Enhancing Animal Disease Resistance, Production Efficiency, and Welfare through Precise Genome Editing
Abstract
:1. Introduction
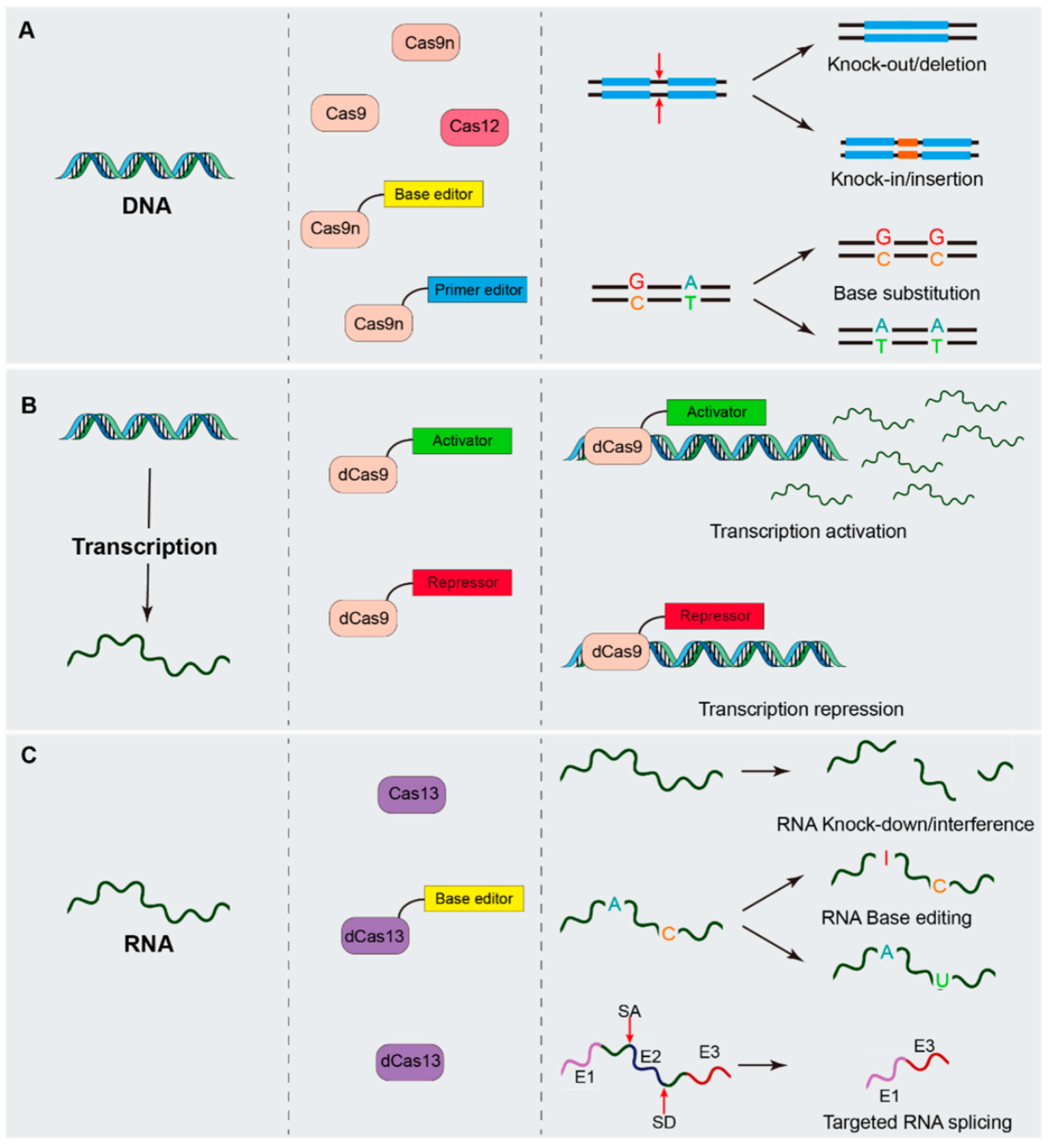
2. CRISPR/Cas System Mediated Genetic Manipulation
3. Disease Resistance Breeding
4. Improving Production Performance
5. Improving Animal Welfare
6. Regulations on GEAs
7. Prospects and Challenges
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kantor, A.; McClements, M.E.; MacLaren, R.E. CRISPR-Cas9 DNA Base-Editing and Prime-Editing. Int. J. Mol. Sci. 2020, 21, 6240. [Google Scholar] [CrossRef] [PubMed]
- Zeballos, C.M.; Gaj, T. Next-Generation CRISPR Technologies and Their Applications in Gene and Cell Therapy. Trends Biotechnol. 2021, 39, 692–705. [Google Scholar] [CrossRef] [PubMed]
- Kurt, I.C.; Zhou, R.; Iyer, S.; Garcia, S.P.; Miller, B.R.; Langner, L.M.; Grünewald, J.; Joung, J.K. CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. 2021, 39, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhu, B.; Chen, L.; Xie, L.; Yu, W.; Wang, Y.; Li, L.; Yin, S.; Yang, L.; Hu, H.; et al. Dual base editor catalyzes both cytosine and adenine base conversions in human cells. Nat. Biotechnol. 2020, 38, 856–860. [Google Scholar] [CrossRef] [PubMed]
- Grunewald, J.; Zhou, R.; Lareau, C.A.; Garcia, S.P.; Iyer, S.; Miller, B.R.; Langner, L.M.; Hsu, J.Y.; Aryee, M.J.; Joung, J.K. A dual-deaminase CRISPR base editor enables concurrent adenine and cytosine editing. Nat. Biotechnol. 2020, 38, 861–864. [Google Scholar] [CrossRef]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A.; et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef]
- Gilbert, L.A.; Larson, M.H.; Morsut, L.; Liu, Z.; Brar, G.A.; Torres, S.E.; Stern-Ginossar, N.; Brandman, O.; Whitehead, E.H.; Doudna, J.A.; et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 2013, 154, 442–451. [Google Scholar] [CrossRef] [Green Version]
- Gilbert, L.A.; Horlbeck, M.A.; Adamson, B.; Villalta, J.E.; Chen, Y.; Whitehead, E.H.; Guimaraes, C.; Panning, B.; Ploegh, H.L.; Bassik, M.C.; et al. Genome-Scale CRISPR-Mediated Control of Gene Repression and Activation. Cell 2014, 159, 647–661. [Google Scholar] [CrossRef] [Green Version]
- Vojta, A.; Dobrinić, P.; Tadić, V.; Bočkor, L.; Korać, P.; Julg, B.; Klasić, M.; Zoldoš, V. Repurposing the CRISPR-Cas9 system for targeted DNA methylation. Nucleic Acids Res. 2016, 44, 5615–5628. [Google Scholar] [CrossRef] [Green Version]
- Nunez, J.K.; Chen, J.; Pommier, G.C.; Cogan, J.Z.; Replogle, J.M.; Adriaens, C.; Ramadoss, G.N.; Shi, Q.; Hung, K.L.; Samelson, A.J.; et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 2021, 184, 2503–2519. [Google Scholar] [CrossRef]
- Engreitz, J.; Abudayyeh, O.; Gootenberg, J.; Zhang, F. CRISPR Tools for Systematic Studies of RNA Regulation. Cold Spring Harb. Perspect. Biol. 2019, 11, a035386. [Google Scholar] [CrossRef] [Green Version]
- Konermann, S.; Lotfy, P.; Brideau, N.J.; Oki, J.; Shokhirev, M.N.; Hsu, P.D. Transcriptome Engineering with RNA-Targeting Type VI-D CRISPR Effectors. Cell 2018, 173, 665–676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cox, D.B.T.; Gootenberg, J.S.; Abudayyeh, O.O.; Franklin, B.; Kellner, M.J.; Joung, J.; Zhang, F. RNA editing with CRISPR-Cas13. Science 2017, 358, 1019–1027. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abudayyeh, O.O.; Gootenberg, J.S.; Essletzbichler, P.; Han, S.; Joung, J.; Belanto, J.J.; Verdine, V.; Cox, D.B.T.; Kellner, M.J.; Regev, A.; et al. RNA targeting with CRISPR-Cas13. Nature 2017, 550, 280–284. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, M.; Jillette, N.; Zhu, J.J.; Li, S.; Cheng, A.W. CRISPR artificial splicing factors. Nat. Commun. 2020, 11, 2973. [Google Scholar] [CrossRef] [PubMed]
- Abudayyeh, O.O.; Gootenberg, J.S.; Franklin, B.; Koob, J.; Kellner, M.J.; Ladha, A.; Joung, J.; Kirchgatterer, P.; Cox, D.B.T.; Zhang, F. A cytosine deaminase for programmable single-base RNA editing. Science 2019, 365, 382–386. [Google Scholar] [CrossRef]
- Sinnamon, J.R.; Kim, S.Y.; Corson, G.M.; Song, Z.; Nakai, H.; Adelman, J.P.; Mandel, G. Site-directed RNA repair of endogenous Mecp2 RNA in neurons. Proc. Natl. Acad. Sci. USA 2017, 114, E9395–E9402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kushawah, G.; Hernandez-Huertas, L.; Abugattas-Nunez Del Prado, J.; Martinez-Morales, J.R.; DeVore, M.L.; Hassan, H.; Moreno-Sanchez, I.; Tomas-Gallardo, L.; Diaz-Moscoso, A.; Monges, D.E.; et al. CRISPR-Cas13d Induces Efficient mRNA Knockdown in Animal Embryos. Dev. Cell 2020, 54, 805–817. [Google Scholar] [CrossRef]
- He, B.; Peng, W.; Huang, J.; Zhang, H.; Zhou, Y.; Yang, X.; Liu, J.; Li, Z.; Xu, C.; Xue, M.; et al. Modulation of metabolic functions through Cas13d-mediated gene knockdown in liver. Protein Cell 2020, 11, 518–524. [Google Scholar] [CrossRef] [Green Version]
- Zhao, X.; Liu, L.; Lang, J.; Cheng, K.; Wang, Y.; Li, X.; Shi, J.; Wang, Y.; Nie, G. A CRISPR-Cas13a system for efficient and specific therapeutic targeting of mutant KRAS for pancreatic cancer treatment. Cancer Lett. 2018, 431, 171–181. [Google Scholar] [CrossRef]
- Richt, J.A.; Kasinathan, P.; Hamir, A.N.; Castilla, J.; Sathiyaseelan, T.; Vargas, F.; Sathiyaseelan, J.; Wu, H.; Matsushita, H.; Koster, J.; et al. Production of cattle lacking prion protein. Nat. Biotechnol. 2007, 25, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Qiao, J.; Fu, Q.; Chen, C.; Ni, W.; Wujiafu, S.; Ma, S.; Zhang, H.; Sheng, J.; Wang, P.; et al. Transgenic shRNA pigs reduce susceptibility to foot and mouth disease virus infection. Elife 2015, 4, e06951. [Google Scholar] [CrossRef] [PubMed]
- Shanthalingam, S.; Srikumaran, S. Intact signal peptide of CD18, the beta-subunit of beta2-integrins, renders ruminants susceptible to Mannheimia haemolytica leukotoxin. Proc. Natl. Acad. Sci. USA 2009, 106, 15448–15453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shanthalingam, S.; Tibary, A.; Beever, J.E.; Kasinathan, P.; Brown, W.C.; Srikumaran, S. Precise gene editing paves the way for derivation of Mannheimia haemolytica leukotoxin-resistant cattle. Proc. Natl. Acad. Sci. USA 2016, 113, 13186–13190. [Google Scholar] [CrossRef] [Green Version]
- Van Breedam, W.; Delputte, P.L.; Van Gorp, H.; Misinzo, G.; Vanderheijden, N.; Duan, X.; Nauwynck, H.J. Porcine reproductive and respiratory syndrome virus entry into the porcine macrophage. J. Gen. Virol. 2010, 91, 1659–1667. [Google Scholar] [CrossRef]
- Van Gorp, H.; Van Breedam, W.; Van Doorsselaere, J.; Delputte, P.L.; Nauwynck, H.J. Identification of the CD163 protein domains involved in infection of the porcine reproductive and respiratory syndrome virus. J. Virol. 2010, 84, 3101–3105. [Google Scholar] [CrossRef] [Green Version]
- Prather, R.S.; Rowland, R.R.; Ewen, C.; Trible, B.; Kerrigan, M.; Bawa, B.; Teson, J.M.; Mao, J.; Lee, K.; Samuel, M.S.; et al. An intact sialoadhesin (Sn/SIGLEC1/CD169) is not required for attachment/internalization of the porcine reproductive and respiratory syndrome virus. J. Virol. 2013, 87, 9538–9546. [Google Scholar] [CrossRef] [Green Version]
- Whitworth, K.M.; Lee, K.; Benne, J.A.; Beaton, B.P.; Spate, L.D.; Murphy, S.L.; Samuel, M.S.; Mao, J.; O’Gorman, C.; Walters, E.M.; et al. Use of the CRISPR/Cas9 system to produce genetically engineered pigs from in vitro-derived oocytes and embryos. Biol. Reprod. 2014, 91, 78. [Google Scholar] [CrossRef] [Green Version]
- Wei, Y.; Liu, Z.; Xu, K.; Evanna, H.; Dyce, P.; Li, J.; Zhou, W.; Dong, S.; Feng, B.; Mu, Y.; et al. Generation and Propagation of Cluster of Differentiation 163 Biallelic Gene Editing Pigs. Sci. Agric. Sin. 2018, 51, 770–777. [Google Scholar] [CrossRef]
- Whitworth, K.M.; Rowland, R.R.; Ewen, C.L.; Trible, B.R.; Kerrigan, M.A.; Cino-Ozuna, A.G.; Samuel, M.S.; Lightner, J.E.; McLaren, D.G.; Mileham, A.J.; et al. Gene-edited pigs are protected from porcine reproductive and respiratory syndrome virus. Nat. Biotechnol. 2016, 34, 20–22. [Google Scholar] [CrossRef]
- Yang, H.; Zhang, J.; Zhang, X.; Shi, J.; Pan, Y.; Zhou, R.; Li, G.; Li, Z.; Cai, G.; Wu, Z. CD163 knockout pigs are fully resistant to highly pathogenic porcine reproductive and respiratory syndrome virus. Antiviral Res. 2018, 151, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Wells, K.D.; Bardot, R.; Whitworth, K.M.; Trible, B.R.; Fang, Y.; Mileham, A.; Kerrigan, M.A.; Samuel, M.S.; Prather, R.S.; Rowland, R.R.R. Replacement of Porcine CD163 Scavenger Receptor Cysteine-Rich Domain 5 with a CD163-Like Homolog Confers Resistance of Pigs to Genotype 1 but Not Genotype 2 Porcine Reproductive and Respiratory Syndrome Virus. J. Virol. 2017, 91, e01521-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Wang, H.; Bai, J.; Liu, W.; Liu, X.; Yu, D.; Feng, T.; Sun, Z.; Zhang, L.; Ma, L.; et al. Generation of Pigs Resistant to Highly Pathogenic-Porcine Reproductive and Respiratory Syndrome Virus through Gene Editing of CD163. Int. J. Biol. Sci. 2019, 15, 481–492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burkard, C.; Lillico, S.G.; Reid, E.; Jackson, B.; Mileham, A.J.; Ait-Ali, T.; Whitelaw, C.B.; Archibald, A.L. Precision engineering for PRRSV resistance in pigs: Macrophages from genome edited pigs lacking CD163 SRCR5 domain are fully resistant to both PRRSV genotypes while maintaining biological function. PLoS Pathog. 2017, 13, e1006206. [Google Scholar] [CrossRef] [PubMed]
- Burkard, C.; Opriessnig, T.; Mileham, A.J.; Stadejek, T.; Ait-Ali, T.; Lillico, S.G.; Whitelaw, C.B.A.; Archibald, A.L. Pigs Lacking the Scavenger Receptor Cysteine-Rich Domain 5 of CD163 Are Resistant to Porcine Reproductive and Respiratory Syndrome Virus 1 Infection. J. Virol. 2018, 92, e00415–e00418. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H.; Shen, L.; Chen, J.; Liu, X.; Tan, T.; Hu, Y.; Bai, X.; Li, Y.; Tian, K.; Li, N.; et al. Deletion of CD163 Exon 7 Confers Resistance to Highly Pathogenic Porcine Reproductive and Respiratory Viruses on Pigs. Int. J. Biol. Sci. 2019, 15, 1993–2005. [Google Scholar] [CrossRef] [Green Version]
- Guo, C.; Wang, M.; Zhu, Z.; He, S.; Liu, H.; Liu, X.; Shi, X.; Tang, T.; Yu, P.; Zeng, J.; et al. Highly Efficient Generation of Pigs Harboring a Partial Deletion of the CD163 SRCR5 Domain, Which Are Fully Resistant to Porcine Reproductive and Respiratory Syndrome Virus 2 Infection. Front. Immunol. 2019, 10, 1846. [Google Scholar] [CrossRef] [Green Version]
- Zhu, X.; Liu, S.; Wang, X.; Luo, Z.; Shi, Y.; Wang, D.; Peng, G.; Chen, H.; Fang, L.; Xiao, S. Contribution of porcine aminopeptidase N to porcine deltacoronavirus infection. Emerg Microbes Infect. 2018, 7, 65. [Google Scholar] [CrossRef]
- Ji, C.M.; Wang, B.; Zhou, J.; Huang, Y.W. Aminopeptidase-N-independent entry of porcine epidemic diarrhea virus into Vero or porcine small intestine epithelial cells. Virology 2018, 517, 16–23. [Google Scholar] [CrossRef]
- Whitworth, K.M.; Rowland, R.R.R.; Petrovan, V.; Sheahan, M.; Cino-Ozuna, A.G.; Fang, Y.; Hesse, R.; Mileham, A.; Samuel, M.S.; Wells, K.D.; et al. Resistance to coronavirus infection in amino peptidase N-deficient pigs. Transgenic Res. 2019, 28, 21–32. [Google Scholar] [CrossRef] [Green Version]
- Luo, L.; Wang, S.; Zhu, L.; Fan, B.; Liu, T.; Wang, L.; Zhao, P.; Dang, Y.; Sun, P.; Chen, J.; et al. Aminopeptidase N-null neonatal piglets are protected from transmissible gastroenteritis virus but not porcine epidemic diarrhea virus. Sci. Rep. 2019, 9, 13186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, K.; Zhou, Y.; Mu, Y.; Liu, Z.; Hou, S.; Xiong, Y.; Fang, L.; Ge, C.; Wei, Y.; Zhang, X.; et al. CD163 and pAPN double-knockout pigs are resistant to PRRSV and TGEV and exhibit decreased susceptibility to PDCoV while maintaining normal production performance. Elife 2020, 9, e57132. [Google Scholar] [CrossRef] [PubMed]
- Tu, C.F.; Chuang, C.K.; Hsiao, K.H.; Chen, C.H.; Chen, C.M.; Peng, S.H.; Su, Y.H.; Chiou, M.T.; Yen, C.H.; Hung, S.W.; et al. Lessening of porcine epidemic diarrhoea virus susceptibility in piglets after editing of the CMP-N-glycolylneuraminic acid hydroxylase gene with CRISPR/Cas9 to nullify N-glycolylneuraminic acid expression. PLoS ONE 2019, 14, e0217236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, H.; Yan, B.-S.; Rojas, M.; Shebzukhov, Y.V.; Zhou, H.; Kobzik, L.; Higgins, D.E.; Daly, M.J.; Bloom, B.R.; Kramnik, I. Ipr1 gene mediates innate immunity to tuberculosis. Nature 2005, 434, 767–772. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Wang, Y.; Zhang, Y.; Yang, M.; Lv, J.; Liu, J.; Zhang, Y. TALE nickase-mediated SP110 knockin endows cattle with increased resistance to tuberculosis. Proc. Natl. Acad. Sci. USA 2015, 112, E1530–E1539. [Google Scholar] [CrossRef] [Green Version]
- Gao, Y.; Wu, H.; Wang, Y.; Liu, X.; Chen, L.; Li, Q.; Cui, C.; Liu, X.; Zhang, J.; Zhang, Y. Single Cas9 nickase induced generation of NRAMP1 knockin cattle with reduced off-target effects. Genome Biol. 2017, 18, 13. [Google Scholar] [CrossRef] [Green Version]
- Lu, T.; Song, Z.; Li, Q.; Li, Z.; Wang, M.; Liu, L.; Tian, K.; Li, N. Overexpression of Histone Deacetylase 6 Enhances Resistance to Porcine Reproductive and Respiratory Syndrome Virus in Pigs. PLoS ONE 2017, 12, e0169317. [Google Scholar] [CrossRef]
- Tang, Y.-D.; Liu, J.-T.; Wang, T.-Y.; Sun, M.-X.; Tian, Z.-J.; Cai, X.-H. CRISPR/Cas9-mediated multiple single guide RNAs potently abrogate pseudorabies virus replication. Arch. Virol. 2017, 162, 3881–3886. [Google Scholar] [CrossRef]
- Hubner, A.; Petersen, B.; Keil, G.M.; Niemann, H.; Mettenleiter, T.C.; Fuchs, W. Efficient inhibition of African swine fever virus replication by CRISPR/Cas9 targeting of the viral p30 gene (CP204L). Sci. Rep. 2018, 8, 1449. [Google Scholar] [CrossRef] [Green Version]
- McPherron, A.C.; Lawler, A.M.; Lee, S.-J. Regulation of skeletal muscle mass in mice by a new TGF-beta superfamily member. Nature 1997, 387, 83–90. [Google Scholar] [CrossRef]
- Grobet, L.; Royo Martin, L.J.; Poncelet, D.; Pirottin, D.; Brouwers, B.; Riquet, J.; Schoeberlein, A.; Dunner, S.; Ménissier, F.; Massabanda, J.; et al. A deletion in the bovine myostatin gene causes the double-muscled phenotype in cattle. Nat. Genet. 1997, 17, 71–74. [Google Scholar] [CrossRef]
- Grobet, L.; Poncelet, D.; Royo, L.J.; Brouwers, B.; Pirottin, D.; Michaux, C.; Ménissier, F.; Zanotti, M.; Dunner, S.; Georges, M. Molecular definition of an allelic series of mutations disrupting the myostatin function and causing double-muscling in cattle. Mamm. Genome 1998, 9, 210–213. [Google Scholar] [CrossRef] [PubMed]
- Boman, I.A.; Klemetsdal, G.; Blichfeldt, T.; Nafstad, O.; Våge, D.I. A frameshift mutation in the coding region of the myostatin gene (MSTN) affects carcass conformation and fatness in Norwegian White Sheep (Ovis aries). Anim. Genet. 2009, 40, 418–422. [Google Scholar] [CrossRef] [PubMed]
- Mosher, D.S.; Quignon, P.; Bustamante, C.D.; Sutter, N.B.; Mellersh, C.S.; Parker, H.G.; Ostrander, E.A. A mutation in the myostatin gene increases muscle mass and enhances racing performance in heterozygote dogs. PLoS Genet. 2007, 3, e79. [Google Scholar] [CrossRef]
- Matika, O.; Robledo, D.; Pong-Wong, R.; Bishop, S.C.; Riggio, V.; Finlayson, H.; Lowe, N.R.; Hoste, A.E.; Walling, G.A.; del-Pozo, J.; et al. Balancing selection at a premature stop mutation in the myostatin gene underlies a recessive leg weakness syndrome in pigs. PLoS Genet. 2019, 15, e1007759. [Google Scholar] [CrossRef] [Green Version]
- Stinckens, A.; Luyten, T.; Bijttebier, J.; Van den Maagdenberg, K.; Dieltiens, D.; Janssens, S.; De Smet, S.; Georges, M.; Buys, N. Characterization of the complete porcine MSTN gene and expression levels in pig breeds differing in muscularity. Anim. Genet. 2008, 39, 586–596. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Tang, H.; Wang, J.; Wu, Y.; Zou, L.; Jiang, Y.; Wu, C.; Li, N. Polymorphisms in the 5′ regulatory region of myostatin gene are associated with early growth traits in Yorkshire pigs. Sci. China Ser. C 2007, 50, 642–647. [Google Scholar] [CrossRef]
- Schuelke, M.; Wagner, K.R.; Stolz, L.E.; Hübner, C.; Riebel, T.; Kömen, W.; Braun, T.; Tobin, J.F.; Lee, S.-J. Myostatin Mutation Associated with Gross Muscle Hypertrophy in a Child. N. Engl. J. Med. 2004, 350, 2682–2688. [Google Scholar] [CrossRef] [Green Version]
- Roberts, S.B.; Goetz, F.W. Myostatin protein and RNA transcript levels in adult and developing brook trout. Mol. Cell. Endocrinol. 2003, 210, 9–20. [Google Scholar] [CrossRef]
- Ding, Y.; Zhou, S.-W.; Ding, Q.; Cai, B.; Zhao, X.-E.; Zhong, S.; Jin, M.-H.; Wang, X.-L.; Ma, B.-H.; Chen, Y.-L. The CRISPR/Cas9 induces large genomic fragment deletions of MSTN and phenotypic changes in sheep. J. Integr. Agric. 2020, 19, 1065–1073. [Google Scholar] [CrossRef]
- He, Z.; Zhang, T.; Jiang, L.; Zhou, M.; Wu, D.; Mei, J.; Cheng, Y. Use of CRISPR/Cas9 technology efficiently targetted goat myostatin through zygotes microinjection resulting in double-muscled phenotype in goats. Biosci. Rep. 2018, 38, BSR20180742. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, Z.; Liu, Z.; Xu, K.; Wu, T.; Ruan, J.; Zheng, X.; Bao, S.; Mu, Y.; Sonstegard, T.; Li, K. Long-term, multidomain analyses to identify the breed and allelic effects in MSTN-edited pigs to overcome lameness and sustainably improve nutritional meat production. Sci. China Life Sci. 2022, 65, 362–375. [Google Scholar] [CrossRef]
- Ren, H.; Xiao, W.; Qin, X.; Cai, G.; Chen, H.; Hua, Z.; Cheng, C.; Li, X.; Hua, W.; Xiao, H.; et al. Myostatin regulates fatty acid desaturation and fat deposition through MEF2C/miR222/SCD5 cascade in pigs. Commun. Biol. 2020, 3, 612. [Google Scholar] [CrossRef]
- Wang, K.; Ouyang, H.; Xie, Z.; Yao, C.; Guo, N.; Li, M.; Jiao, H.; Pang, D. Efficient Generation of Myostatin Mutations in Pigs Using the CRISPR/Cas9 System. Sci. Rep. 2015, 5, 16623. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Tang, X.; Liu, Y.; Xie, Z.; Zou, X.; Li, M.; Yuan, H.; Ouyang, H.; Jiao, H.; Pang, D. Efficient Generation of Orthologous Point Mutations in Pigs via CRISPR-assisted ssODN-mediated Homology-directed Repair. Mol. Ther. Nucleic Acids 2016, 5, e396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zou, Y.; Li, Z.; Zou, Y.; Hao, H.; Hu, J.; Li, N.; Li, Q. Generation of pigs with a Belgian Blue mutation in MSTN using CRISPR/Cpf1-assisted ssODN-mediated homologous recombination. J. Integr. Agric. 2019, 18, 1329–1336. [Google Scholar] [CrossRef]
- Li, M.; Tang, X.; You, W.; Wang, Y.; Chen, Y.; Liu, Y.; Yuan, H.; Gao, C.; Chen, X.; Xiao, Z.; et al. HMEJ-mediated site-specific integration of a myostatin inhibitor increases skeletal muscle mass in porcine. Mol. Ther. Nucleic Acids 2021, 26, 49–62. [Google Scholar] [CrossRef]
- Younis, S.; Schonke, M.; Massart, J.; Hjortebjerg, R.; Sundstrom, E.; Gustafson, U.; Bjornholm, M.; Krook, A.; Frystyk, J.; Zierath, J.R.; et al. The ZBED6-IGF2 axis has a major effect on growth of skeletal muscle and internal organs in placental mammals. Proc. Natl. Acad. Sci. USA 2018, 115, E2048–E2057. [Google Scholar] [CrossRef] [Green Version]
- Xiang, G.; Ren, J.; Hai, T.; Fu, R.; Yu, D.; Wang, J.; Li, W.; Wang, H.; Zhou, Q. Editing porcine IGF2 regulatory element improved meat production in Chinese Bama pigs. Cell. Mol. Life Sci. 2018, 75, 4619–4628. [Google Scholar] [CrossRef]
- Liu, X.; Liu, H.; Wang, M.; Li, R.; Zeng, J.; Mo, D.; Cong, P.; Liu, X.; Chen, Y.; He, Z. Disruption of the ZBED6 binding site in intron 3 of IGF2 by CRISPR/Cas9 leads to enhanced muscle development in Liang Guang Small Spotted pigs. Transgenic Res. 2019, 28, 141–150. [Google Scholar] [CrossRef]
- Rupp, R.; Senin, P.; Sarry, J.; Allain, C.; Tasca, C.; Ligat, L.; Portes, D.; Woloszyn, F.; Bouchez, O.; Tabouret, G.; et al. A Point Mutation in Suppressor of Cytokine Signalling 2 (Socs2) Increases the Susceptibility to Inflammation of the Mammary Gland while Associated with Higher Body Weight and Size and Higher Milk Production in a Sheep Model. PLoS Genet. 2015, 11, e1005629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, S.; Cai, B.; He, C.; Wang, Y.; Ding, Q.; Liu, J.; Liu, Y.; Ding, Y.; Zhao, X.; Li, G.; et al. Programmable Base Editing of the Sheep Genome Revealed No Genome-Wide Off-Target Mutations. Front. Genet. 2019, 10, 215. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Ouyang, H.; Yuan, H.; Li, J.; Xie, Z.; Wang, K.; Yu, T.; Liu, M.; Chen, X.; Tang, X.; et al. Site-Specific Fat-1 Knock-In Enables Significant Decrease of n-6PUFAs/n-3PUFAs Ratio in Pigs. G3-Genes Genom Genet. 2018, 8, 1747–1754. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Cui, M.L.; Nie, Y.W.; Dai, B.; Li, F.R.; Liu, D.J.; Liang, H.; Cang, M. CRISPR/Cas9-mediated specific integration of fat-1 at the goat MSTN locus. FEBS J. 2018, 285, 2828–2839. [Google Scholar] [CrossRef] [Green Version]
- You, W.; Li, M.; Qi, Y.; Wang, Y.; Chen, Y.; Liu, Y.; Li, L.; Ouyang, H.; Pang, D. CRISPR/Cas9-Mediated Specific Integration of Fat-1 and IGF-1 at the pRosa26 Locus. Genes 2021, 12, 1027. [Google Scholar] [CrossRef]
- Zheng, Q.; Lin, J.; Huang, J.; Zhang, H.; Zhang, R.; Zhang, X.; Cao, C.; Hambly, C.; Qin, G.; Yao, J.; et al. Reconstitution of UCP1 using CRISPR/Cas9 in the white adipose tissue of pigs decreases fat deposition and improves thermogenic capacity. Proc. Natl. Acad. Sci. USA 2017, 114, E9474–E9482. [Google Scholar] [CrossRef] [Green Version]
- Carlson, D.F.; Lancto, C.A.; Zang, B.; Kim, E.S.; Walton, M.; Oldeschulte, D.; Seabury, C.; Sonstegard, T.S.; Fahrenkrug, S.C. Production of hornless dairy cattle from genome-edited cell lines. Nat. Biotechnol. 2016, 34, 479–481. [Google Scholar] [CrossRef]
- Sonstegard, T.S.; Carlson, D.; Lancto, C.A.; Fahrenkrug, S.C. Precision animal breeding as a sustainable, non-GMO solution for improving animal production and welfare. ASAP Anim. Prod. 2016, 31, 316–317. [Google Scholar]
- McCleary, S.; Strong, R.; McCarthy, R.R.; Edwards, J.C.; Howes, E.L.; Stevens, L.M.; Sanchez-Cordon, P.J.; Nunez, A.; Watson, S.; Mileham, A.J.; et al. Substitution of warthog NF-κB motifs into RELA of domestic pigs is not sufficient to confer resilience to African swine fever virus. Sci. Rep. 2020, 10, 8951. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Wei, Y.; Zhan, Q.; Yan, A.; Feng, J.; Liu, L.; Tang, D. CRISPR/Cas9-Mediated Biallelic Knockout of IRX3 Reduces the Production and Survival of Somatic Cell-Cloned Bama Minipigs. Animals 2020, 10, 501. [Google Scholar] [CrossRef] [Green Version]
- Zou, Y.; Li, Z.; Zou, Y.; Hao, H.; Li, N.; Li, Q. An FBXO40 knockout generated by CRISPR/Cas9 causes muscle hypertrophy in pigs without detectable pathological effects. Biochem. Biophys. Res. Commun. 2018, 498, 940–945. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Wang, A.; Huang, C.; Sun, Y.; Song, B.; Zhou, R.; Li, L. Generation of Marker-Free pbd-2 Knock-in Pigs Using the CRISPR/Cas9 and Cre/loxP Systems. Genes 2020, 11, 951. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Jiao, H.; Xiao, H.; Jiang, Y.; Liu, Z.; Qi, C.; Zhao, D.; Jiao, S.; Yu, T.; Tang, X.; et al. Generation of pRSAD2 gene knock-in pig via CRISPR/Cas9 technology. Antivir. Res. 2020, 174, 104696. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Matsuyama, S.; Akagi, S.; Ohkoshi, K.; Nakamura, S.; Minabe, S.; Kimura, K.; Hosoe, M. Correction of a Disease Mutation using CRISPR/Cas9-assisted Genome Editing in Japanese Black Cattle. Sci. Rep. 2017, 7, 17827. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, S.; Yu, H.; Zhao, X.; Cai, B.; Ding, Q.; Huang, Y.; Li, Y.; Li, Y.; Niu, Y.; Lei, A.; et al. Generation of gene-edited sheep with a defined Booroola fecundity gene (FecBB) mutation in bone morphogenetic protein receptor type 1B (BMPR1B) via clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated (Cas) 9. Reprod. Fertil. Dev. 2018, 30, 1616–1621. [Google Scholar] [CrossRef]
- Niu, Y.; Zhao, X.; Zhou, J.; Li, Y.; Huang, Y.; Cai, B.; Liu, Y.; Ding, Q.; Zhou, S.; Zhao, J.; et al. Efficient generation of goats with defined point mutation (I397V) in GDF9 through CRISPR/Cas9. Reprod. Fertil. Dev. 2018, 30, 307–312. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. FDA Approves First-of-its-Kind Intentional Genomic Alteration in Line of Domestic Pigs for Both Human Food, Potential Therapeutic Uses. Available online: https://www.fda.gov/news-events/press-announcements/fda-approves-first-its-kind-intentional-genomic-alteration-line-domestic-pigs-both-human-food (accessed on 14 December 2020).
- U.S. Food and Drug Administration. FDA Makes Low-Risk Determination for Marketing of Products from Genome-Edited Beef Cattle After Safety Review. Available online: https://www.fda.gov/news-events/press-announcements/fda-makes-low-risk-determination-marketing-products-genome-edited-beef-cattle-after-safety-review (accessed on 7 March 2022).
- Ministry of Agriculture and Rural Affairs of China. Guidelines for Safety Evaluation of Gene Edited Plants for Agricultural Use (for Trial Implementation). Available online: http://www.moa.gov.cn/ztzl/zjyqwgz/sbzn/202201/t20220124_6387561.htm (accessed on 24 January 2022).
- Lema, M.A. Regulatory aspects of gene editing in Argentina. Transgenic Res. 2019, 28, 147–150. [Google Scholar] [CrossRef]
- Thygesen, P. Clarifying the regulation of genome editing in Australia: Situation for genetically modified organisms. Transgenic Res. 2019, 28, 151–159. [Google Scholar] [CrossRef]
- Fan, Z.; Mu, Y.; Sonstegard, T.; Zhai, X.; Li, K.; Hackett, P.B.; Zhu, Z. Social Acceptance for Commercialization of Genetically Modified Food Animals. Natl. Sci. Rev. 2021, 8, nwab067. [Google Scholar] [CrossRef]
- Fan, Z.; Wu, T.; Wu, K.; Mu, Y.; Li, K. Reflections on the system of evaluation of gene-edited livestock. Front. Agr. Sci. Eng. 2020, 7, 211–217. [Google Scholar] [CrossRef]
- Xu, K.; Zhang, X.; Liu, Z.; Ruan, J.; Xu, C.; Che, J.; Fan, Z.; Mu, Y.; Li, K. A transgene-free method for rapid and efficient generation of precisely edited pigs without monoclonal selection. Sci. China Life Sci. 2022, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Bi, D.; Qin, G.; Song, R.; Yao, J.; Cao, C.; Zheng, Q.; Hou, N.; Wang, Y.; Zhao, J. Cytosine Base Editor (hA3A-BE3-NG)-Mediated Multiple Gene Editing for Pyramid Breeding in Pigs. Front. Genet. 2020, 11, 592623. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Yu, T.; Wang, L.; Yang, L.; Zhang, Y.; Liu, H.; Li, M.; Tang, X.; Liu, Z.; Li, Z.; et al. Efficient base editing by RNA-guided cytidine base editors (CBEs) in pigs. Cell. Mol. Life Sci. 2020, 77, 719–733. [Google Scholar] [CrossRef] [PubMed]

| Species | Gene | Modification * | Method * | Applications | References |
|---|---|---|---|---|---|
| Pig | RELA | KO | ZFN | Disease resistance | [79] |
| CD169 | KO | HR | Disease resistance | [27] | |
| CD163 | KO | CRISPR/Cas9 | Disease resistance | [29,30,31] | |
| CD163 (SRCR 5 domain) | hCD163L1 SRCR domain 8 replacement | CRISPR/Cas9 | Disease resistance | [32,33] | |
| CD163 (SRCR 5 domain) | Domain deletion | CRISPR/Cas9 | Disease resistance | [34,35,36] | |
| HDAC6 | TG | SCNT | Disease resistance | [47] | |
| CD163 | Partial domain deletion | CRISPR/Cas9 | Disease resistance | [37] | |
| pAPN | KO | CRISPR/Cas9 | Disease resistance | [40] | |
| CD163, pAPN | Double KO | CRISPR/Cas9 | Disease resistance | [42] | |
| shRNA | TG | Injection | Disease resistance | [22] | |
| CMAH | KO | CRISPR/Cas9 | Disease resistance | [43] | |
| IGF2 | Regulatory element mutation | CRISPR/Cas9 | Meat production | [69,70] | |
| IRX3 | KO | CRISPR/Cas9 | Fat content | [80] | |
| FBXO40 | KO | CRISPR/Cas9 | Meat production | [81] | |
| PBD-2 | KI | CRISPR/Cas9 | Disease resistance | [82] | |
| MSTN | Point mutation | CRISPR/Cas9 | Meat production | [65] | |
| MSTN | Partial deletion | CRISPR/Cpf1 | Meat production | [66] | |
| FST | KI | CRISPR/Cas9 | Meat production | [67] | |
| MSTN | KO | CRISPR/Cas9 | Meat production | [62] | |
| RSAD2 | KI | CRISPR/Cas9 | Disease resistance | [83] | |
| fat-1 | KI | CRISPR/Cas9 | Meat quality | [73] | |
| fat-1, IGF1 | Double KI | CRISPR/Cas9 | Meat quality and meat production | [75] | |
| UCP1 | KI | CRISPR/Cas9 | Fat content and animal welfare | [76] | |
| KISSR | KO | CRISPR/Cas9 | Animal welfare | [78] | |
| Cattle | SP110 | KI | TALENs | Disease resistance | [45] |
| NRAMP1/SLC11A1 | KI | CRISPR/Cas9 | Disease resistance | [46] | |
| CD18 | Point mutation | ZFN | Disease resistance | [24] | |
| IARS | KI | CRISPR/Cas9 | Animal welfare | [84] | |
| POLLED allele | KI | TALENs | Animal welfare | [77] | |
| Sheep | BMPR1B | Point mutation | CRISPR/Cas9 | Reproductive traits | [85] |
| MSTN | KO | CRISPR/Cas9 | Meat production | [60] | |
| SOCS2 | Point mutation | Base Editor | Growth rate | [72] | |
| Goat | GDF9 | KI | CRISPR/Cas9 | Reproductive traits | [86] |
| fat-1, MSTN | KI and KO | CRISPR/Cas9 | Meat quality and meat production | [74] | |
| MSTN | KO | CRISPR/Cas9 | Meat production | [61] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Z.; Wu, T.; Xiang, G.; Wang, H.; Wang, B.; Feng, Z.; Mu, Y.; Li, K. Enhancing Animal Disease Resistance, Production Efficiency, and Welfare through Precise Genome Editing. Int. J. Mol. Sci. 2022, 23, 7331. https://doi.org/10.3390/ijms23137331
Liu Z, Wu T, Xiang G, Wang H, Wang B, Feng Z, Mu Y, Li K. Enhancing Animal Disease Resistance, Production Efficiency, and Welfare through Precise Genome Editing. International Journal of Molecular Sciences. 2022; 23(13):7331. https://doi.org/10.3390/ijms23137331
Chicago/Turabian StyleLiu, Zhiguo, Tianwen Wu, Guangming Xiang, Hui Wang, Bingyuan Wang, Zheng Feng, Yulian Mu, and Kui Li. 2022. "Enhancing Animal Disease Resistance, Production Efficiency, and Welfare through Precise Genome Editing" International Journal of Molecular Sciences 23, no. 13: 7331. https://doi.org/10.3390/ijms23137331
APA StyleLiu, Z., Wu, T., Xiang, G., Wang, H., Wang, B., Feng, Z., Mu, Y., & Li, K. (2022). Enhancing Animal Disease Resistance, Production Efficiency, and Welfare through Precise Genome Editing. International Journal of Molecular Sciences, 23(13), 7331. https://doi.org/10.3390/ijms23137331







