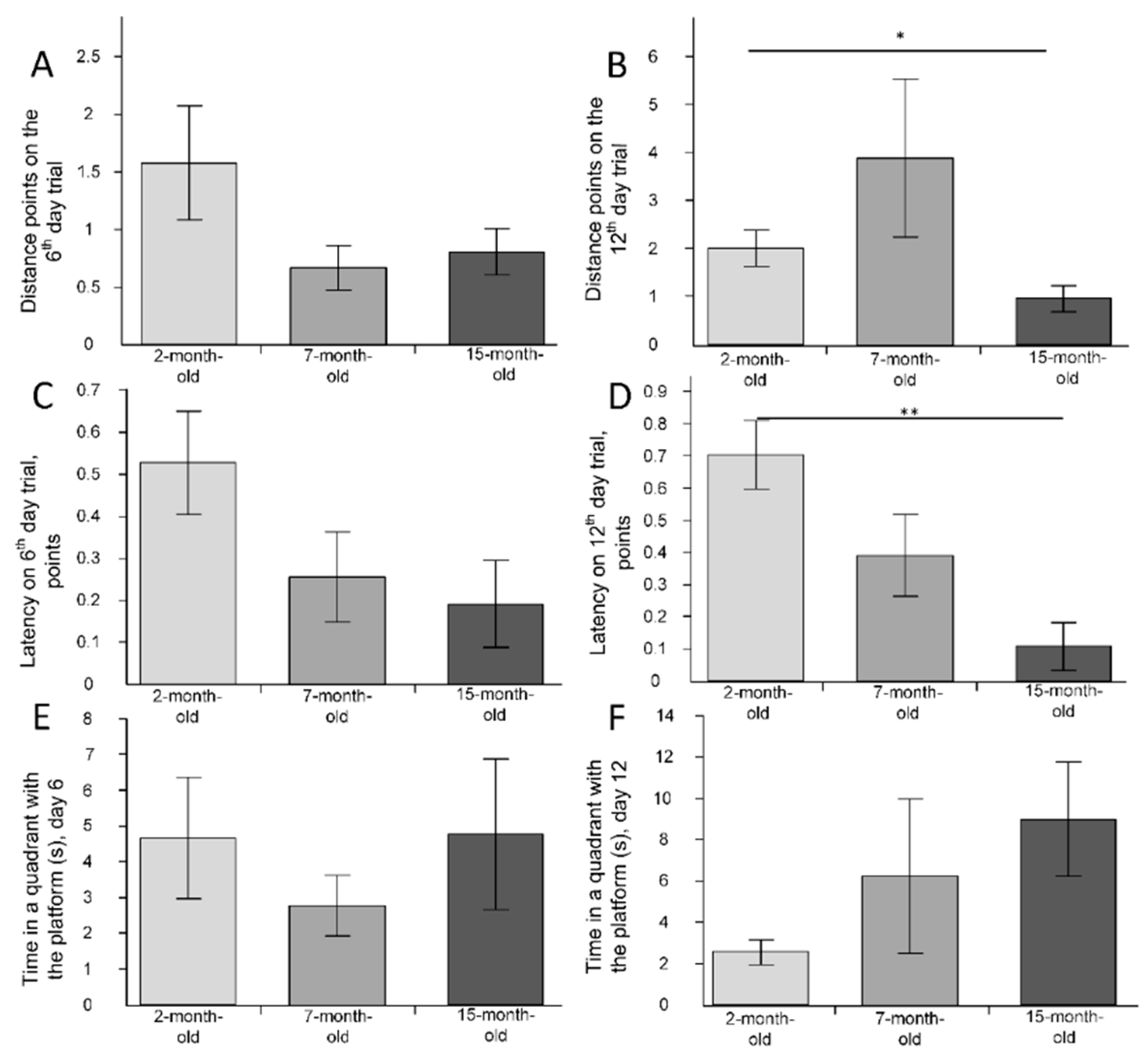
2.5. The Behavioral Changes
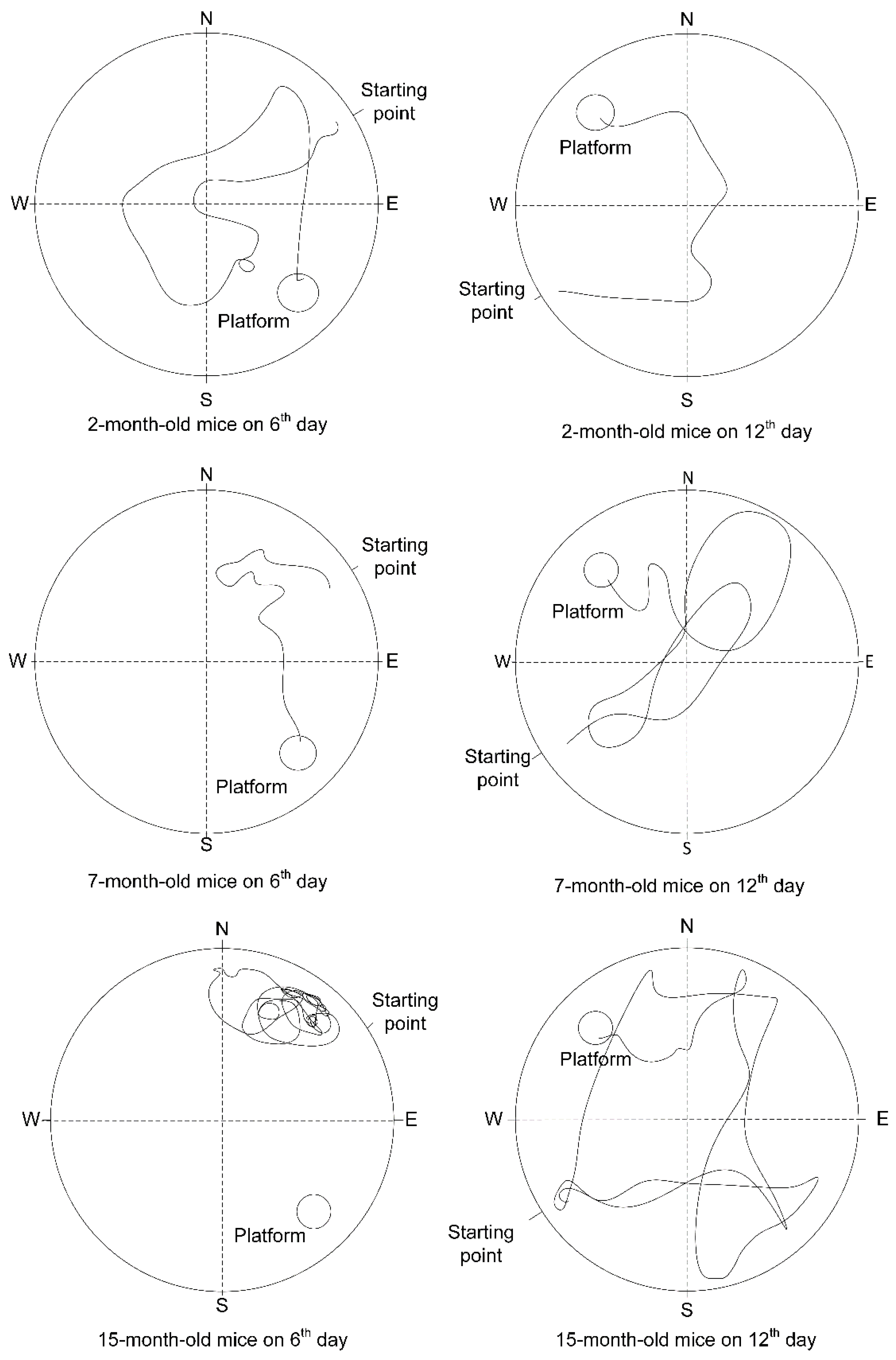
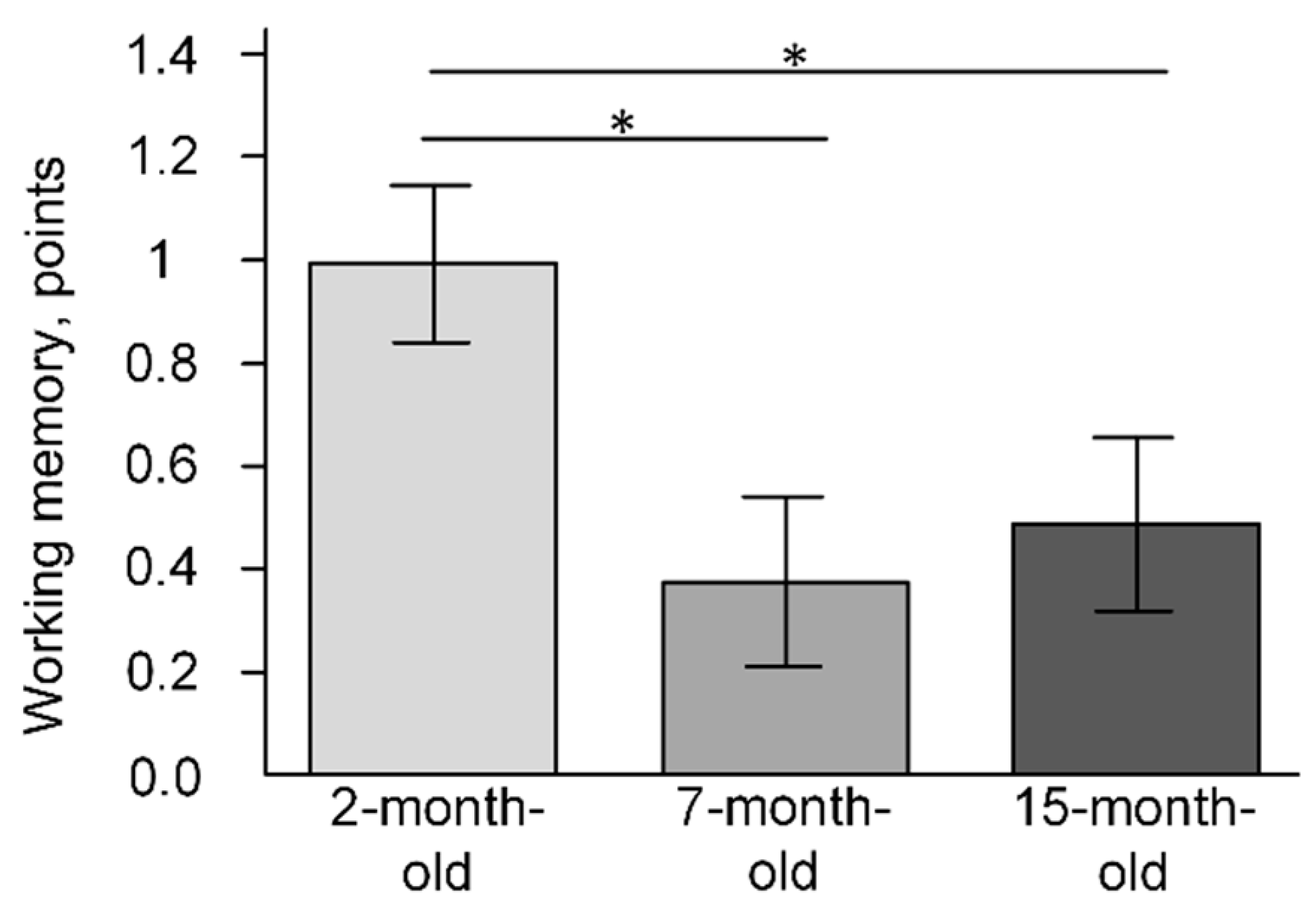
2.6. The Change of Cognitive Function
2.7. The Changes of Strength and Endurance
3. Discussion
4. Materials and Methods
4.1. Animals
4.2. The Open Field Test
4.3. The String Test
4.4. The Dark-Light Box Test
4.5. The Morris Water Maze Test
4.6. DNA and RNA Extraction
4.7. Reverse Transcription
4.8. Quantitative PCR Analysis
4.9. Measurement of mtDNA Damage
4.10. MtDNA Copy Measurement
4.11. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Patrushev, M.V.; Mazunin, I.O.; Vinogradova, E.N.; Kamenski, P.A. Mitochondrial Fission and Fusion. Biochemistry 2015, 80, 1457–1464. [Google Scholar] [CrossRef] [PubMed]
- Herculano-Houzel, S. Scaling of brain metabolism with a fixed energy budget per neuron: Implications for neuronal activity, plasticity and evolution. PLoS ONE 2011, 6, e17514. [Google Scholar] [CrossRef] [Green Version]
- Van Bulck, M.; Sierra-Magro, A.; Alarcon-Gil, J.; Perez-Castillo, A.; Morales-Garcia, J.A. Novel Approaches for the Treatment of Alzheimer’s and Parkinson’s Disease. Int. J. Mol. Sci. 2019, 20, 719. [Google Scholar] [CrossRef] [Green Version]
- Park, D.C.; Bischof, G.N. The aging mind: Neuroplasticity in response to cognitive training. Dialogues Clin. Neurosci. 2013, 15, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Gureev, A.P.; Shaforostova, E.A.; Popov, V.N. Regulation of Mitochondrial Biogenesis as a Way for Active Longevity: Interaction Between the Nrf2 and PGC-1a Signaling Pathways. Front. Genet. 2019, 10, 435. [Google Scholar] [CrossRef] [Green Version]
- de Grey, A.D. Reactive oxygen species production in the mitochondrial matrix: Implications for the mechanism of mitochondrial mutation accumulation. Rejuvenation Res. 2005, 8, 13–17. [Google Scholar] [CrossRef]
- Bogenhagen, D.F. Mitochondrial DNA nucleoid structure. Biochim. Biophys. Acta 2012, 1819, 914–920. [Google Scholar] [CrossRef]
- Zinovkina, L.A. Mechanisms of Mitochondrial DNA Repair in Mammals. Biochemistry 2018, 83, 233–249. [Google Scholar] [CrossRef]
- Muramatsu, H.; Katsuoka, F.; Toide, K.; Shimizu, Y.; Furusako, S.; Yamamoto, M. Nrf2 deficiency leads to behavioral, neurochemical and transcriptional changes in mice. Genes Cells 2013, 18, 899–908. [Google Scholar] [CrossRef] [PubMed]
- Schratt, G.M.; Nigh, E.A.; Chen, W.G.; Hu, L.; Greenberg, M.E. BDNF regulates the translation of a select group of mRNAs by a mammalian target of rapamycin- phosphatidylinositol 3-kinase-dependent pathway during neuronal development. J. Neurosci. 2004, 24, 7366–7377. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stoica, L.; Zhu, P.J.; Huang, W.; Zhou, H.; Kozma, S.C.; Costa-Mattioli, M. Selective pharmacogenetic inhibition of mammalian target of Rapamycin complex I (mTORC1) blocks long-term synaptic plasticity and memory storage. Proc. Natl. Acad. Sci. USA 2011, 108, 3791–3796. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gureev, A.P.; Popov, V.N.; Starkov, A.A. Crosstalk between the mTOR and Nrf2/ARE signaling pathways as a target in the improvement of long-term potentiation. Exp. Neurol. 2020, 328, 113285. [Google Scholar] [CrossRef] [PubMed]
- Dinkova-Kostova, A.T.; Abramov, A.Y. The emerging role of Nrf2 in mitochondrial function. Free. Radic. Biol. Med. 2015, 88, 179–188. [Google Scholar] [CrossRef] [Green Version]
- Schmidlin, C.J.; Dodson, M.B.; Madhavan, L.; Zhang, D.D. Redox regulation by NRF2 in aging and disease. Free. Radic. Biol. Med. 2019, 134, 702–707. [Google Scholar] [CrossRef]
- Kasai, S.; Shimizu, S.; Tatara, Y.; Mimura, J.; Itoh, K. Regulation of Nrf2 by Mitochondrial Reactive Oxygen Species in Physiology and Pathology. Biomolecules 2020, 10, 320. [Google Scholar] [CrossRef] [Green Version]
- Neuhaus, J.F.; Baris, O.R.; Kittelmann, A.; Becker, K.; Rothschild, M.A.; Wiesner, R.J. Catecholamine Metabolism Induces Mitochondrial DNA Deletions and Leads to Severe Adrenal Degeneration during Aging. Neuroendocrinology 2017, 104, 72–84. [Google Scholar] [CrossRef]
- Genova, M.L.; Bovina, C.; Marchetti, M.; Pallotti, F.; Tietz, C.; Biagini, G.; Pugnaloni, A.; Viticchi, C.; Gorini, A.; Villa, R.F.; et al. Decrease of rotenone inhibition is a sensitive parameter of complex I damage in brain non-synaptic mitochondria of aged rats. FEBS Lett. 1997, 410, 467–469. [Google Scholar] [CrossRef] [Green Version]
- Lenaz, G.; D’Aurelio, M.; Merlo Pich, M.; Genova, M.L.; Ventura, B.; Bovina, C.; Formiggini, G.; Parenti Castelli, G. Mitochondrial bioenergetics in aging. Biochim. Biophys. Acta 2000, 1459, 397–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tatarkova, Z.; Kovalska, M.; Timkova, V.; Racay, P.; Lehotsky, J.; Kaplan, P. The Effect of Aging on Mitochondrial Complex I and the Extent of Oxidative Stress in the Rat Brain Cortex. Neurochem. Res. 2016, 41, 2160–2172. [Google Scholar] [CrossRef]
- Crescenzo, R.; Spagnuolo, M.S.; Cancelliere, R.; Iannotta, L.; Mazzoli, A.; Gatto, C.; Iossa, S.; Cigliano, L. Effect of Initial Aging and High-Fat/High-Fructose Diet on Mitochondrial Bioenergetics and Oxidative Status in Rat Brain. Mol. Neurobiol. 2019, 56, 7651–7663. [Google Scholar] [CrossRef]
- Davies, S.M.; Poljak, A.; Duncan, M.W.; Smythe, G.A.; Murphy, M.P. Measurements of protein carbonyls, ortho- and meta-tyrosine and oxidative phosphorylation complex activity in mitochondria from young and old rats. Free. Radic. Biol. Med. 2001, 31, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Nicoletti, V.G.; Tendi, E.A.; Lalicata, C.; Reale, S.; Costa, A.; Villa, R.F.; Ragusa, N.; Giuffrida Stella, A.M. Changes of mitochondrial cytochrome c oxidase and FoF1 ATP synthase subunits in rat cerebral cortex during aging. Neurochem. Res. 1995, 20, 1465–1470. [Google Scholar] [CrossRef] [PubMed]
- Li, P.A.; Hou, X.; Hao, S. Mitochondrial biogenesis in neurodegeneration. J. Neurosci. Res. 2017, 95, 2025–2029. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Kroemer, G.; Kepp, O. Mitophagy: An Emerging Role in Aging and Age-Associated Diseases. Front. Cell Dev. Biol. 2020, 8, 200. [Google Scholar] [CrossRef] [Green Version]
- Gureev, A.P.; Sadovnikova, I.S.; Starkov, N.N.; Starkov, A.A.; Popov, V.N. p62-Nrf2-p62 Mitophagy Regulatory Loop as a Target for Preventive Therapy of Neurodegenerative Diseases. Brain Sci. 2020, 10, 847. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Dai, C.; Fan, Y.; Guo, B.; Ren, K.; Sun, T.; Wang, W. From autophagy to mitophagy: The roles of P62 in neurodegenerative diseases. J. Bioenerg. Biomembr. 2017, 49, 413–422. [Google Scholar] [CrossRef]
- Vorhees, C.V.; Williams, M.T. Morris water maze: Procedures for assessing spatial and related forms of learning and memory. Nat. Protoc. 2006, 1, 848–858. [Google Scholar] [CrossRef] [Green Version]
- Bellush, L.L.; Wright, A.M.; Walker, J.P.; Kopchick, J.; Colvin, R.A. Caloric restriction and spatial learning in old mice. Physiol. Behav. 1996, 60, 541–547. [Google Scholar] [CrossRef]
- Magnusson, K.R.; Scruggs, B.; Aniya, J.; Wright, K.C.; Ontl, T.; Xing, Y.D.; Bai, L. Age-related deficits in mice performing working memory tasks in a water maze. Behav. Neurosci. 2003, 117, 485–495. [Google Scholar] [CrossRef]
- van Praag, H.M. Can stress cause depression? World J. Biol. Psychiatr. 2005, 6, 5–22. [Google Scholar] [CrossRef]
- Harburger, L.L.; Lambert, T.J.; Frick, K.M. Age-dependent effects of environmental enrichment on spatial reference memory in male mice. Behav. Brain Res. 2007, 185, 43–48. [Google Scholar] [CrossRef] [Green Version]
- Wong, A.A.; Brown, R.E. Age-related changes in visual acuity, learning and memory in C57BL/6J and DBA/2J mice. Neurobiol. Aging 2007, 28, 1577–1593. [Google Scholar] [CrossRef]
- Lovell, M.A.; Markesbery, W.R. Oxidative DNA damage in mild cognitive impairment and late-stage Alzheimer’s disease. Nucleic Acids Res. 2007, 35, 7497–7504. [Google Scholar] [CrossRef] [Green Version]
- Bendavit, G.; Aboulkassim, T.; Hilmi, K.; Shah, S.; Batist, G. Nrf2 transcription factor can directly regulate mTOR: Linking cytoprotective gene expression to a major metabolic regulator that generates redox activity. J. Biol. Chem. 2016, 291, 25476–25488. [Google Scholar] [CrossRef] [Green Version]
- Hasegawa, I.; Hayashi, T.; Miyashita, Y. Memory retrieval under the control of the prefrontal cortex. Ann. Med. 1999, 31, 380–387. [Google Scholar] [CrossRef]
- Ferrucci, L.; Fabbri, E. Inflammageing: Chronic inflammation in ageing, cardiovascular disease, and frailty. Nat. Rev. Cardiol. 2018, 15, 505–522. [Google Scholar] [CrossRef]
- Dinarello, C.A. Overview of the IL-1 family in innate inflammation and acquired immunity. Immunol. Rev. 2018, 281, 8–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kobayashi, E.H.; Suzuki, T.; Funayama, R.; Nagashima, T.; Hayashi, M.; Sekine, H.; Tanaka, N.; Moriguchi, T.; Motohashi, H.; Nakayama, K.; et al. Nrf2 suppresses macrophage inflammatory response by blocking proinflammatory cytokine transcription. Nat. Commun. 2016, 23, 11624. [Google Scholar] [CrossRef] [Green Version]
- Khairova, R.A.; Machado-Vieira, R.; Du, J.; Manji, H.K. A potential role for pro- inflammatory cytokines in regulating synaptic plasticity in major depressive disorder. Int. J. Neuropsychopharmacol. 2009, 12, 561–578. [Google Scholar] [CrossRef] [PubMed]
- Brenes, J.C.; Padilla, M.; Fornaguera, J. A detailed analysis of open-field habituation and behavioral and neurochemical antidepressant-like effects in postweaning enriched rats. Behav. Brain Res. 2009, 197, 125–137. [Google Scholar] [CrossRef] [PubMed]
- Malatynska, E.; Steinbusch, H.W.; Redkozubova, O.; Bolkunov, A.; Kubatiev, A.; Yeritsyan, N.B.; Vignisse, J.; Bachurin, S.; Strekalova, T. Anhedonic-like traits and lack of affective deficits in 18-month-old C57BL/6 mice: Implications for modeling elderly depression. Exp. Gerontol. 2012, 47, 552–564. [Google Scholar] [CrossRef] [PubMed]
- Benice, T.S.; Rizk, A.; Kohama, S.; Pfankuch, T.; Raber, J. Sex-differences in age- related cognitive decline in C57BL/6J mice associated with increased brain microtubule- associated protein 2 and synaptophysin immunoreactivity. Neuroscience 2006, 137, 413–423. [Google Scholar] [CrossRef] [PubMed]
- Lalonde, R.; Strazielle, C. The relation between open-field and emergence tests in a hyperactive mouse model. Neuropharmacology 2009, 57, 722–724. [Google Scholar] [CrossRef]
- Shoji, H.; Takao, K.; Hattori, S.; Miyakawa, T. Age-related changes in behavior in C57BL/6J mice from young adulthood to middle age. Mol. Brain 2016, 28, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sturman, O.; Germain, P.L.; Bohacek, J. Exploratory rearing: A context- and stress- sensitive behavior recorded in the open-field test. Stress 2018, 21, 443–452. [Google Scholar] [CrossRef]
- Bryant, C.D. The blessings and curses of C57BL/6 substrains in mouse genetic studies. Ann. N. Y. Acad. Sci. 2011, 1245, 31–33. [Google Scholar] [CrossRef] [Green Version]
- Bryant, C.D.; Zhang, N.N.; Sokoloff, G.; Fanselow, M.S.; Ennes, H.S.; Palmer, A.A.; McRoberts, J.A. Behavioral differences among C57BL/6 substrains: Implications for transgenic and knockout studies. J. Neurogenet. 2008, 22, 315–331. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, N.; Takao, K.; Nakanishi, K.; Yamasaki, N.; Tanda, K.; Miyakawa, T. Behavioral profiles of three C57BL/6 substrains. Front. Behav. Neurosci. 2010, 14, 29. [Google Scholar] [CrossRef] [Green Version]
- An, X.L.; Zou, J.X.; Wu, R.Y.; Yang, Y.; Tai, F.D.; Zeng, S.Y.; Jia, R.; Zhang, X.; Liu, E.Q.; Broders, H. Strain and sex differences in anxiety-like and social behaviors in C57BL/6J and BALB/cJ mice. Exp. Anim. 2011, 60, 111–123. [Google Scholar] [CrossRef] [Green Version]
- Gureev, A.P.; Syromyatnikov, M.Y.; Gorbacheva, T.M.; Starkov, A.A.; Popov, V.N. Methylene blue improves sensorimotor phenotype and decreases anxiety in parallel with activating brain mitochondria biogenesis in mid-age mice. Neurosci. Res. 2016, 113, 19–27. [Google Scholar] [CrossRef]
- Paxinos, G.; Franklin, K. The Mouse Brain in Stereotaxic Coordinates; Academic: San Diego, CA, USA, 2001. [Google Scholar]
- Cardozo-Pelaez, F.; Song, S.; Parthasarathy, A.; Hazzi, C.; Naidu, K.; Sanchez-Ramos, J. Oxidative DNA damage in the aging mouse brain. Mov. Disord. 1999, 14, 972–980. [Google Scholar] [CrossRef] [PubMed]
- Gureev, A.P.; Shaforostova, E.A.; Vitkalova, I.Y.; Sadovnikova, I.S.; Kalinina, Y.I.; Cherednichenko, V.R.; Reznikova, K.A.; Valuyskikh, V.V.; Popov, V.N. Long-term mildronate treatment increased Proteobacteria level in gut microbiome, and caused behavioral deviations and transcriptome change in liver, heart and brain of healthy mice. Toxicol. Appl. Pharmacol. 2020, 398, 115031. [Google Scholar] [CrossRef] [PubMed]
- Antunes, M.; Biala, G. The novel object recognition memory: Neurobiology, test procedure, and its modifications. Cogn. Process. 2012, 13, 93–110. [Google Scholar] [CrossRef] [Green Version]
- Gureev, A.P.; Shaforostova, E.A.; Starkov, A.A.; Popov, V.N. Simplified qPCR method for detecting excessive mtDNA damage induced by exogenous factors. Toxicology 2017, 382, 67–74. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khorolskaya, V.G.; Gureev, A.P.; Shaforostova, E.A.; Laver, D.A.; Popov, V.N. The Fenofibrate Effect on Genotoxicity in Brain and Liver and on the Expression of Genes Regulating Fatty Acids Metabolism of Mice. Biochem. (Mosc.) Suppl. Ser. B Biomed. Chem. 2020, 14, 23–32. [Google Scholar] [CrossRef]





| Fragment | Number of mtDNA Lesions (per 10 kb) | ||
|---|---|---|---|
| 2-Month-Old | 7-Month-Old | 15-Month-Old | |
| 12–16s (1st) | 2.36 ± 0.39 | 4.01 ± 0.37 ** | 3.34 ± 0.30 |
| 16s-Nd1 (2nd) | 2.45 ± 0.41 | 3.59 ± 0.71 | 4.11 ± 0.29 * |
| Nd1-Nd2 (3th) | 1.03 ± 0.36 | 2.86 ± 0.62 * | 2.05 ± 0.15 |
| Nd5 (4th) | 1.97 ± 0.43 | 3.12 ± 0.51 | 2.65 ± 0.11 |
| Nd6-CytB (5th) | 2.16 ± 0.34 | 1.89 ± 0.59 | 2.73 ± 0.19 |
| D-loop (6th) | 3.89 ± 0.54 | 3.02 ± 0.67 | 5.44 ± 0.07 ***†† |
| Acquisition. Start Located in the SW | ||||
|---|---|---|---|---|
| Day | Trial 1 | Trial 2 | Trial 3 | Trial 4 |
| 1 | N | E | SE | NW |
| 2 | SE | N | NW | E |
| 3 | NW | SE | E | N |
| 4 | E | NW | N | SE |
| 5 | N | SE | E | NW |
| 6 (Probe) | NE | - | - | - |
| Reversal. Start Located in the NE | ||||
| Day | Trial 1 | Trial 2 | Trial 3 | Trial 4 |
| 1 | S | W | NW | SE |
| 2 | NW | S | SE | W |
| 3 | SE | NW | W | S |
| 4 | W | SE | S | NW |
| 5 | S | NW | W | SE |
| 6 (Probe) | SW | - | - | - |
| Day | Start | Goal | Day | Start | Goal |
|---|---|---|---|---|---|
| 1 | N | SE | 11 | E | SW |
| 2 | E | NE | 12 | N | SW |
| 3 | S | SW | 13 | E | NW |
| 4 | W | SE | 14 | W | NE |
| 5 | S | NE | 15 | N | SE |
| 6 | N | NW | 16 | S | SW |
| 7 | W | NE | 17 | N | NE |
| 8 | E | SE | 18 | S | NW |
| 9 | W | NW | 19 | E | NW |
| 10 | S | SE | 20 | W | SW |
| 21 | N | SE |
| Gene Name | Transcript Accession Number | Forward Primer 5′–3′ | Reverse Primer 5′–3′ |
|---|---|---|---|
| 18s | NR_003278.3 | CGGCTACCACATCCAAGGAA | GCTGGAATTACTGTGGCT |
| Gapdh | NC_000072.7 | GGCTCCCTAGGCCCCTCCTG | TCCCAACTCGGCCCCCAACA |
| Nfe2l2 | NM_010902.4 | CTCTCTGAACTCCTGGACGG | GGGTCTCCGTAAATGGAAG |
| Bdnf | NM_007540.4 | AAGGACGCGGACTTGTACAC | CGCTAATACTGTCACACACGC |
| Mtor | NM_020009.2 | AGATAAGCTCACTGGTCGGG | GTGGTTTTCCAGGCCTCAGT |
| Akt1 | NM_009652.3 | TGATCAAGATGACAGCATGGAGTG | GATGATCCATGCGGGGCTT |
| Pten | NM_008960.2 | AGGGACGAACTGGTGTAATGA | GGGAATAGTTACTCCCTTTTTGTCT |
| Il1b | NM_008361.4 | TTGACGGACCCCAAAAGATG | AGAAGGTGCTCATGTCCTCA |
| Nrf1 | NM_001164226.1 | AGCACGGAGTGACCCAAA | TGTACGTGGCTACATGGACCT |
| Cox1 | NC_005089.1 | TCGCAATTCCTACCGGTCTC | CGTGTAGGGTTGCAAGTCAGC |
| p62 | U17961.1 | GCCAGAGGAACAGATGGAGT | TCCGATTCTGGCATCTGTAG |
| Sod2 | NM_013671.3 | CAGACCTGCCTTACGACTATGG | CTCGGTGGCGTTGAGATTGTT |
| Tfam | NC_000076.7 | ATTCCGAAGTGTTTTTCCAGCA | TCTGAAAGTTTTCGATCTGGGT |
| Fragment Name | Forward Primer 5′–3′ | Reverse Primer 5′–3′ | Length Fragment |
|---|---|---|---|
| Short | ACGAGGGTCCAACTGTCTCTTA | AGCTCCATAGGGTCTTCTCGT | 97 bp |
| 12–16s (1st) | TAAATTTCGTGCCAGCCACC | ATGCTACCTTTGCACGGTCA | 1739 bp |
| 16s-Nd1 (2nd) | ACGAGGGTCCAACTGTCTCTTA | CCGGCTGCGTATTCTACGTT | 1326 bp |
| Nd1-Nd2 (3th) | CTAGCAGAAACAAACCGGGC | TTAGGGCTTTGAAGGCTCGC | 1675 bp |
| Nd5 (4th) | TCATTCTTCTACTATCCCCAATCC | TGGTTTGGGAGATTGGTTGATG | 1942 bp |
| Nd6-CytB (5th) | CCCCAATCCCTCCTTCCAAC | GGTGGGGAGTAGCTCCTTCTT | 1732 bp |
| D-loop (6th) | AAGAAGGAGCTACTCCCCACC | GTTGACACGTTTTACGCCGA | 1308 bp |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gureev, A.P.; Khorolskaya, V.G.; Sadovnikova, I.S.; Shaforostova, E.A.; Cherednichenko, V.R.; Burakova, I.Y.; Plotnikov, E.Y.; Popov, V.N. Age-Related Decline in Nrf2/ARE Signaling Is Associated with the Mitochondrial DNA Damage and Cognitive Impairments. Int. J. Mol. Sci. 2022, 23, 15197. https://doi.org/10.3390/ijms232315197
Gureev AP, Khorolskaya VG, Sadovnikova IS, Shaforostova EA, Cherednichenko VR, Burakova IY, Plotnikov EY, Popov VN. Age-Related Decline in Nrf2/ARE Signaling Is Associated with the Mitochondrial DNA Damage and Cognitive Impairments. International Journal of Molecular Sciences. 2022; 23(23):15197. https://doi.org/10.3390/ijms232315197
Chicago/Turabian StyleGureev, Artem P., Victoria G. Khorolskaya, Irina S. Sadovnikova, Ekaterina A. Shaforostova, Vadim R. Cherednichenko, Inna Y. Burakova, Egor Y. Plotnikov, and Vasily N. Popov. 2022. "Age-Related Decline in Nrf2/ARE Signaling Is Associated with the Mitochondrial DNA Damage and Cognitive Impairments" International Journal of Molecular Sciences 23, no. 23: 15197. https://doi.org/10.3390/ijms232315197
APA StyleGureev, A. P., Khorolskaya, V. G., Sadovnikova, I. S., Shaforostova, E. A., Cherednichenko, V. R., Burakova, I. Y., Plotnikov, E. Y., & Popov, V. N. (2022). Age-Related Decline in Nrf2/ARE Signaling Is Associated with the Mitochondrial DNA Damage and Cognitive Impairments. International Journal of Molecular Sciences, 23(23), 15197. https://doi.org/10.3390/ijms232315197








