The Patterning and Proportion of Charged Residues in the Arginine-Rich Mixed-Charge Domain Determine the Membrane-Less Organelle Targeted by the Protein
Abstract
1. Introduction
2. Results
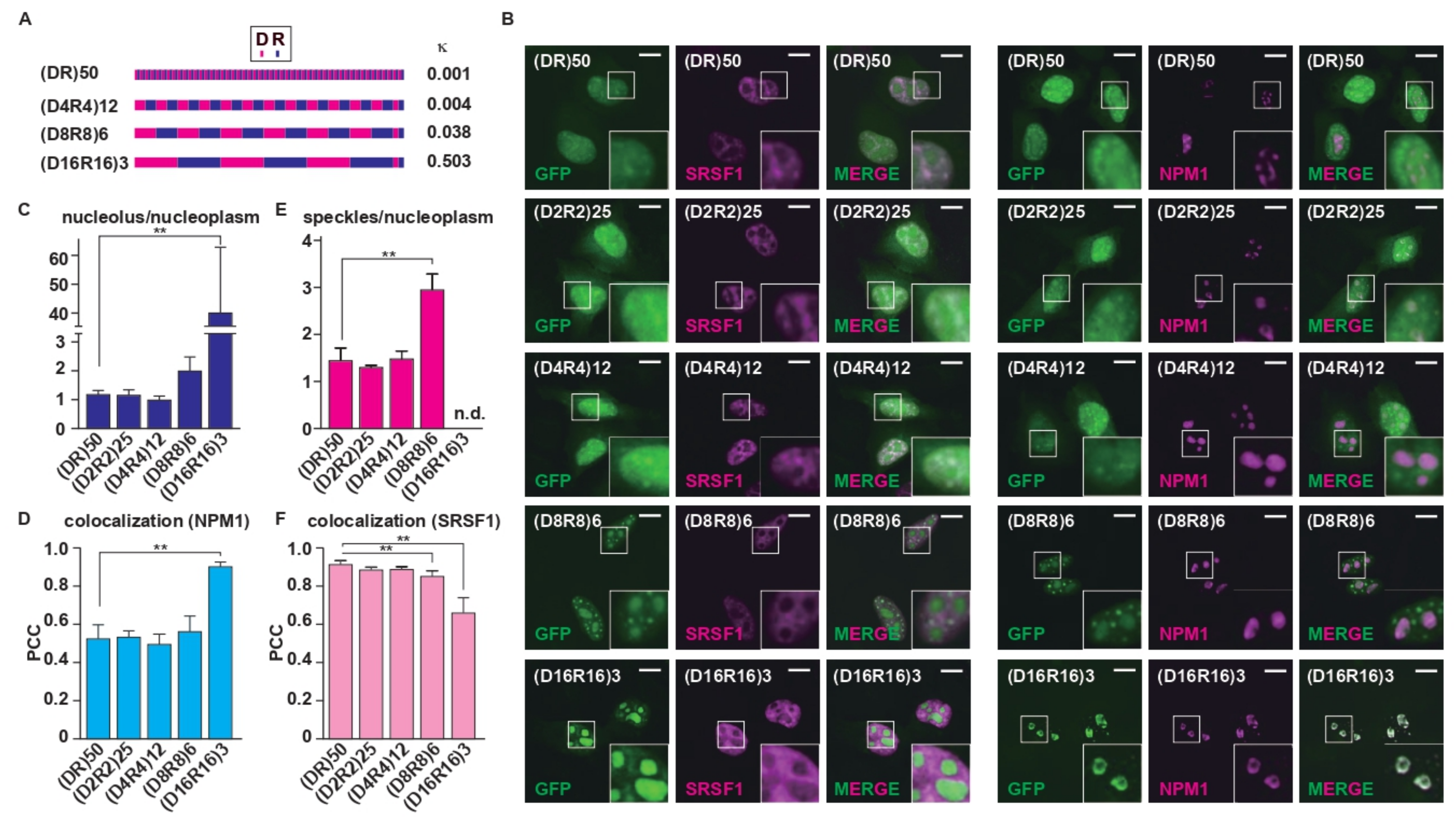
2.1. Charge Patterning in the R-MCDs Influences Their Distribution to MLOs
2.2. Charge Patterning Affects the Interactome of R-MCDs
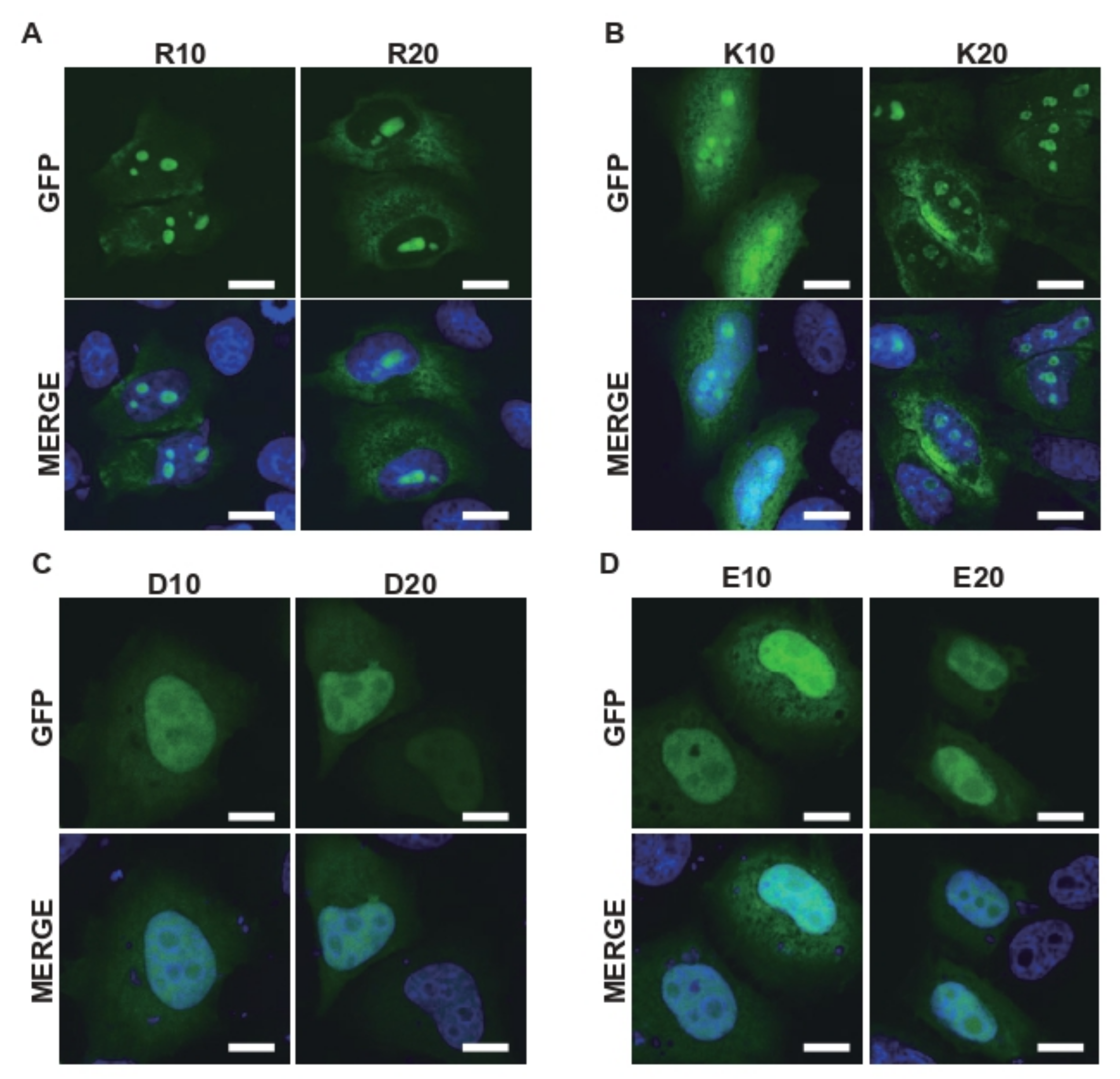
2.3. A Net Positive Charge in R-MCDs Enhances Their Incorporation into MLOs
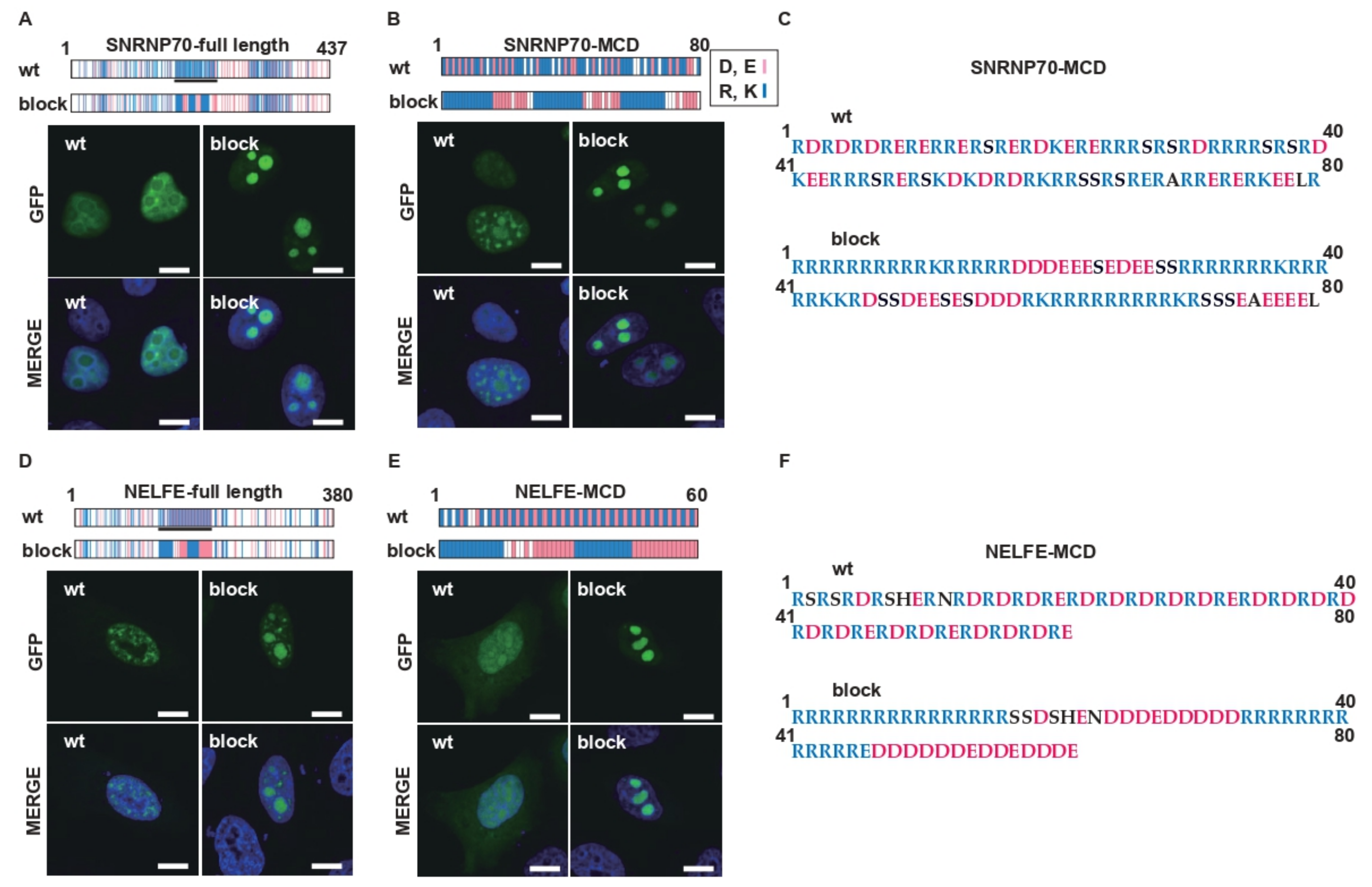
2.4. Charge Patterning in Natural R-MCDs Determines Protein Distribution to Specific MLOs
2.5. Charge Patterning Determines the Phase-Separating Properties of R-MCD Peptides
3. Discussion
4. Materials and Methods
4.1. Recombinant Plasmids Construction
4.2. Transfection of HeLa Cells and Image Acquisition by Confocal Microscopy
4.3. Peptide Synthesis and Purification of Recombinant mScarletI-NPM1
4.4. Phase Separation of Peptides
4.5. Proximity Labeling and Quantitative LC–MS
4.6. Colocalization Analysis
4.7. Bioinformatic Analysis of Biochemical Properties of Peptides and Proteins
4.8. Statistic Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Su, Q.; Mehta, S.; Zhang, J. Liquid-liquid phase separation: Orchestrating cell signaling through time and space. Mol. Cell 2021, 81, 4137–4146. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 2015, 16, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Alberti, S.; Gladfelter, A.; Mittag, T. Considerations and Challenges in Studying Liquid-Liquid Phase Separation and Biomolecular Condensates. Cell 2019, 176, 419–434. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.-H.; Zhang, P.; Kim, H.J.; Mitrea, D.M.; Sarkar, M.; Freibaum, B.D.; Cika, J.; Coughlin, M.; Messing, J.; Molliex, A.; et al. C9orf72 Dipeptide Repeats Impair the Assembly, Dynamics, and Function of Membrane-Less Organelles. Cell 2016, 167, 774–788.e17. [Google Scholar] [CrossRef]
- Babinchak, W.M.; Surewicz, W.K. Liquid-Liquid Phase Separation and Its Mechanistic Role in Pathological Protein Aggregation. J. Mol. Biol. 2020, 432, 1910–1925. [Google Scholar] [CrossRef]
- Holehouse, A.S.; Das, R.K.; Ahad, J.N.; Richardson, M.O.; Pappu, R.V. CIDER: Resources to Analyze Sequence-Ensemble Relationships of Intrinsically Disordered Proteins. Biophys. J. 2017, 112, 16–21. [Google Scholar] [CrossRef]
- Martin, R.M.; Ter-Avetisyan, G.; Herce, H.D.; Ludwig, A.K.; Lattig-Tunnemann, G.; Cardoso, M.C. Principles of protein targeting to the nucleolus. Nucleus 2015, 6, 314–325. [Google Scholar] [CrossRef]
- Musinova, Y.R.; Kananykhina, E.Y.; Potashnikova, D.M.; Lisitsyna, O.M.; Sheval, E.V. A charge-dependent mechanism is responsible for the dynamic accumulation of proteins inside nucleoli. Biochim. Biophys. Acta 2015, 1853, 101–110. [Google Scholar] [CrossRef]
- Emmott, E.; Hiscox, J.A. Nucleolar targeting: The hub of the matter. EMBO Rep. 2009, 10, 231–238. [Google Scholar] [CrossRef]
- Stenstrom, L.; Mahdessian, D.; Gnann, C.; Cesnik, A.J.; Ouyang, W.; Leonetti, M.D.; Uhlen, M.; Cuylen-Haering, S.; Thul, P.J.; Lundberg, E. Mapping the nucleolar proteome reveals a spatiotemporal organization related to intrinsic protein disorder. Mol. Syst. Biol. 2020, 16, e9469. [Google Scholar] [CrossRef]
- Galganski, L.; Urbanek, M.O.; Krzyzosiak, W.J. Nuclear speckles: Molecular organization, biological function and role in disease. Nucleic Acids Res. 2017, 45, 10350–10368. [Google Scholar] [CrossRef]
- Ilik, I.A.; Malszycki, M.; Lubke, A.K.; Schade, C.; Meierhofer, D.; Aktas, T. SON and SRRM2 are essential for nuclear speckle formation. Elife 2020, 9, e60579. [Google Scholar] [CrossRef]
- Greig, J.A.; Nguyen, T.A.; Lee, M.; Holehouse, A.S.; Posey, A.E.; Pappu, R.V.; Jedd, G. Arginine-Enriched Mixed-Charge Domains Provide Cohesion for Nuclear Speckle Condensation. Mol. Cell 2020, 77, 1237–1250.e4. [Google Scholar] [CrossRef]
- Das, R.K.; Pappu, R.V. Conformations of intrinsically disordered proteins are influenced by linear sequence distributions of oppositely charged residues. Proc. Natl. Acad. Sci. USA 2013, 110, 13392–13397. [Google Scholar] [CrossRef]
- Tripathi, V.; Song, D.Y.; Zong, X.; Shevtsov, S.P.; Hearn, S.; Fu, X.D.; Dundr, M.; Prasanth, K.V. SRSF1 regulates the assembly of pre-mRNA processing factors in nuclear speckles. Mol. Biol. Cell 2012, 23, 3694–3706. [Google Scholar] [CrossRef]
- Yun, C.; Wang, Y.; Mukhopadhyay, D.; Backlund, P.; Kolli, N.; Yergey, A.; Wilkinson, K.D.; Dasso, M. Nucleolar protein B23/nucleophosmin regulates the vertebrate SUMO pathway through SENP3 and SENP5 proteases. J. Cell Biol. 2008, 183, 589–595. [Google Scholar] [CrossRef]
- Branon, T.C.; Bosch, J.A.; Sanchez, A.D.; Udeshi, N.D.; Svinkina, T.; Carr, S.A.; Feldman, J.L.; Perrimon, N.; Ting, A.Y. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 2018, 36, 880–887. [Google Scholar] [CrossRef]
- Liu, F.; Morderer, D.; Wren, M.C.; Vettleson-Trutza, S.A.; Wang, Y.; Rabichow, B.E.; Salemi, M.R.; Phinney, B.S.; Oskarsson, B.; Dickson, D.W.; et al. Proximity proteomics of C9orf72 dipeptide repeat proteins identifies molecular chaperones as modifiers of poly-GA aggregation. Acta Neuropathol. Commun. 2022, 10, 22. [Google Scholar] [CrossRef]
- Ueda, K.; Saichi, N.; Takami, S.; Kang, D.; Toyama, A.; Daigo, Y.; Ishikawa, N.; Kohno, N.; Tamura, K.; Shuin, T.; et al. A Comprehensive Peptidome Profiling Technology for the Identification of Early Detection Biomarkers for Lung Adenocarcinoma. PLoS ONE 2011, 6, e18567. [Google Scholar] [CrossRef]
- Decker, C.J.; Burke, J.M.; Mulvaney, P.K.; Parker, R. RNA is required for the integrity of multiple nuclear and cytoplasmic membrane-less RNP granules. EMBO J. 2022, 41, e110137. [Google Scholar] [CrossRef]
- Boulon, S.; Westman, B.J.; Hutten, S.; Boisvert, F.M.; Lamond, A.I. The nucleolus under stress. Mol. Cell 2010, 40, 216–227. [Google Scholar] [CrossRef] [PubMed]
- Shav-Tal, Y.; Blechman, J.; Darzacq, X.; Montagna, C.; Dye, B.T.; Patton, J.G.; Singer, R.H.; Zipori, D. Dynamic sorting of nuclear components into distinct nucleolar caps during transcriptional inhibition. Mol. Biol. Cell 2005, 16, 2395–2413. [Google Scholar] [CrossRef] [PubMed]
- Raina, K.; Rao, B.J. Mammalian nuclear speckles exhibit stable association with chromatin: A biochemical study. Nucleus 2022, 13, 58–73. [Google Scholar] [CrossRef] [PubMed]
- Bursac, S.; Brdovcak, M.C.; Pfannkuchen, M.; Orsolic, I.; Golomb, L.; Zhu, Y.; Katz, C.; Daftuar, L.; Grabusic, K.; Vukelic, I.; et al. Mutual protection of ribosomal proteins L5 and L11 from degradation is essential for p53 activation upon ribosomal biogenesis stress. Proc. Natl. Acad. Sci. USA 2012, 109, 20467–20472. [Google Scholar] [CrossRef] [PubMed]
- Spector, D.L.; Lamond, A.I. Nuclear speckles. Cold Spring Harb. Perspect. Biol. 2011, 3, a000646. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Peng, Q.; Wang, L.; Zhang, X.; Huang, L.; Wang, J.; Qin, Z. Serine/arginine-rich splicing factors: The bridge linking alternative splicing and cancer. Int. J. Biol. Sci. 2020, 16, 2442–2453. [Google Scholar] [CrossRef] [PubMed]
- Cazalla, D.; Zhu, J.; Manche, L.; Huber, E.; Krainer, A.R.; Caceres, J.F. Nuclear export and retention signals in the RS domain of SR proteins. Mol. Cell. Biol. 2002, 22, 6871–6882. [Google Scholar] [CrossRef]
- Staknis, D.; Reed, R. Members of a family of proteins (the RD family) detected by a U1 70K monoclonal antibody are present in spliceosomal complexes. Nucleic Acids Res. 1995, 23, 4081–4086. [Google Scholar] [CrossRef][Green Version]
- Renton, A.E.; Majounie, E.; Waite, A.; Simón-Sánchez, J.; Rollinson, S.; Gibbs, J.R.; Schymick, J.C.; Laaksovirta, H.; van Swieten, J.C.; Myllykangas, L.; et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 2011, 72, 257–268. [Google Scholar] [CrossRef]
- DeJesus-Hernandez, M.; Mackenzie, I.R.; Boeve, B.F.; Boxer, A.L.; Baker, M.; Rutherford, N.J.; Nicholson, A.M.; Finch, N.A.; Flynn, H.; Adamson, J.; et al. Expanded GGGGCC Hexanucleotide Repeat in Noncoding Region of C9ORF72 Causes Chromosome 9p-Linked FTD and ALS. Neuron 2011, 72, 245–256. [Google Scholar] [CrossRef]
- Kanekura, K.; Harada, Y.; Fujimoto, M.; Yagi, T.; Hayamizu, Y.; Nagaoka, K.; Kuroda, M. Characterization of membrane penetration and cytotoxicity of C9orf72-encoding arginine-rich dipeptides. Sci. Rep. 2018, 8, 12740. [Google Scholar] [CrossRef]
- Ederth, T.; Lerm, M.; Orihuela, B.; Rittschof, D. Resistance of Zwitterionic Peptide Monolayers to Biofouling. Langmuir 2019, 35, 1818–1827. [Google Scholar] [CrossRef]
- Beyer, C.D.; Thavalingam, S.; Guseva, T.; Schardt, L.; Zimmermann, R.; Werner, C.; Dietze, P.; Bandow, J.E.; Metzler-Nolte, N.; Rosenhahn, A. Zwitterionic Peptides Reduce Accumulation of Marine and Freshwater Biofilm Formers. ACS Appl. Mater. Interfaces 2021, 13, 49682–49691. [Google Scholar] [CrossRef]
- Chen, C.; Yamanaka, Y.; Ueda, K.; Li, P.; Miyagi, T.; Harada, Y.; Tezuka, S.; Narumi, S.; Sugimoto, M.; Kuroda, M.; et al. Phase separation and toxicity of C9orf72 poly(PR) depends on alternate distribution of arginine. J. Cell Biol. 2021, 220, e202103160. [Google Scholar] [CrossRef]
- Klein, I.A.; Boija, A.; Afeyan, L.K.; Hawken, S.W.; Fan, M.; Dall’Agnese, A.; Oksuz, O.; Henninger, J.E.; Shrinivas, K.; Sabari, B.R.; et al. Partitioning of cancer therapeutics in nuclear condensates. Science 2020, 368, 1386–1392. [Google Scholar] [CrossRef]
- Stauffer, W.; Sheng, H.; Lim, H.N. EzColocalization: An ImageJ plugin for visualizing and measuring colocalization in cells and organisms. Sci. Rep. 2018, 8, 15764. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miyagi, T.; Yamazaki, R.; Ueda, K.; Narumi, S.; Hayamizu, Y.; Uji-i, H.; Kuroda, M.; Kanekura, K. The Patterning and Proportion of Charged Residues in the Arginine-Rich Mixed-Charge Domain Determine the Membrane-Less Organelle Targeted by the Protein. Int. J. Mol. Sci. 2022, 23, 7658. https://doi.org/10.3390/ijms23147658
Miyagi T, Yamazaki R, Ueda K, Narumi S, Hayamizu Y, Uji-i H, Kuroda M, Kanekura K. The Patterning and Proportion of Charged Residues in the Arginine-Rich Mixed-Charge Domain Determine the Membrane-Less Organelle Targeted by the Protein. International Journal of Molecular Sciences. 2022; 23(14):7658. https://doi.org/10.3390/ijms23147658
Chicago/Turabian StyleMiyagi, Tamami, Rio Yamazaki, Koji Ueda, Satoshi Narumi, Yuhei Hayamizu, Hiroshi Uji-i, Masahiko Kuroda, and Kohsuke Kanekura. 2022. "The Patterning and Proportion of Charged Residues in the Arginine-Rich Mixed-Charge Domain Determine the Membrane-Less Organelle Targeted by the Protein" International Journal of Molecular Sciences 23, no. 14: 7658. https://doi.org/10.3390/ijms23147658
APA StyleMiyagi, T., Yamazaki, R., Ueda, K., Narumi, S., Hayamizu, Y., Uji-i, H., Kuroda, M., & Kanekura, K. (2022). The Patterning and Proportion of Charged Residues in the Arginine-Rich Mixed-Charge Domain Determine the Membrane-Less Organelle Targeted by the Protein. International Journal of Molecular Sciences, 23(14), 7658. https://doi.org/10.3390/ijms23147658





