Microfluidic Approaches for Affinity-Based Exosome Separation
Abstract
:1. Introduction
2. Separation Mechanisms
2.1. General Considerations for the Selection of an Exosome Separation Process
2.2. Separation Systems Depending on the Applications of Exosomes
2.3. Conventional Separation Methods for Exosomes
2.3.1. Centrifugation-Based Methods
2.3.2. Ultrafiltration Methods
2.3.3. Precipitation Methods
2.3.4. Field Flow Fractionation Methods
2.3.5. Chromatographic Methods
2.3.6. Affinity Binding-Based Separations
3. Microfluidics—How to Build an Affinity Exosome Separation Chip
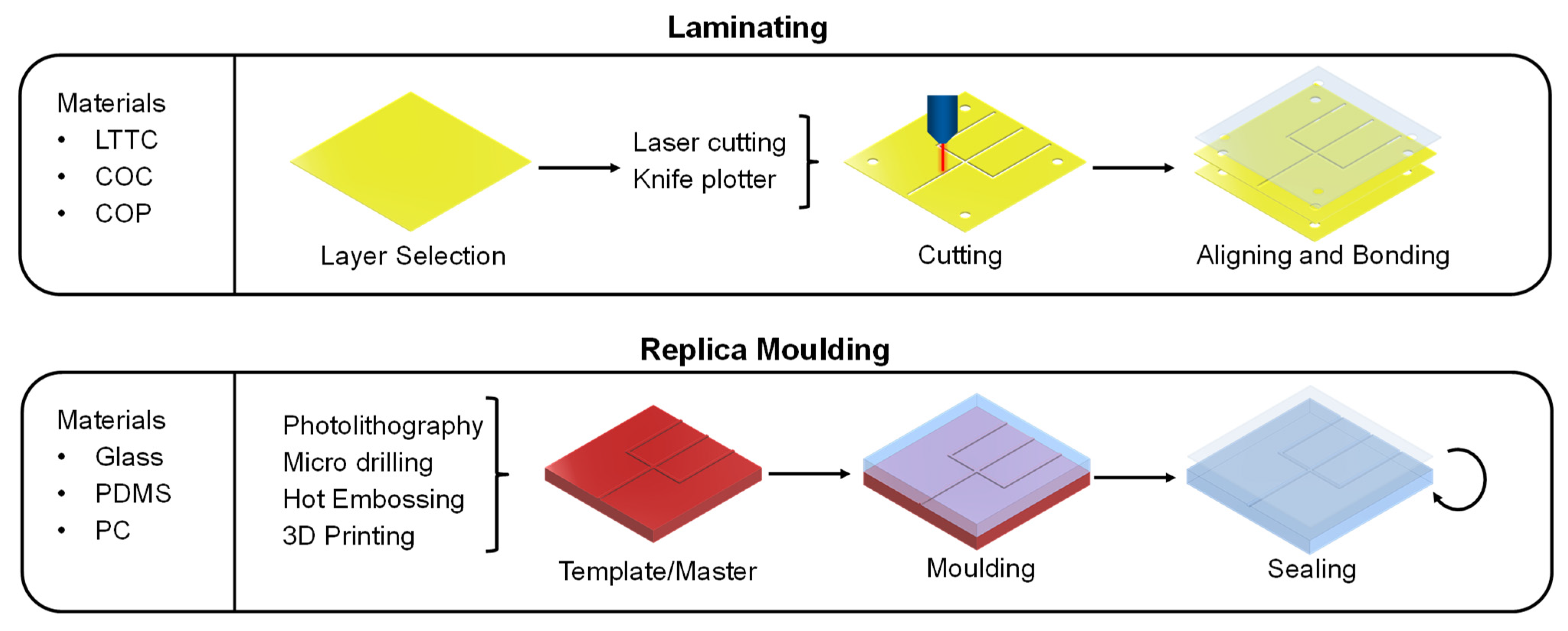
3.1. Manufacturing Methods of Microfluidic Devices
3.2. Structures and Modifications for Microfluidic Devices
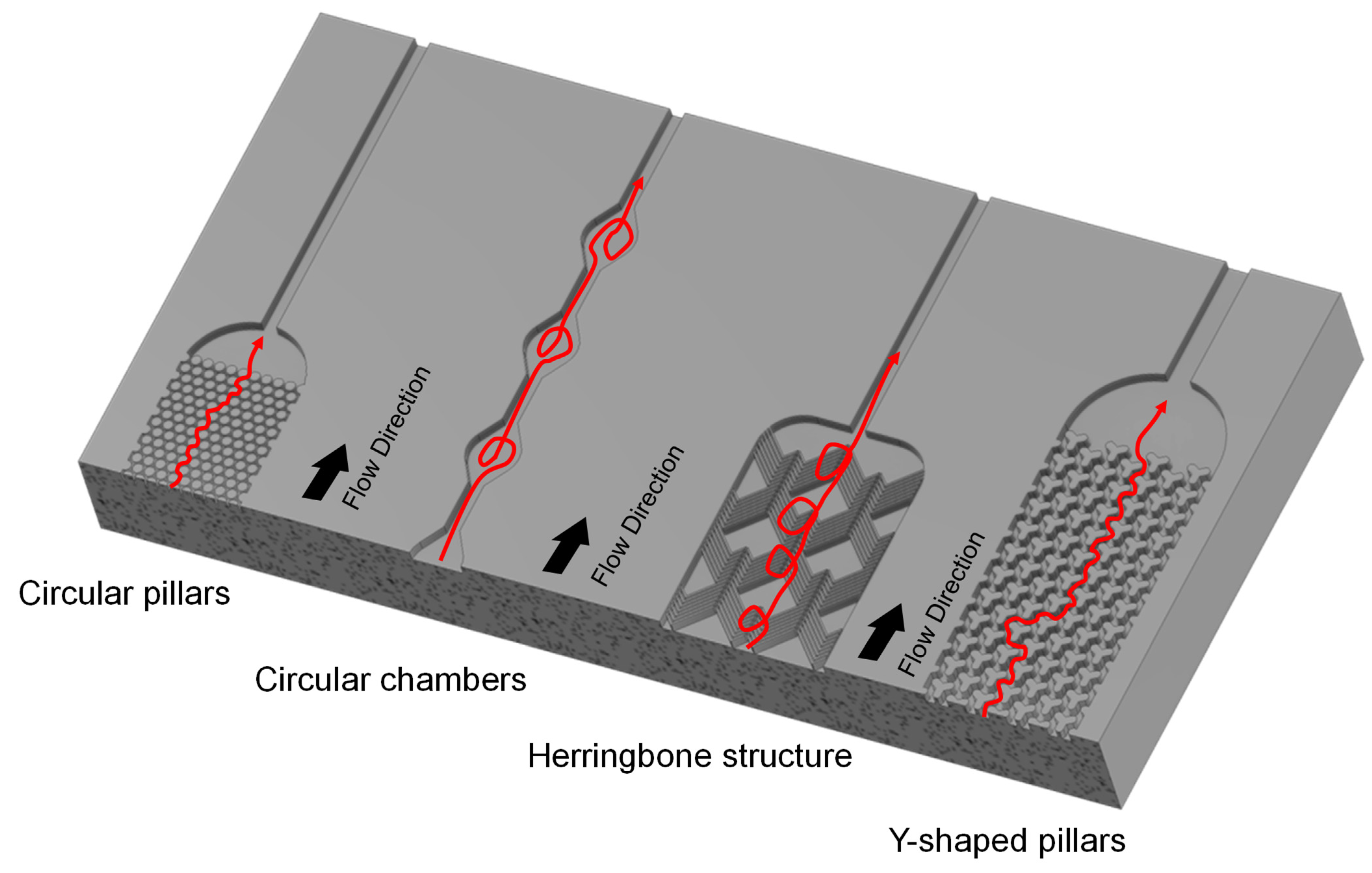
3.2.1. Design of Physical Property-Based Microfluidic Devices
3.2.2. Design of Affinity-Based Microfluidic Devices
3.2.3. Enhancing the Affinity Separation Effect in Microfluidics
3.2.4. Microfluidic Design for Affinity Approaches Utilizing Beads
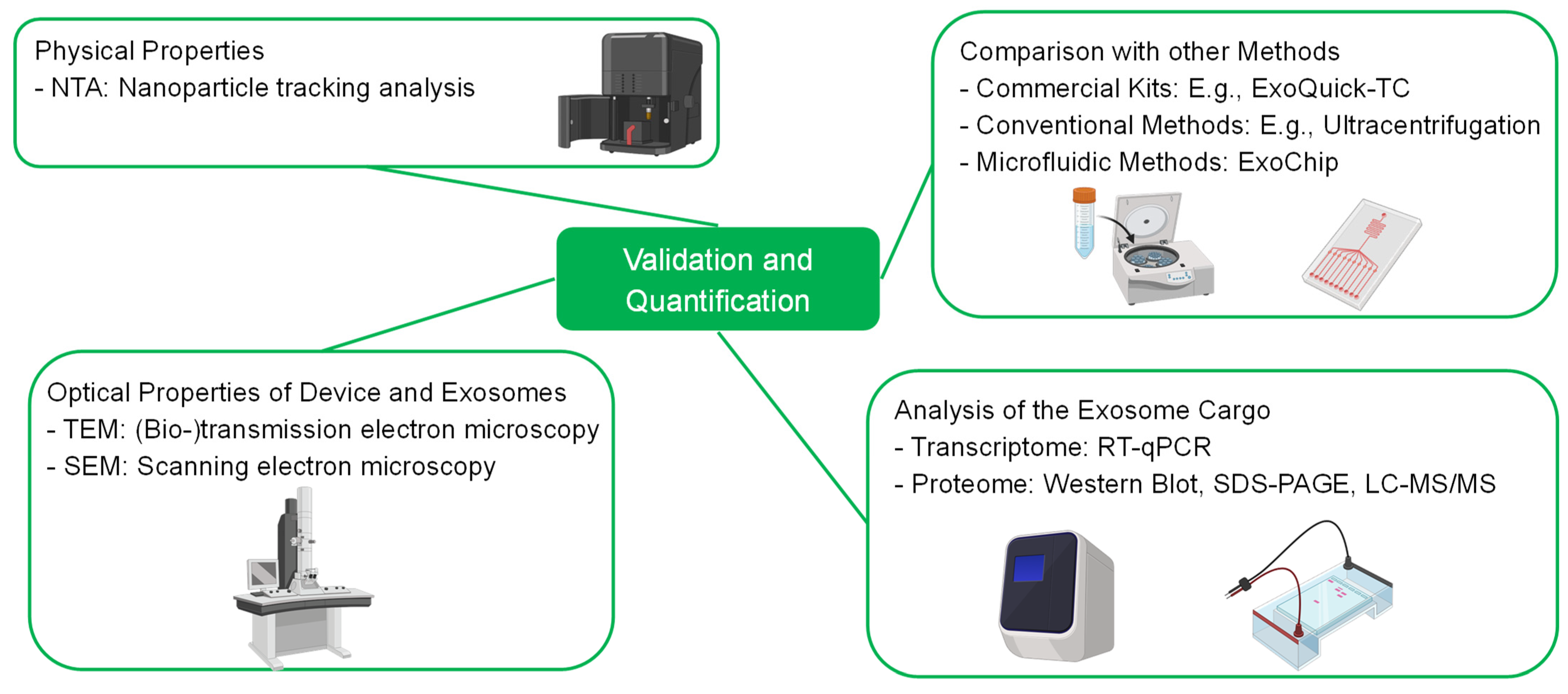
4. Bioassays—Validation of the Separation
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hromada, C.; Mühleder, S.; Grillari, J.; Redl, H.; Holnthoner, W. Endothelial extracellular vesicles-promises and challenges. Front. Physiol. 2017, 8, 275. [Google Scholar] [CrossRef]
- Johnstone, R.M. The Jeanne Manery-Fisher Memorial Lecture 1991. Maturation of reticulocytes: Formation of exosomes as a mechanism for shedding membrane proteins. Biochem. Cell Biol. 1992, 70, 179–190. [Google Scholar] [CrossRef] [PubMed]
- Rashed, M.H.; Bayraktar, E.; Helal, G.K.; Abd-Ellah, M.F.; Amero, P.; Chavez-Reyes, A.; Rodriguez-Aguayo, C. Exosomes: From Garbage Bins to Promising Therapeutic Targets. Int. J. Mol. Sci. 2017, 18, 538. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Yu, X.; Han, W.; Kong, Y.; Bao, W.; Zhang, J.; Zhang, W.; Gu, Y. Ultrasensitive and reversible nanoplatform of urinary exosomes for prostate cancer diagnosis. ACS Sens. 2019, 4, 1433–1441. [Google Scholar] [CrossRef]
- Rasuleva, K.; Elamurugan, S.; Bauer, A.; Khan, M.; Wen, Q.; Li, Z.; Steen, P.; Guo, A.; Xia, W.; Mathew, S.; et al. β-sheet richness of the circulating tumor-derived extracellular vesicles for noninvasive pancreatic cancer screening. ACS Sens. 2021, 6, 4489–4498. [Google Scholar] [CrossRef]
- Ko, J.; Bhagwat, N.; Yee, S.S.; Ortiz, N.; Sahmoud, A.; Black, T.; Aiello, N.M.; McKenzie, L.; O’Hara, M.; Redlinger, C.; et al. Combining machine learning and nanofluidic technology to diagnose pancreatic cancer using exosomes. ACS Nano 2017, 11, 11182–11193. [Google Scholar] [CrossRef]
- Sun, Y.; Huo, C.; Qiao, Z.; Shang, Z.; Uzzaman, A.; Liu, S.; Jiang, X.; Fan, L.-Y.; Ji, L.; Guan, X.; et al. Comparative proteomic analysis of exosomes and microvesicles in human saliva for lung cancer. J. Proteome Res. 2018, 17, 1101–1107. [Google Scholar] [CrossRef]
- Jayaseelan, V.P.; Arumugam, P. Dissecting the theranostic potential of exosomes in autoimmune disorders. Cell. Mol. Immunol. 2019, 16, 935–936. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Gu, Y.; Du, Y.; Tang, X.; Wu, X.; Liu, J. Engineering exosomes endowed with targeted delivery of triptolide for malignant melanoma therapy. ACS Appl. Mater. Interfaces 2021, 13, 42411–42428. [Google Scholar] [CrossRef]
- Kibria, G.; Ramos, E.K.; Wan, Y.; Gius, D.R.; Liu, H. Exosomes as a drug delivery system in cancer therapy: Potential and challenges. Mol. Pharm. 2018, 15, 3625–3633. [Google Scholar] [CrossRef] [PubMed]
- Bray, E.R.; Oropallo, A.R.; Grande, D.A.; Kirsner, R.S.; Badiavas, E.V. Extracellular vesicles as therapeutic tools for the treatment of chronic wounds. Pharmaceutics 2021, 13, 1543. [Google Scholar] [CrossRef]
- Jia, X.; Tang, J.; Yao, C.; Yang, D. Recent progress of extracellular vesicle engineering. ACS Biomater. Sci. Eng. 2021, 7, 4430–4438. [Google Scholar] [CrossRef]
- Yi, K.; Rong, Y.; Huang, L.; Tang, X.; Zhang, Q.; Wang, W.; Wu, J.; Wang, F. Aptamer-exosomes for tumor theranostics. ACS Sens. 2021, 6, 1418–1429. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, Y.; Li, Z.; Zhou, K.; Feng, N. Exosomes as carriers for antitumor therapy. ACS Biomater. Sci. Eng. 2019, 5, 4870–4881. [Google Scholar] [CrossRef] [PubMed]
- Cordonnier, M.; Chanteloup, G.; Isambert, N.; Seigneuric, R.; Fumoleau, P.; Garrido, C.; Gobbo, J. Exosomes in cancer theranostic: Diamonds in the rough. Cell Adhes. Migr. 2017, 11, 151–163. [Google Scholar] [CrossRef]
- Yang, B.; Chen, Y.; Shi, J. Exosome biochemistry and advanced nanotechnology for next-generation theranostic platforms. Adv. Mater. 2019, 31, e1802896. [Google Scholar] [CrossRef] [PubMed]
- Tewari Kumar, P.; Decrop, D.; Safdar, S.; Passaris, I.; Kokalj, T.; Puers, R.; Aertsen, A.; Spasic, D.; Lammertyn, J. Digital microfluidics for single bacteria capture and selective retrieval using optical tweezers. Micromachines 2020, 11, 308. [Google Scholar] [CrossRef]
- Yildizhan, Y.; Vajrala, V.S.; Geeurickx, E.; Declerck, C.; Duskunovic, N.; de Sutter, D.; Noppen, S.; Delport, F.; Schols, D.; Swinnen, J.V.; et al. FO-SPR biosensor calibrated with recombinant extracellular vesicles enables specific and sensitive detection directly in complex matrices. J. Extracell. Vesicles 2021, 10, e12059. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef] [PubMed]
- Royo, F.; Théry, C.; Falcón-Pérez, J.M.; Nieuwland, R.; Witwer, K.W. Methods for separation and characterization of extracellular VESICLES: Results of a worldwide survey performed by the ISEV rigor and standardization subcommittee. Cells 2020, 9, 1955. [Google Scholar] [CrossRef] [PubMed]
- Spada, S.; Galluzzi, L. Extracellular Vesicles; Academic Press: Amsterdam, The Netherlands, 2020; ISBN 9780128206638. [Google Scholar]
- Willms, E.; Cabañas, C.; Mäger, I.; Wood, M.J.A.; Vader, P. Extracellular vesicle heterogeneity: Subpopulations, isolation techniques, and diverse functions in cancer progression. Front. Immunol. 2018, 9, 738. [Google Scholar] [CrossRef]
- Record, M.; Carayon, K.; Poirot, M.; Silvente-Poirot, S. Exosomes as new vesicular lipid transporters involved in cell-cell communication and various pathophysiologies. Biochim. Biophys. Acta 2014, 1841, 108–120. [Google Scholar] [CrossRef]
- Yáñez-Mó, M.; Siljander, P.R.-M.; Andreu, Z.; Zavec, A.B.; Borràs, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef]
- Chen, Y.; Zhu, Q.; Cheng, L.; Wang, Y.; Li, M.; Yang, Q.; Hu, L.; Lou, D.; Li, J.; Dong, X.; et al. Exosome detection via the ultrafast-isolation system: EXODUS. Nat. Methods 2021, 18, 212–218. [Google Scholar] [CrossRef]
- Wang, X.; Shang, H.; Ma, C.; Chen, L. A fluorescence assay for exosome detection based on bivalent cholesterol anchor triggered target conversion and enzyme-free signal amplification. Anal. Chem. 2021, 93, 8493–8500. [Google Scholar] [CrossRef]
- Yu, Z.-L.; Zhao, Y.; Miao, F.; Wu, M.; Xia, H.-F.; Chen, Z.-K.; Liu, H.-M.; Zhao, Y.-F.; Chen, G. In situ membrane biotinylation enables the direct labeling and accurate kinetic analysis of small extracellular vesicles in circulation. Anal. Chem. 2021, 93, 10862–10870. [Google Scholar] [CrossRef]
- Hoshino, A.; Kim, H.S.; Bojmar, L.; Gyan, K.E.; Cioffi, M.; Hernandez, J.; Zambirinis, C.P.; Rodrigues, G.; Molina, H.; Heissel, S.; et al. Extracellular vesicle and particle biomarkers define multiple human cancers. Cell 2020, 182, 1044–1061. [Google Scholar] [CrossRef]
- Ibsen, S.D.; Wright, J.; Lewis, J.M.; Kim, S.; Ko, S.-Y.; Ong, J.; Manouchehri, S.; Vyas, A.; Akers, J.; Chen, C.C.; et al. Rapid isolation and detection of exosomes and associated biomarkers from plasma. ACS Nano 2017, 11, 6641–6651. [Google Scholar] [CrossRef]
- He, M.; Crow, J.; Roth, M.; Zeng, Y.; Godwin, A.K. Integrated immunoisolation and protein analysis of circulating exosomes using microfluidic technology. Lab Chip 2014, 14, 3773–3780. [Google Scholar] [CrossRef]
- Keerthikumar, S.; Chisanga, D.; Ariyaratne, D.; Al Saffar, H.; Anand, S.; Zhao, K.; Samuel, M.; Pathan, M.; Jois, M.; Chilamkurti, N.; et al. ExoCarta: A web-based compendium of exosomal cargo. J. Mol. Biol. 2016, 428, 688–692. [Google Scholar] [CrossRef]
- Nakai, W.; Yoshida, T.; Diez, D.; Miyatake, Y.; Nishibu, T.; Imawaka, N.; Naruse, K.; Sadamura, Y.; Hanayama, R. A novel affinity-based method for the isolation of highly purified extracellular vesicles. Sci. Rep. 2016, 6, 33935. [Google Scholar] [CrossRef] [PubMed]
- Kastelowitz, N.; Yin, H. Exosomes and microvesicles: Identification and targeting by particle size and lipid chemical probes. ChemBioChem 2014, 15, 923–928. [Google Scholar] [CrossRef]
- Kang, Y.-T.; Purcell, E.; Palacios-Rolston, C.; Lo, T.-W.; Ramnath, N.; Jolly, S.; Nagrath, S. Isolation and profiling of circulating tumor-associated exosomes using extracellular vesicular lipid-protein binding affinity based microfluidic device. Small 2019, 15, e1903600. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Liu, G.; Li, J. Recent progress on the isolation and detection methods of exosomes. Chem. Asian J. 2020, 15, 3973–3982. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Malla, R.R.; Pandrangi, S.; Kumari, S.; Gavara, M.M.; Badana, A.K. Exosomal tetraspanins as regulators of cancer progression and metastasis and novel diagnostic markers. Asia Pac. J. Clin. Oncol. 2018, 14, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Rana, S.; Yue, S.; Stadel, D.; Zöller, M. Toward tailored exosomes: The exosomal tetraspanin web contributes to target cell selection. Int. J. Biochem. Cell Biol. 2012, 44, 1574–1584. [Google Scholar] [CrossRef]
- He, F.; Liu, H.; Guo, X.; Yin, B.-C.; Ye, B.-C. Direct exosome quantification via bivalent-cholesterol-labeled DNA anchor for signal amplification. Anal. Chem. 2017, 89, 12968–12975. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Raposo, G.; Théry, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell Dev. Biol. 2014, 30, 255–289. [Google Scholar] [CrossRef] [PubMed]
- Rana, S.; Claas, C.; Kretz, C.C.; Nazarenko, I.; Zoeller, M. Activation-induced internalization differs for the tetraspanins CD9 and Tspan8: Impact on tumor cell motility. Int. J. Biochem. Cell Biol. 2011, 43, 106–119. [Google Scholar] [CrossRef] [PubMed]
- Suwatthanarak, T.; Thiodorus, I.A.; Tanaka, M.; Shimada, T.; Takeshita, D.; Yasui, T.; Baba, Y.; Okochi, M. Microfluidic-based capture and release of cancer-derived exosomes via peptide-nanowire hybrid interface. Lab Chip 2021, 21, 597–607. [Google Scholar] [CrossRef] [PubMed]
- Heijnen, H.F.; Schiel, A.E.; Fijnheer, R.; Geuze, H.J.; Sixma, J.J. Activated platelets release two types of membrane vesicles: Microvesicles by surface shedding and exosomes derived From exocytosis of multivesicular bodies and α-granules. Blood 1999, 94, 3791–3799. [Google Scholar] [CrossRef] [PubMed]
- Wubbolts, R.; Leckie, R.S.; Veenhuizen, P.T.M.; Schwarzmann, G.; Möbius, W.; Hoernschemeyer, J.; Slot, J.-W.; Geuze, H.J.; Stoorvogel, W. Proteomic and biochemical analyses of human B cell-derived exosomes. Potential implications for their function and multivesicular body formation. J. Biol. Chem. 2003, 278, 10963–10972. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Yang, X.; Gao, Z.; Effah, C.Y.; Zhang, X.; Wu, Y.; Qu, L. A holistic review of the state-of-the-art microfluidics for exosome separation: An overview of the current status, existing obstacles, and future outlook. Small 2021, 17, e2007174. [Google Scholar] [CrossRef] [PubMed]
- Soung, Y.H.; Ford, S.; Zhang, V.; Chung, J. Exosomes in cancer diagnostics. Cancers 2017, 9, 8. [Google Scholar] [CrossRef] [PubMed]
- Munson, P.; Shukla, A. Exosomes: Potential in cancer diagnosis and therapy. Medicines 2015, 2, 310–327. [Google Scholar] [CrossRef]
- Harris, J.R.; Marles-Wright, J. (Eds.) Macromolecular Protein Complexes: Structure and Function; Springer: Cham, Switzerland, 2017; ISBN 9783319465012. [Google Scholar]
- Haque, S.; Vaiselbuh, S.R. Exosomes molecular diagnostics: Direct conversion of exosomes into the cDNA for gene amplification by two-step polymerase chain reaction. J. Biol. Methods 2018, 5, e96. [Google Scholar] [CrossRef]
- Cheng, S.; Li, Y.; Yan, H.; Wen, Y.; Zhou, X.; Friedman, L.; Zeng, Y. Advances in microfluidic extracellular vesicle analysis for cancer diagnostics. Lab Chip 2021, 21, 3219–3243. [Google Scholar] [CrossRef]
- Gulei, D.; Irimie, A.I.; Cojocneanu-Petric, R.; Schultze, J.L.; Berindan-Neagoe, I. Exosomes-small players, big sound. Bioconjugate Chem. 2018, 29, 635–648. [Google Scholar] [CrossRef]
- Hassanpour Tamrin, S.; Sanati Nezhad, A.; Sen, A. Label-free Isolation of exosomes using microfluidic technologies. ACS Nano 2021, 15, 17047–17079. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Ma, P.; Kim, D.H.; Liu, B.-F.; Demirci, U. Towards microfluidic-based exosome isolation and detection for tumor therapy. Nano Today 2021, 37, 101066. [Google Scholar] [CrossRef] [PubMed]
- Patel, G.K.; Khan, M.A.; Zubair, H.; Srivastava, S.K.; Khushman, M.; Singh, S.; Singh, A.P. Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci. Rep. 2019, 9, 5335. [Google Scholar] [CrossRef] [PubMed]
- Nicoliche, C.Y.N.; de Oliveira, R.A.G.; Da Silva, G.S.; Ferreira, L.F.; Rodrigues, I.L.; Faria, R.C.; Fazzio, A.; Carrilho, E.; de Pontes, L.G.; Schleder, G.R.; et al. Converging multidimensional sensor and machine learning toward high-throughput and biorecognition element-free multidetermination of extracellular vesicle biomarkers. ACS Sens. 2020, 5, 1864–1871. [Google Scholar] [CrossRef] [PubMed]
- Colao, I.L.; Corteling, R.; Bracewell, D.; Wall, I. Manufacturing exosomes: A promising therapeutic platform. Trends Mol. Med. 2018, 24, 242–256. [Google Scholar] [CrossRef] [PubMed]
- Liga, A.; Vliegenthart, A.D.B.; Oosthuyzen, W.; Dear, J.W.; Kersaudy-Kerhoas, M. Exosome isolation: A microfluidic road-map. Lab Chip 2015, 15, 2388–2394. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Pan, B.; Zeng, F.; He, B.; Gao, Y.; Liu, X.; Song, Y. Magnetic colloid antibodies accelerate small extracellular vesicles isolation for point-of-care diagnostics. Nano Lett. 2021, 21, 2001–2009. [Google Scholar] [CrossRef] [PubMed]
- Yager, P.; Domingo, G.J.; Gerdes, J. Point-of-care diagnostics for global health. Annu. Rev. Biomed. Eng. 2008, 10, 107–144. [Google Scholar] [CrossRef] [PubMed]
- Shirejini, S.Z.; Inci, F. The Yin and Yang of exosome isolation methods: Conventional practice, microfluidics, and commercial kits. Biotechnol. Adv. 2022, 54, 107814. [Google Scholar] [CrossRef]
- Batrakova, E.V.; Kim, M.S. Using exosomes, naturally-equipped nanocarriers, for drug delivery. J. Control. Release 2015, 219, 396–405. [Google Scholar] [CrossRef]
- Liang, Y.; Duan, L.; Lu, J.; Xia, J. Engineering exosomes for targeted drug delivery. Theranostics 2021, 11, 3183–3195. [Google Scholar] [CrossRef] [PubMed]
- Greening, D.W.; Xu, R.; Ji, H.; Tauro, B.J.; Simpson, R.J. A protocol for exosome isolation and characterization: Evaluation of ultracentrifugation, density-gradient separation, and immunoaffinity capture methods. Methods Mol. Biol. 2015, 1295, 179–209. [Google Scholar] [CrossRef] [PubMed]
- Livshits, M.A.; Livshts, M.A.; Khomyakova, E.; Evtushenko, E.G.; Lazarev, V.N.; Kulemin, N.A.; Semina, S.E.; Generozov, E.V.; Govorun, V.M. Isolation of exosomes by differential centrifugation: Theoretical analysis of a commonly used protocol. Sci. Rep. 2015, 5, 17319. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Wong, D.K.; Hong, K.Y.; Raffai, R.L. Cushioned-density gradient ultracentrifugation (C-DGUC): A refined and high performance method for the isolation, characterization, and use of exosomes. Methods Mol. Biol. 2018, 1740, 69–83. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.-Y.; Sung, C.W.-H.; Chen, C.; Cheng, C.-M.; Lin, D.P.-C.; Huang, C.-T.; Hsu, M.-Y. Advances in exosomes technology. Clin. Chim. Acta 2019, 493, 14–19. [Google Scholar] [CrossRef] [PubMed]
- Kwon, D.; Vigneswaran, S.; Fane, A.; Aim, R. Experimental determination of critical flux in cross-flow microfiltration. Sep. Purif. Technol. 2000, 19, 169–181. [Google Scholar] [CrossRef]
- Ludwig, A.-K.; de Miroschedji, K.; Doeppner, T.R.; Börger, V.; Ruesing, J.; Rebmann, V.; Durst, S.; Jansen, S.; Bremer, M.; Behrmann, E.; et al. Precipitation with polyethylene glycol followed by washing and pelleting by ultracentrifugation enriches extracellular vesicles from tissue culture supernatants in small and large scales. J. Extracell. Vesicles 2018, 7, 1528109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef]
- Carta, G.; Jungbauer, A. Protein Chromatography; Wiley: New York, NY, USA, 2020; ISBN 9783527346660. [Google Scholar]
- Li, Y.; Yang, K.; Yuan, H.; Zhang, W.; Sui, Z.; Wang, N.; Lin, H.; Zhang, L.; Zhang, Y. Surface nanosieving polyether sulfone particles with graphene oxide encapsulation for the negative isolation toward extracellular vesicles. Anal. Chem. 2021, 93, 16835–16844. [Google Scholar] [CrossRef] [PubMed]
- Bellotti, C.; Lang, K.; Kuplennik, N.; Sosnik, A.; Steinfeld, R. High-grade extracellular vesicles preparation by combined size-exclusion and affinity chromatography. Sci. Rep. 2021, 11, 10550. [Google Scholar] [CrossRef]
- Wang, T.; Turko, I.V. Proteomic toolbox to standardize the separation of extracellular vesicles and lipoprotein particles. J. Proteome Res. 2018, 17, 3104–3113. [Google Scholar] [CrossRef]
- Wehmeyer, K.R.; White, R.J.; Kissinger, P.T.; Heineman, W.R. Electrochemical affinity assays/sensors: Brief history and current status. Annu. Rev. Anal. Chem. 2021, 14, 109–131. [Google Scholar] [CrossRef]
- Azimzadeh, A.; van Regenmortel, M.H. Antibody affinity measurements. J. Mol. Recognit. 1990, 3, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Sackmann, E.K.; Wypisniak, K.; Hornsby, M.; Datwani, S.S.; Herr, A.E. Determination of equilibrium dissociation constants for recombinant antibodies by high-throughput affinity electrophoresis. Sci. Rep. 2016, 6, 39774. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Yang, L.; He, L. Overview of the detection methods for equilibrium dissociation constant KD of drug-receptor interaction. J. Pharm. Anal. 2018, 8, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Lin, L.; Zhao, K.; Song, Y.; Huang, M.; Zhu, Z.; Zhou, L.; Yang, C. Auto-affitech: An automated ligand binding affinity evaluation platform using digital microfluidics with a bidirectional magnetic separation method. Lab Chip 2020, 20, 1577–1585. [Google Scholar] [CrossRef]
- Boriachek, K.; Masud, M.K.; Palma, C.; Phan, H.-P.; Yamauchi, Y.; Hossain, M.S.A.; Nguyen, N.-T.; Salomon, C.; Shiddiky, M.J.A. Avoiding pre-isolation step in exosome analysis: Direct isolation and sensitive detection of exosomes using gold-loaded nanoporous ferric oxide nanozymes. Anal. Chem. 2019, 91, 3827–3834. [Google Scholar] [CrossRef]
- Wu, X.; Li, L.; Iliuk, A.; Tao, W.A. Highly efficient phosphoproteome capture and analysis from urinary extracellular vesicles. J. Proteome Res. 2018, 17, 3308–3316. [Google Scholar] [CrossRef]
- Akbarinejad, A.; Hisey, C.L.; Brewster, D.; Ashraf, J.; Chang, V.; Sabet, S.; Nursalim, Y.; Lucarelli, V.; Blenkiron, C.; Chamley, L.; et al. Novel electrochemically switchable, flexible, microporous cloth that selectively captures, releases, and concentrates intact extracellular vesicles. ACS Appl. Mater. Interfaces 2020, 12, 39005–39013. [Google Scholar] [CrossRef]
- Convery, N.; Gadegaard, N. 30 years of microfluidics. Micro Nano Eng. 2019, 2, 76–91. [Google Scholar] [CrossRef]
- Erickson, D.; Li, D. Integrated microfluidic devices. Anal. Chim. Acta 2004, 507, 11–26. [Google Scholar] [CrossRef]
- Lin, B.; Lei, Y.; Wang, J.; Zhu, L.; Wu, Y.; Zhang, H.; Wu, L.; Zhang, P.; Yang, C. Microfluidic-based exosome analysis for liquid biopsy. Small Methods 2021, 5, e2001131. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.-M.; Lv, S.; Zhang, W.; Cui, Y. Microfluidic point-of-care (POC) devices in early diagnosis: A review of opportunities and challenges. Sensors 2022, 22, 1620. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wei, X.; Zeng, R.; Xu, F.; Li, X. Stem cell culture and differentiation in microfluidic devices toward organ-on-a-chip. Future Sci. OA 2017, 3, FSO187. [Google Scholar] [CrossRef] [PubMed]
- Scott, S.M.; Ali, Z. Fabrication methods for microfluidic devices: An overview. Micromachines 2021, 12, 319. [Google Scholar] [CrossRef]
- Niculescu, A.-G.; Chircov, C.; Bîrcă, A.C.; Grumezescu, A.M. Fabrication and applications of microfluidic devices: A review. Int. J. Mol. Sci. 2021, 22, 2011. [Google Scholar] [CrossRef] [PubMed]
- Tsao, C.-W. Polymer microfluidics: Simple, low-cost fabrication process bridging academic lab research to commercialized production. Micromachines 2016, 7, 225. [Google Scholar] [CrossRef]
- Gray, B.; Jaeggi, D.; Mourlas, N.; van Drieënhuizen, B.; Williams, K.; Maluf, N.; Kovacs, G. Novel interconnection technologies for integrated microfluidic systems. Sens. Actuators Phys. 1999, 77, 57–65. [Google Scholar] [CrossRef]
- Thompson, B.L.; Ouyang, Y.; Duarte, G.R.M.; Carrilho, E.; Krauss, S.T.; Landers, J.P. Inexpensive, rapid prototyping of microfluidic devices using overhead transparencies and a laser print, cut and laminate fabrication method. Nat. Protoc. 2015, 10, 875–886. [Google Scholar] [CrossRef]
- Walsh, D.I.; Kong, D.S.; Murthy, S.K.; Carr, P.A. Enabling microfluidics: From clean rooms to makerspaces. Trends Biotechnol. 2017, 35, 383–392. [Google Scholar] [CrossRef]
- Ma, X.; Li, R.; Jin, Z.; Fan, Y.; Zhou, X.; Zhang, Y. Injection molding and characterization of PMMA-based microfluidic devices. Microsyst. Technol. 2020, 26, 1317–1324. [Google Scholar] [CrossRef]
- Sticker, D.; Rothbauer, M.; Lechner, S.; Hehenberger, M.-T.; Ertl, P. Multi-layered, membrane-integrated microfluidics based on replica molding of a thiol-ene epoxy thermoset for organ-on-a-chip applications. Lab Chip 2015, 15, 4542–4554. [Google Scholar] [CrossRef]
- Guo, L.J. Nanoimprint lithography: Methods and material requirements. Adv. Mater. 2007, 19, 495–513. [Google Scholar] [CrossRef]
- Tucher, N.; Höhn, O.; Hauser, H.; Müller, C.; Bläsi, B. Characterizing the degradation of PDMS stamps in nanoimprint lithography. Microelectron. Eng. 2017, 180, 40–44. [Google Scholar] [CrossRef]
- Gale, B.; Jafek, A.; Lambert, C.; Goenner, B.; Moghimifam, H.; Nze, U.; Kamarapu, S. A review of current methods in microfluidic device fabrication and future commercialization prospects. Inventions 2018, 3, 60. [Google Scholar] [CrossRef]
- Sanchez Noriega, J.L.; Chartrand, N.A.; Valdoz, J.C.; Cribbs, C.G.; Jacobs, D.A.; Poulson, D.; Viglione, M.S.; Woolley, A.T.; van Ry, P.M.; Christensen, K.A.; et al. Spatially and optically tailored 3D printing for highly miniaturized and integrated microfluidics. Nat. Commun. 2021, 12, 5509. [Google Scholar] [CrossRef]
- Mehta, V.; Rath, S.N. 3D printed microfluidic devices: A review focused on four fundamental manufacturing approaches and implications on the field of healthcare. Bio. Des. Manuf. 2021, 4, 311–343. [Google Scholar] [CrossRef]
- Amin, R.; Knowlton, S.; Hart, A.; Yenilmez, B.; Ghaderinezhad, F.; Katebifar, S.; Messina, M.; Khademhosseini, A.; Tasoglu, S. 3D-printed microfluidic devices. Biofabrication 2016, 8, 22001. [Google Scholar] [CrossRef]
- Tasoglu, S.; Folch, A. 3D Printed Microfluidic Devices; MDPI: Basel, Switzerland, 2018; ISBN 978-3-03897-468-0. [Google Scholar]
- Lee, J.; Kim, H.; Heo, Y.; Yoo, Y.K.; Han, S.I.; Kim, C.; Hur, D.; Kim, H.; Kang, J.Y.; Lee, J.H. Enhanced paper-based ELISA for simultaneous EVs/exosome isolation and detection using streptavidin agarose-based immobilization. Analyst 2019, 145, 157–164. [Google Scholar] [CrossRef]
- Liu, H.; Xiang, Y.; Lu, Y.; Crooks, R.M. Aptamer-based origami paper analytical device for electrochemical detection of adenosine. Angew. Chem. 2012, 124, 7031–7034. [Google Scholar] [CrossRef]
- Ozer, T.; Henry, C.S. Paper-based analytical devices for virus detection: Recent strategies for current and future pandemics. Trends Analyt. Chem. 2021, 144, 116424. [Google Scholar] [CrossRef]
- Davim, J.; Santana, H.S.; Da Lameu Silva, J., Jr.; Taranto, O.P. Process. Analysis, Design, and Intensification in Microfluidics and Chemical Engineering; IGI Global: Hershey, PA, USA, 2019; ISBN 9781522571384. [Google Scholar]
- Pamme, N. Continuous flow separations in microfluidic devices. Lab Chip 2007, 7, 1644–1659. [Google Scholar] [CrossRef]
- Rogers, C.I.; Qaderi, K.; Woolley, A.T.; Nordin, G.P. 3D printed microfluidic devices with integrated valves. Biomicrofluidics 2015, 9, 16501. [Google Scholar] [CrossRef] [PubMed]
- Bao, B.; Wang, Z.; Thushara, D.; Liyanage, A.; Gunawardena, S.; Yang, Z.; Zhao, S. Recent advances in microfluidics-based chromatography—A mini review. Separations 2021, 8, 3. [Google Scholar] [CrossRef]
- Wang, Z.; Wu, H.; Fine, D.; Schmulen, J.; Hu, Y.; Godin, B.; Zhang, J.X.J.; Liu, X. Ciliated micropillars for the microfluidic-based isolation of nanoscale lipid vesicles. Lab Chip 2013, 13, 2879–2882. [Google Scholar] [CrossRef]
- Liu, F.; Vermesh, O.; Mani, V.; Ge, T.J.; Madsen, S.J.; Sabour, A.; Hsu, E.-C.; Gowrishankar, G.; Kanada, M.; Jokerst, J.V.; et al. The exosome total isolation chip. ACS Nano 2017, 11, 10712–10723. [Google Scholar] [CrossRef] [PubMed]
- Hattori, Y.; Shimada, T.; Yasui, T.; Kaji, N.; Baba, Y. Micro- and nanopillar chips for continuous separation of extracellular vesicles. Anal. Chem. 2019, 91, 6514–6521. [Google Scholar] [CrossRef]
- Liu, C.; Guo, J.; Tian, F.; Yang, N.; Yan, F.; Ding, Y.; Wei, J.; Hu, G.; Nie, G.; Sun, J. Field-free isolation of exosomes from extracellular vesicles by microfluidic viscoelastic flows. ACS Nano 2017, 11, 6968–6976. [Google Scholar] [CrossRef]
- Wu, M.; Ouyang, Y.; Wang, Z.; Zhang, R.; Huang, P.-H.; Chen, C.; Li, H.; Li, P.; Quinn, D.; Dao, M.; et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc. Natl. Acad. Sci. USA 2017, 114, 10584–10589. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Brownlow, J.W.; Walker, D.L.; Lu, J. Effect of surfactant-assisted wettability alteration on immiscible displacement: A microfluidic study. Water Resour. Res. 2021, 57, e2020WR029522. [Google Scholar] [CrossRef]
- Salva, M.L.; Rocca, M.; Niemeyer, C.M.; Delamarche, E. Methods for immobilizing receptors in microfluidic devices: A review. Micro Nano Eng. 2021, 11, 100085. [Google Scholar] [CrossRef]
- Zhou, J.; Ellis, A.V.; Voelcker, N.H. Recent developments in PDMS surface modification for microfluidic devices. Electrophoresis 2010, 31, 2–16. [Google Scholar] [CrossRef] [PubMed]
- Shakeri, A.; Imani, S.M.; Chen, E.; Yousefi, H.; Shabbir, R.; Didar, T.F. Plasma-induced covalent immobilization and patterning of bioactive species in microfluidic devices. Lab Chip 2019, 19, 3104–3115. [Google Scholar] [CrossRef] [PubMed]
- Hisey, C.L.; Dorayappan, K.D.P.; Cohn, D.E.; Selvendiran, K.; Hansford, D.J. Microfluidic affinity separation chip for selective capture and release of label-free ovarian cancer exosomes. Lab Chip 2018, 18, 3144–3153. [Google Scholar] [CrossRef]
- Nair, S.V.; Witek, M.A.; Jackson, J.M.; Lindell, M.A.M.; Hunsucker, S.A.; Sapp, T.; Perry, C.E.; Hupert, M.L.; Bae-Jump, V.; Gehrig, P.A.; et al. Enzymatic cleavage of uracil-containing single-stranded DNA linkers for the efficient release of affinity-selected circulating tumor cells. Chem. Commun. 2015, 51, 3266–3269. [Google Scholar] [CrossRef]
- Wijerathne, H.; Witek, M.A.; Jackson, J.M.; Brown, V.; Hupert, M.L.; Herrera, K.; Kramer, C.; Davidow, A.E.; Li, Y.; Baird, A.E.; et al. Affinity enrichment of extracellular vesicles from plasma reveals mRNA changes associated with acute ischemic stroke. Commun. Biol. 2020, 3, 613. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Li, W.; Pappas, D. Recent advances in microfluidic cell separations. Analyst 2013, 138, 4714–4721. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lyons, V.; Pappas, D. Fundamentals of affinity cell separations. Electrophoresis 2018, 39, 732–741. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhou, X.; Zeng, Y. Multiplexed immunophenotyping of circulating exosomes on nano-engineered ExoProfile chip towards early diagnosis of cancer. Chem. Sci. 2019, 10, 5495–5504. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Kaslan, M.; Lee, S.H.; Yao, J.; Gao, Z. Progress in exosome isolation techniques. Theranostics 2017, 7, 789–804. [Google Scholar] [CrossRef]
- Zhang, M.; Jin, K.; Gao, L.; Zhang, Z.; Li, F.; Zhou, F.; Zhang, L. Methods and technologies for exosome isolation and characterization. Small Methods 2018, 2, 1800021. [Google Scholar] [CrossRef]
- Oksvold, M.P.; Neurauter, A.; Pedersen, K.W. Magnetic bead-based isolation of exosomes. Methods Mol. Biol. 2015, 1218, 465–481. [Google Scholar] [CrossRef]
- Tayebi, M.; Zhou, Y.; Tripathi, P.; Chandramohanadas, R.; Ai, Y. Exosome purification and analysis using a facile microfluidic hydrodynamic trapping device. Anal. Chem. 2020, 92, 10733–10742. [Google Scholar] [CrossRef]
- Xu, H.; Liao, C.; Zuo, P.; Liu, Z.; Ye, B.-C. Magnetic-Based microfluidic device for on-chip isolation and detection of tumor-derived exosomes. Anal. Chem. 2018, 90, 13451–13458. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Li, Q.; Wang, C.; Pang, Y.; Sun, Z.; Xiao, R. In situ exosomal MicroRNA determination by target-triggered SERS and Fe3O4@TiO2-based exosome accumulation. ACS Sens. 2021, 6, 852–862. [Google Scholar] [CrossRef]
- Wang, J.; Wuethrich, A.; Lobb, R.J.; Antaw, F.; Sina, A.A.I.; Lane, R.E.; Zhou, Q.; Zieschank, C.; Bell, C.; Bonazzi, V.F.; et al. Characterizing the heterogeneity of small extracellular vesicle populations in multiple cancer types via an ultrasensitive chip. ACS Sens. 2021, 6, 3182–3194. [Google Scholar] [CrossRef] [PubMed]
- Cavallaro, S.; Horak, J.; Hååg, P.; Gupta, D.; Stiller, C.; Sahu, S.S.; Görgens, A.; Gatty, H.K.; Viktorsson, K.; El Andaloussi, S.; et al. Label-free surface protein profiling of extracellular vesicles by an electrokinetic sensor. ACS Sens. 2019, 4, 1399–1408. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Xu, X.; Li, B.; Situ, B.; Pan, W.; Hu, Y.; An, T.; Yao, S.; Zheng, L. Single-exosome-counting immunoassays for cancer diagnostics. Nano Lett. 2018, 18, 4226–4232. [Google Scholar] [CrossRef] [PubMed]
- Chin, L.K.; Son, T.; Hong, J.-S.; Liu, A.-Q.; Skog, J.; Castro, C.M.; Weissleder, R.; Lee, H.; Im, H. Plasmonic sensors for extracellular vesicle analysis: From scientific development to translational research. ACS Nano 2020, 14, 14528–14548. [Google Scholar] [CrossRef]
- Jeong, S.; Park, J.; Pathania, D.; Castro, C.M.; Weissleder, R.; Lee, H. Integrated magneto-electrochemical sensor for exosome analysis. ACS Nano 2016, 10, 1802–1809. [Google Scholar] [CrossRef]
- Saha, N.; Brunetti, G.; Kumar, A.; Armenise, M.N.; Ciminelli, C. Highly sensitive refractive index sensor based on polymer bragg grating: A case study on extracellular vesicles detection. Biosensors 2022, 12, 415. [Google Scholar] [CrossRef]
- Im, H.; Lee, K.; Weissleder, R.; Lee, H.; Castro, C.M. Novel nanosensing technologies for exosome detection and profiling. Lab Chip 2017, 17, 2892–2898. [Google Scholar] [CrossRef]


| Cancer Cell Type | Biofluid | Biomarkers |
|---|---|---|
| Bladder | Whole blood | CD9, CD81 |
| Bladder | Urine | CD63 |
| Bladder and liver | Plasma | CD63 |
| Liver | Serum | CD63 |
| Lung | Plasma | CD9, CD63, CD81, CD82, CA125, EpCAM, IGF-1R |
| Prostate | Serum | EpCAM |
| Breast | Serum | PSA |
| Ovarian and breast | Plasma | CD24, EpCAM, FRα, HER2, MMP14 |
| Ovarian | Plasma | CD9, CD63, CD81, EpCAM |
| Ovarian | Ascites | CD24, EpCAM |
| Glioblastoma multiforme | Blood | CD63, EGFR, EGFRvIII, EPphA2, IDH1, PDGFR, PDPN, R132H |
| Melanoma | Whole blood | CD9 |
| Colorectal and gastric | Ascites | EV glycans |
| Immune Cell Type | Biomarkers |
|---|---|
| Dendritic cells | MHC complexes, CD89, CD86, ICAM-1 |
| B cells | D19, CD37, MHC II |
| T cells: Treg | D25, CD73, CTKA-4 |
| T cells: CD4+ | CD4, HLA class 1, TfR, TCR, integrin β2, Fas ligand |
| T cells: CD8+ | CR Fas ligand CD8 |
| Natural killer cells | D56, perforin, granzyme B, Fas ligand |
| Macrophages | Bacterial antigens, cofilin-1, GNB1, actin, cyclophilin A |
| Mast cells | MHC II, c-Kit, LFA1, ICAM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Theel, E.K.; Schwaminger, S.P. Microfluidic Approaches for Affinity-Based Exosome Separation. Int. J. Mol. Sci. 2022, 23, 9004. https://doi.org/10.3390/ijms23169004
Theel EK, Schwaminger SP. Microfluidic Approaches for Affinity-Based Exosome Separation. International Journal of Molecular Sciences. 2022; 23(16):9004. https://doi.org/10.3390/ijms23169004
Chicago/Turabian StyleTheel, Eike K., and Sebastian P. Schwaminger. 2022. "Microfluidic Approaches for Affinity-Based Exosome Separation" International Journal of Molecular Sciences 23, no. 16: 9004. https://doi.org/10.3390/ijms23169004
APA StyleTheel, E. K., & Schwaminger, S. P. (2022). Microfluidic Approaches for Affinity-Based Exosome Separation. International Journal of Molecular Sciences, 23(16), 9004. https://doi.org/10.3390/ijms23169004







