N-Methyltransferase CaASHH3 Acts as a Positive Regulator of Immunity against Bacterial Pathogens in Pepper
Abstract
:1. Introduction
2. Results
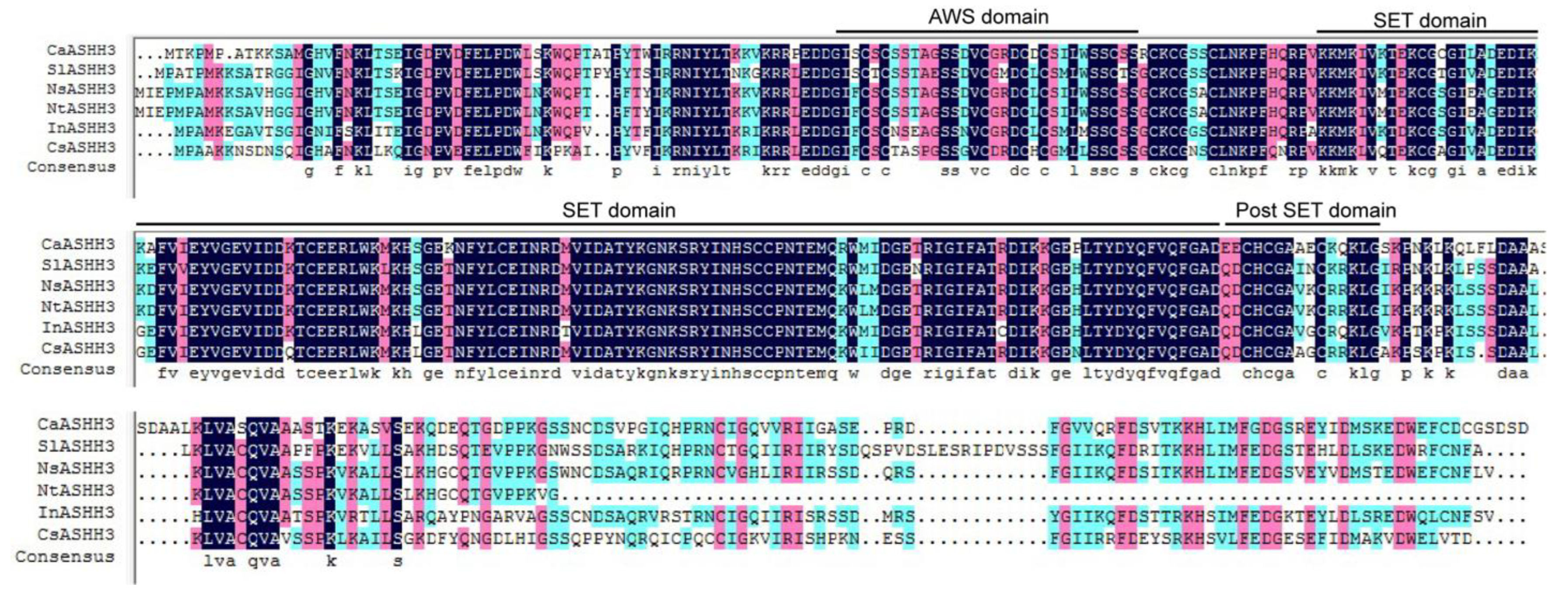
2.1. Cloning and Sequence Analysis of the Full-Length cDNA of CaASHH3
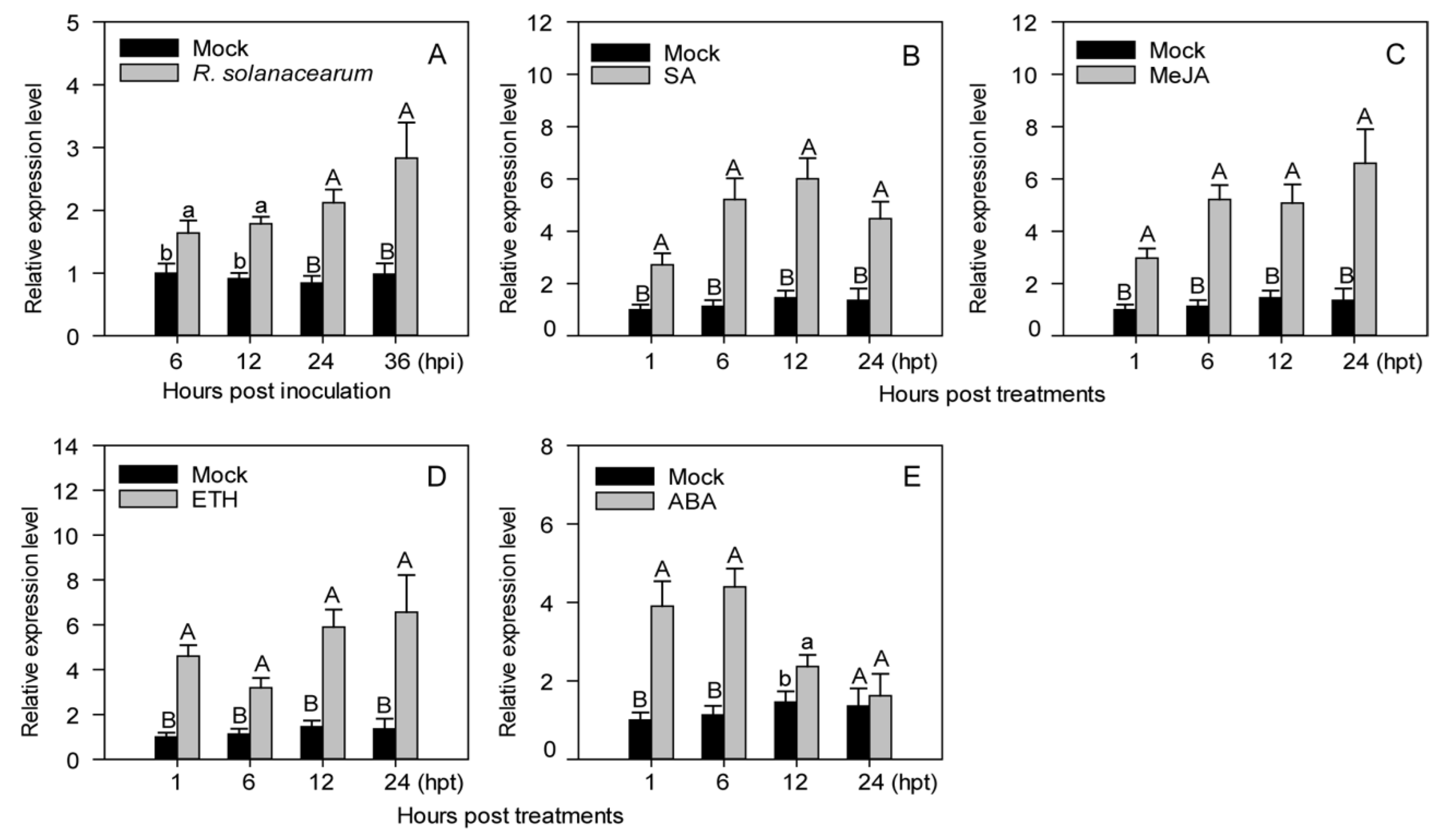
2.2. Transcriptional Expression Levels of CaASHH3 Were Upregulated by R. solanacearum Inoculation and Exogenously Supplied Phytohormones including SA, MeJA, ETH, and ABA
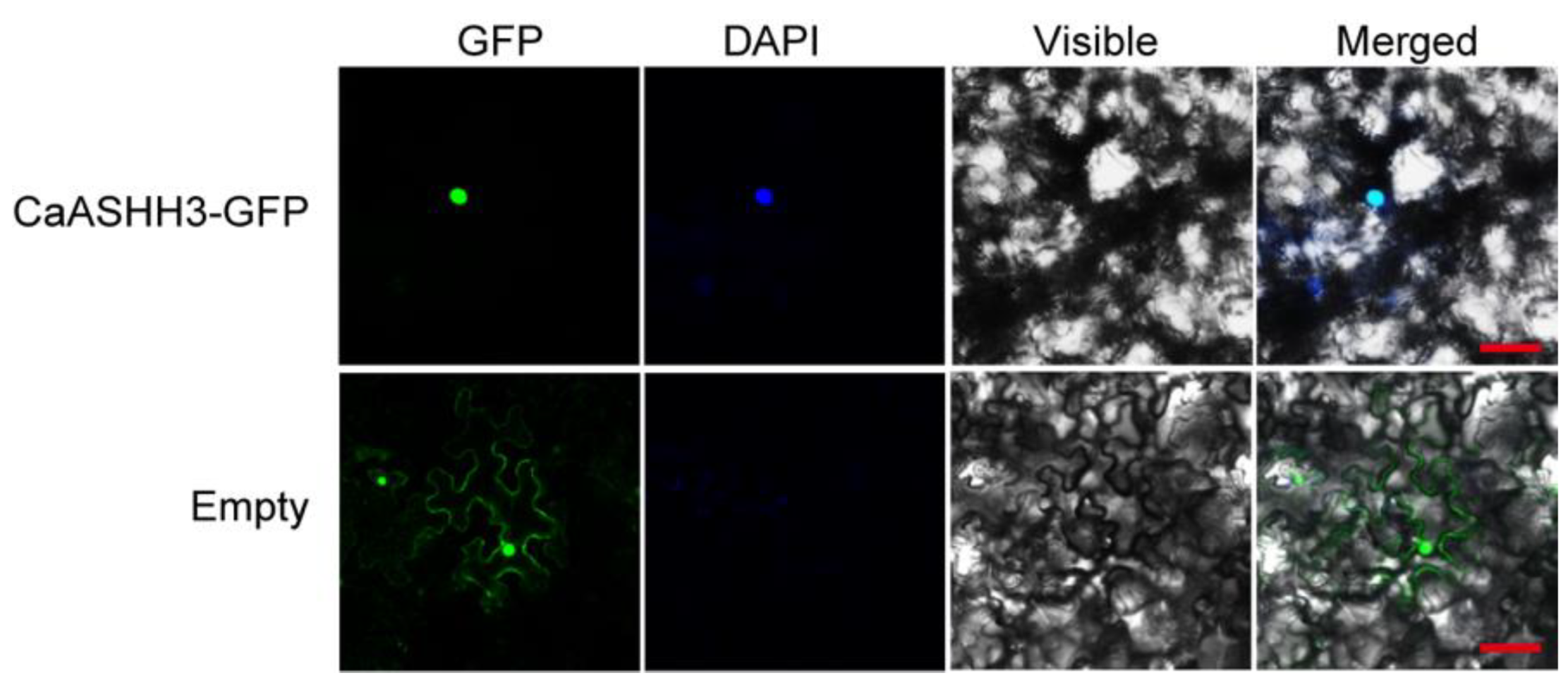
2.3. CaASHH3 Was Localized to the Nucleus
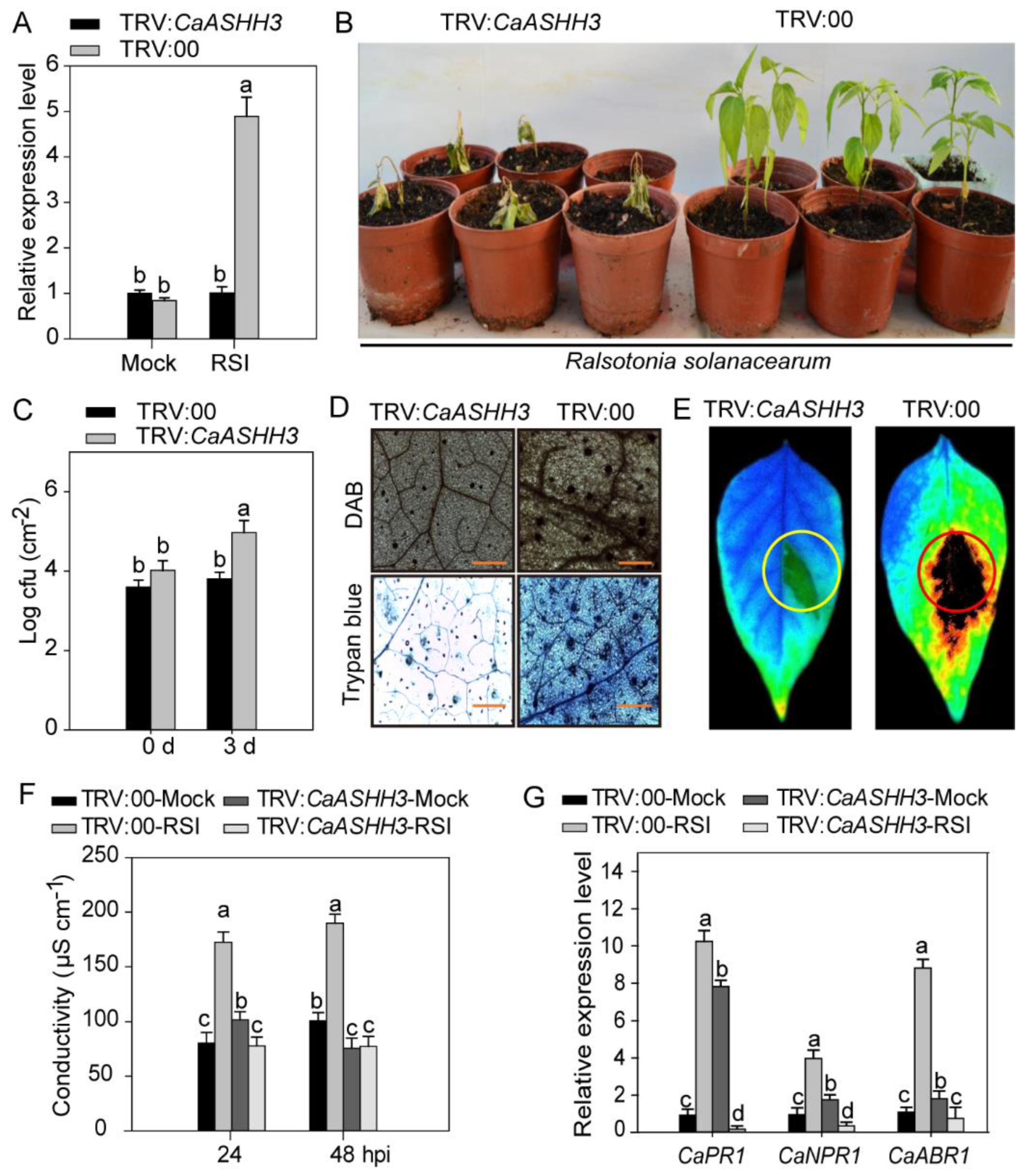
2.4. CaASHH3-Silencing Compromised the Resistance of Pepper against R. solanacearum Infection
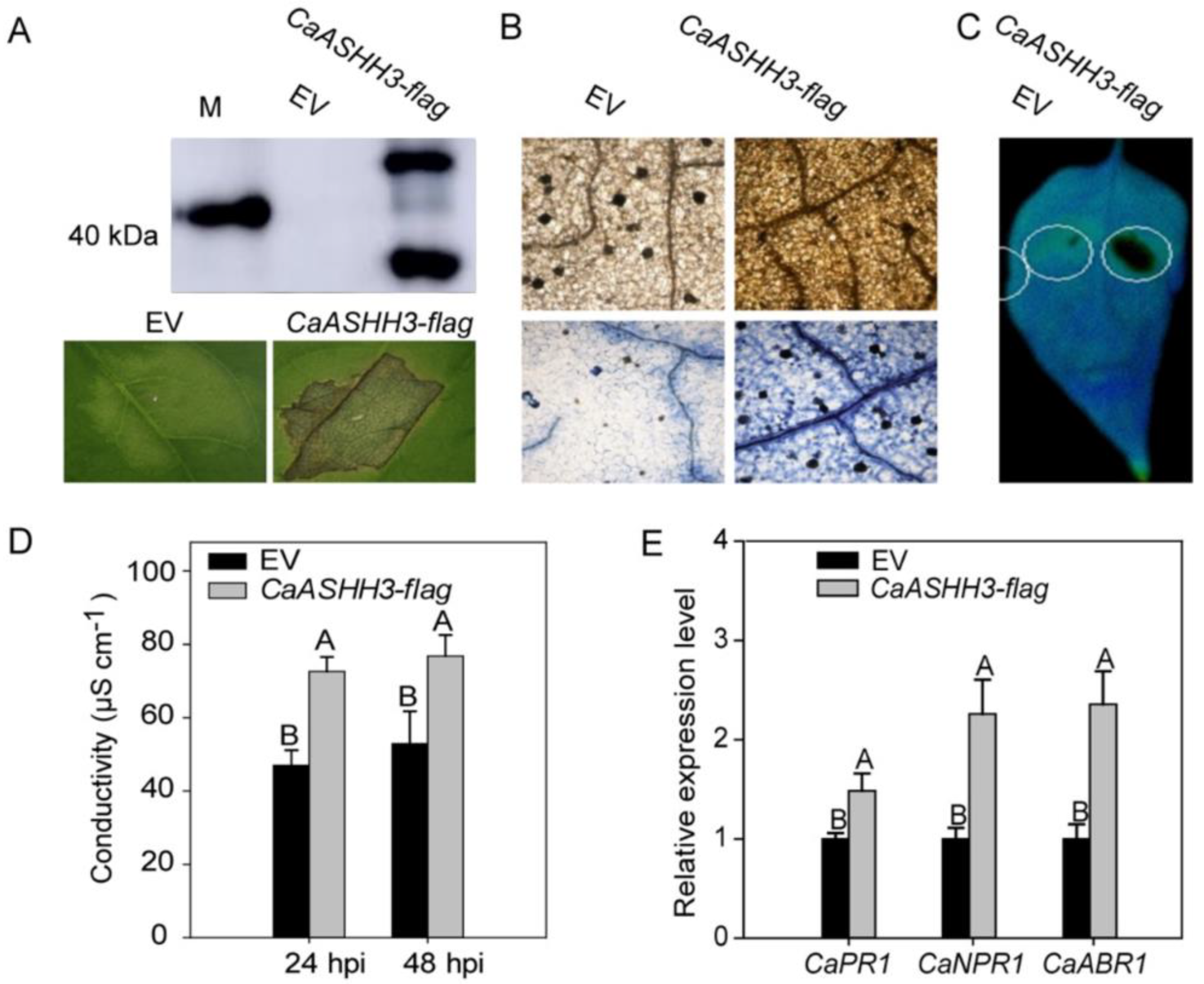
2.5. Transient Over-Expression of CaASHH3 Caused HR-like Cell Death and Accumulation of H2O2 in the Leaves of Pepper Plants
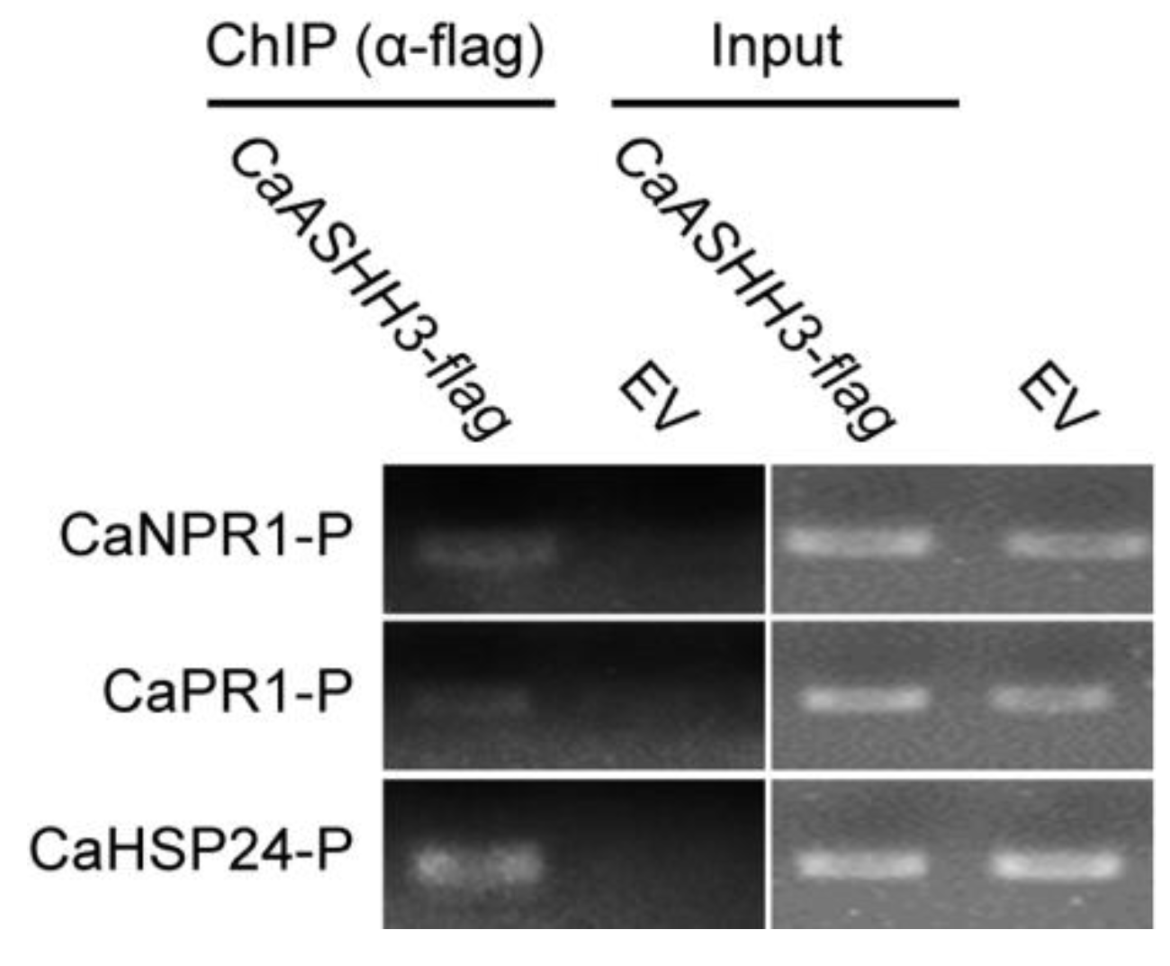
2.6. ChIP-PCR Analysis to Test Chromatin Modification in Pepper
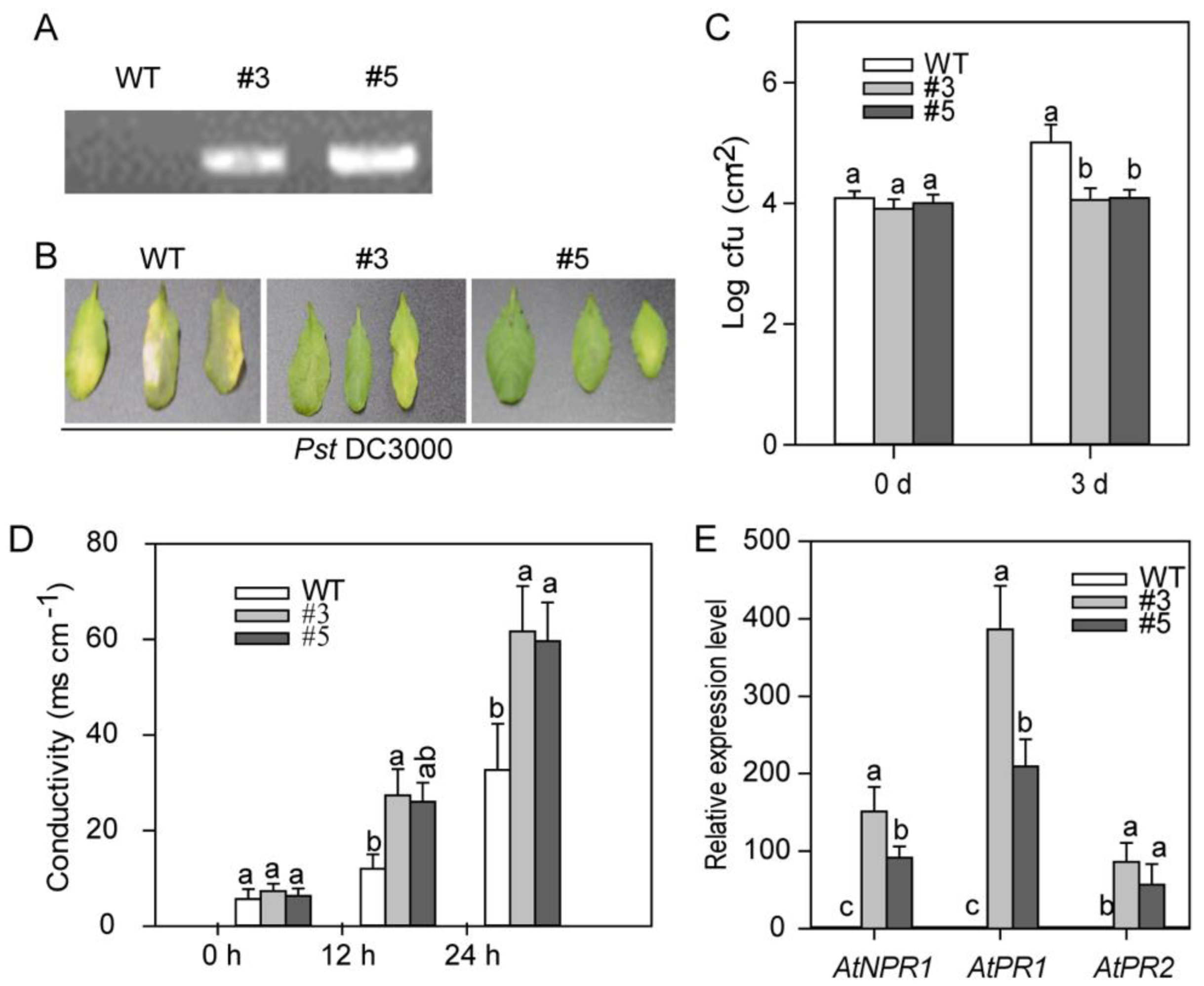
2.7. CaASHH3-OX (Over-Expression) in Transgenic Arabidopsis Plants Enhanced the Resistance to Bacterial Pathogen
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Vectors Construction
4.3. Pathogens and Inoculation Procedures
4.4. Treatment of Pepper Plants with Exogenously Applied Phytohormones
4.5. Subcellular Localization Experiment of CaASHH3
4.6. Virus-Induced Gene Silencing (VIGS) in Pepper Plants
4.7. Agrobacterium-Mediated Transient CaASHH3 Over-Expression Assay
4.8. Histochemical Staining
4.9. RNA Extraction and Real-Time qRT-PCR
4.10. Measurement of Ion Conductivity
4.11. Immunoblotting
4.12. ChIP Assays
4.13. Generation of CaASHH3-OX-Overexpressing Transgenic Arabidopsis Plants
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pastori, G.M.; Foyer, C.H. Common components, networks, and pathways of cross-tolerance to stress. The central role of “redox” and abscisic acid-mediated controls. Plant Physiol. 2002, 129, 460–468. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hussain, A.; Li, X.; Weng, Y.; Liu, Z.; Ashraf, M.F.; Noman, A.; Yang, S.; Ifnan, M.; Qiu, S.; Yang, Y. CaWRKY22 Acts as a Positive Regulator in Pepper Response to Ralstonia solanacearum by Constituting Networks with CaWRKY6, CaWRKY27, CaWRKY40, and CaWRKY58. Int. J. Mol. Sci. 2018, 19, 1426. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sung, D.-Y.; Kaplan, F.; Lee, K.-J.; Guy, C.L. Acquired tolerance to temperature extremes. Trends Plant Sci. 2003, 8, 179–187. [Google Scholar] [CrossRef]
- Zhu, J.; Dong, C.-H.; Zhu, J.-K. Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr. Opin. Plant Biol. 2007, 10, 290–295. [Google Scholar] [CrossRef]
- Schramm, F.; Larkindale, J.; Kiehlmann, E.; Ganguli, A.; Englich, G.; Vierling, E.; Koskull-Döring, V. A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J. 2008, 53, 264–274. [Google Scholar] [CrossRef] [Green Version]
- Larkindale, J.; Vierling, E. Core genome responses involved in acclimation to high temperature. Plant Physiol. 2008, 146, 748–761. [Google Scholar] [CrossRef] [Green Version]
- Boller, T.; Felix, G. A renaissance of elicitors: Perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef]
- Ifnan Khan, M.; Zhang, Y.; Liu, Z.; Hu, J.; Liu, C.; Yang, S.; Hussain, A.; Furqan Ashraf, M.; Noman, A.; Shen, L. CaWRKY40b in Pepper Acts as a Negative Regulator in Response to Ralstonia solanacearum by Directly Modulating Defense Genes Including CaWRKY40. Int. J. Mol. Sci. 2018, 19, 1403. [Google Scholar] [CrossRef] [Green Version]
- Cai, H.; Yang, S.; Yan, Y.; Xiao, Z.; Cheng, J.; Wu, J.; Qiu, A.; Lai, Y.; Mou, S.; Guan, D. CaWRKY6 transcriptionally activates CaWRKY40, regulates Ralstonia solanacearum resistance, and confers high-temperature and high-humidity tolerance in pepper. J. Exp. Bot. 2015, 66, 3163–3174. [Google Scholar] [CrossRef] [Green Version]
- Gómez-Gómez, L.; Felix, G.; Boller, T. A single locus determines sensitivity to bacterial flagellin in Arabidopsis thaliana. Plant J. 1999, 18, 277–284. [Google Scholar] [CrossRef]
- Tsuda, K.; Katagiri, F. Comparing signaling mechanisms engaged in pattern-triggered and effector-triggered immunity. Curr. Opin. Plant Biol. 2010, 13, 459–465. [Google Scholar] [CrossRef] [PubMed]
- Boudsocq, M.; Willmann, M.R.; McCormack, M.; Lee, H.; Shan, L.; He, P.; Bush, J.; Cheng, S.-H.; Sheen, J. Differential innate immune signalling via Ga 2+ sensor protein kinases. Nature 2010, 464, 418. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zimmerli, L.; Métraux, J.-P.; Mauch-Mani, B. β-Aminobutyric acid-induced protection of Arabidopsis against the necrotrophic fungus Botrytis cinerea. Plant Physiol. 2001, 126, 517–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gruner, K.; Griebel, T.; Návarová, H.; Attaran, E.; Zeier, J. Reprogramming of plants during systemic acquired resistance. Front. Plant Sci. 2013, 4, 252. [Google Scholar] [CrossRef] [Green Version]
- Návarová, H.; Bernsdorff, F.; Döring, A.-C.; Zeier, J. Pipecolic acid, an endogenous mediator of defense amplification and priming, is a critical regulator of inducible plant immunity. Plant Cell 2012, 24, 5123–5141. [Google Scholar] [CrossRef] [Green Version]
- van Hulten, M.; Pelser, M.; Van Loon, L.; Pieterse, C.M.; Ton, J. Costs and benefits of priming for defense in Arabidopsis. Proc. Natl. Acad. Sci. 2006, 103, 5602–5607. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.; Lu, F.; Cui, X.; Cao, X. Histone methylation in higher plants. Annu. Rev. Plant Biol. 2010, 61, 395–420. [Google Scholar] [CrossRef]
- Xu, C.; Bian, C.; Yang, W.; Galka, M.; Ouyang, H.; Chen, C.; Qiu, W.; Liu, H.; Jones, A.E.; MacKenzie, F. Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc. Natl. Acad. Sci. 2010, 107, 19266–19271. [Google Scholar] [CrossRef] [Green Version]
- Wagner, E.J.; Carpenter, P.B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 2012, 13, 115–126. [Google Scholar] [CrossRef] [Green Version]
- Turner, B.M. Histone acetylation and an epigenetic code. Bioessays 2000, 22, 836–845. [Google Scholar] [CrossRef]
- Martin, C.; Zhang, Y. The diverse functions of histone lysine methylation. Nat. Rev. Mol. Cell Biol. 2005, 6, 838. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Reinberg, D. Transcription regulation by histone methylation: Interplay between different covalent modifications of the core histone tails. Genes Dev. 2001, 15, 2343–2360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lachner, M.; Jenuwein, T. The many faces of histone lysine methylation. Curr. Opin. Cell Biol. 2002, 14, 286–298. [Google Scholar] [CrossRef]
- Dang, F.F.; Wang, Y.N.; Yu, L.; Eulgem, T.; Lai, Y.; Liu, Z.Q.; Wang, X.; Qiu, A.L.; Zhang, T.X.; Lin, J. CaWRKY40, a WRKY protein of pepper, plays an important role in the regulation of tolerance to heat stress and resistance to Ralstonia solanacearum infection. Plant Cell Environ. 2013, 36, 757–774. [Google Scholar] [CrossRef]
- Ng, D.W.; Wang, T.; Chandrasekharan, M.B.; Aramayo, R.; Kertbundit, S.; Hall, T.C. Plant SET domain-containing proteins: Structure, function and regulation. Biochim. Biophys. Acta (BBA)-Gene Struct. Expr. 2007, 1769, 316–329. [Google Scholar] [CrossRef] [Green Version]
- Berr, A.; McCallum, E.J.; Alioua, A.; Heintz, D.; Heitz, T.; Shen, W.-H. Arabidopsis histone methyltransferase SET DOMAIN GROUP8 mediates induction of the jasmonate/ethylene pathway genes in plant defense response to necrotrophic fungi. Plant Physiol. 2010, 154, 1403–1414. [Google Scholar] [CrossRef] [Green Version]
- Roudier, F.; Teixeira, F.K.; Colot, V. Chromatin indexing in Arabidopsis: An epigenomic tale of tails and more. Trends Genet. 2009, 11, 511–517. [Google Scholar] [CrossRef]
- Jaskiewicz, M.; Conrath, U.; Peterhänsel, C. Chromatin modification acts as a memory for systemic acquired resistance in the plant stress response. EMBO Rep. 2011, 12, 50–55. [Google Scholar] [CrossRef] [Green Version]
- Luna, E.; Bruce, T.J.; Roberts, M.R.; Flors, V.; Ton, J. Next-generation systemic acquired resistance. Plant Physiol. 2012, 158, 844–853. [Google Scholar] [CrossRef] [Green Version]
- Po-Wen, C.; Singh, P.; Zimmerli, L. Priming of the Arabidopsis pattern-triggered immunity response upon infection by necrotrophic Pectobacterium carotovorum bacteria. Mol. Plant Pathol. 2013, 14, 58–70. [Google Scholar] [CrossRef]
- Bateman, A.; Coin, L.; Durbin, R.; Finn, R.D.; Hollich, V.; Griffiths-Jones, S.; Khanna, A.; Marshall, M.; Moxon, S.; Sonnhammer, E.L. The Pfam protein families database. Nucleic Acids Res. 2004, 32, D138–D141. [Google Scholar] [CrossRef] [PubMed]
- Noman, A.; Liu, Z.; Yang, S.; Shen, L.; Hussain, A.; Ashraf, M.F.; Khan, M.I.; He, S. Expression and functional evaluation of CaZNF830 during pepper response to Ralstonia solanacearum or high temperature and humidity. Microb. Pathog. 2018, 118, 336–346. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Liu, Z.; Yang, S.; Yang, T.; Liang, J.; Wen, J.; Liu, Y.; Li, J.; Shi, L.; Tang, Q. Pepper CabZIP63 acts as a positive regulator during Ralstonia solanacearum or high temperature–high humidity challenge in a positive feedback loop with CaWRKY40. J. Exp. Bot. 2016, 67, 2439–2451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.; Shi, L.; Yang, S.; Lin, Y.; Weng, Y.; Li, X.; Hussain, A.; Noman, A.; He, S. Functional and Promoter Analysis of ChiIV3, a Chitinase of Pepper Plant, in Response to Phytophthora capsici Infection. Int. J. Mol. Sci. 2017, 18, 1661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rizhsky, L.; Liang, H.; Mittler, R. The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol. 2002, 130, 1143–1151. [Google Scholar] [CrossRef] [Green Version]
- Dang, F.; Wang, Y.; She, J.; Lei, Y.; Liu, Z.; Eulgem, T.; Lai, Y.; Lin, J.; Yu, L.; Lei, D. Overexpression of CaWRKY27, a subgroup IIe WRKY transcription factor of Capsicum annuum, positively regulates tobacco resistance to Ralstonia solanacearum infection. Physiol. Plant. 2014, 150, 397–411. [Google Scholar] [CrossRef]
- Poueymiro, M.; Cunnac, S.; Barberis, P.; Deslandes, L.; Peeters, N.; Cazale-Noel, A.-C.; Boucher, C.; Genin, S. Two type III secretion system effectors from Ralstonia solanacearum GMI1000 determine host-range specificity on tobacco. Mol. Plant-Microbe Interact. 2009, 22, 538–550. [Google Scholar] [CrossRef] [Green Version]
- De-La-Peña, C.; Rangel-Cano, A.; Alvarez-Venegas, R. Regulation of disease-responsive genes mediated by epigenetic factors: Interaction of Arabidopsis–Pseudomonas. Mol. Plant Pathol. 2012, 13, 388–398. [Google Scholar] [CrossRef]
- Noman, A.; Hussain, A.; Ashraf, M.F.; Khan, M.I.; Liu, Z.; He, S. CabZIP53 is targeted by CaWRKY40 and act as positive regulator in pepper defense against Ralstonia solanacearum and thermotolerance. Environ. Exp. Bot. 2019, 159, 138–148. [Google Scholar] [CrossRef]
- Noman, A.; Hussain, A.; Adnan, M.; Khan, M.I.; Ashraf, M.F.; Zainab, M.; Khan, K.A.; Ghramh, H.A.; He, S. A novel MYB transcription factor CaPHL8 provide clues about evolution of pepper immunity againstsoil borne pathogen. Microb. Pathog. 2019, 137, 103758. [Google Scholar] [CrossRef]
- Liu, Z.; Shi, L.; Weng, Y.; Zou, H.; Li, X.; Yang, S.; Qiu, S.; Huang, X.; Huang, J.; Hussain, A. ChiIV3 acts as a novel target of WRKY40 to mediate pepper immunity against Ralstonia solanacearum infection. Mol. Plant-Microbe Interact. 2019, 32, 1121–1133. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.-Q.; Liu, Y.-Y.; Shi, L.-P.; Yang, S.; Shen, L.; Yu, H.-X.; Wang, R.-Z.; Wen, J.-Y.; Tang, Q.; Hussain, A. SGT1 is required in PcINF1/SRC2-1 induced pepper defense response by interacting with SRC2-1. Sci. Rep. 2016, 6, 21651. [Google Scholar] [CrossRef] [PubMed]
- Adnan, M.; Islam, W.; Noman, A.; Hussain, A.; Anwar, M.; Khan, M.U.; Akram, W.; Ashraf, M.F.; Raza, M.F. Q-SNARE protein FgSyn8 plays important role in growth, DON production and pathogenicity of Fusarium graminearum. Microb. Pathog. 2020, 140, 103948. [Google Scholar] [CrossRef] [PubMed]
- Smale, S.T.; Tarakhovsky, A.; Natoli, G. Chromatin contributions to the regulation of innate immunity. Annu. Rev. Immunol. 2014, 32, 489–511. [Google Scholar] [CrossRef] [PubMed]
- Obata, Y.; Furusawa, Y.; Hase, K. Epigenetic modifications of the immune system in health and disease. Immunol. Cell Biol. 2015, 93, 226–232. [Google Scholar] [CrossRef] [Green Version]
- Ramirez-Prado, J.S.; Abulfaraj, A.A.; Rayapuram, N.; Benhamed, M.; Hirt, H. Plant immunity: From signaling to epigenetic control of defense. Trends Plant Sci. 2018, 23, 833–844. [Google Scholar] [CrossRef]
- Ding, B.; Wang, G.-L. Chromatin versus pathogens: The function of epigenetics in plant immunity. Front. Plant Sci. 2015, 6, 675. [Google Scholar] [CrossRef] [Green Version]
- Antunez-Sanchez, J.; Naish, M.; Ramirez-Prado, J.S.; Ohno, S.; Huang, Y.; Dawson, A.; Manza-Mianza, D.; Ariel, F.; Raynaud, C.; Wibowo, A. A new role for histone demethylases in the maintenance of plant genome integrity. Elife 2020, 9, e58533. [Google Scholar] [CrossRef]
- Ramirez-Prado, J.S.; Piquerez, S.J.; Bendahmane, A.; Hirt, H.; Raynaud, C.; Benhamed, M. Modify the histone to win the battle: Chromatin dynamics in plant–pathogen interactions. Front. Plant Sci. 2018, 9, 355. [Google Scholar] [CrossRef]
- Berr, A.; Ménard, R.; Heitz, T.; Shen, W.H. Chromatin modification and remodelling: A regulatory landscape for the control of Arabidopsis defence responses upon pathogen attack. Cell. Microbiol. 2012, 14, 829–839. [Google Scholar] [CrossRef]
- Lu, F.; Li, G.; Cui, X.; Liu, C.; Wang, X.J.; Cao, X. Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J. Integr. Plant Biol. 2008, 50, 886–896. [Google Scholar] [CrossRef] [PubMed]
- Dutta, A.; Choudhary, P.; Caruana, J.; Raina, R. JMJ 27, an Arabidopsis H3K9 histone demethylase, modulates defense against Pseudomonas syringae and flowering time. Plant J. 2017, 91, 1015–1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alvarez-Venegas, R.; Avramova, Z. Methylation patterns of histone H3 Lys 4, Lys 9 and Lys 27 in transcriptionally active and inactive Arabidopsis genes and in atx1 mutants. Nucleic Acids Res. 2005, 33, 5199–5207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, S.; Fu, F.; Xu, S.; Lee, S.Y.; Yun, D.-J.; Mengiste, T. Global regulation of plant immunity by histone lysine methyl transferases. Plant Cell 2016, 28, 1640–1661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsu, F.-C.; Chou, M.-Y.; Chou, S.-J.; Li, Y.-R.; Peng, H.-P.; Shih, M.-C. Submergence confers immunity mediated by the WRKY22 transcription factor in Arabidopsis. Plant Cell 2013, 25, 2699–2713. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, J.; Huh, S.U.; Kojima, M.; Sakakibara, H.; Paek, K.-H.; Hwang, I. The cytokinin-activated transcription factor ARR2 promotes plant immunity via TGA3/NPR1-dependent salicylic acid signaling in Arabidopsis. Dev. Cell 2010, 19, 284–295. [Google Scholar] [CrossRef] [Green Version]
- Choi, D.S.; Hwang, I.S.; Hwang, B.K. Requirement of the cytosolic interaction between PATHOGENESIS-RELATED PROTEIN10 and LEUCINE-RICH REPEAT PROTEIN1 for cell death and defense signaling in pepper. Plant Cell 2012, 24, 1675–1690. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.S.; Hwang, B.K. The pepper MLO gene, CaMLO2, is involved in the susceptibility cell-death response and bacterial and oomycete proliferation. Plant J. 2012, 72, 843–855. [Google Scholar] [CrossRef]
- Hwang, I.S.; Kim, N.H.; Choi, D.S.; Hwang, B.K. Overexpression of Xanthomonas campestris pv. vesicatoria effector AvrBsT in Arabidopsis triggers plant cell death, disease and defense responses. Planta 2012, 236, 1191–1204. [Google Scholar] [CrossRef]
- Üstün, S.; Bartetzko, V.; Börnke, F. The Xanthomonas campestris type III effector XopJ targets the host cell proteasome to suppress salicylic-acid mediated plant defence. PLoS Pathog. 2013, 9, e1003427. [Google Scholar] [CrossRef]
- Choi, D.S.; Hwang, B.K. Proteomics and functional analyses of pepper abscisic acid–responsive 1 (ABR1), which is involved in cell death and defense signaling. Plant Cell 2011, 23, 823–842. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kunkel, B.N.; Brooks, D.M. Cross talk between signaling pathways in pathogen defense. Curr. Opin. Plant Biol. 2002, 5, 325–331. [Google Scholar] [CrossRef]
- Jacobo-Velázquez, D.A.; González-Agüero, M.; Cisneros-Zevallos, L. Cross-talk between signaling pathways: The link between plant secondary metabolite production and wounding stress response. Sci. Rep. 2015, 5, 8608. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuda, K.; Sato, M.; Stoddard, T.; Glazebrook, J.; Katagiri, F. Network properties of robust immunity in plants. PLoS Genet. 2009, 5, e1000772. [Google Scholar] [CrossRef] [Green Version]
- Bagnaresi, P.; Biselli, C.; Orrù, L.; Urso, S.; Crispino, L.; Abbruscato, P.; Piffanelli, P.; Lupotto, E.; Cattivelli, L.; Valè, G. Comparative transcriptome profiling of the early response to Magnaporthe oryzae in durable resistant vs susceptible rice (Oryza sativa L.) genotypes. PLoS ONE 2012, 7, e51609. [Google Scholar] [CrossRef] [Green Version]
- Matić, S.; Bagnaresi, P.; Biselli, C.; Carneiro, G.A.; Siciliano, I.; Valé, G.; Gullino, M.L.; Spadaro, D. Comparative transcriptome profiling of resistant and susceptible rice genotypes in response to the seedborne pathogen Fusarium fujikuroi. BMC Genom. 2016, 17, 608. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Xia, B.; Jiang, Y.; Wu, Q.; Wang, C.; He, L.; Peng, F.; Wang, R. A new pathogenesis-related protein, LrPR4, from Lycoris radiata, and its antifungal activity against Magnaporthe grisea. Mol. Biol. Rep. 2010, 37, 995–1001. [Google Scholar] [CrossRef]
- Zhang, H.; Huang, Z.; Xie, B.; Chen, Q.; Tian, X.; Zhang, X.; Zhang, H.; Lu, X.; Huang, D.; Huang, R. The ethylene-, jasmonate-, abscisic acid-and NaCl-responsive tomato transcription factor JERF1 modulates expression of GCC box-containing genes and salt tolerance in tobacco. Planta 2004, 220, 262–270. [Google Scholar] [CrossRef]
- Gu, Y.-Q.; Wildermuth, M.C.; Chakravarthy, S.; Loh, Y.-T.; Yang, C.; He, X.; Han, Y.; Martin, G.B. Tomato transcription factors Pti4, Pti5, and Pti6 activate defense responses when expressed in Arabidopsis. Plant Cell 2002, 14, 817–831. [Google Scholar] [CrossRef] [Green Version]
- Bhattarai, K.K.; Xie, Q.-G.; Mantelin, S.; Bishnoi, U.; Girke, T.; Navarre, D.A.; Kaloshian, I. Tomato susceptibility to root-knot nematodes requires an intact jasmonic acid signaling pathway. Mol. Plant-Microbe Interact. 2008, 21, 1205–1214. [Google Scholar] [CrossRef] [Green Version]
- Vicedo, B.; Flors, V.; de la O Leyva, M.; Finiti, I.; Kravchuk, Z.; Real, M.D.; García-Agustín, P.; González-Bosch, C. Hexanoic acid-induced resistance against Botrytis cinerea in tomato plants. Mol. Plant-Microbe Interact. 2009, 22, 1455–1465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O'Donnell, P.J.; Schmelz, E.; Block, A.; Miersch, O.; Wasternack, C.; Jones, J.B.; Klee, H.J. Multiple hormones act sequentially to mediate a susceptible tomato pathogen defense response. Plant Physiol. 2003, 133, 1181–1189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, S.-H.; Sharrocks, A.D.; Whitmarsh, A.J. Transcriptional regulation by the MAP kinase signaling cascades. Gene 2003, 320, 3–21. [Google Scholar] [CrossRef]
- Kelman, A. The relationship of pathogenicity of Pseudomonas solanacearum to colony appearance in a tetrazolium medium. Phytopathology 1954, 44, 693–695. [Google Scholar]
- Wang, Y.; Dang, F.; Liu, Z.; Wang, X.; Eulgem, T.; Lai, Y.; Yu, L.; She, J.; Shi, Y.; Lin, J. C a WRKY 58, encoding a group I WRKY transcription factor of C apsicum annuum, negatively regulates resistance to R alstonia solanacearum infection. Mol. Plant Pathol. 2013, 14, 131–144. [Google Scholar] [CrossRef]
- Shen, L.; Yang, S.; Yang, T.; Liang, J.; Cheng, W.; Wen, J.; Liu, Y.; Li, J.; Shi, L.; Tang, Q. CaCDPK15 positively regulates pepper responses to Ralstonia solanacearum inoculation and forms a positive-feedback loop with CaWRKY40 to amplify defense signaling. Sci. Rep. 2016, 6, 22439. [Google Scholar] [CrossRef] [Green Version]
- Korasick, D.A.; McMichael, C.; Walker, K.A.; Anderson, J.C.; Bednarek, S.Y.; Heese, A. Novel functions of Stomatal Cytokinesis-Defective 1 (SCD1) in innate immune responses against bacteria. J. Biol. Chem. 2010, 285, 23342–23350. [Google Scholar] [CrossRef] [Green Version]
- He, S. Genome-wide identification and transcriptional expression analysis of mitogen-activated protein kinase and mitogen-activated protein kinase kinase genes in Capsicum annuum. Front. Plant Sci. 2015, 6, 780. [Google Scholar]
- Hwang, I.S.; Hwang, B.K. The pepper mannose-binding lectin gene CaMBL1 is required to regulate cell death and defense responses to microbial pathogens. Plant Physiol. 2011, 155, 447–463. [Google Scholar] [CrossRef] [Green Version]
- Singh, P.; Yekondi, S.; Chen, P.-W.; Tsai, C.-H.; Yu, C.-W.; Wu, K.; Zimmerli, L. Environmental history modulates Arabidopsis pattern-triggered immunity in a HISTONE ACETYLTRANSFERASE1–dependent manner. Plant Cell 2014, 26, 2676–2688. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Wei, G.; Shi, J.; Jin, J.; Shen, T.; Ni, T.; Shen, W.H.; Yu, Y.; Dong, A. SET DOMAIN GROUP 708, a histone H3 lysine 36-specific methyltransferase, controls flowering time in rice (Oryza sativa). New Phytol. 2016, 210, 577–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method forAgrobacterium-mediated transformation ofArabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hussain, A.; Kaisheng, L.; Noman, A.; Ashraf, M.F.; Albaqami, M.; Khan, M.I.; Liu, Z.; He, S. N-Methyltransferase CaASHH3 Acts as a Positive Regulator of Immunity against Bacterial Pathogens in Pepper. Int. J. Mol. Sci. 2022, 23, 6492. https://doi.org/10.3390/ijms23126492
Hussain A, Kaisheng L, Noman A, Ashraf MF, Albaqami M, Khan MI, Liu Z, He S. N-Methyltransferase CaASHH3 Acts as a Positive Regulator of Immunity against Bacterial Pathogens in Pepper. International Journal of Molecular Sciences. 2022; 23(12):6492. https://doi.org/10.3390/ijms23126492
Chicago/Turabian StyleHussain, Ansar, Liu Kaisheng, Ali Noman, Muhammad Furqan Ashraf, Mohammed Albaqami, Muhammad Ifnan Khan, Zhiqin Liu, and Shuilin He. 2022. "N-Methyltransferase CaASHH3 Acts as a Positive Regulator of Immunity against Bacterial Pathogens in Pepper" International Journal of Molecular Sciences 23, no. 12: 6492. https://doi.org/10.3390/ijms23126492
APA StyleHussain, A., Kaisheng, L., Noman, A., Ashraf, M. F., Albaqami, M., Khan, M. I., Liu, Z., & He, S. (2022). N-Methyltransferase CaASHH3 Acts as a Positive Regulator of Immunity against Bacterial Pathogens in Pepper. International Journal of Molecular Sciences, 23(12), 6492. https://doi.org/10.3390/ijms23126492





