Microbe–Immune–Stress Interactions Impact Behaviour during Postnatal Development
Abstract
1. Introduction
2. Results
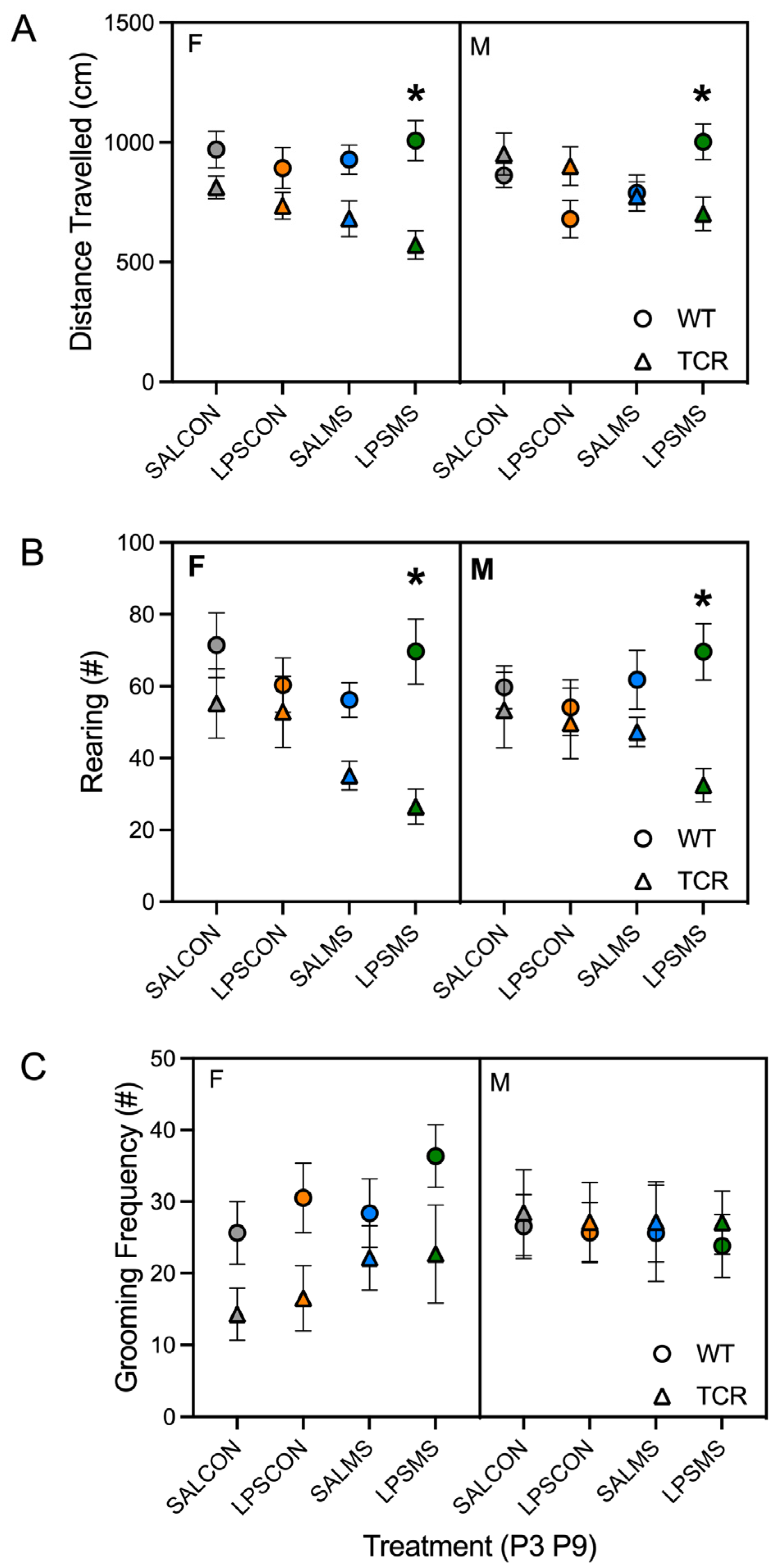
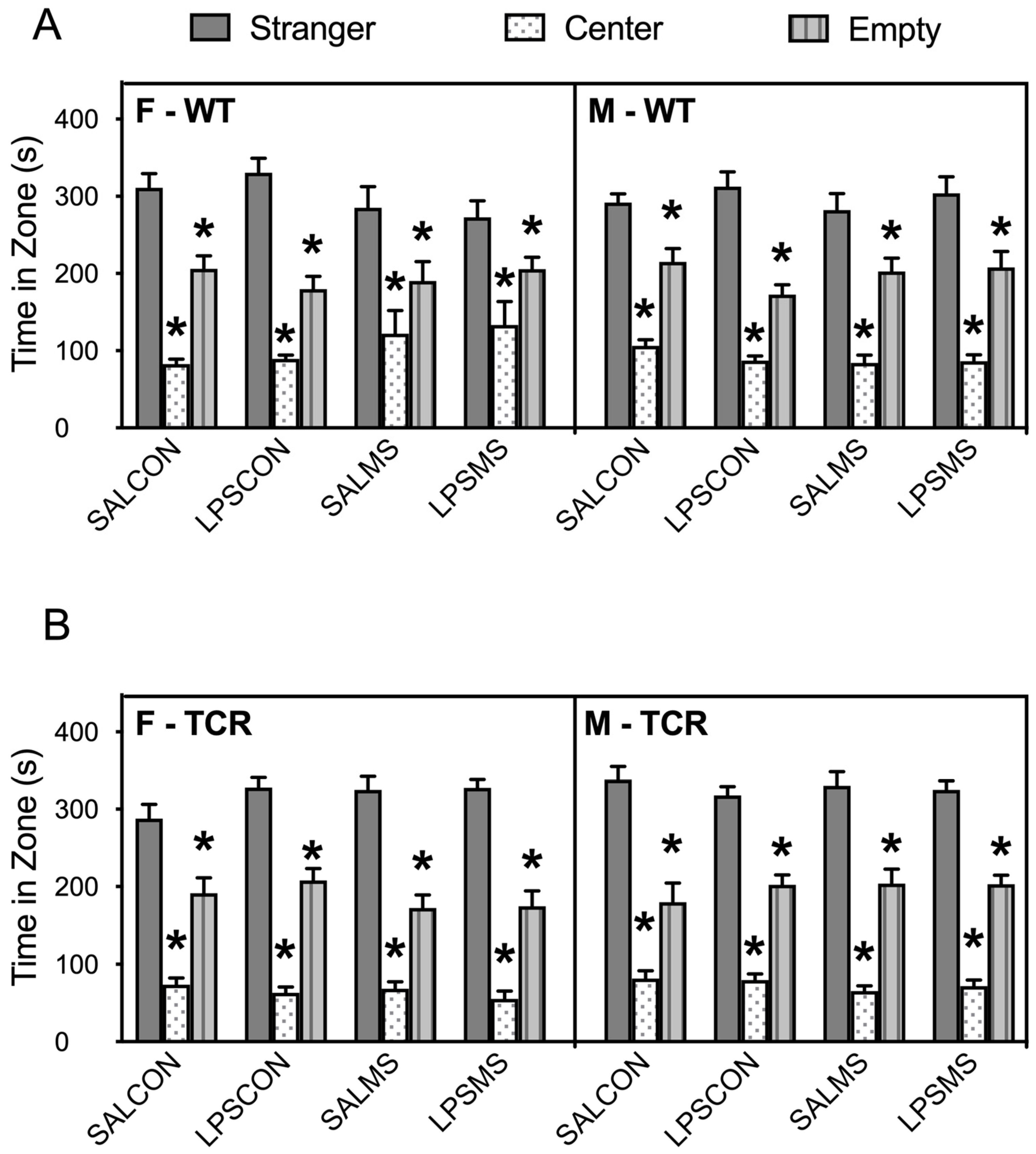
2.1. T-Cell-Deficient Mice Showed Altered Postnatal Behaviour in Comparison to B6 Mice
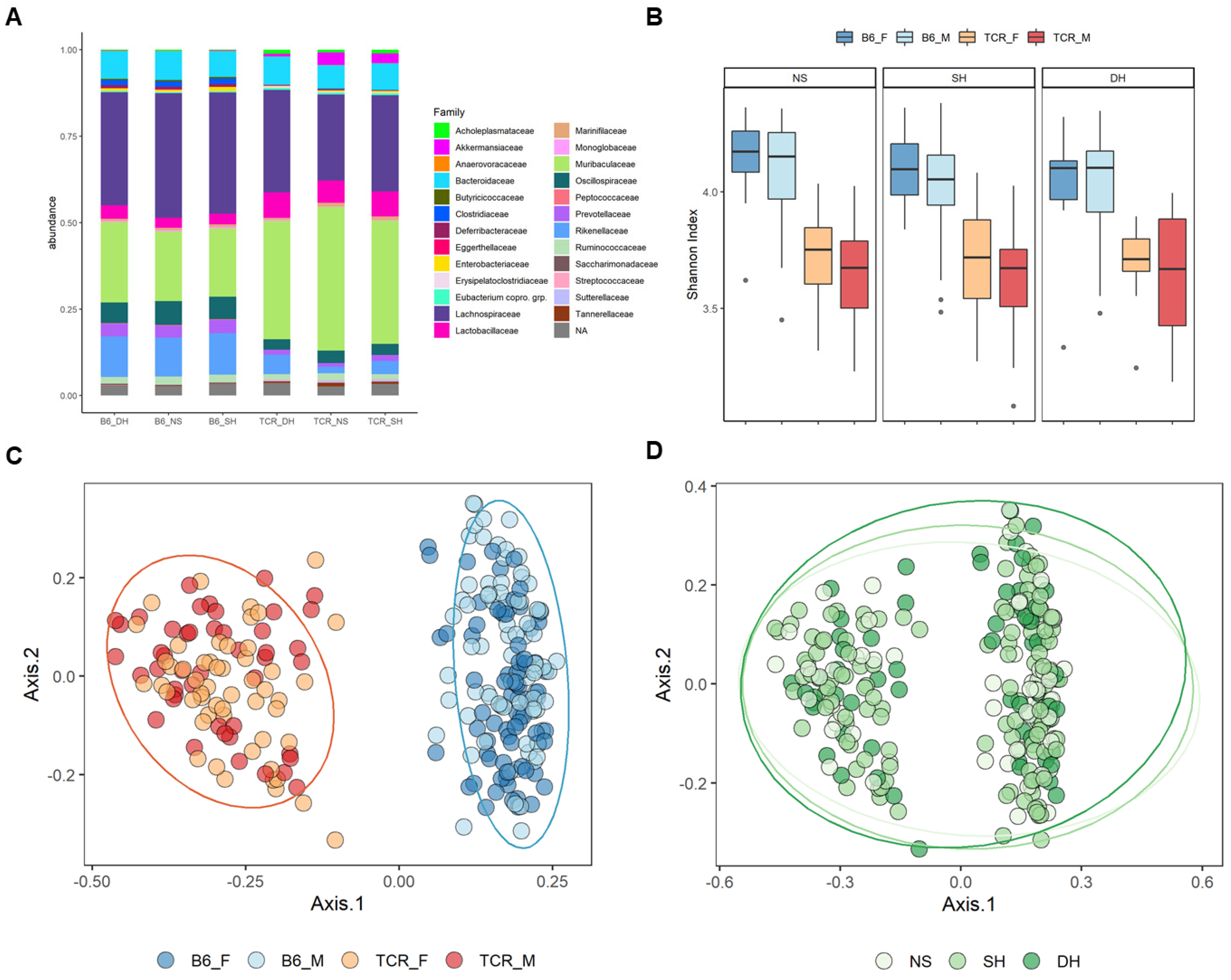
2.2. Microbial Diversity and Compositional Differences between B6 and TCRβ-/-δ-/-
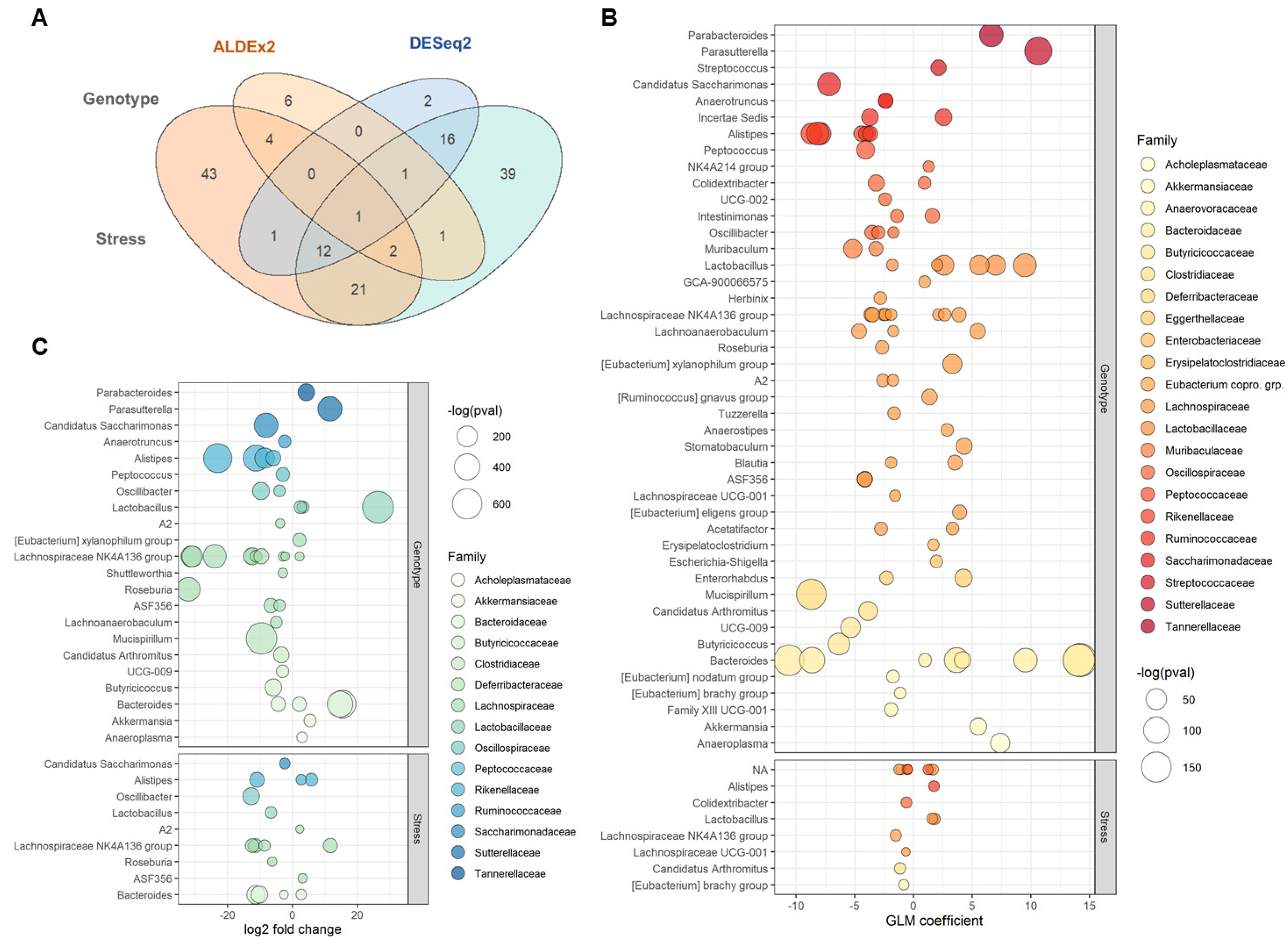
2.3. Differential Taxonomic Abundance between B6 and TCRβ-/-δ-/-
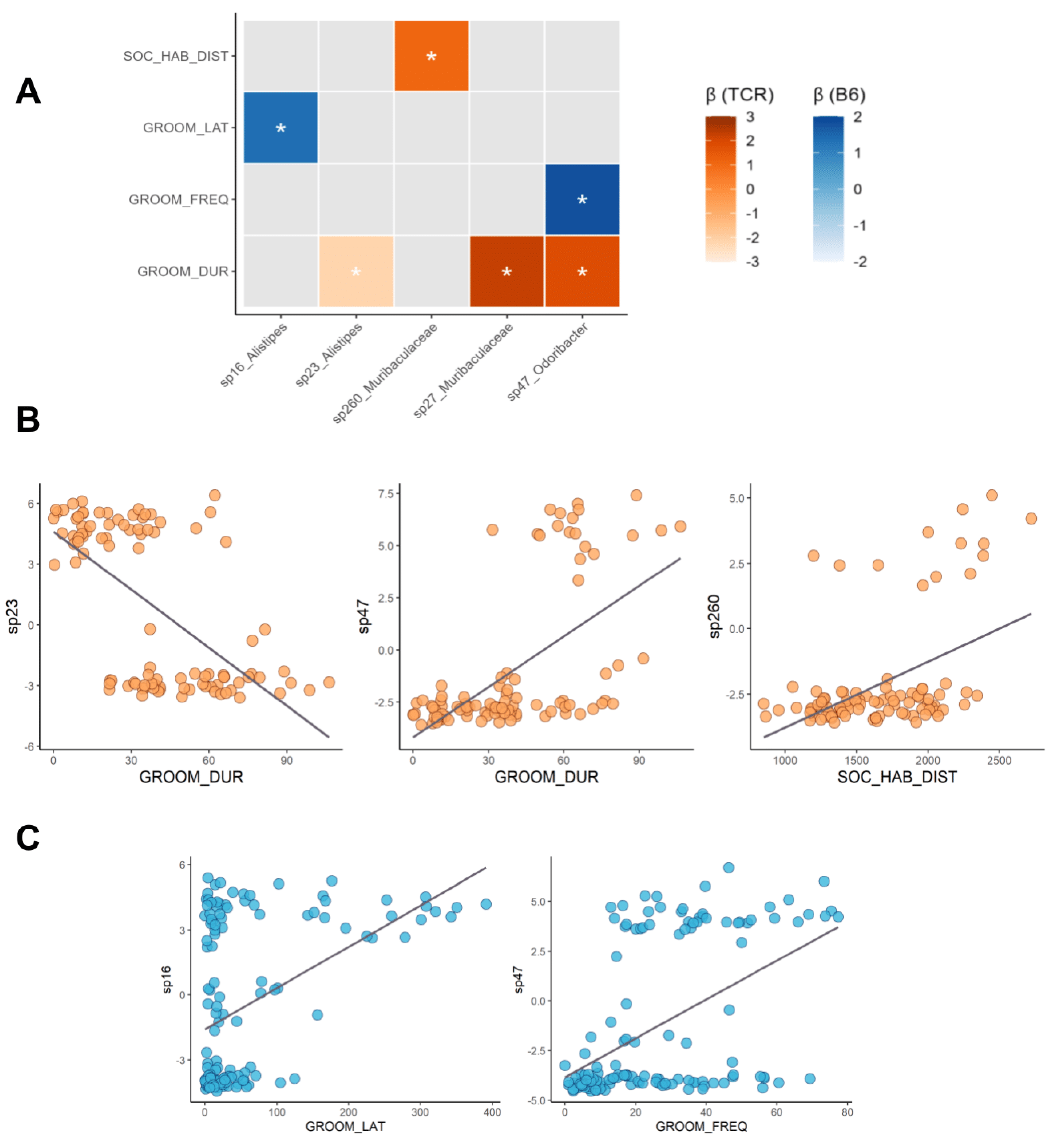
2.4. Microbe–Behaviour Relationships in B6 and TCRβ-/-δ-/- Mice
3. Discussion
4. Materials and Methods
4.1. Animals and Experimental Design
4.2. Behavioural Testing
4.2.1. Righting Reflex
4.2.2. USV Recordings
4.2.3. Open Field
4.2.4. Sociability
4.2.5. Self-Grooming
4.2.6. Behavioural Data Analysis
4.3. 16S rRNA Gene Sequencing and Amplicon Sequence Variant (ASV) Processing
4.4. 16S rRNA Data Statistical Analysis
4.4.1. QC of Sequencing Data
4.4.2. Alpha Diversity Analysis
4.4.3. Beta Diversity Analysis
4.4.4. Retain-Resolve Preprocessing of Microbiome Data
4.4.5. Bacterial Abundance Analysis: Assessing Differences between B6 and TCRβ-/-δ-/- Mice
4.5. Microbe–Behaviour Correlation Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jasarevic, E.; Bale, T.L. Prenatal and postnatal contributions of the maternal microbiome on offspring programming. Front. Neuroendocrinol. 2019, 55, 100797. [Google Scholar] [CrossRef] [PubMed]
- Jaggar, M.; Rea, K.; Spichak, S.; Dinan, T.G.; Cryan, J.F. You’ve got male: Sex and the microbiota-gut-brain axis across the lifespan. Front. Neuroendocrinol. 2020, 56, 100815. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.; Kim, W.-U. Microbiota in T-cell homeostasis and inflammatory diseases. Exp. Mol. Med. 2017, 49, e340. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Guan, X.; Chen, D.; Ma, W. The Th17/Treg Cell Balance: A Gut Microbiota-Modulated Story. Microorganisms 2019, 7, 583. [Google Scholar] [CrossRef] [PubMed]
- Dollé, L.; Tran, H.Q.; Etienne-Mesmin, L.; Chassaing, B. Policing of gut microbiota by the adaptive immune system. BMC Med. 2016, 14, 27. [Google Scholar] [CrossRef]
- Zhang, H.; Sparks, J.B.; Karyala, S.V.; Settlage, R.; Luo, X.M. Host adaptive immunity alters gut microbiota. ISME J. 2015, 9, 770–781. [Google Scholar] [CrossRef]
- Tanabe, S.; Yamashita, T. The role of immune cells in brain development and neurodevelopmental diseases. Int. Immunol. 2018, 30, 437–444. [Google Scholar] [CrossRef]
- Quinnies, K.M.; Cox, K.H.; Rissman, E.F. Immune deficiency influences juvenile social behavior and maternal behavior. Behav. Neurosci. 2015, 129, 331–338. [Google Scholar] [CrossRef]
- McGowan, P.O.; Hope, T.A.; Meck, W.H.; Kelsoe, G.; Williams, C.L. Impaired social recognition memory in recombination activating gene 1-deficient mice. Brain Res. 2011, 1383, 187–195. [Google Scholar] [CrossRef]
- Derecki, N.C.; Cardani, A.N.; Yang, C.H.; Quinnies, K.M.; Crihfield, A.; Lynch, K.R.; Kipnis, J. Regulation of learning and memory by meningeal immunity: A key role for IL-4. J. Exp. Med. 2010, 207, 1067–1080. [Google Scholar] [CrossRef]
- Choi, B.; Yim Yeong, S.; Wong, H.; Kim, S.; Kim, H.; Kim Sangwon, V.; Hoeffer Charles, A.; Littman Dan, R.; Huh Jun, R. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 2016, 351, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, S.; Yamashita, T. B-1a lymphocytes promote oligodendrogenesis during brain development. Nat. Neurosci. 2018, 21, 506–516. [Google Scholar] [CrossRef] [PubMed]
- Alves de Lima, K.; Rustenhoven, J.; Da Mesquita, S.; Wall, M.; Salvador, A.F.; Smirnov, I.; Martelossi Cebinelli, G.; Mamuladze, T.; Baker, W.; Papadopoulos, Z.; et al. Meningeal γδ T cells regulate anxiety-like behavior via IL-17a signaling in neurons. Nat. Immunol. 2020, 21, 1421–1429. [Google Scholar] [CrossRef] [PubMed]
- Filiano, A.J.; Xu, Y.; Tustison, N.J.; Marsh, R.L.; Baker, W.; Smirnov, I.; Overall, C.C.; Gadani, S.P.; Turner, S.D.; Weng, Z.; et al. Unexpected role of interferon-γ in regulating neuronal connectivity and social behaviour. Nature 2016, 535, 425–429. [Google Scholar] [CrossRef]
- Clark, S.M.; Vaughn, C.N.; Soroka, J.A.; Li, X.; Tonelli, L.H. Neonatal adoptive transfer of lymphocytes rescues social behaviour during adolescence in immune-deficient mice. Eur. J. Neurosci. 2018, 47, 968–978. [Google Scholar] [CrossRef]
- Rilett, K.C.; Friedel, M.; Ellegood, J.; MacKenzie, R.N.; Lerch, J.P.; Foster, J.A. Loss of T cells influences sex differences in behavior and brain structure. Brain Behav. Immun. 2015, 46, 249–260. [Google Scholar] [CrossRef]
- Rilett, K.C.; Luo, O.D.; McVey-Neufeld, K.-A.; MacKenzie, R.N.; Foster, J.A. Loss of T cells influences sex differences in stress-related gene expression. J. Neuroimmunol. 2020, 343, 577213. [Google Scholar] [CrossRef]
- Caspani, G.; Green, M.; Swann, J.R.; Foster, J.A. Microbe-Immune Crosstalk: Evidence That T Cells Influence the Development of the Brain Metabolome. Int. J. Mol. Sci. 2022, 23, 3259. [Google Scholar] [CrossRef]
- Asbury, S.; Lai, J.K.Y.; Rilett, K.C.; Haqqee, Z.; Darwin, B.C.; Ellegood, J.; Lerch, J.P.; Foster, J.A. Host genetics maps to behaviour and brain structure in mice. bioRxiv 2022. [Google Scholar] [CrossRef]
- Park, H.J.; Kim, S.A.; Kang, W.S.; Kim, J.W. Early-Life Stress Modulates Gut Microbiota and Peripheral and Central Inflammation in a Sex-Dependent Manner. Int. J. Mol. Sci. 2021, 22, 1899. [Google Scholar] [CrossRef]
- De Palma, G.; Blennerhassett, P.; Lu, J.; Deng, Y.; Park, A.J.; Green, W.; Denou, E.; Silva, M.A.; Santacruz, A.; Sanz, Y.; et al. Microbiota and host determinants of behavioural phenotype in maternally separated mice. Nat. Commun. 2015, 6, 7735. [Google Scholar] [CrossRef] [PubMed]
- Rincel, M.; Darnaudéry, M. Maternal separation in rodents: A journey from gut to brain and nutritional perspectives. Proc. Nutr. Soc. 2020, 79, 113–132. [Google Scholar] [CrossRef] [PubMed]
- Amath, A.; Foster, J.A.; Sidor, M.M. Developmental alterations in CNS stress-related gene expression following postnatal immune activation. Neuroscience 2012, 220, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Sidor, M.M.; Halgren, C.R.; Foster, J.A. The impact of early life immune challenge on behavior and microglia during postnatal development. Inflamm. Cell Signal. 2014, 1. [Google Scholar] [CrossRef]
- Binder, E.; Malki, K.; Paya-Cano, J.L.; Fernandes, C.; Aitchison, K.J.; Mathe, A.A.; Sluyter, F.; Schalkwyk, L.C. Antidepressants and the resilience to early-life stress in inbred mouse strains. Pharm. Genom. 2011, 21, 779–789. [Google Scholar] [CrossRef] [PubMed]
- Akillioglu, K.; Yilmaz, M.B.; Boga, A.; Binokay, S.; Kocaturk-Sel, S. Environmental enrichment does not reverse the effects of maternal deprivation on NMDAR and Balb/c mice behaviors. Brain Res. 2015, 1624, 479–488. [Google Scholar] [CrossRef]
- Bouet, V.; Percelay, S.; Leroux, E.; Diarra, B.; Leger, M.; Delcroix, N.; Andrieux, A.; Dollfus, S.; Freret, T.; Boulouard, M. A new 3-hit mouse model of schizophrenia built on genetic, early and late factors. Schizophr. Res. 2021, 228, 519–528. [Google Scholar] [CrossRef]
- Huo, R.; Zeng, B.; Zeng, L.; Cheng, K.; Li, B.; Luo, Y.; Wang, H.; Zhou, C.; Fang, L.; Li, W.; et al. Microbiota Modulate Anxiety-Like Behavior and Endocrine Abnormalities in Hypothalamic-Pituitary-Adrenal Axis. Front. Cell. Infect. Microbiol. 2017, 7, 489. [Google Scholar] [CrossRef]
- Luczynski, P.; Whelan, S.O.; O’Sullivan, C.; Clarke, G.; Shanahan, F.; Dinan, T.G.; Cryan, J.F. Adult microbiota-deficient mice have distinct dendritic morphological changes: Differential effects in the amygdala and hippocampus. Eur. J. Neurosci. 2016, 44, 2654–2666. [Google Scholar] [CrossRef]
- Heijtz, R.D.; Wang, S.; Anuar, F.; Qian, Y.; Björkholm, B.; Samuelsson, A.; Hibberd, M.L.; Forssberg, H.; Pettersson, S. Normal gut microbiota modulates brain development and behavior. Proc. Natl. Acad. Sci. USA 2011, 108, 3047. [Google Scholar] [CrossRef]
- Neufeld, K.M.; Kang, N.; Bienenstock, J.; Foster, J.A. Reduced anxiety-like behavior and central neurochemical change in germ-free mice. Neurogastroenterol. Motil. 2011, 23, 255-e119. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.; Dikiy, S.; Bhattarai, S.K.; Chinen, T.; Matheis, F.; Calafiore, M.; Hoyos, B.; Hanash, A.; Mucida, D.; Bucci, V.; et al. Extrathymically Generated Regulatory T Cells Establish a Niche for Intestinal Border-Dwelling Bacteria and Affect Physiologic Metabolite Balance. Immunity 2018, 48, 1245–1257.e1249. [Google Scholar] [CrossRef] [PubMed]
- Kawamoto, S.; Maruya, M.; Kato, L.M.; Suda, W.; Atarashi, K.; Doi, Y.; Tsutsui, Y.; Qin, H.; Honda, K.; Okada, T.; et al. Foxp3+ T Cells Regulate Immunoglobulin A Selection and Facilitate Diversification of Bacterial Species Responsible for Immune Homeostasis. Immunity 2014, 41, 152–165. [Google Scholar] [CrossRef] [PubMed]
- Plöger, S.; Stumpff, F.; Penner, G.B.; Schulzke, J.-D.; Gäbel, G.; Martens, H.; Shen, Z.; Günzel, D.; Aschenbach, J.R. Microbial butyrate and its role for barrier function in the gastrointestinal tract. Ann. N. Y. Acad. Sci. 2012, 1258, 52–59. [Google Scholar] [CrossRef]
- Wang, H.-B.; Wang, P.-Y.; Wang, X.; Wan, Y.-L.; Liu, Y.-C. Butyrate Enhances Intestinal Epithelial Barrier Function via Up-Regulation of Tight Junction Protein Claudin-1 Transcription. Dig. Dis. Sci. 2012, 57, 3126–3135. [Google Scholar] [CrossRef]
- Burger-van Paassen, N.; Vincent, A.; Puiman, P.J.; van der Sluis, M.; Bouma, J.; Boehm, G.; Van Goudoever, J.B.; Van Seuningen, I.; Renes, I.B. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: Implications for epithelial protection. Biochem. J. 2009, 420, 211–219. [Google Scholar] [CrossRef]
- Kelly, C.J.; Zheng, L.; Campbell, E.L.; Saeedi, B.; Scholz, C.C.; Bayless, A.J.; Wilson, K.E.; Glover, L.E.; Kominsky, D.J.; Magnuson, A.; et al. Crosstalk between Microbiota-Derived Short-Chain Fatty Acids and Intestinal Epithelial HIF Augments Tissue Barrier Function. Cell Host Microbe 2015, 17, 662–671. [Google Scholar] [CrossRef]
- Atarashi, K. Control of Intestinal Regulatory T Cells by Human Commensal Bacteria. In Chronic Inflammation: Mechanisms and Regulation; Miyasaka, M., Takatsu, K., Eds.; Springer: Tokyo, Japan, 2016; pp. 591–601. [Google Scholar]
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013, 504, 451–455. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Hase, K. Commensal microbiota regulates T cell fate decision in the gut. Semin. Immunopathol. 2015, 37, 17–25. [Google Scholar] [CrossRef]
- Ansaldo, E.; Slayden, L.C.; Ching, K.L.; Koch, M.A.; Wolf, N.K.; Plichta, D.R.; Brown, E.M.; Graham, D.B.; Xavier, R.J.; Moon, J.J.; et al. Akkermansia muciniphila induces intestinal adaptive immune responses during homeostasis. Science 2019, 364, 1179–1184. [Google Scholar] [CrossRef]
- Moens, F.; Van den Abbeele, P.; Basit, A.W.; Dodoo, C.; Chatterjee, R.; Smith, B.; Gaisford, S. A four-strain probiotic exerts positive immunomodulatory effects by enhancing colonic butyrate production in vitro. Int. J. Pharm. 2019, 555, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Patman, G. Lactobacillus acidophilus opens the door to butyrate. Nat. Rev. Gastroenterol. Hepatol. 2015, 12, 552. [Google Scholar] [CrossRef] [PubMed]
- Livingston, M.; Loach, D.; Wilson, M.; Tannock, G.W.; Baird, M. Gut commensal Lactobacillus reuteri 100-23 stimulates an immunoregulatory response. Immunol. Cell Biol. 2010, 88, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Jiang, P.; Yang, W.; Jin, Y.; Huang, H.; Shi, C.; Jiang, Y.; Wang, J.; Kang, Y.; Wang, C.; Yang, G. Lactobacillus reuteri protects mice against Salmonella typhimurium challenge by activating macrophages to produce nitric oxide. Microb. Pathog. 2019, 137, 103754. [Google Scholar] [CrossRef] [PubMed]
- Seifert, S.; Rodriguez Gómez, M.; Watzl, B.; Holzapfel, W.H.; Franz, C.M.A.P.; Vizoso Pinto, M.G. Differential Effect of Lactobacillus johnsonii BFE 6128 on Expression of Genes Related to TLR Pathways and Innate Immunity in Intestinal Epithelial Cells. Probiotics Antimicrob. Proteins 2010, 2, 211–217. [Google Scholar] [CrossRef]
- Wang, M.; Wu, H.; Lu, L.; Jiang, L.; Yu, Q. Lactobacillus reuteri Promotes Intestinal Development and Regulates Mucosal Immune Function in Newborn Piglets. Front. Vet. Sci. 2020, 7, 42. [Google Scholar] [CrossRef]
- Forsberg, A.; Abrahamsson, T.R.; Björkstén, B.; Jenmalm, M.C. Pre- and postnatal administration of Lactobacillus reuteri decreases TLR2 responses in infants. Clin. Transl. Allergy 2014, 4, 21. [Google Scholar] [CrossRef]
- Fonseca, W.; Lucey, K.; Jang, S.; Fujimura, K.E.; Rasky, A.; Ting, H.A.; Petersen, J.; Johnson, C.C.; Boushey, H.A.; Zoratti, E.; et al. Lactobacillus johnsonii supplementation attenuates respiratory viral infection via metabolic reprogramming and immune cell modulation. Mucosal Immunol. 2017, 10, 1569–1580. [Google Scholar] [CrossRef]
- Liu, Y.; Fatheree, N.Y.; Dingle, B.M.; Tran, D.Q.; Rhoads, J.M. Lactobacillus reuteri DSM 17938 Changes the Frequency of Foxp3+ Regulatory T Cells in the Intestine and Mesenteric Lymph Node in Experimental Necrotizing Enterocolitis. PLoS ONE 2013, 8, e56547. [Google Scholar] [CrossRef]
- Cervantes-Barragan, L.; Chai Jiani, N.; Tianero Ma, D.; Di Luccia, B.; Ahern Philip, P.; Merriman, J.; Cortez Victor, S.; Caparon Michael, G.; Donia Mohamed, S.; Gilfillan, S.; et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+ T cells. Science 2017, 357, 806–810. [Google Scholar] [CrossRef]
- Bharwani, A.; Mian, M.F.; Foster, J.A.; Surette, M.G.; Bienenstock, J.; Forsythe, P. Structural & functional consequences of chronic psychosocial stress on the microbiome & host. Psychoneuroendocrinology 2016, 63, 217–227. [Google Scholar] [PubMed]
- Boukerb, A.M.; Cambronel, M.; Rodrigues, S.; Mesguida, O.; Knowlton, R.; Feuilloley, M.G.J.; Zommiti, M.; Connil, N. Inter-Kingdom Signaling of Stress Hormones: Sensing, Transport and Modulation of Bacterial Physiology. Front. Microbiol. 2021, 12, 690942. [Google Scholar] [CrossRef] [PubMed]
- Bonaz, B.; Bazin, T.; Pellissier, S. The Vagus Nerve at the Interface of the Microbiota-Gut-Brain Axis. Front. Neurosci. 2018, 12, 49. [Google Scholar] [CrossRef] [PubMed]
- Maltz, R.M.; Keirsey, J.; Kim, S.C.; Mackos, A.R.; Gharaibeh, R.Z.; Moore, C.C.; Xu, J.; Somogyi, A.; Bailey, M.T. Social Stress Affects Colonic Inflammation, the Gut Microbiome, and Short-chain Fatty Acid Levels and Receptors. J. Pediatr. Gastroenterol. Nutr. 2019, 68, 533–540. [Google Scholar] [CrossRef]
- Maltz, R.M.; Keirsey, J.; Kim, S.C.; Mackos, A.R.; Gharaibeh, R.Z.; Moore, C.C.; Xu, J.; Bakthavatchalu, V.; Somogyi, A.; Bailey, M.T. Prolonged restraint stressor exposure in outbred CD-1 mice impacts microbiota, colonic inflammation, and short chain fatty acids. PLoS ONE 2018, 13, e0196961. [Google Scholar] [CrossRef] [PubMed]
- Madison, A.; Kiecolt-Glaser, J.K. Stress, depression, diet, and the gut microbiota: Human-bacteria interactions at the core of psychoneuroimmunology and nutrition. Curr. Opin. Behav. Sci. 2019, 28, 105–110. [Google Scholar] [CrossRef]
- Galley, J.D.; Nelson, M.C.; Yu, Z.; Dowd, S.E.; Walter, J.; Kumar, P.S.; Lyte, M.; Bailey, M.T. Exposure to a social stressor disrupts the community structure of the colonic mucosa-associated microbiota. BMC Microbiol. 2014, 14, 189. [Google Scholar] [CrossRef]
- Gomez-Arango, L.F.; Barrett, H.L.; McIntyre, H.D.; Callaway, L.K.; Morrison, M.; Dekker Nitert, M. Increased Systolic and Diastolic Blood Pressure Is Associated with Altered Gut Microbiota Composition and Butyrate Production in Early Pregnancy. Hypertension 2016, 68, 974–981. [Google Scholar] [CrossRef]
- Zhou, L.; Xiao, X.; Zhang, Q.; Zheng, J.; Li, M.; Wang, X.; Deng, M.; Zhai, X.; Liu, J. Gut microbiota might be a crucial factor in deciphering the metabolic benefits of perinatal genistein consumption in dams and adult female offspring. Food Funct. 2019, 10, 4505–4521. [Google Scholar] [CrossRef]
- Parker, B.J.; Wearsch, P.A.; Veloo, A.C.M.; Rodriguez-Palacios, A. The Genus Alistipes: Gut Bacteria with Emerging Implications to Inflammation, Cancer, and Mental Health. Front. Immunol. 2020, 11, 906. [Google Scholar] [CrossRef]
- Li, H.; Xiang, Y.; Zhu, Z.; Wang, W.; Jiang, Z.; Zhao, M.; Cheng, S.; Pan, F.; Liu, D.; Ho, R.C.M.; et al. Rifaximin-mediated gut microbiota regulation modulates the function of microglia and protects against CUMS-induced depression-like behaviors in adolescent rat. J. Neuroinflamm. 2021, 18, 254. [Google Scholar] [CrossRef] [PubMed]
- Kong, Q.; Tian, P.; Zhao, J.; Zhang, H.; Wang, G.; Chen, W. The autistic-like behaviors development during weaning and sexual maturation in VPA-induced autistic-like rats is accompanied by gut microbiota dysbiosis. PeerJ 2021, 9, e11103. [Google Scholar] [CrossRef] [PubMed]
- Sgritta, M.; Dooling, S.W.; Buffington, S.A.; Momin, E.N.; Francis, M.B.; Britton, R.A.; Costa-Mattioli, M. Mechanisms Underlying Microbial-Mediated Changes in Social Behavior in Mouse Models of Autism Spectrum Disorder. Neuron 2019, 101, 246–259.e246. [Google Scholar] [CrossRef] [PubMed]
- Buffington, S.A.; Di Prisco, G.V.; Auchtung, T.A.; Ajami, N.J.; Petrosino, J.F.; Costa-Mattioli, M. Microbial Reconstitution Reverses Maternal Diet-Induced Social and Synaptic Deficits in Offspring. Cell 2016, 165, 1762–1775. [Google Scholar] [CrossRef]
- Silverman, J.L.; Yang, M.; Lord, C.; Crawley, J.N. Behavioural phenotyping assays for mouse models of autism. Nat. Rev. Neurosci. 2010, 11, 490–502. [Google Scholar] [CrossRef]
- Bartram, A.K.; Lynch, M.D.; Stearns, J.C.; Moreno-Hagelsieb, G.; Neufeld, J.D. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end illumina reads. Appl. Env. Microbiol. 2011, 77, 3846–3852. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Mukherjee, C.; Beall, C.J.; Griffen, A.L.; Leys, E.J. High-resolution ISR amplicon sequencing reveals personalized oral microbiome. Microbiome 2018, 6, 153. [Google Scholar] [CrossRef]
- Mallick, H.; Rahnavard, A.; McIver, L.J.; Ma, S.; Zhang, Y.; Nguyen, L.H.; Tickle, T.L.; Weingart, G.; Ren, B.; Schwager, E.H.; et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 2021, 17, e1009442. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Francella, C.; Green, M.; Caspani, G.; Lai, J.K.Y.; Rilett, K.C.; Foster, J.A. Microbe–Immune–Stress Interactions Impact Behaviour during Postnatal Development. Int. J. Mol. Sci. 2022, 23, 15064. https://doi.org/10.3390/ijms232315064
Francella C, Green M, Caspani G, Lai JKY, Rilett KC, Foster JA. Microbe–Immune–Stress Interactions Impact Behaviour during Postnatal Development. International Journal of Molecular Sciences. 2022; 23(23):15064. https://doi.org/10.3390/ijms232315064
Chicago/Turabian StyleFrancella, Cassandra, Miranda Green, Giorgia Caspani, Jonathan K. Y. Lai, Kelly C. Rilett, and Jane A. Foster. 2022. "Microbe–Immune–Stress Interactions Impact Behaviour during Postnatal Development" International Journal of Molecular Sciences 23, no. 23: 15064. https://doi.org/10.3390/ijms232315064
APA StyleFrancella, C., Green, M., Caspani, G., Lai, J. K. Y., Rilett, K. C., & Foster, J. A. (2022). Microbe–Immune–Stress Interactions Impact Behaviour during Postnatal Development. International Journal of Molecular Sciences, 23(23), 15064. https://doi.org/10.3390/ijms232315064




