Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes
Abstract
:1. Introduction
2. Results
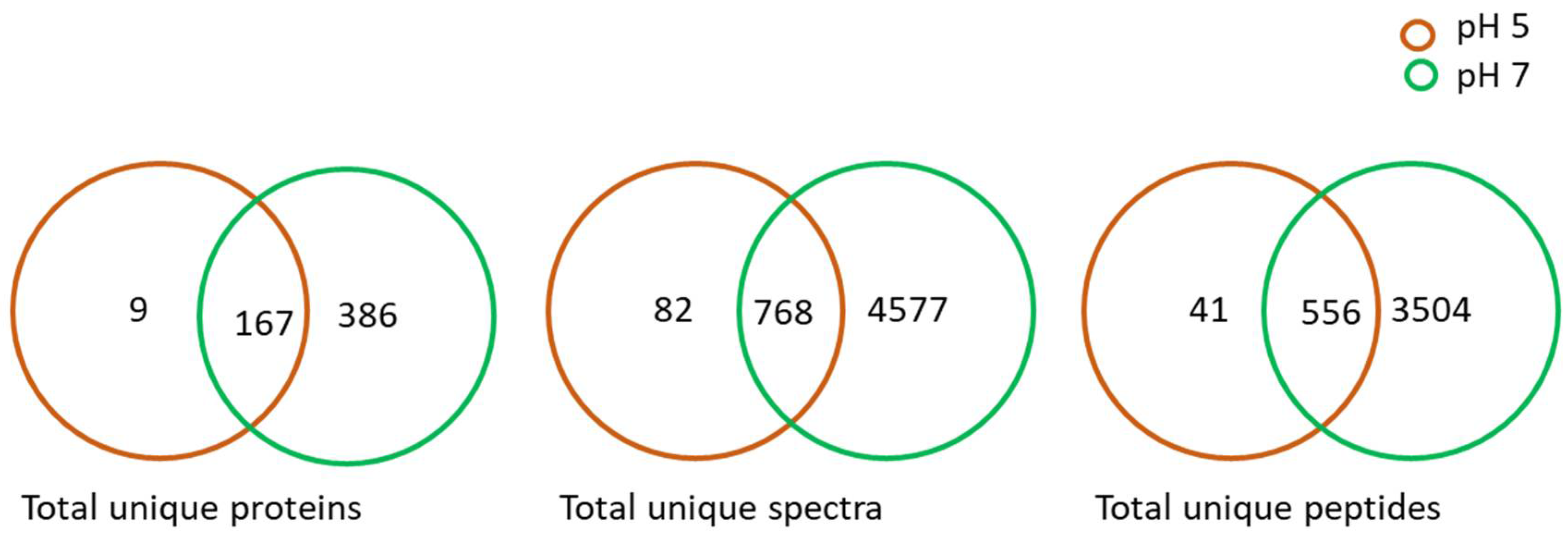
2.1. Proteome Analysis of L. helveticus (EL2006H) CFS
2.2. Groups of Identified Proteins from L. helveticus (EL2006H) CFS and Their Functional Annotation
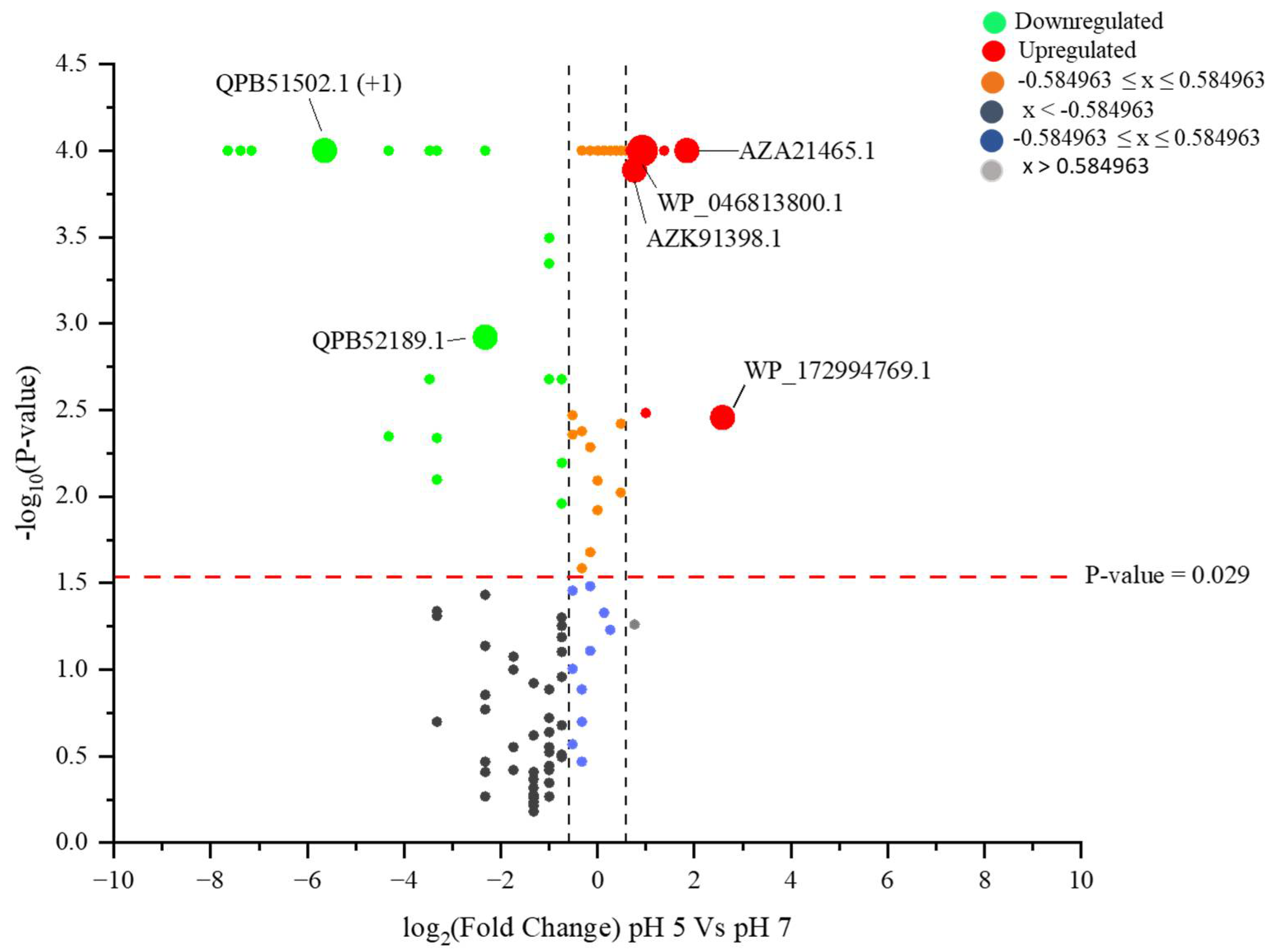
2.3. Identified Proteins from L. helveticus (EL2006H) CFS
2.4. Proteome Analysis of B. subtilis (EB2004S) CFS
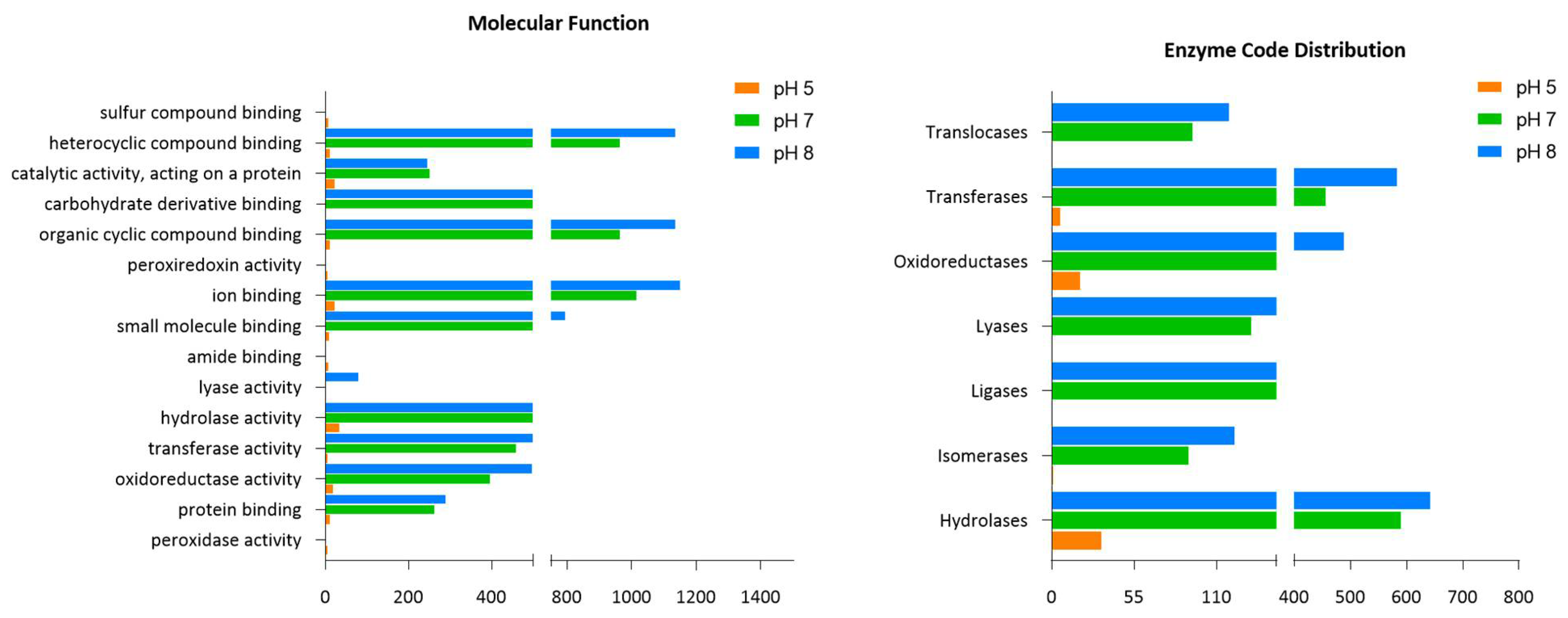
2.5. Groups of Identified Proteins from B. subtilis (EB2004S) CFS and Their Functional Annotation
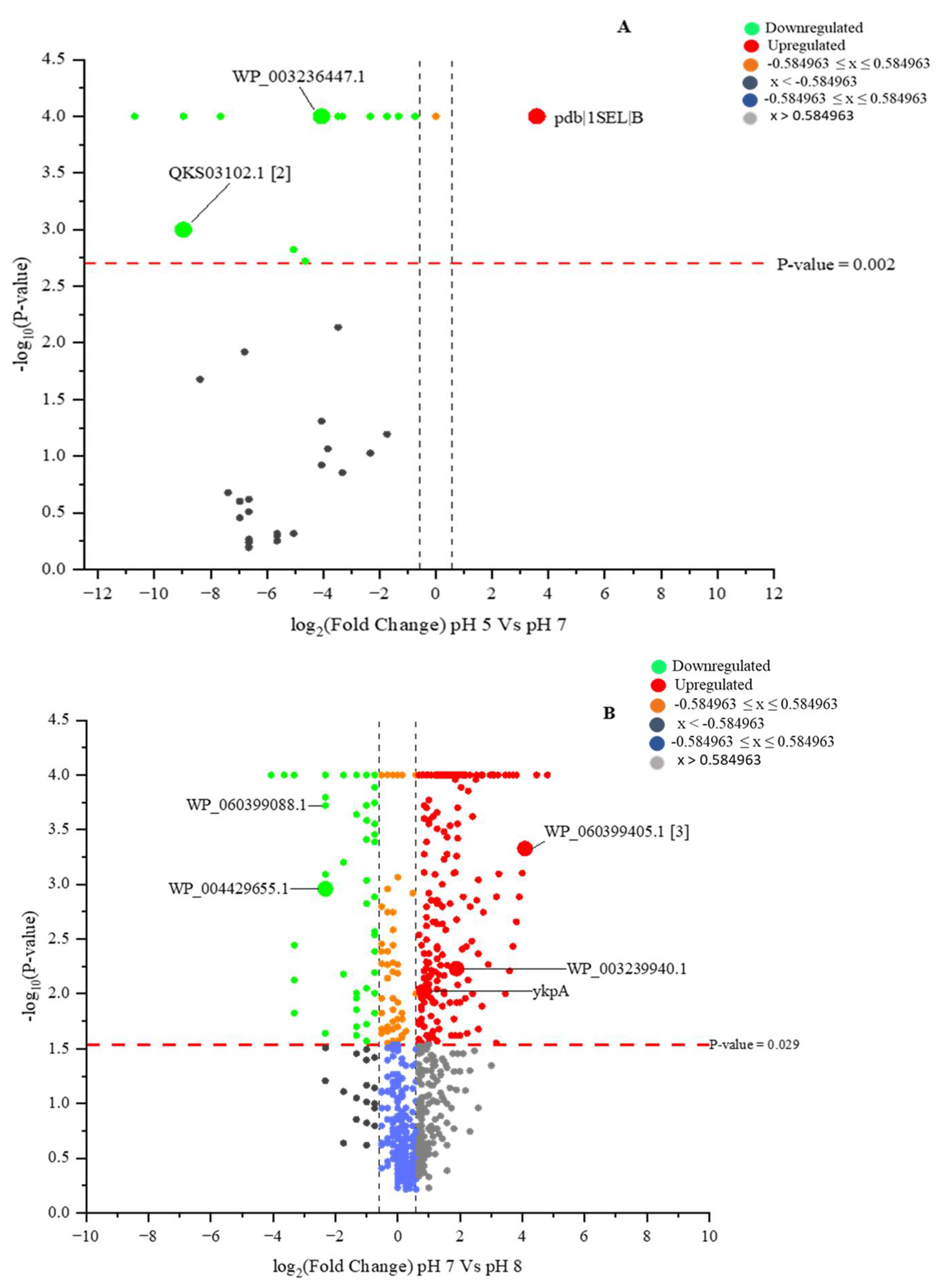
2.6. Identified Proteins with Differential Identities from B. subtilis (EB2004S) CFS
3. Discussion
4. Materials and Methods
4.1. Microbial Strain Growth Conditions and Media
Proteomics Study from the CFSs
4.2. Protein Profiling
4.3. Protein Identification Confidence Level
4.4. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Abdulhussain Kareem, R.; Razavi, S.H. Plantaricin bacteriocins: As safe alternative antimicrobial peptides in food preservation—A review. J. Food Saf. 2020, 40, e12735. [Google Scholar] [CrossRef]
- Ahrne, S.; Nobaek, S.; Jeppsson, B.; Adlerberth, I.; Wold, A.; Molin, G. The normal Lactobacillus flora of healthy human rectal and oral mucosa. J. Appl. Microbiol. 1998, 85, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Alnakli, A.A.A.; Jabeen, A.; Chakraborty, R.; Mohamedali, A.; Ranganathan, S. A Bioinformatics Approach to Mine the Microbial Proteomic Profile of COVID-19 Mass Spectrometry Data. Appl. Microbiol. 2022, 2, 150–164. [Google Scholar] [CrossRef]
- Altermann, E.; Buck, L.B.; Cano, R.; Klaenhammer, T.R. Identification and phenotypic characterization of the cell-division protein CdpA. Gene 2004, 342, 189–197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alyousef, A.; Alqasim, A.; Aloahd, M. Isolation and characterization of lectin with antibacterial, antibiofilm and antiproliferative activities from Acinetobacter baumannii of environmental origin. J. Appl. Microbiol. 2018, 124, 1139–1146. [Google Scholar] [CrossRef]
- Angeles, D.M.; Scheffers, D.-J. The cell wall of Bacillus subtilis. Curr. Issues Mol. Biol. 2021, 41, 539–596. [Google Scholar] [CrossRef] [PubMed]
- Antelmann, H.; Tjalsma, H.; Voigt, B.; Ohlmeier, S.; Bron, S.; Van Dijl, J.M.; Hecker, M. A proteomic view on genome-based signal peptide predictions. Genome Res. 2001, 11, 1484–1502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Antikainen, J.; Anton, L.; Sillanpää, J.; Korhonen, T.K. Domains in the S-layer protein CbsA of Lactobacillus crispatus involved in adherence to collagens, laminin and lipoteichoic acids and in self-assembly. Mol. Microbiol. 2002, 46, 381–394. [Google Scholar] [CrossRef]
- Antikainen, J.; Kupannen, V.; Läteenmäki, K.; Korhonen, T.K. pH-dependent association of enolase and glyceraldehyde-3-phosphate dehydrogenase of Lactobacillus crispatus with the cell wall and lipoteichoic acids. J. Bacteriol. 2007, 189, 4539–4543. [Google Scholar] [CrossRef] [Green Version]
- Backer, R.; Rokem, J.S.; Ilangumaran, G.; Lamont, J.; Praslickova, D.; Ricci, E.; Subramanian, S.; Smith, D.L. Plant growth-promoting rhizobacteria: Context, mechanisms of action, and roadmap to commercialization of biostimulants for sustainable agriculture. Front. Plant Sci. 2018, 1473. [Google Scholar] [CrossRef]
- Bagnoli, F.; Rappuoli, R. Protein and Sugar Export and Assembly in Gram-Positive Bacteria; Springer: Basel, Switzerland, 2017. [Google Scholar]
- Bandow, J.E.; Brötz, H.; Hecker, M. Bacillus subtilis tolerance of moderate concentrations of rifampin involves the σB-dependent general and multiple stress response. J. Bacteriol. 2002, 184, 459–467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beganović, J.; Frece, J.; Kos, B.; Leboš Pavunc, A.; Habjanič, K.; Šušković, J. Functionality of the S-layer protein from the probiotic strain Lactobacillus helveticus M92. Antonie Van Leeuwenhoek 2011, 100, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Beis, K.; Rebuffat, S. Multifaceted ABC transporters associated to microcin and bacteriocin export. Res. Microbiol. 2019, 170, 399–406. [Google Scholar] [CrossRef] [PubMed]
- Boël, G.; Pichereau, V.; Mijakovic, I.; Mazé, A.; Poncet, S.; Gillet, S.; Giard, J.-C.; Hartke, A.; Auffray, Y.; Deutscher, J. Is 2-phosphoglycerate-dependent automodification of bacterial enolases implicated in their export? J. Mol. Biol. 2004, 337, 485–496. [Google Scholar] [CrossRef] [PubMed]
- Borriss, R.; Danchin, A.; Harwood, C.R.; Médigue, C.; Rocha, E.P.; Sekowska, A.; Vallenet, D. Bacillus subtilis, the model Gram-positive bacterium: 20 years of annotation refinement. Microb. Biotechnol. 2018, 11, 3–17. [Google Scholar] [CrossRef]
- Brown, L.; Wolf, J.M.; Prados-Rosales, R.; Casadevall, A. Through the wall: Extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat. Rev. Microbiol. 2015, 13, 620–630. [Google Scholar] [CrossRef] [Green Version]
- Buck, B.L.; Altermann, E.; Svingerud, T.; Klaenhammer, T.R. Functional analysis of putative adhesion factors in Lactobacillus acidophilus NCFM. Appl. Environ. Microbiol. 2005, 71, 8344–8351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cai, X.; Zheng, W.; Li, Z. High-throughput screening strategies for the development of anti-virulence inhibitors against Staphylococcus aureus. Curr. Med. Chem. 2019, 26, 2297–2312. [Google Scholar] [CrossRef]
- Call, E.K.; Goh, Y.J.; Selle, K.; Klaenhammer, T.R.; O’flaherty, S. Sortase-deficient lactobacilli: Effect on immunomodulation and gut retention. Microbiology 2015, 161, 311. [Google Scholar] [CrossRef] [Green Version]
- Cassio Barreto De Oliveira, M.; Balan, A. The ATP-binding cassette (ABC) transport systems in Mycobacterium tuberculosis: Structure, function, and possible targets for therapeutics. Biology 2020, 9, 443. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhao, L.; Fu, G.; Zhou, W.; Sun, Y.; Zheng, P.; Sun, J.; Zhang, D. A novel strategy for protein production using non-classical secretion pathway in Bacillus subtilis. Microb. Cell Factories 2016, 15, 69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, J.; Guffanti, A.A.; Krulwich, T.A. The chromosomal tetracycline resistance locus of Bacillus subtilis encodes a Na+/H+ antiporter that is physiologically important at elevated pH. J. Biol. Chem. 1994, 269, 27365–27371. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Guffanti, A.A.; Wang, W.; Krulwich, T.A.; Bechhofer, D.H. Chromosomal tetA (L) gene of Bacillus subtilis: Regulation of expression and physiology of a tetA (L) deletion strain. J. Bacteriol. 1996, 178, 2853–2860. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, J.; Hicks, D.B.; Krulwich, T.A. The purified Bacillus subtilis tetracycline efflux protein TetA (L) reconstitutes both tetracycline–cobalt/H+ and Na+ (K+)/H+ exchange. Proc. Natl. Acad. Sci. USA 1996, 93, 14446–14451. [Google Scholar] [CrossRef] [Green Version]
- Chignell, J.F.; Park, S.; Lacerda, C.; De Long, S.K.; Reardon, K.F. Label-free proteomics of a defined, binary co-culture reveals diversity of competitive responses between members of a model soil microbial system. Microb. Ecol. 2018, 75, 701–719. [Google Scholar] [CrossRef]
- Chiu, C.H.; Paszkowski, U. Receptor-like kinases sustain symbiotic scrutiny. Plant Physiol. 2020, 182, 1597–1612. [Google Scholar] [CrossRef] [Green Version]
- Da Costa, W.L.O.; Araújo, C.L.D.A.; Dias, L.M.; Pereira, L.C.D.S.; Alves, J.T.C.; Araujo, F.A.; Folador, E.L.; Henriques, I.; Silva, A.; Folador, A.R.C. Functional annotation of hypothetical proteins from the Exiguobacterium antarcticum strain B7 reveals proteins involved in adaptation to extreme environments, including high arsenic resistance. PLoS ONE 2018, 13, e0198965. [Google Scholar] [CrossRef] [Green Version]
- De Souza Vandenberghe, L.P.; Karp, S.G.; Pagnoncelli, M.G.B.; Von Linsingen Tavares, M.; Junior, N.L.; Diestra, K.V.; Viesser, J.A.; Soccol, C.R. Classification of Enzymes and Catalytic Properties. In Biomass, Biofuels, Biochemicals; Elsevier: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Derman, Y.; Söderholm, H.; Lindström, M.; Korkeala, H. Role of csp genes in NaCl, pH, and ethanol stress response and motility in Clostridium botulinum ATCC 3502. Food Microbiol. 2015, 46, 463–470. [Google Scholar] [CrossRef]
- Dirix, G.; Monsieurs, P.; Dombrecht, B.; Daniels, R.; Marchal, K.; Vanderleyden, J.; Michiels, J. Peptide signal molecules and bacteriocins in Gram-negative bacteria: A genome-wide in silico screening for peptides containing a double-glycine leader sequence and their cognate transporters. Peptides 2004, 25, 1425–1440. [Google Scholar] [CrossRef]
- Errington, J.; Van Der Aart, L.T. Microbe Profile: Bacillus subtilis: Model organism for cellular development, and industrial workhorse. Microbiology 2020, 166, 425. [Google Scholar] [CrossRef]
- Etzold, S.; Kober, O.I.; Mackenzie, D.A.; Tailford, L.E.; Gunning, A.P.; Walshaw, J.; Hemmings, A.M.; Juge, N. Structural basis for adaptation of lactobacilli to gastrointestinal mucus. Environ. Microbiol. 2014, 16, 888–903. [Google Scholar] [CrossRef] [PubMed]
- Ferro-Luzzi Ames, G.; Mimura, C.S.; Shyamala, V. Bacterial periplasmic permeases belong to a family of transport proteins operating from Escherichia coli to human: Traffic ATPases. FEMS Microbiol. Rev. 1990, 6, 429–446. [Google Scholar] [CrossRef] [PubMed]
- Gagné-Bourque, F.; Bertrand, A.; Claessens, A.; Aliferis, K.A.; Jabaji, S. Alleviation of drought stress and metabolic changes in timothy (Phleum pratense L.) colonized with Bacillus subtilis B26. Front. Plant Sci. 2016, 7, 584. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gajbhiye, M.H.; Kapadnis, B.P. Antifungal-activity-producing lactic acid bacteria as biocontrol agents in plants. Biocontrol Sci. Technol. 2016, 26, 1451–1470. [Google Scholar] [CrossRef]
- Gray, E.; Lee, K.; Souleimanov, A.; Di Falco, M.; Zhou, X.; Ly, A.; Charles, T.; Driscoll, B.; Smith, D. A novel bacteriocin, thuricin 17, produced by plant growth promoting rhizobacteria strain Bacillus thuringiensis NEB17: Isolation and classification. J. Appl. Microbiol. 2006, 100, 545–554. [Google Scholar] [CrossRef]
- Gurumallesh, P.; Alagu, K.; Ramakrishnan, B.; Muthusamy, S. A systematic reconsideration on proteases. Int. J. Biol. Macromol. 2019, 128, 254–267. [Google Scholar] [CrossRef] [PubMed]
- Hardt, P.; Engels, I.; Rausch, M.; Gajdiss, M.; Ulm, H.; Sass, P.; Ohlsen, K.; Sahl, H.-G.; Bierbaum, G.; Schneider, T. The cell wall precursor lipid II acts as a molecular signal for the Ser/Thr kinase PknB of Staphylococcus aureus. Int. J. Med. Microbiol. 2017, 307, 1–10. [Google Scholar] [CrossRef]
- Harwood, C.R.; Kikuchi, Y. The ins and outs of Bacillus proteases: Activities, functions and commercial significance. FEMS Microbiol. Rev. 2022, 46, fuab046. [Google Scholar] [CrossRef] [PubMed]
- Hassani, M.; Durán, P.; Hacquard, S. Microbial interactions within the plant holobiont. Microbiome 2018, 6, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Hodge, K.; Ten Have, S.; Hutton, L.; Lamond, A.I. Cleaning up the masses: Exclusion lists to reduce contamination with HPLC-MS/MS. J. Proteom. 2013, 88, 92–103. [Google Scholar] [CrossRef]
- Hooper, L.V.; Gordon, J.I. Glycans as legislators of host–microbial interactions: Spanning the spectrum from symbiosis to pathogenicity. Glycobiology 2001, 11, 1R–10R. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hynönen, U.; Kant, R.; Lähteinen, T.; Pietilä, T.E.; Beganović, J.; Smidt, H.; Uroić, K.; Åvall-Jääskeläinen, S.; Palva, A. Functional characterization of probiotic surface layer protein-carrying Lactobacillus amylovorus strains. BMC Microbiol. 2014, 14, 199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnson, B.R.; Hymes, J.; Sanozky-Dawes, R.; Henriksen, E.D.; Barrangou, R.; Klaenhammer, T.R. Conserved S-layer-associated proteins revealed by exoproteomic survey of S-layer-forming lactobacilli. Appl. Environ. Microbiol. 2016, 82, 134–145. [Google Scholar] [CrossRef] [Green Version]
- Johnson, B.R.; Klaenhammer, T.R. AcmB is an S-layer-associated β-N-acetylglucosaminidase and functional autolysin in Lactobacillus acidophilus NCFM. Appl. Environ. Microbiol. 2016, 82, 5687–5697. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keller, A.; Nesvizhskii, A.I.; Kolker, E.; Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 2002, 74, 5383–5392. [Google Scholar] [CrossRef]
- Keto-Timonen, R.; Hietala, N.; Palonen, E.; Hakakorpi, A.; Lindström, M.; Korkeala, H. Cold shock proteins: A minireview with special emphasis on Csp-family of enteropathogenic Yersinia. Front. Microbiol. 2016, 7, 1151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.; Wu, K.; Lee, C. Stress-responsive periplasmic chaperones in bacteria. Front. Mol. Biosci. 2021, 357. [Google Scholar] [CrossRef]
- Kim, M.K.; Kang, M.-S.; Srinivasan, S.; Lee, D.H.; Lee, S.-Y.; Jung, H.-Y. Complete genome sequence of Hymenobacter sedentarius DG5BT, a bacterium resistant to gamma radiation. Mol. Cell. Toxicol. 2017, 13, 199–205. [Google Scholar] [CrossRef]
- Klein, G.; Pack, A.; Bonaparte, C.; Reuter, G. Taxonomy and physiology of probiotic lactic acid bacteria. Int. J. Food Microbiol. 1998, 41, 103–125. [Google Scholar] [CrossRef]
- Klotz, C.; Goh, Y.J.; O’flaherty, S.; Barrangou, R. S-layer associated proteins contribute to the adhesive and immunomodulatory properties of Lactobacillus acidophilus NCFM. BMC Microbiol. 2020, 20, 248. [Google Scholar] [CrossRef]
- Konstantinov, S.R.; Smidt, H.; De Vos, W.M.; Bruijns, S.C.; Singh, S.K.; Valence, F.; Molle, D.; Lortal, S.; Altermann, E.; Klaenhammer, T.R. S layer protein A of Lactobacillus acidophilus NCFM regulates immature dendritic cell and T cell functions. Proc. Natl. Acad. Sci. USA 2008, 105, 19474–19479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koo, B.-M.; Kritikos, G.; Farelli, J.D.; Todor, H.; Tong, K.; Kimsey, H.; Wapinski, I.; Galardini, M.; Cabal, A.; Peters, J.M. Construction and analysis of two genome-scale deletion libraries for Bacillus subtilis. Cell Syst. 2017, 4, 291–305.e7. [Google Scholar] [CrossRef] [PubMed]
- Kurian, D.; Phadwal, K.; Mäenpää, P. Proteomic characterization of acid stress response in Synechocystis sp. PCC 6803. Proteomics 2006, 6, 3614–3624. [Google Scholar] [CrossRef] [PubMed]
- Lamont, J.R.; Wilkins, O.; Bywater-Ekegärd, M.; Smith, D.L. From yogurt to yield: Potential applications of lactic acid bacteria in plant production. Soil Biol. Biochem. 2017, 111, 1–9. [Google Scholar] [CrossRef]
- Lee, K.; Rho, B.-S.; Pi, K.; Kim, H.-J.; Choi, Y.-J. Proteomic analysis of protein expression in Lactobacillus plantarum in response to alkaline stress. J. Biotechnol. 2011, 153, 1–7. [Google Scholar] [CrossRef]
- Li, J.; Liu, X.; Ji, Y.; Qi, Z.; Ge, Y.; Xu, J.; Liu, J.; Luo, G.; Shen, J. Biosynthesis of selenosubtilisin: A novel way to target selenium into the active site of subtilisin. Chin. Sci. Bull. 2008, 53, 2454–2461. [Google Scholar] [CrossRef] [Green Version]
- Ling, H.L.; Rahmat, Z.; Bakar, F.D.A.; Murad, A.M.A.; Illias, R.M. Secretome analysis of alkaliphilic bacterium Bacillus lehensis G1 in response to pH changes. Microbiol. Res. 2018, 215, 46–54. [Google Scholar] [CrossRef]
- Liu, L.; Guan, N.; Li, J.; Shin, H.-D.; Du, G.; Chen, J. Development of GRAS strains for nutraceutical production using systems and synthetic biology approaches: Advances and prospects. Crit. Rev. Biotechnol. 2017, 37, 139–150. [Google Scholar] [CrossRef]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [Google Scholar] [CrossRef]
- Lund, P.; Tramonti, A.; De Biase, D. Coping with low pH: Molecular strategies in neutralophilic bacteria. FEMS Microbiol. Rev. 2014, 38, 1091–1125. [Google Scholar] [CrossRef] [PubMed]
- Lund, P.A.; De Biase, D.; Liran, O.; Scheler, O.; Mira, N.P.; Cetecioglu, Z.; Fernández, E.N.; Bover-Cid, S.; Hall, R.; Sauer, M. Understanding how microorganisms respond to acid pH is central to their control and successful exploitation. Front. Microbiol. 2020, 11, 556140. [Google Scholar] [CrossRef] [PubMed]
- Lyu, D.; Zajonc, J.; Pagé, A.; Tanney, C.A.; Shah, A.; Monjezi, N.; Msimbira, L.A.; Antar, M.; Nazari, M.; Backer, R. Plant holobiont theory: The phytomicrobiome plays a central role in evolution and success. Microorganisms 2021, 9, 675. [Google Scholar] [CrossRef] [PubMed]
- Mahapatra, S.; Yadav, R.; Ramakrishna, W. Bacillus subtilis impact on plant growth, soil health and environment: Dr. Jekyll and Mr. Hyde. J. Appl. Microbiol. 2022, 132, 3543–3562. [Google Scholar] [CrossRef] [PubMed]
- Marschner, H. Marschner’s Mineral Nutrition of Higher Plants; Academic Press: Cambridge, MA, USA, 2011. [Google Scholar]
- Mazmanian, S.K.; Liu, G.; Ton-That, H.; Schneewind, O. Staphylococcus aureus sortase, an enzyme that anchors surface proteins to the cell wall. Science 1999, 285, 760–763. [Google Scholar] [CrossRef]
- Morinaga, K.; Kusada, H.; Watanabe, M.; Tamaki, H. Complete Genome Sequence of Lactobacillus helveticus JCM 1004, an Aminopeptidase-Producing Lactic Acid Bacterium. Microbiol. Resour. Announc. 2021, 10, e00641-21. [Google Scholar] [CrossRef]
- Msimbira, L.A.; Naamala, J.; Antar, M.; Subramanian, S.; Smith, D.L. Effect of Microbial Cell-Free Supernatants Extracted From a Range of pH Levels on Corn (Zea mays L.) and Tomato (Solanum lycopersicum L.) Seed Germination and Seedling Growth. Front. Sustain. Food Syst. 2022, 6, 789335. [Google Scholar] [CrossRef]
- Msimbira, L.A.; Smith, D.L. The roles of plant growth promoting microbes in enhancing plant tolerance to acidity and alkalinity stresses. Front. Sustain. Food Syst. 2020, 4, 106. [Google Scholar] [CrossRef]
- Naamala, J.; Msimbira, L.A.; Antar, M.; Subramanian, S.; Smith, D.L. Cell-Free Supernatant Obtained From a Salt Tolerant Bacillus amyloliquefaciens Strain Enhances Germination and Radicle Length under NaCl Stressed and Optimal Conditions. Front. Sustain. Food Syst. 2022, 6, 788939. [Google Scholar] [CrossRef]
- Nesvizhskii, A.I.; Keller, A.; Kolker, E.; Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 2003, 75, 4646–4658. [Google Scholar] [CrossRef]
- Niazi, A.; Manzoor, S.; Asari, S.; Bejai, S.; Meijer, J.; Bongcam-Rudloff, E. Genome analysis of Bacillus amyloliquefaciens subsp. plantarum UCMB5113: A rhizobacterium that improves plant growth and stress management. PLoS ONE 2014, 9, e104651. [Google Scholar] [CrossRef]
- Raman, J.; Kim, J.-S.; Choi, K.R.; Eun, H.; Yang, D.; Ko, Y.-J.; Kim, S.-J. Application of lactic acid bacteria (LAB) in sustainable agriculture: Advantages and limitations. Int. J. Mol. Sci. 2022, 23, 7784. [Google Scholar] [CrossRef] [PubMed]
- Ratzke, C.; Gore, J. Modifying and reacting to the environmental pH can drive bacterial interactions. PLoS Biol. 2018, 16, e2004248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rehman, H.M.; Cheung, W.-L.; Wong, K.-S.; Xie, M.; Luk, C.-Y.; Wong, F.-L.; Li, M.-W.; Tsai, S.-N.; To, W.-T.; Chan, L.-Y. High-throughput mass spectrometric analysis of the whole proteome and secretome from Sinorhizobium fredii strains CCBAU25509 and CCBAU45436. Front. Microbiol. 2019, 10, 2569. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.S.; Ragg, N.L.; Cummings, V.J.; Zhang, J. Ocean acidification and dynamic energy budget models: Parameterisation and simulations for the green-lipped mussel. Ecol. Model. 2020, 426, 109069. [Google Scholar] [CrossRef]
- Rodríguez Ayala, F.; Bartolini, M.; Grau, R. The stress-responsive alternative sigma factor SigB of Bacillus subtilis and its relatives: An old friend with new functions. Front. Microbiol. 2020, 11, 1761. [Google Scholar] [CrossRef]
- Rousk, J.; Bååth, E.; Brookes, P.C.; Lauber, C.L.; Lozupone, C.; Caporaso, J.G.; Knight, R.; Fierer, N. Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J. 2010, 4, 1340–1351. [Google Scholar] [CrossRef] [PubMed]
- Satapute, P.; Olekar, H.; Shetti, A.; Kulkarni, A.; Hiremath, G.; Patagundi, B.; Shivsharan, C.; Kaliwal, B. Isolation and characterization of nitrogen fixing Bacillus subtilis strain as-4 from agricultural soil. Int. J. Recent. Sci. Res. 2012, 3, 762–765. [Google Scholar]
- Schleifer, K.H.; Ludwig, W. Phylogeny of the genus Lactobacillus and related genera. Syst. Appl. Microbiol. 1995, 18, 461–467. [Google Scholar] [CrossRef]
- Schneider, E.; Hunke, S. ATP-binding-cassette (ABC) transport systems: Functional and structural aspects of the ATP-hydrolyzing subunits/domains. FEMS Microbiol. Rev. 1998, 22, 1–20. [Google Scholar] [CrossRef]
- Sessitsch, A.; Mitter, B. 21st century agriculture: Integration of plant microbiomes for improved crop production and food security. Microb. Biotechnol. 2015, 8, 32. [Google Scholar] [CrossRef]
- Shabayek, S.; Spellerberg, B. Acid stress response mechanisms of group B streptococci. Front. Cell. Infect. Microbiol. 2017, 7, 395. [Google Scholar] [CrossRef] [PubMed]
- Shrivastava, P.; Kumar, R. Soil salinity: A serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saudi J. Biol. Sci. 2015, 22, 123–131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simonen, M.; Palva, I. Protein secretion in Bacillus species. Microbiol. Rev. 1993, 57, 109–137. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.P.; Runthala, A.; Khan, S.; Jha, P.N. Quantitative proteomics analysis reveals the tolerance of wheat to salt stress in response to Enterobacter cloacae SBP-8. PLoS ONE 2017, 12, e0183513. [Google Scholar] [CrossRef] [Green Version]
- Slattery, L.; O’callaghan, J.; Fitzgerald, G.; Beresford, T.; Ross, R. Invited review: Lactobacillus helveticus—A thermophilic dairy starter related to gut bacteria. J. Dairy Sci. 2010, 93, 4435–4454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sofi, M.H.; Gudi, R.; Karumuthil-Melethil, S.; Perez, N.; Johnson, B.M.; Vasu, C. pH of drinking water influences the composition of gut microbiome and type 1 diabetes incidence. Diabetes 2014, 63, 632–644. [Google Scholar] [CrossRef] [Green Version]
- Souza, P.M.; Werneck, G.; Aliakbarian, B.; Siqueira, F.; Ferreira Filho, E.X.; Perego, P.; Converti, A.; Magalhães, P.O.; Junior, A.P. Production, purification and characterization of an aspartic protease from Aspergillus foetidus. Food Chem. Toxicol. 2017, 109, 1103–1110. [Google Scholar] [CrossRef]
- Stadtman, T.C. Selenium-dependent enzymes. Annu. Rev. Biochem. 1980, 49, 93–110. [Google Scholar] [CrossRef]
- Stephenson, K.; Bron, S.; Harwood, C. Cellular lysis in Bacillus subtilis; the affect of multiple extracellular protease deficiencies. Lett. Appl. Microbiol. 1999, 29, 141–145. [Google Scholar] [CrossRef]
- Subramanian, S.; Smith, D.L. Bacteriocins from the rhizosphere microbiome–from an agriculture perspective. Front. Plant Sci. 2015, 6, 909. [Google Scholar] [CrossRef] [Green Version]
- Subramanian, S.; Souleimanov, A.; Smith, D.L. Proteomic studies on the effects of lipo-chitooligosaccharide and thuricin 17 under unstressed and salt stressed conditions in Arabidopsis thaliana. Front. Plant Sci. 2016, 7, 1314. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, S.; Souleimanov, A.; Smith, D.L. Thuricin17 Production and Proteome Differences in Bacillus thuringiensis NEB17 Cell-Free Supernatant Under NaCl Stress. Front. Sustain. Food Syst. 2021, 5, 69. [Google Scholar] [CrossRef]
- Sumby, K.M.; Grbin, P.R.; Jiranek, V. Implications of new research and technologies for malolactic fermentation in wine. Appl. Microbiol. Biotechnol. 2014, 98, 8111–8132. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y. F1F0-ATPase functions under markedly acidic conditions in bacteria. In Regulation of Ca2+-ATPases, V-ATPases and F-ATPases; Springer: Cham, Switzerland, 2016. [Google Scholar]
- Suzuki, S.; Yokota, K.; Igimi, S.; Kajikawa, A. Comparative analysis of immunological properties of S-layer proteins isolated from Lactobacillus strains. Microbiology 2019, 165, 188–196. [Google Scholar] [CrossRef]
- Takishita, Y.; Souleimanov, A.; Bourguet, C.; Ohlund, L.B.; Arnold, A.A.; Sleno, L.; Smith, D.L. Pseudomonas entomophila 23S produces a novel antagonistic compound against Clavibacter michiganensis subsp. michiganensis, a pathogen of tomato bacterial canker. Appl. Microbiol. 2021, 1, 60–73. [Google Scholar] [CrossRef]
- Terrasse, R. La Glycéraldéhyde-3-Phosphate Déshydrogénase, Une Protéine De La Glycolyse Présente À La Surface Cellulaire, Est Impliquée Dans La Reconnaissance Par Le Système Du Complément Chez Streptococcus Pneumoniae. Ph.D. Thesis, Université Grenoble Alpes, Grenoble, France, 2013. [Google Scholar]
- Tjalsma, H.; Antelmann, H.; Jongbloed, J.D.; Braun, P.G.; Darmon, E.; Dorenbos, R.; Dubois, J.-Y.F.; Westers, H.; Zanen, G.; Quax, W.J. Proteomics of protein secretion by Bacillus subtilis: Separating the “secrets” of the secretome. Microbiol. Mol. Biol. Rev. 2004, 68, 207–233. [Google Scholar] [CrossRef] [Green Version]
- Tjalsma, H.; Bolhuis, A.; Jongbloed, J.D.; Bron, S.; Van Dijl, J.M. Signal peptide-dependent protein transport in Bacillus subtilis: A genome-based survey of the secretome. Microbiol. Mol. Biol. Rev. 2000, 64, 515–547. [Google Scholar] [CrossRef] [Green Version]
- Toyofuku, M.; Nomura, N.; Eberl, L. Types and origins of bacterial membrane vesicles. Nat. Rev. Microbiol. 2019, 17, 13–24. [Google Scholar] [CrossRef]
- Tyler, B.; Collmer, A.; Collmer, C.; Dean, R. The PAMGO consortium: Unifying themes in microbe-host associations identified through the gene ontology. BMC Microbiol. 2009, 9. [Google Scholar]
- Vandenkoornhuyse, P.; Quaiser, A.; Duhamel, M.; Le Van, A.; Dufresne, A. The importance of the microbiome of the plant holobiont. New Phytol. 2015, 206, 1196–1206. [Google Scholar] [CrossRef]
- Varki, A.; Cummings, R.D.; Esko, J.D.; Stanley, P.; Hart, G.W.; Aebi, M.; Darvill, A.G.; Kinoshita, T.; Packer, N.H.; Prestegard, J.H. Essentials of Glycobiology [Internet]; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2015. [Google Scholar]
- Wang, C.; Cui, Y.; Qu, X. Mechanisms and improvement of acid resistance in lactic acid bacteria. Arch. Microbiol. 2018, 200, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Xia, Y.; Gu, Z.; Zhang, H.; Chen, Y.Q.; Chen, H.; Ai, L.; Chen, W. A new potential secretion pathway for recombinant proteins in Bacillus subtilis. Microb. Cell Factories 2015, 14, 179. [Google Scholar] [CrossRef] [PubMed]
- Wiegert, T.; Homuth, G.; Versteeg, S.; Schumann, W. Alkaline shock induces the Bacillus subtilisσW regulon. Mol. Microbiol. 2001, 41, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Wilks, J.C.; Kitko, R.D.; Cleeton, S.H.; Lee, G.E.; Ugwu, C.S.; Jones, B.D.; Bondurant, S.S.; Slonczewski, J.L. Acid and base stress and transcriptomic responses in Bacillus subtilis. Appl. Environ. Microbiol. 2009, 75, 981–990. [Google Scholar] [CrossRef] [Green Version]
- Wong, S.-L. Advances in the use of Bacillus subtilis for the expression and secretion of heterologous proteins. Curr. Opin. Biotechnol. 1995, 6, 517–522. [Google Scholar] [CrossRef]
- Wooldridge, K. Bacterial Secreted Proteins; Caister Academic Press: Wymondham, UK, 2009. [Google Scholar]
- Wu, Z.P.; Hilvert, D. Conversion of a protease into an acyl transferase: Selenolsubtilisin. J. Am. Chem. Soc. 1989, 111, 4513–4514. [Google Scholar] [CrossRef]
- Wu, Z.P.; Hilvert, D. Selenosubtilisin as a glutathione peroxidase mimic. J. Am. Chem. Soc. 1990, 112, 5647–5648. [Google Scholar] [CrossRef]
- Yadav, R.; Ror, P.; Rathore, P.; Ramakrishna, W. Bacteria from native soil in combination with arbuscular mycorrhizal fungi augment wheat yield and biofortification. Plant Physiol. Biochem. 2020, 150, 222–233. [Google Scholar] [CrossRef]
- Yaghoubian, I.; Msimbira, L.A.; Smith, D.L. Cell-free supernatant of Bacillus strains (CFS) can improve seed vigor index of corn under salinity stress. Front. Sustain. Food Syst. 2022, 92. [Google Scholar]
- Yang, C.-K.; Ewis, H.E.; Zhang, X.; Lu, C.-D.; Hu, H.-J.; Pan, Y.; Abdelal, A.T.; Tai, P.C. Nonclassical protein secretion by Bacillus subtilis in the stationary phase is not due to cell lysis. J. Bacteriol. 2011, 193, 5607–5615. [Google Scholar] [CrossRef]
- Zheng, W.; Cai, X.; Xie, M.; Liang, Y.; Wang, T.; Li, Z. Structure-based identification of a potent inhibitor targeting Stp1-mediated virulence regulation in Staphylococcus aureus. Cell Chem. Biol. 2016, 23, 1002–1010. [Google Scholar] [CrossRef] [PubMed]



| Parameter | B. subtilis | L. helveticus |
|---|---|---|
| Genome size (bp) | 4,214,630 [23,32] | 21,000 [30] |
| Optimum growth pH range | 7.0 [18,29] | 5.5 to 5.8 [18,30] |
| Proteins protein-coding genes | 4610 [33] | 2462 [34] |
| Enzyme Code Classes | B. subtilis (EB2004S) | L. helveticus (EL2006H) | |||
|---|---|---|---|---|---|
| pH 7 | pH 5 | pH 8 | pH 7 | pH 5 | |
| Hydrolases | 591 | 34 (↓94%) | 644 (↑9%) | 431 | 118 (↓73%) |
| Isomerases | 92 | 2 (↓98%) | 123 (↑33%) | 66.3 | 13.7 (↓79%) |
| Ligases | 155 | 0 (↓100%) | 188 (↑21%) | 103.3 | 1.7 (↓98%) |
| Lyases | 134 | 0 (↓100%) | 199 (↑49%) | 44.7 | 6.7 (↓85%) |
| Oxidoreductases | 392 | 20 (↓95%) | 490 (↑25%) | 72 | 0 (↓100%) |
| Transferases | 457 | 6 (↓98 %) | 584 (↑28%) | 194.7 | 3 (↓99%) |
| Translocases | 95 | 0 (↓100%) | 119 (↑26%) | 31.3 | 6.7 (↓79%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Msimbira, L.A.; Subramanian, S.; Naamala, J.; Antar, M.; Smith, D.L. Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes. Int. J. Mol. Sci. 2022, 23, 15144. https://doi.org/10.3390/ijms232315144
Msimbira LA, Subramanian S, Naamala J, Antar M, Smith DL. Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes. International Journal of Molecular Sciences. 2022; 23(23):15144. https://doi.org/10.3390/ijms232315144
Chicago/Turabian StyleMsimbira, Levini A., Sowmyalakshmi Subramanian, Judith Naamala, Mohammed Antar, and Donald L. Smith. 2022. "Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes" International Journal of Molecular Sciences 23, no. 23: 15144. https://doi.org/10.3390/ijms232315144
APA StyleMsimbira, L. A., Subramanian, S., Naamala, J., Antar, M., & Smith, D. L. (2022). Secretome Analysis of the Plant Biostimulant Bacteria Strains Bacillus subtilis (EB2004S) and Lactobacillus helveticus (EL2006H) in Response to pH Changes. International Journal of Molecular Sciences, 23(23), 15144. https://doi.org/10.3390/ijms232315144









