What Can We Learn from -Omics Approaches to Understand Clubroot Disease?
Abstract
:1. Introduction
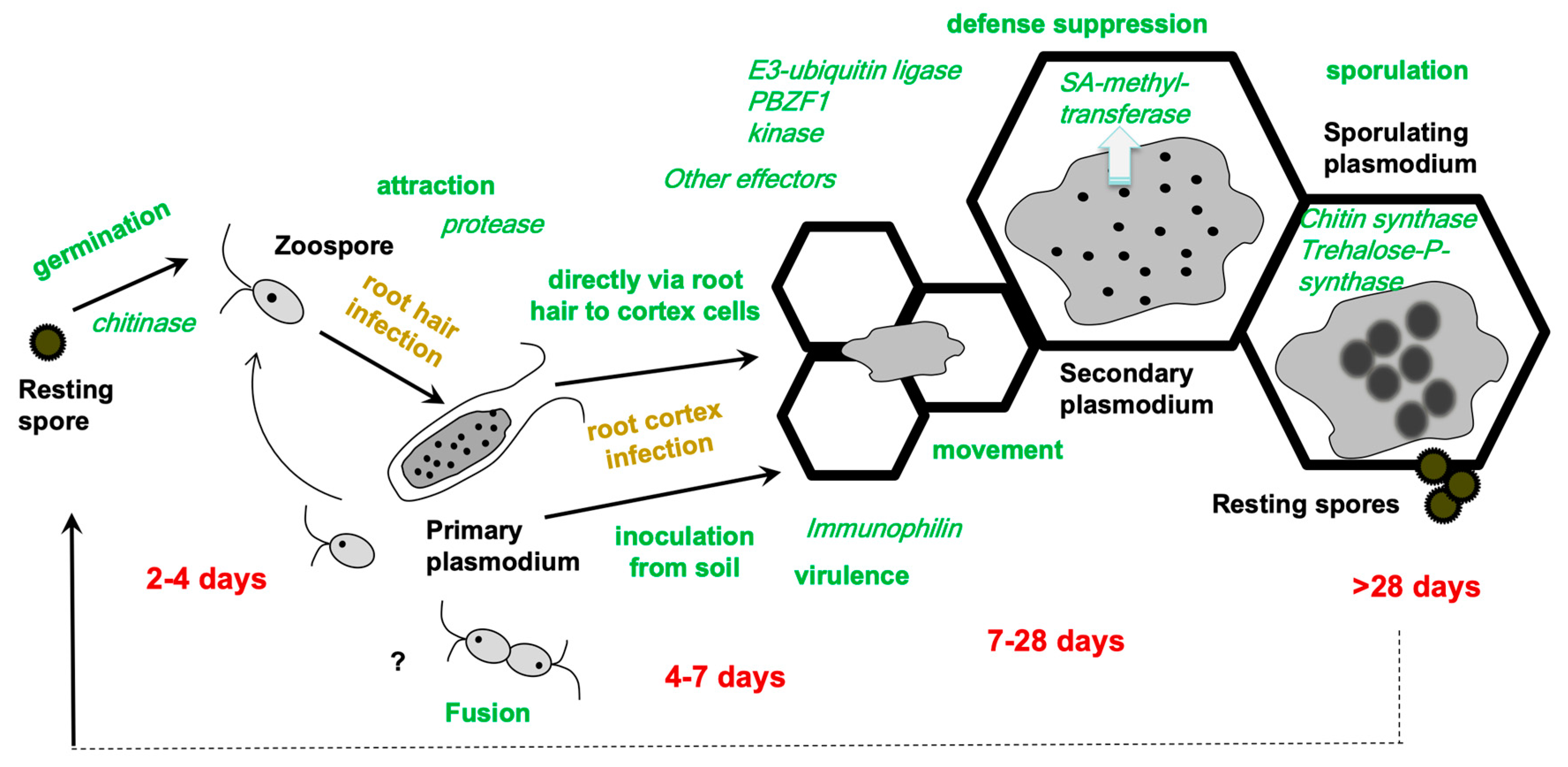
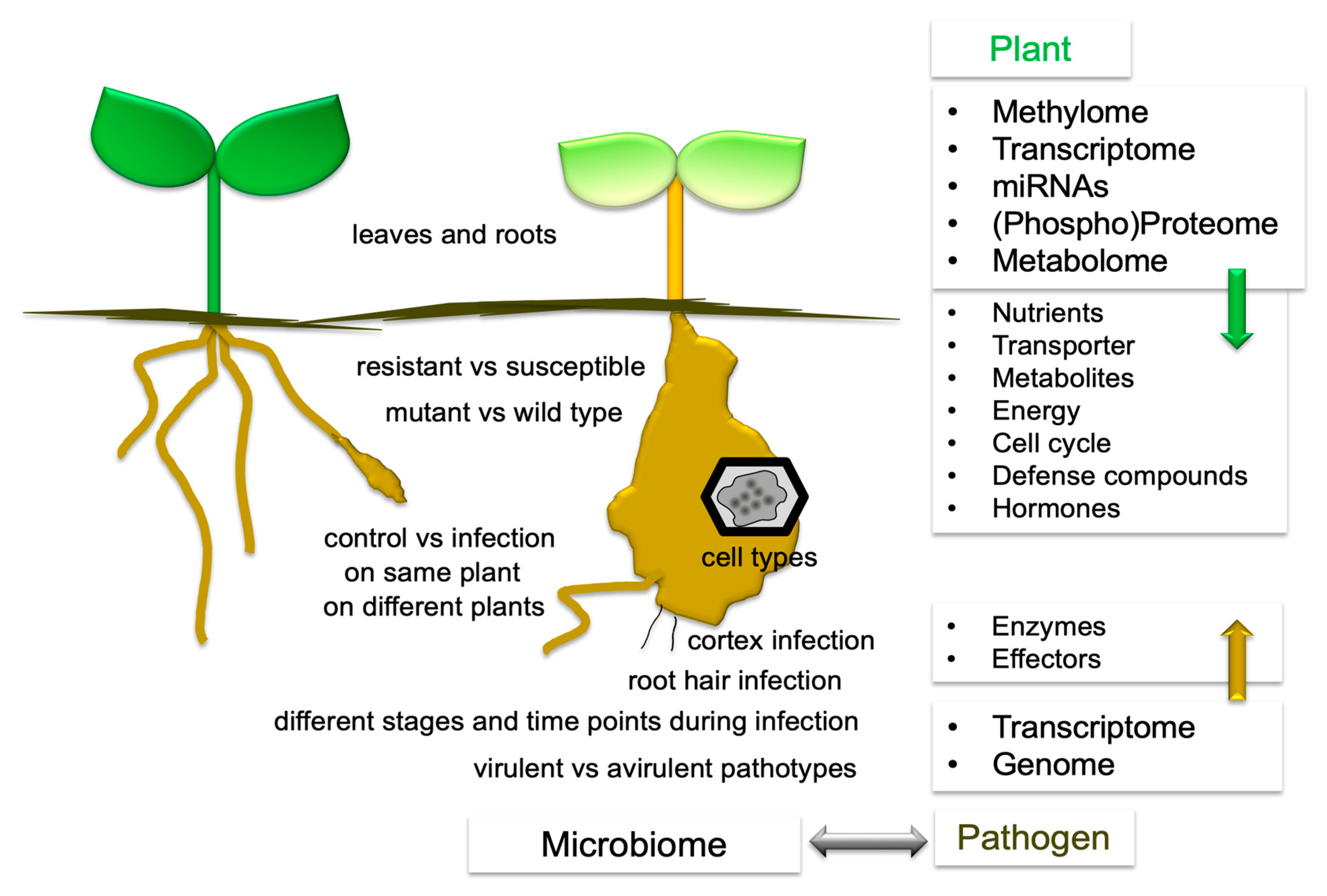
2. How to Integrate -Omics Data into the Life Cycle of Plasmodiophora brassicae
3. What Has Been Compared?
4. Plasmodiophora brassicae Genomes across the Globe
4.1. Effector Candidates
| Effector | Putative Annotated Function | Pathotype | Experimental Verification | Reference |
|---|---|---|---|---|
| Pro 1 (?) 1 | Protease | SSI 2 Williams 3 P3 | In vitro protease activity shown Treatment of plants resulted in better colonization | [70] |
| PbBSMT | SABATH-type methyltransferase | SSI e3 NZ field isolate SSI Williams P3 | In vitro conversion of SA, BA, and AA to their methylester Transgenic A. thaliana plants are more susceptible to pathogens including P. brassicae and have higher Me-SA vs. SA content | [71,73,74] |
| PbCyp3 (?) | Immunophilin | SSI e3 | Heterologous expression in Magnaporthe oryzae mutant restored virulence on rice | [72] |
| PbRING1 | E3-ubiquitin ligase | SSI e3 | E3-ubiquitin ligase activity confirmed in vitro Heterologous expression in yeast confirmed signal peptide function | [76] |
| PBZF1 | RxLR effector | Chinese field isolate Present in SSI e3 and other isolates from databases | Physical interaction with kinase SnRK1.1 Heterologous expression in A. thaliana caused plants to be more susceptible to P. brassicae | [77] |
| SSPbP22 | Kinase | SSI Williams P3 | Kinase activity determined in vitro and protein modeling | [81] |
| SSPbP53 | Cysteine protease inhibitor | SSI Williams P3 | Interaction with and inhibition of cruciferous papain-like cysteine protease A. thaliana mutant in CYSTEINE PEPTIDASE 1 more resistant to clubroot | [82] |
| PbChiB2 PbChiB4 | Chitin-binding domain carbohydrate-binding module family 18 | SSI Williams P3 | Co-precipitation showed in vitro binding to spores of P. brassicae and chitin | [83] |
| GPCR pathway | G-protein coupled receptor pathway | SSI ZJ-1 | Treatment with GPCR inhibitor resulted in reduced symptoms | [64] |
| NUDIX_hydrolase | NUDIX effector | eH, Somé et al. based P1 | No experimental confirmation | [43] |
| PbGH3 | IAA conjugating enzyme | SSI e3 | In vitro conjugation of IAA to various amino acids | [27] |
| Indole-3-acetic acid dehydrogenase | Indole-3-acetic acid dehydrogenase | CCD based P5X | No experimental confirmation; Predicted function in IAA synthesis | [79] |
| Chitin synthase | Chitin synthase | SSI e3 | Sequence prediction | [27] |
| PbTPS 4 | Trehalose-P-synthase | SSI e3 | Indirect by identification of trehalose in resting spores | [87] |
4.2. Genomes from Host Plants Can Be Used for Functional Analyses
4.3. Methylome and Epigenetic Regulations
5. Transcriptome and Posttranscriptional Regulations
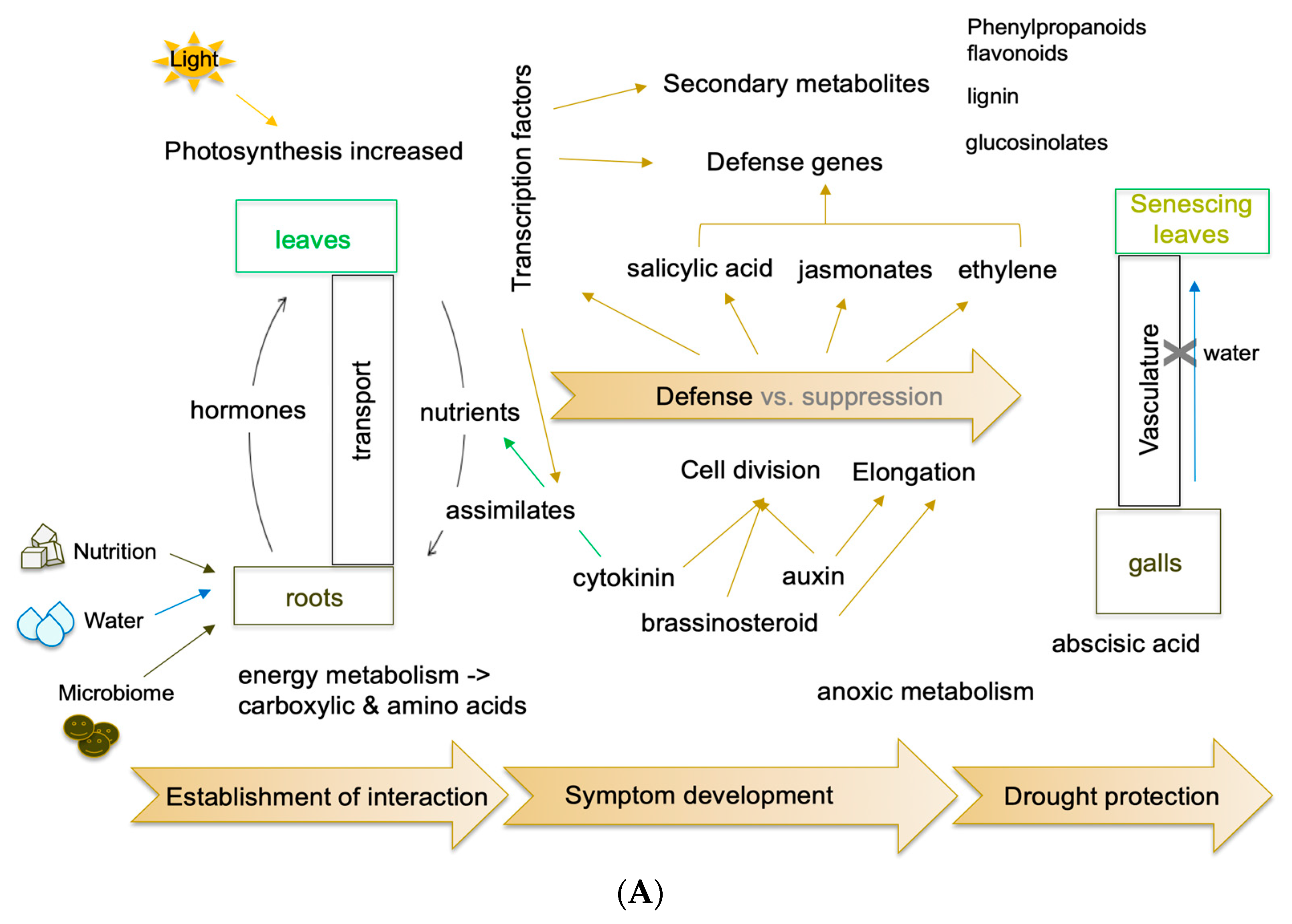
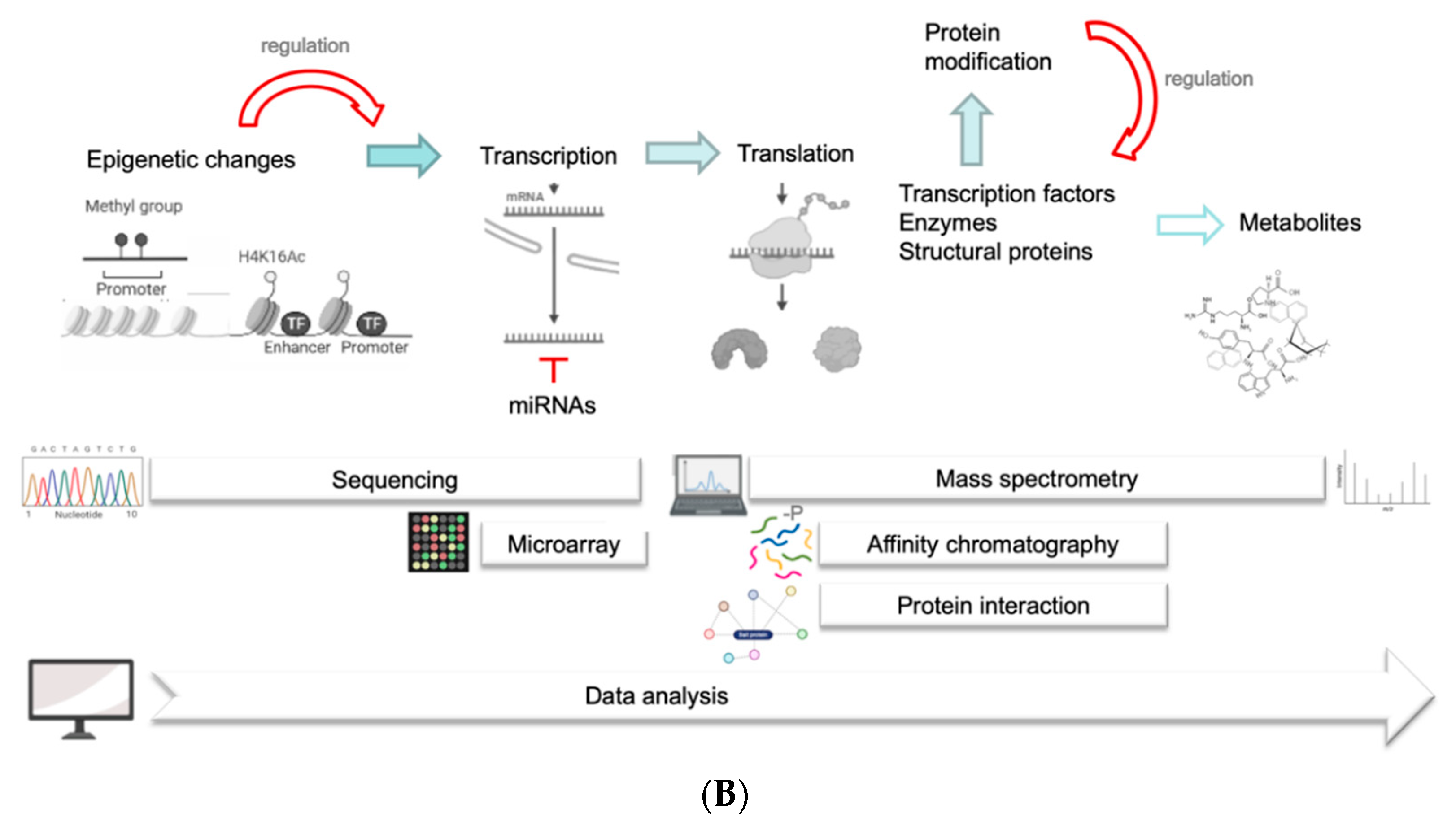
5.1. miRNAs
5.2. Nutrition
5.3. Cell biology and Growth Promoting Hormones
5.4. Defense Responses and Defense Signals
6. Proteomes of Plasmodiophora brassicae Inoculated Tissues
7. Metabolome Analyses of Plasmodiophora brassicae Inoculated Tissues
8. Potential Role for the Microbiome
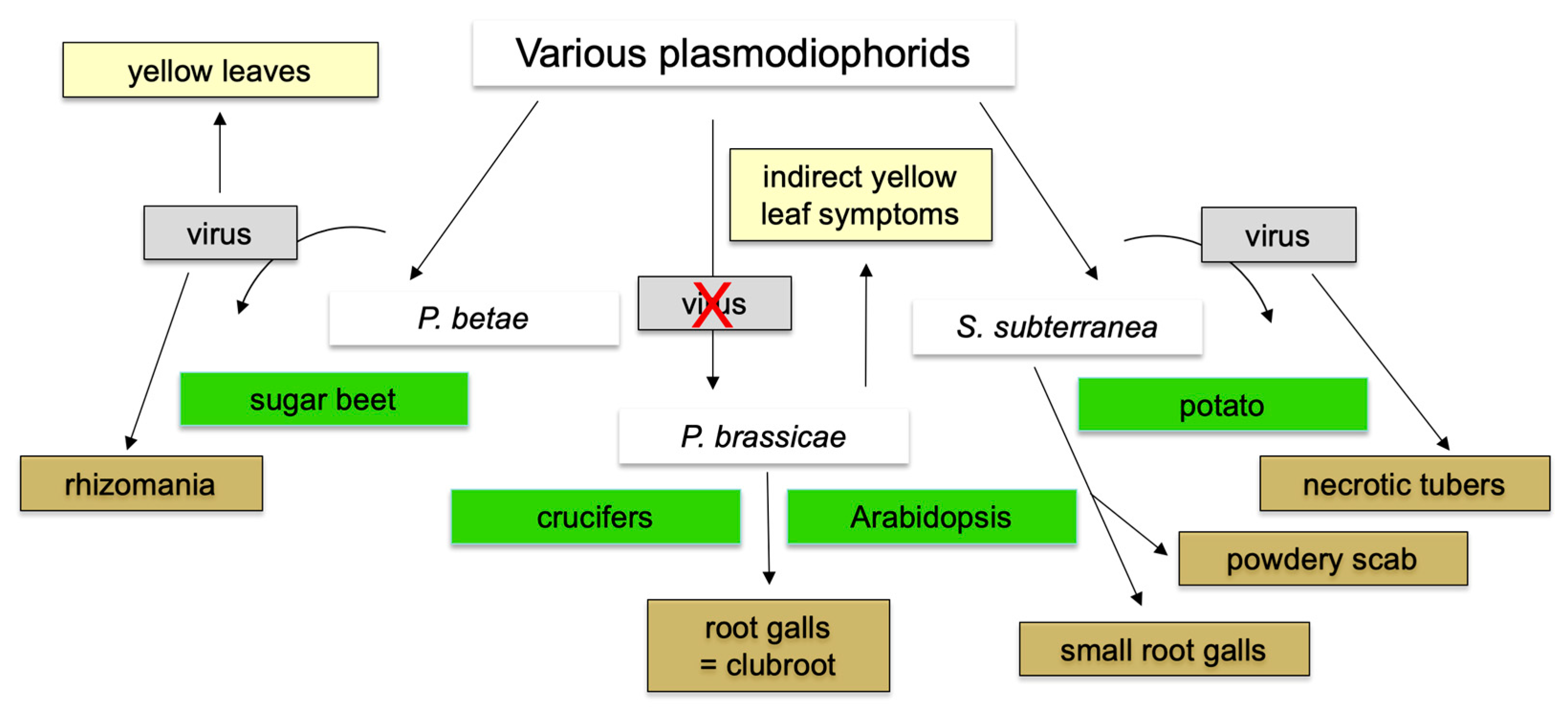
9. Status of Other Plasmodiophorid Genomes
10. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dixon, G.R. The Occurrence and Economic Impact of Plasmodiophora brassicae and Clubroot Disease. J. Plant Growth Regul. 2009, 28, 194–202. [Google Scholar] [CrossRef]
- Siemens, J.; Keller, I.; Sarx, J.; Kunz, S.; Schuller, A.; Nagel, W.; Schmülling, T.; Parniske, M.; Ludwig-Müller, J. Transcriptome Analysis of Arabidopsis Clubroots Indicate a Key Role for Cytokinins in Disease Development. Mol. Plant Microbe Interact. 2006, 19, 480–494. [Google Scholar] [CrossRef] [Green Version]
- Devos, S.; Laukens, K.; Deckers, P.; Van Der Straeten, D.; Beeckman, T.; Inzé, D.; Van Onckelen, H.; Witters, E.; Prinsen, E. A Hormone and Proteome Approach to Picturing the Initial Metabolic Events during Plasmodiophora brassicae Infection on Arabidopsis. Mol. Plant Microbe Interact. 2006, 19, 1431–1443. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Pang, W.; Chen, B.; Zhang, C.; Piao, Z. Transcriptome Analysis of Brassica rapa Near-Isogenic Lines Carrying Clubroot-Resistant and –Susceptible Alleles in Response to Plasmodiophora brassicae during Early Infection. Front. Plant Sci. 2016, 6, 1183. [Google Scholar] [CrossRef] [Green Version]
- Ning, Y.; Wang, Y.; Fang, Z.; Zhuang, M. Comparative Transcriptome Analysis of Cabbage (Brassica oleracea var. capitata) Infected by Plasmodiophora brassicae Reveals Drastic Defense Response at Secondary Infection Stage. Plant Soil 2019, 443, 167–183. [Google Scholar] [CrossRef]
- Moon, J.Y.; Tae, S.; Gyung, K.; Choi, J.; Yoon, S.; Cho, H.S.; Kim, H.S. Comparative Proteomic Analysis of Host Responses to Plasmodiophora brassicae Infection in Susceptible and Resistant Brassica oleracea. Plant Biotechnol. Rep. 2020, 14, 263–274. [Google Scholar] [CrossRef]
- Gossen, B.D.; Strelkov, S.E.; Todd, C.D. Clubroot Disease in Latin America: Distribution and Management Strategies. Plant Pathol. 2019, 68, 827–833. [Google Scholar] [CrossRef] [Green Version]
- Tommerup, I.; Ingram, D. The Life-Cycle of Plasmodiophora brassicae Woron. In Brassica Tissue Cultures and in Intact Roots. New Phytol. 1971, 70, 327–332. [Google Scholar] [CrossRef]
- Kageyama, K.; Asano, T. Life Cycle of Plasmodiophora brassicae. J. Plant Growth Regul. 2009, 28, 203–211. [Google Scholar] [CrossRef]
- Liu, L.; Qin, L.; Zhou, Z.; Hendriks, W.G.H.M.; Liu, S.; Wei, Y. Refining the Life Cycle of Plasmodiophora brassicae. Phytopathology 2020, 110, 1704–1712. [Google Scholar] [CrossRef]
- Donald, E.C.; Jaudzems, G.; Porter, I.J. Pathology of Cortical Invasion by Plasmodiophora brassicae in Clubroot Resistant and Susceptible Brassica oleracea Hosts. Plant Pathol. 2008, 57, 201–209. [Google Scholar] [CrossRef]
- Kobelt, P.; Siemens, J.; Sacristán, M.D. Histological Characterisation of the Incompatible Interaction between Arabidopsis thaliana and the Obligate Biotrophic Pathogen Plasmodiophora brassicae. Mycol. Res. 2000, 104, 220–225. [Google Scholar] [CrossRef]
- Fähling, M.; Graf, H.; Siemens, J. Characterization of a Single-Spore Isolate Population of Plasmodiophora brassicae Resulting from a Single Club. J. Phytopathol. 2004, 152, 438–444. [Google Scholar] [CrossRef]
- Klewer, A.; Luerßen, H.; Graf, H.; Siemens, J. Restriction Fragment Length Polymorphism Markers to Characterize Plasmodiophora brassicae Single-Spore Isolates with Different Virulence Patterns. J. Phytopathol. 2001, 149, 121–128. [Google Scholar] [CrossRef]
- Buczacki, S.T.; Clay, C.M. Some Observations on Secondary Zoospore Development in Plasmodiophora brassicae. Trans. Br. Mycol. Soc. 1984, 82, 339–342. [Google Scholar] [CrossRef]
- Asano, T.; Kageyama, K. Growth and Movement of Secondary Plasmodia of Plasmodiophora brassicae in Turnip Suspension-Culture Cells. Plant Pathol. 2006, 55, 145–151. [Google Scholar] [CrossRef]
- Badstöber, J.; Gachon, C.M.M.; Ludwig-Müller, J.; Sandbichler, A.M.; Neuhauser, S. Demystifying Biotrophs: FISHing for mRNAs to Decipher Plant and Algal Pathogen–Host Interaction at the Single Cell Level. Sci. Rep. 2020, 10, 14269. [Google Scholar] [CrossRef]
- Ludwig-Müller, J.; Bennett, R.; Kiddle, G.; Ihmig, S.; Ruppel, M.; Hilgenberg, W. The Host Range of Plasmodiophora brassicae and Its Relationship to Endogenous Glucosinolate Content. New Phytol. 1999, 141, 443–458. [Google Scholar] [CrossRef] [Green Version]
- Mühlenberg, I.; Schuller, A.; Siemens, J.; Kobelt, P.; Ludwig-Müller, J. Plasmodiophora brassicae, the Causal Agent of Clubroot Disease, May Penetrate Plant Cell Walls via Cellulase. Plant Prot. Sci. 2003, 38, 69–72. [Google Scholar] [CrossRef] [Green Version]
- Olszak, M.; Truman, W.; Stefanowicz, K.; Sliwinska, E.; Ito, M.; Walerowski, P.; Rolfe, S.; Malinowski, R. Transcriptional Profiling Identifies Critical Steps of Cell Cycle Reprogramming Necessary for Plasmodiophora brassicae-Driven Gall Formation in Arabidopsis. Plant J. 2019, 97, 715–729. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schuller, A.; Ludwig-Müller, J. Histological Methods to Detect the Clubroot Pathogen Plasmodiophora brassicae during Its Complex Life Cycle. Plant Pathol. 2016, 65, 1223–1237. [Google Scholar] [CrossRef]
- Ito, S.; Ichinose, H.; Yanagi, C.; Tanaka, S.; Kameya-Iwaki, M.; Kishi, F. Identification of an in Planta-Induced mRNA of Plasmodiophora brassicae. J. Phytopathol. 1999, 147, 79–82. [Google Scholar] [CrossRef]
- Bulman, S.; Siemens, J.; Ridgway, H.J.; Eady, C.; Conner, A.J. Identification of Genes from the Obligate Intracellular Plant Pathogen, Plasmodiophora brassicae. FEMS Microbiol. Lett. 2006, 264, 198–204. [Google Scholar] [CrossRef] [Green Version]
- Bulman, S.; Candy, J.M.; Fiers, M.; Lister, R.; Conner, A.J.; Eady, C.C. Genomics of Biotrophic, Plant-Infecting Plasmodiophorids Using In Vitro Dual Cultures. Protist 2011, 162, 449–461. [Google Scholar] [CrossRef]
- Siemens, J.; Graf, H.; Bulman, S.; In, O.; Ludwig-Müller, J. Monitoring Expression of Selected Plasmodiophora brassicae Genes during Clubroot Development in Arabidopsis thaliana. Plant Pathol. 2009, 58, 130–136. [Google Scholar] [CrossRef]
- Sundelin, T.; Jensen, D.F.; Lübeck, M. Identification of Expressed Genes during Infection of Chinese Cabbage (Brassica rapa subsp. pekinensis) by Plasmodiophora brassicae. J. Eukaryot. Microbiol. 2011, 58, 310–314. [Google Scholar] [CrossRef] [PubMed]
- Schwelm, A.; Fogelqvist, J.; Knaust, A.; Jülke, S.; Lilja, T.; Bonilla-Rosso, G.; Karlsson, M.; Shevchenko, A.; Dhandapani, V.; Choi, S.R.; et al. The Plasmodiophora brassicae Genome Reveals Insights in Its Life Cycle and Ancestry of Chitin Synthases. Sci. Rep. 2015, 5, 11153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bulman, S.; Ridgway, H.J.; Eady, C.; Conner, A.J. Intron-Rich Gene Structure in the Intracellular Plant Parasite Plasmodiophora brassicae. Protist 2007, 158, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Hwang, S.F.; Strelkov, S.E. Assessment of Gene Expression Profiles in Primary and Secondary Zoospores of Plasmodiophora brassicae by Dot Blot and Real-Time PCR. Microbiol. Res. 2013, 168, 518–524. [Google Scholar] [CrossRef]
- Agarwal, A.; Kaul, V.; Faggian, R.; Rookes, J.E.; Ludwig-Müller, J.; Cahill, D.M. Analysis of Global Host Gene Expression during the Primary Phase of the Arabidopsis thaliana–Plasmodiophora brassicae Interaction. Funct. Plant Biol. 2011, 38, 462–478. [Google Scholar] [CrossRef] [Green Version]
- Jubault, M.; Lariagon, C.; Taconnat, L.; Renou, J.P.; Gravot, A.; Delourme, R.; Manzanares-Dauleux, M.J. Partial Resistance to Clubroot in Arabidopsis Is Based on Changes in the Host Primary Metabolism and Targeted Cell Division and Expansion Capacity. Funct. Integr. Genom. 2013, 13, 191–205. [Google Scholar] [CrossRef] [Green Version]
- Schuller, A.; Kehr, J.; Ludwig-Müller, J. Laser Microdissection Coupled to Transcriptional Profiling of Arabidopsis Roots Inoculated by Plasmodiophora brassicae Indicates a Role for Brassinosteroids in Clubroot Formation. Plant Cell Physiol. 2014, 55, 392–411. [Google Scholar] [CrossRef]
- Ciaghi, S.; Schwelm, A.; Neuhauser, S. Transcriptomic Response in Symptomless Roots of Clubroot Infected Kohlrabi (Brassica oleracea var. gongylodes) Mirrors Resistant Plants. BMC Plant Biol. 2019, 19, 288. [Google Scholar]
- Zhang, X.; Liu, Y.; Fang, Z.; Li, Z.; Yang, L.; Zhuang, M.; Zhang, Y.; Lv, H. Comparative Transcriptome Analysis between Broccoli (Brassica oleracea var. italica) and Wild Cabbage (Brassica macrocarpa Guss.) in Response to Plasmodiophora brassicae during Different Infection Stages. Front. Plant Sci. 2016, 7, 1929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Bi, K.; Gao, Z.; Chen, T.; Liu, H.; Xie, J.; Cheng, J.; Fu, Y.; Jiang, D. Transcriptome Analysis of Arabidopsis thaliana in Response to Plasmodiophora brassicae during Early Infection. Front. Microbiol. 2017, 8, 673. [Google Scholar] [CrossRef]
- Verma, S.S.; Rahman, M.H.; Deyholos, M.K.; Basu, U.; Kav, N.N.V. Differential Expression of miRNAs in Brassica napus Root Following Infection with Plasmodiophora brassicae. PLoS ONE 2014, 9, e86648. [Google Scholar] [CrossRef] [PubMed]
- Irani, S.; Trost, B.; Waldner, M.; Nayidu, N.; Tu, J.; Kusalik, A.J.; Todd, C.D.; Wei, Y.; Bonham-Smith, P.C. Transcriptome Analysis of Response to Plasmodiophora brassicae Infection in the Arabidopsis Shoot and Root. BMC Genom. 2018, 19, 23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Irani, S.; Todd, C.D.; Wei, Y.; Bonham-Smith, P.C. Changes in Phenylpropanoid Pathway Gene Expression in Roots and Leaves of Susceptible and Resistant Brassica napus Lines in Response to Plasmodiophora brassicae Inoculation. Physiol. Mol. Plant Pathol. 2019, 106, 196–203. [Google Scholar] [CrossRef]
- Prerostova, S.; Dobrev, P.I.; Konradyova, V.; Knirsch, V.; Gaudinova, A.; Kramna, B.; Kazda, J.; Ludwig-Müller, J.; Vankova, R. Hormonal Responses to Plasmodiophora brassicae Infection in Brassica napus Cultivars Differing in Their Pathogen Resistance. Int. J. Mol. Sci. 2018, 19, 4024. [Google Scholar] [CrossRef] [Green Version]
- Liégard, B.; Baillet, V.; Etcheverry, M.; Joseph, E.; Lariagon, C.; Lemoine, J.; Evrard, A.; Colot, V.; Gravot, A.; Manzanares-Dauleux, M.J.; et al. Quantitative Resistance to Clubroot Infection Mediated by Transgenerational Epigenetic Variation in Arabidopsis. New Phytol. 2019, 222, 468–479. [Google Scholar] [CrossRef] [PubMed]
- Lebreton, L.; Guillerm-Erckelboudt, A.Y.; Gazengel, K.; Linglin, J.; Ourry, M.; Glory, P.; Sarniguet, A.; Daval, S.; Manzanares-Dauleux, M.J.; Mougel, C. Temporal Dynamics of Bacterial and Fungal Communities during the Infection of Brassica rapa Roots by the Protist Plasmodiophora brassicae. PLoS ONE 2019, 14, e0204195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Gao, Z.; Tian, B.; Bi, K.; Chen, T.; Liu, H.; Xie, J.; Cheng, J.; Fu, Y.; Jiang, D. Endosphere Microbiome Comparison between Symptomatic and Asymptomatic Roots of Brassica napus Infected with Plasmodiophora brassicae. PLoS ONE 2017, 12, e0185907. [Google Scholar] [CrossRef] [Green Version]
- Daval, S.; Gazengel, K.; Belcour, A.; Linglin, J.; Sarniguet, A.; Manzanares-Dauleux, M.J.; Mougel, C. Soil Microbiota Influences Clubroot Disease by Modulating Plasmodiophora brassicae and Brassica napus Transcriptomes. Microb. Biotechnol. 2020, 13, 1648–1672. [Google Scholar] [CrossRef]
- Yang, H.; Zheng, J.; Fu, Y.D.; Zhang, Y.H.; Yi, C.L. Specific Genes and Sequence Variation in Pathotype 7 of the Clubroot Pathogen Plasmodiophora brassicae. Eur. J. Plant Pathol. 2020, 157, 17–28. [Google Scholar] [CrossRef]
- Han, Z.; Di, C.; Khashi, M.; Gao, D.; Wu, F.; Pan, K. Repeated Application of Rice Straw Stabilizes Soil Bacterial Community Composition and Inhibits Clubroot Disease. Agriculture 2021, 11, 108. [Google Scholar] [CrossRef]
- Hu, Y.; Qiu, L.; Zhang, Z.; Liu, K.; Xia, X.; Xiong, S.; Zhao, S.; Zhao, Z.; Hu, Y.; Liang, Y. Control of Streptomyces Alfalfae XY25 T Over Clubroot Disease and Its Effect on Rhizosphere Microbial Community in Chinese Cabbage Field Trials. Front. Microbiol. 2021, 12, 641556. [Google Scholar] [CrossRef] [PubMed]
- Liao, J.; Luo, L.; Zhang, L.; Wang, L.; Shi, X.; Yang, H.; Tan, S.; Tan, L. Comparison of the Effects of Three Fungicides on Clubroot Disease of Tumorous Stem Mustard and Soil Bacterial Community. J. Soils Sediments 2022, 22, 256–271. [Google Scholar] [CrossRef]
- Jiang, J.; Fredua-Agyeman, R.; Hwang, S.-F.; Strelkov, S.E. Differentially expressed genes in canola (Brassica napus) during infection by virulent and avirulent Plasmodiophora brassicae pathotypes. Plant Pathol. 2021, 70, 50–60. [Google Scholar] [CrossRef]
- Bi, K.; He, Z.; Gao, Z.; Zhao, Y.; Fu, Y.; Cheng, J.; Xie, J.; Jiang, D.; Chen, T. Integrated Omics Study of Lipid Droplets from Plasmodiophora brassicae. Sci. Rep. 2016, 6, 36965. [Google Scholar] [CrossRef] [Green Version]
- Schwelm, A.; Dixelius, C.; Ludwig-Müller, J. New Kid on the Block—The Clubroot Pathogen Genome Moves the Plasmodiophorids into the Genomic Era. Eur. J. Plant Pathol. 2016, 145, 531–542. [Google Scholar] [CrossRef]
- Mencia, R.; Welchen, E.; Auer, S.; Ludwig-Müller, J. A Novel Target (Oxidation Resistant 2) in Arabidopsis thaliana to Reduce Clubroot Disease Symptoms via the Salicylic Acid Pathway without Growth Penalties. Horticulturae 2022, 8, 9. [Google Scholar] [CrossRef]
- Neuhauser, S.; Bulman, S.; Kirchmair, M. Plasmodiophorids: The challenge to understand soil-borne, obligate biotrophs with a multiphasic life cycle. In Molecular Identification of Fungi; Gherbawy, Y., Voigt, K., Eds.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 51–78. [Google Scholar]
- Burki, F.; Keeling, P.J. Rhizaria. Curr. Biol. 2014, 24, 103–107. [Google Scholar] [CrossRef] [Green Version]
- Sibbald, S.J.; Archibald, J.M. More protist genomes needed. Nat. Ecol. Evol. 2017, 1, 145. [Google Scholar] [CrossRef]
- Curtis, B.; Tanifuji, G.; Burki, F.; Gruber, A.; Irimia, M.; Maruyama, S.; Arias, M.C.; Ball, S.G.; Gile, G.H.; Hirakawa, Y.; et al. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature 2012, 492, 59–65. [Google Scholar] [CrossRef]
- Glöckner, G.; Hülsmann, N.; Schleicher, M.; Noegel, A.A.; Eichinger, L.; Gallinger, C.; Pawlowski, J.; Sierra, R.; Euteneuer, U.; Pillet, L.; et al. The genome of the foraminiferan Reticulomyxa filosa. Curr. Biol. 2014, 24, 11–18. [Google Scholar] [CrossRef] [Green Version]
- Lhee, D.; Lee, J.; Cho, C.H.; Ha, J.-S.; Jeong, S.E.; Jeon, C.O.; Zelzion, U.; Price, D.C.; Chan, Y.-F.; Gabr, A.; et al. Paulinella micropora KR01 holobiont genome assembly for studying primary plastid evolution. bioRxiv 2019. [Google Scholar] [CrossRef] [Green Version]
- Fähling, M.; Graf, H.; Siemens, J. Pathotype Separation of Plasmodiophora brassicae by the Host Plant. J. Phytopathol. 2003, 151, 425–430. [Google Scholar] [CrossRef]
- Graf, H.; Sokolowski, F.; Klewer, A.; Diederichsen, E.; Luerßen, H.; Siemens, J. Electrophoretic Karyotype of the Obligate Biotrophic Parasite Plasmodiophora brassicae Wor. J. Phytopathol. 2001, 149, 313–318. [Google Scholar] [CrossRef]
- Graf, H.; Fähling, M.; Siemens, J. Chromosome Polymorphism of the Obligate Biotrophic Parasite Plasmodiophora brassicae. J. Phytopathol. 2004, 152, 86–91. [Google Scholar] [CrossRef]
- Stjelja, S.; Fogelqvist, J.; Tellgren-roth, C.; Dixelius, C. The Architecture of the Plasmodiophora brassicae Nuclear and Mitochondrial Genomes. Sci. Rep. 2019, 9, 15753. [Google Scholar] [CrossRef] [Green Version]
- Rolfe, S.A.; Strelkov, S.E.; Links, M.G.; Clarke, W.E.; Robinson, S.J.; Djavaheri, M.; Malinowski, R.; Haddadi, P.; Kagale, S.; Parkin, I.A.P.; et al. The Compact Genome of the Plant Pathogen Plasmodiophora brassicae Is Adapted to Intracellular Interactions with Host Brassica spp. BMC Genom. 2016, 17, 272. [Google Scholar] [CrossRef] [Green Version]
- Bi, K.; Chen, T.; He, Z.; Gao, Z.; Zhao, Y.; Fu, Y.; Cheng, J.; Xie, J.; Jiang, D. Proto-Oncogenes in a Eukaryotic Unicellular Organism Play Essential Roles in Plasmodial Growth in Host Cells. BMC Genom. 2018, 19, 881. [Google Scholar] [CrossRef] [Green Version]
- Bi, K.; Chen, T.; He, Z.; Gao, Z.; Zhao, Y.; Liu, H.; Fu, Y.; Xie, J.; Cheng, J.; Jiang, D. Comparative Genomics Reveals the Unique Evolutionary Status of Plasmodiophora brassicae and the Essential Role of GPCR Signaling Pathways. Phytopathol. Res. 2019, 1, 12. [Google Scholar] [CrossRef]
- Sedaghatkish, A.; Gossen, B.D.; Yu, F.; Torkamaneh, D.; Mcdonald, M.R. Whole-Genome DNA Similarity and Population Structure of Plasmodiophora brassicae Strains from Canada. BMC Genom. 2019, 20, 744. [Google Scholar] [CrossRef] [PubMed]
- Schwelm, A.; Ludwig-Müller, J. Molecular Pathotyping of Plasmodiophora brassicae—Genomes, Marker Genes, and Obstacles. Pathogens 2021, 10, 259. [Google Scholar] [CrossRef]
- Daval, S.; Belcour, A.; Gazengel, K.; Legrand, L.; Gouzy, J.; Cottret, L.; Lebreton, L.; Aigu, Y.; Mougel, C.; Manzanares-Dauleux, M.J. Computational Analysis of the Plasmodiophora brassicae Genome: Mitochondrial Sequence Description and Metabolic Pathway Database Design. Genomics 2019, 111, 1629–1640. [Google Scholar] [CrossRef]
- Pérez-López, E.; Waldner, M.; Hossain, M.; Kusalik, A.J.; Wei, Y.; Bonham-Smith, P.C.; Todd, C.D. Identification of Plasmodiophora brassicae Effectors—A Challenging Goal. Virulence 2018, 9, 1344–1353. [Google Scholar] [CrossRef]
- Toruno, T.Y.; Stergiopoulos, I.; Coaker, G. Plant-Pathogen Effectors: Cellular Probes Interfering with Plant Defenses in Spatial and Temporal Manners. Annu. Rev. Phytopathol. 2016, 54, 419–441. [Google Scholar] [CrossRef] [Green Version]
- Feng, J.; Hwang, R.; Hwang, S.-F.; Strelkov, S.E.; Gossen, B.D.; Zhou, Q.I.X.; Peng, G. Molecular Characterization of a Serine Protease Pro1 from Plasmodiophora brassicae That Stimulates Resting Spore Germination. Mol. Plant Pathol. 2010, 11, 503–512. [Google Scholar] [CrossRef]
- Ludwig-Müller, J.; Jülke, S.; Geiß, K.; Richter, F.; Mithöfer, A.; Sola, I.; Rusak, G.; Keenan, S.; Bulman, S. A Novel Methyltransferase from the Intracellular Pathogen Plasmodiophora brassicae Methylates Salicylic Acid. Mol. Plant Pathol. 2015, 16, 349–364. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.; Tzelepis, G.; Zouhar, M.; Ryšánek, P.; Dixelius, C. The Immunophilin Repertoire of Plasmodiophora brassicae and Functional Analysis of PbCYP3 Cyclophilin. Mol. Genet. Genom. 2017, 293, 381–390. [Google Scholar] [CrossRef] [Green Version]
- Djavaheri, M.; Ma, L.; Klessig, D.F.; Mithöfer, A.; Gropp, G.; Borhan, H. Mimicking the Host Regulation of Salicylic Acid: A Virulence Strategy by the Clubroot Pathogen Plasmodiophora brassicae. Mol. Plant Microbe Interact. 2019, 32, 296–305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bulman, S.; Richter, F.; Marschollek, S.; Benade, F.; Jülke, S.; Ludwig-Müller, J. Arabidopsis thaliana Expressing PbBSMT, a Gene Encoding a SABATH-Type Methyltransferase from the Plant Pathogenic Protist Plasmodiophora brassicae, Show Leaf Chlorosis and Altered Host Susceptibility. Plant Biol. 2019, 21, 120–130. [Google Scholar] [CrossRef]
- Mencia, R.; Céccoli, G.; Fabro, G.; Torti, P.; Colombatti, F. OXR2 Increases Plant Defense against a Hemibiotrophic Pathogen via the Salicylic Acid Pathway 1. Plant Physiol. 2020, 184, 1112–1127. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Wang, S.; Zhang, W.; Tang, J.; Wang, H.; Yu, L.; Zhang, X.; Fei, Z.; Li, J. Genome-Wide Identification of Genes Encoding Putative Secreted E3 Ubiquitin Ligases and Functional Characterization of PbRING1 in the Biotrophic Protist Plasmodiophora brassicae. Curr. Genet. 2019, 65, 1355–1365. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Li, Y.; Yan, R.; Ren, L.; Liu, F.; Zeng, L.; Sun, S.; Yang, H.; Chen, K.; Xu, L.; et al. SnRK1.1-Mediated Resistance of Arabidopsis thaliana to Clubroot Disease Is Inhibited by the Novel Plasmodiophora brassicae Effector PBZF1. Mol. Plant Pathol. 2021, 22, 1057–1069. [Google Scholar] [CrossRef]
- Baena-Gonzalez, E.; Rolland, F.; Thevelein, J.; Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 2007, 448, 938–942. [Google Scholar] [CrossRef] [PubMed]
- Galindo-González, L.; Hwang, S.; Strelkov, S.E. Candidate Effectors of Plasmodiophora brassicae Pathotype 5X During Infection of Two Brassica napus Genotypes. Front. Microbiol. 2021, 12, 742268. [Google Scholar] [CrossRef] [PubMed]
- Malinowski, R.; Novák, O.; Borhan, M.H.; Spíchal, L.; Strnad, M.; Rolfe, S.A. The Role of Cytokinins in Clubroot Disease. Eur. J. Plant Pathol. 2016, 145, 543–557. [Google Scholar] [CrossRef] [Green Version]
- Dong, S.; Wang, Y. Nudix effectors: A common weapon in the arsenal of plant pathogens. PLoS Pathog. 2016, 12, e1005704. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pérez-López, E.; Hossain, M.; Tu, J.; Waldner, M.; Todd, C.D.; Kusalik, A.J.; Wei, Y.; Bonham-Smith, P.C. Transcriptome Analysis Identifies Plasmodiophora brassicae Secondary Infection Effector Candidates. J. Eukaryot. Microbiol. 2020, 67, 337–351. [Google Scholar] [CrossRef] [Green Version]
- Pérez-López, E.; Hossain, M.; Wei, Y.; Todd, C.D.; Bonham-Smith, P.C. A Clubroot Pathogen Effector Targets Cruciferous Cysteine Proteases to Suppress Plant Immunity. Virulence 2021, 12, 2327–2340. [Google Scholar] [CrossRef]
- Muirhead, K.; Pérez-López, E. Plasmodiophora brassicae chitin-binding effectors guard and mask spores during infection. bioRxiv 2020. preprint. [Google Scholar] [CrossRef]
- Hossain, M.; Pérez-López, E.; Todd, C.D.; Wei, Y.; Bonham-Smith, P.C. Endomembrane-Targeting Plasmodiophora brassicae Effectors Modulate PAMP Triggered Immune Responses in Plants. Front. Microbiol. 2021, 12, 651279. [Google Scholar] [CrossRef]
- Chen, W.; Li, Y.; Yan, R.; Xu, L.; Ren, L.; Liu, F.; Zeng, L.; Yang, H.; Chi, P.; Wang, X.; et al. Identification and Characterization of Plasmodiophora brassicae Primary Infection Effector Candidates That Suppress or Induce Cell Death in Host and Nonhost Plants. Phytopathology 2019, 109, 1689–1697. [Google Scholar] [CrossRef]
- Brodmann, D.; Schuller, A.; Ludwig-Müller, J.; Aeschbacher, R.A.; Wiemken, A.; Boller, T.; Wingler, A. Induction of Trehalase in Arabidopsis Plants Infected with the Trehalose-Producing Pathogen Plasmodiophora brassicae. Mol. Plant Microbe Interact. 2002, 15, 693–700. [Google Scholar] [CrossRef] [Green Version]
- González-García, M.; Pérez-López, E. Looking for a Cultured Surrogate for Effectome Studies of the Clubroot Pathogen. Front. Microbiol. 2021, 12, 650307. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Galindo-González, L.; Hwang, S.; Strelkov, S.E. Application of Genomics and Transcriptomics to Accelerate Development of Clubroot Resistant Canola Can. J. Plant Pathol. 2021, 43, 189–208. [Google Scholar] [CrossRef]
- Yong, W.S.; Hsu, F.M.; Chen, P.Y. Profiling genome-wide DNA methylation. Epigenet. Chromatin 2016, 9, 26. [Google Scholar] [CrossRef] [Green Version]
- Ludwig-Müller, J.; Pieper, K.; Ruppel, M.; Cohen, J.D.; Epstein, E.; Kiddle, G.; Bennett, R. Indole Glucosinolate and Auxin Biosynthesis in Arabidopsis thaliana (L.) Heynh. Glucosinolate Mutants and the Development of Clubroot Disease. Planta 1999, 208, 409–419. [Google Scholar] [CrossRef]
- Heo, J.B.; Lee, Y.-S.; Sung, S. Epigenetic regulation by long noncoding RNAs in plants. Chromosome Res. 2013, 21, 685–693. [Google Scholar] [CrossRef] [Green Version]
- Summanwar, A.; Basu, U.; Rahman, H.; Kav, N. Identification of LncRNAs Responsive to Infection by Plasmodiophora brassicae in Clubroot-Susceptible and -Resistant Brassica napus Lines Carrying Resistance Introgressed from Rutabaga. Mol. Plant Microbe Interact. 2019, 32, 1360–1377. [Google Scholar] [CrossRef]
- Zhu, H.; Li, X.; Xi, D.; Zhai, W.; Zhang, Z. Integrating Long Noncoding RNAs and MRNAs Expression Profiles of Response to Plasmodiophora brassicae Infection in Pakchoi (Brassica campestris ssp. chinensis Makino). PLoS ONE 2019, 14, e0224927. [Google Scholar] [CrossRef] [Green Version]
- Hruz, T.; Laule, O.; Szabo, G.; Wessendorp, F.; Bleuler, S.; Oertle, L.; Widmayer, P.; Gruissem, W.; Zimmermann, P. Genevestigator V3: A reference expression database for the metaanalysis of transcriptomes. Adv. Bioinform. 2008, 2008, 420747. [Google Scholar] [CrossRef]
- Winter, D.; Vinegar, B.; Nahal, H.; Ammar, R.; Wilson, G.; Provart, N. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2007, 2, e718. [Google Scholar] [CrossRef]
- Rehman, H.M.; Nawaz, M.A.; Shah, Z.H.; Ludwig-Müller, J.; Chung, G.; Ahmad, M.Q.; Yang, S.H.; Lee, S.I. Comparative Genomic and Transcriptomic Analyses of Family-1 UDP Glycosyltransferase in Three Brassica Species and Arabidopsis Indicates Stress-Responsive Regulation. Sci. Rep. 2018, 8, 1875. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Wang, S.; Yu, F.; Tang, J.; Shan, X.; Bao, K.; Yu, L.; Wang, H.; Fei, Z.; Li, J. Genome-Wide Characterization and Expression Profiling of SWEET Genes in Cabbage (Brassica oleracea var. capitata L.) Reveal Their Roles in Chilling and Clubroot Disease Responses. BMC Genom. 2019, 20, 93. [Google Scholar] [CrossRef]
- Zhang, W.; Wang, S.; Yu, F.; Tang, J.; Yu, L.; Wang, H.; Li, J. Genome-Wide Identification and Expression Profiling of Sugar Transporter Protein (STP) Family Genes in Cabbage (Brassica oleracea var. capitata L.) Reveals Their Involvement in Clubroot Disease Responses. Genes 2019, 10, 71. [Google Scholar] [CrossRef] [Green Version]
- Siemens, J.; Gonzales, M.C.; Wolf, S.; Hofmann, C.; Greiner, S.; Du, Y.; Rausch, T.; Roitsch, T.; Ludwig-Müller, J. Extracellular Invertase Is Involved in the Regulation of Clubroot Disease in Arabidopsis thaliana. Mol. Plant Pathol. 2011, 12, 247–262. [Google Scholar] [CrossRef]
- Chen, J.; Piao, Y.; Liu, Y.; Li, X.; Piao, Z. Genome-Wide Identification and Expression Analysis of Chitinase Gene Family in Brassica rapa Reveals Its Role in Clubroot Resistance. Plant Sci. 2018, 270, 257–267. [Google Scholar] [CrossRef]
- Jahn, L.; Mucha, S.; Bergmann, S.; Horn, C.; Staswick, P.; Steffens, B.; Siemens, J.; Ludwig-Müller, J. The Clubroot Pathogen (Plasmodiophora brassicae) Influences Auxin Signaling to Regulate Auxin Homeostasis in Arabidopsis. Plants 2013, 2, 726–749. [Google Scholar] [CrossRef] [Green Version]
- Stefanowicz, K.; Szymanska-chargot, M.; Truman, W.; Walerowski, P. Cell Enlargement and Loss of Cellular Integrity in Root Systems Are Mediated by Pectin Demethylation. Front. Plant Sci. 2021, 12, 711838. [Google Scholar] [CrossRef] [PubMed]
- Päsold, S.; Siegel, I.; Seidel, C.; Ludwig-Müller, J. Flavonoid Accumulation in Arabidopsis thaliana Root Galls Caused by the Obligate Biotrophic Pathogen Plasmodiophora brassicae. Mol. Plant Pathol. 2010, 11, 545–562. [Google Scholar] [CrossRef]
- Li, L.; Long, Y.; Li, H.; Wu, X. Comparative Transcriptome Analysis Reveals Key Pathways and Hub Genes in Rapeseed During the Early Stage of Plasmodiophora brassicae Infection. Front. Genet. 2020, 10, 1275. [Google Scholar] [CrossRef] [Green Version]
- Ludwig-Müller, J. Auxin Homeostasis, Signaling and Interaction with Other Growth Hormones during the Clubroot Disease of Brassicaceae. Plant Signal. Behav. 2014, 9, 9–12. [Google Scholar] [CrossRef] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Wei, X.; Xu, W.; Yuan, Y.; Yao, Q.; Zhao, Y.; Wang, Z.; Jiang, W.; Zhang, X. Genome-Wide Investigation of MicroRNAs and Their Targets in Brassica rapa ssp. pekinensis Root with Plasmodiophora brassicae Infection. Hortic. Plant J. 2016, 2, 209–216. [Google Scholar] [CrossRef]
- Li, Q.; Shah, N.; Zhou, X.; Wang, H.; Yu, W.; Luo, J.; Liu, Y.; Li, G.; Liu, C.; Zhang, C.; et al. Identification of Micro Ribonucleic Acids and Their Targets in Response to Plasmodiophora brassicae Infection in Brassica napus. Front. Plant Sci. 2021, 12, 734419. [Google Scholar] [CrossRef] [PubMed]
- Paul, P.; Chhapekar, S.S.; Rameneni, J.J.; Oh, S.H.; Dhandapani, V.; Subburaj, S.; Shin, S.; Ramchiary, N. MiR1885 Regulates Disease Tolerance Genes in Brassica rapa during Early Infection with Plasmodiophora brassicae. Int. J. Mol. Sci. 2021, 22, 9433. [Google Scholar] [CrossRef]
- Malinowski, R.; Truman, W.; Blicharz, S. Genius Architect or Clever Thief—How Plasmodiophora brassicae Reprograms Host Development to Establish a Pathogen-Oriented Physiological Sink. Mol. Plant Microbe Interact. 2019, 32, 1259–1266. [Google Scholar] [CrossRef] [Green Version]
- Gravot, A.; Richard, G.; Lime, T.; Lemarié, S.; Jubault, M.; Lariagon, C.; Lemoine, J.; Vicente, J.; Robert-Seilaniantz, A.; Holdsworth, M.J.; et al. Hypoxia Response in Arabidopsis Roots Infected by Plasmodiophora brassicae Supports the Development of Clubroot. BMC Plant Biol. 2016, 16, 251. [Google Scholar] [CrossRef] [PubMed]
- Aigu, Y.; Daval, S.; Gazengel, K.; Marnet, N.; Lariagon, C.; Laperche, A.; Legeai, F.; Manzanares-Dauleux, M.J.; Gravot, A. Multi-Omic Investigation of Low-Nitrogen Conditional Resistance to Clubroot Reveals Brassica napus Genes Involved in Nitrate Assimilation. Front. Plant Sci. 2022, 13, 790563. [Google Scholar] [CrossRef]
- Werner, T.; Schmülling, T. Cytokinin action in plant development. Curr. Opin. Plant Biol. 2009, 12, 527–538. [Google Scholar] [CrossRef]
- Shimotohno, A.; Aki, S.S.; Takahashi, N.; Umeda, M. Regulation of the Plant Cell Cycle in Response to Hormones and the Environment. Annu. Rev. Plant Biol. 2021, 72, 273–296. [Google Scholar] [CrossRef]
- Majda, M.; Robert, S. The Role of Auxin in Cell Wall Expansion. Int. J. Mol. Sci. 2018, 19, 951. [Google Scholar] [CrossRef] [Green Version]
- Staswick, P.E.; Serban, B.; Rowe, M.; Tiryaki, I.; Maldonado, M.T.; Maldonado, M.C.; Suza, W. Characterization of an Arabidopsis Enzyme Family That Conjugates Amino Acids to Indole-3-Acetic Acid. Plant Cell 2005, 17, 616–627. [Google Scholar] [CrossRef] [Green Version]
- Marowa, P.; Ding, A.; Kong, Y. Expansins: Roles in plant growth and potential applications in crop improvement. Plant Cell Rep. 2016, 35, 949–965. [Google Scholar] [CrossRef] [Green Version]
- Galindo-Gonzalez, L.; Manolii, V.; Hwang, S.; Strelkov, S.E. Response of Brassica napus to Plasmodiophora brassicae Involves Salicylic Acid-Mediated Immunity: An RNA-Seq-Based Study. Front. Plant Sci. 2020, 11, 1025. [Google Scholar] [CrossRef]
- Ludwig-Müller, J. Plant Defence—What Can We Learn from Clubroots? Australas. Plant Pathol. 2009, 38, 318–324. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef]
- Lemarié, S.; Robert-Seilaniantz, A.; Lariagon, C.; Lemoine, J.; Marnet, N.; Jubault, M.; Manzanares-Dauleux, M.J.; Gravot, A. Both the Jasmonic Acid and the Salicylic Acid Pathways Contribute to Resistance to the Biotrophic Clubroot Agent Plasmodiophora brassicae in Arabidopsis. Plant Cell Physiol. 2015, 56, 2158–2168. [Google Scholar] [CrossRef] [Green Version]
- Park, S.-G.; Noh, E.; Choi, S.; Choi, B.; Shin, I.-G.; Yoo, S.-I.; Lee, D.; Ji, S.; Kim, H.-S.; Hwang, Y.-J.; et al. Draft Genome Assembly and Transcriptome Dataset for European Turnip (Brassica rapa L. ssp. rapifera), ECD4 Carrying Clubroot Resistance. Front. Genet. 2021, 12, 651298. [Google Scholar] [CrossRef]
- Hejna, O.; Havlickova, L.; He, Z.; Bancroft, I.; Curn, V. Analysing the Genetic Architecture of Clubroot Resistance Variation in Brassica napus by Associative Transcriptomics. Mol. Breed. 2019, 39, 112. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Q.; Galindo-Gonzalez, L.; Manolii, V.; Hwang, S.; Strelkov, S.E. Comparative Transcriptome Analysis of Rutabaga (Brassica napus) Cultivars Indicates Activation of Salicylic Acid and Ethylene-Mediated Defenses in Response to Plasmodiophora brassicae. Int. J. Mol. Sci. 2020, 21, 8381. [Google Scholar] [CrossRef]
- Eulgem, T.; Somssich, I.E. Networks of WRKY Transcription Factors in Defense Signaling. Curr. Opin. Plant Biol. 2007, 10, 366–371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, F.; Zhang, X.; Yuan, Y. Root Transcriptome and Metabolome Profiling Reveal Key Phytohormone-Related Genes and Pathways Involved Clubroot Resistance in Brassica rapa L. Front. Plant Sci. 2021, 12, 759623. [Google Scholar] [CrossRef]
- Fu, P.; Piao, Y.; Zhan, Z.; Zhao, Y.; Pang, W.; Li, X. Transcriptome Profile of Brassica Rapa L. Reveals the Involvement of Jasmonic Acid, Ethylene, and Brassinosteroid Signaling Pathways in Clubroot Resistance. Agronomy 2019, 9, 589. [Google Scholar] [CrossRef] [Green Version]
- Knaust, A.; Ludwig-Müller, J. The Ethylene Signaling Pathway Is Needed to Restrict Root Gall Growth in Arabidopsis after Infection with the Obligate Biotrophic Protist Plasmodiophora brassicae. J. Plant Growth Regul. 2013, 32, 9–21. [Google Scholar] [CrossRef]
- Cao, T.; Srivastava, S.; Rahman, M.; Kav, N.; Hotte, N.; Deyholos, M.; Strelkov, S. Proteome-Level Changes in the Roots of Brassica napus as a Result of Plasmodiophora brassicae Infection. Plant Sci. 2007, 174, 97–115. [Google Scholar] [CrossRef]
- Ji, R.; Wang, Y.; Wang, X.; Liu, Y.; Shen, X.; Feng, H. Proteomic Analysis of the Interaction between Plasmodiophora brassicae and Chinese Cabbage (Brassica rapa L. ssp. pekinensis) at the Initial Infection Stage. Sci. Hortic. 2018, 233, 386–393. [Google Scholar] [CrossRef]
- Adhikary, D.; Mehta, D.; Uhrig, R.G.; Rahman, H.; Kav, N.N.V. A Proteome-Level Investigation into Plasmodiophora brassicae Resistance in Brassica napus Canola. Front. Plant Sci. 2022, 13, 860393. [Google Scholar] [CrossRef]
- Song, T.; Chu, M.; Lahlali, R.; Yu, F.; Peng, G. Shotgun Label-Free Proteomic Analysis of Clubroot (Plasmodiophora brassicae) Resistance Conferred by the Gene Rcr1 in Brassica rapa. Front. Plant Sci. 2016, 7, 1013. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lan, M.; Li, G.; Hu, J.; Yang, H.; Zhang, L.; Xu, X.; Liu, J. ITRAQ-Based Quantitative Analysis Reveals Proteomic Changes in Chinese Cabbage (Brassica rapa L.) in Response to Plasmodiophora brassicae Infection. Sci. Rep. 2019, 9, 12058. [Google Scholar] [CrossRef]
- Su, T.; Yu, S.; Wang, W.; Li, P.; Zhang, F.; Yu, Y.; Zhang, D.; Zhao, X. ITRAQ Analysis of Protein Profile during the Secondary Stage of Infection of Plasmodiophora brassicae in Chinese Cabbage (Brassica rapa). J. Plant Pathol. 2018, 100, 533–542. [Google Scholar] [CrossRef]
- Ikegami, H.; Naiki, T.; Ito, T.; Imuro, Y. Ultrastructural Growth Process of Plasmodiophora brassicae in Infected Cells of Chinese Cabbage and Turnip. Res. Bull. Fac. Agric. Gifu Univ. 1982, 46, 9–19. [Google Scholar]
- Jülke, S.; Ludwig-Müller, J. Modulation of Lipid Transfer Proteins Alters Clubroot Development in Arabidopsis thaliana. Acta Hortic. 2010, 867, 165–172. [Google Scholar] [CrossRef]
- Tu, J.; Bush, J.; Bonham-Smith, P.; Wei, Y. Live Cell Imaging of Plasmodiophora brassicae—Host Plant Interactions Based on a Two-Step Axenic Culture System. Microbiologyopen 2018, 8, e00765. [Google Scholar] [CrossRef] [Green Version]
- Pedras, M.S.C.; Zheng, Q.-A.; Strelkov, S. Metabolic Changes in Roots of the Oilseed Canola Infected with the Biotroph Plasmodiophora brassicae: Phytoalexins and Phytoanticipins. J. Agric. Food Chem. 2008, 56, 9949–9961. [Google Scholar] [CrossRef]
- Wagner, G.; Laperche, A.; Lariagon, C.; Marnet, N.; Renault, D.; Guitton, Y.; Bouchereau, A.; Delourme, R.; Manzanares-dauleux, M.J.; Gravot, A. Resolution of Quantitative Resistance to Clubroot into QTL-Specific Metabolic Modules. J. Exp. Bot. 2019, 70, 5375–5390. [Google Scholar] [CrossRef] [PubMed]
- Lan, M.E.I.; Hu, J.I.; Yang, H.O.; Zhang, L.I.; Xu, X.U.; He, J.I. Phytohormonal and Metabolism Analysis of Brassica rapa L. ssp. pekinensis with Different Resistance during Plasmodiophora brassicae Infection. Biocell 2020, 44, 751–767. [Google Scholar] [CrossRef]
- Hassani, M.A.; Durán, P.; Hacquard, S. Microbial interactions within the plant holobiont. Microbiome 2018, 6, 58. [Google Scholar] [CrossRef]
- Lahlali, R.; Peng, G. Suppression of Clubroot by Clonostachys rosea via Antibiosis and Induced Host Resistance. Plant Pathol. 2014, 63, 447–455. [Google Scholar] [CrossRef]
- Zhu, M.; He, Y.; Li, Y.; Ren, T.; Liu, H.; Huang, J.; Jiang, D. Two New Biocontrol Agents Against Clubroot Caused by Plasmodiophora brassicae. Front. Microbiol. 2020, 10, 3099. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Sun, T.; Zhao, S.; Pan, L.; Liu, H.; Tian, X. Physiological Change Alters Endophytic Bacterial Community in Clubroot of Tumorous Stem Mustard Infected by Plasmodiophora brassicae. BMC Microbiol. 2020, 20, 244. [Google Scholar] [CrossRef]
- Hittorf, M.; Letsch-Praxmarer, S.; Windegger, A.; Bass, D.; Kirchmair, M.; Neuhauser, S. Revised Taxonomy and Expanded Biodiversity of the Phytomyxea (Rhizaria, Endomyxa). J. Eukaryot. Microbiol. 2020, 67, 648–659. [Google Scholar] [CrossRef]
- Tamada, T.; Kondo, H. Biological and Genetic Diversity of Plasmodiophorid-Transmitted Viruses and Their Vectors. J Gen. Plant Pathol. 2013, 79, 307–320. [Google Scholar] [CrossRef]
- Balendres, M.A.; Tegg, R.S.; Wilson, C.R. Key Events in Pathogenesis of Spongospora Diseases in Potato: A Review. Australas. Plant Pathol. 2016, 45, 229–240. [Google Scholar] [CrossRef]
- Murúa, P.; Goecke, F.; Westermeier, R.; van West, P.; Küpper, F.C.; Neuhauser, S. Maullinia braseltonii Sp. Nov. (Rhizaria, Phytomyxea, Phagomyxida): A Cyst-Forming Parasite of the Bull Kelp Durvillaea Spp. (Stramenopila, Phaeophyceae, Fucales). Protist 2017, 168, 468–480. [Google Scholar] [CrossRef] [Green Version]
- Maier, I.; Parodi, E.; Westermeier, R.; Müller, D.G. Maullinia ectocarpii Gen. et Sp. Nov. (Plasmodiophorea), an Intracellular Parasite in Ectocarpus siliculosus (Ectocarpales, Phaeophyceae) and Other Filamentous Brown Algae. Protist 2000, 151, 225–238. [Google Scholar] [CrossRef] [Green Version]
- Mutasa-Göttgens, E.S.; Chwarzczynska, D.; Halsey, K.; Asher, M.J.C. Specific Polyclonal Antibodies for the Obligate Plant Parasite Polymyxa—A Targeted Recombinant DNA Approach. Plant Pathol. 2000, 49, 276–287. [Google Scholar] [CrossRef]
- Šubr, Z.W.; Kastirr, U.; Kühne, T. Subtractive cloning of DNA from Polymyxa graminis—An obligate parasitic plasmodiophorid. J. Phytopathol. 2002, 150, 564–568. [Google Scholar] [CrossRef]
- Desoignies, N.; Carbonell, J.; Moreau, J.S.; Conesa, A.; Dopazo, J.; Legrève, A. Molecular Interactions between Sugar Beet and Polymyxa betae during Its Life Cycle. Ann. Appl. Biol. 2014, 164, 244–256. [Google Scholar] [CrossRef]
- Burki, F.; Kudryavtsev, A.; Matz, M.V.; Aglyamova, G.V.; Bulman, S.; Fiers, M.; Keeling, P.J.; Pawlowski, J. Evolution of Rhizaria: New Insights from Phylogenomic Analysis of Uncultivated Protists. BMC Evol. Biol. 2010, 10, 377. [Google Scholar] [CrossRef]
- Ciaghi, S.; Neuhauser, S.; Schwelm, A. Draft Genome Resource for the Potato Powdery Scab Pathogen Spongospora subterranea. Mol. Plant Microbe Interact. 2018, 31, 1227–1229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Decroes, A.; Li, J.; Richardson, L.; Mutasa-Gottgens, E.; Lima-Mendez, G.; Mahillon, M.; Bragard, C.; Finn, R.D.; Legr, A. Genomics Metagenomics Approach for Polymyxa betae Genome Assembly Enables Comparative Analysis towards Deciphering the Intracellular Parasitic Lifestyle of the Plasmodiophorids. Genomics 2022, 114, 9–22. [Google Scholar] [CrossRef]
- Neuhauser, S.; Kirchmair, M.; Bulman, S.; Bass, D. Cross-Kingdom Host Shifts of Phytomyxid Parasites. BMC Evol. Biol. 2014, 14, 33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [Green Version]

| Method | Achievement | Reference |
|---|---|---|
| (Random) PCR cloning | Genomic DNA (fragments) | [28] Bulman 2007 |
| Subtractive cDNA library | 76 gene sequences from P. brassicae | [23,28] |
| Suppressive subtractive cDNA library | Ca 60 P. brassicae sequences | [25,26] |
| Dot blot and qPCR | Larger-scale expression profile of >100 genes during primary and secondary zoospore development | [29] |
| Microarray 1 | Role of cytokinins Early infection Partial resistance | [2,30,31] |
| Laser capture microdissection coupled to microarray | Role of brassinosteroids | [32] |
| RNAseq 1 | Host plant genes P. brassicae genes | [5,20,33,34,35] |
| miRNA expression | Host plant | [36] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ludwig-Müller, J. What Can We Learn from -Omics Approaches to Understand Clubroot Disease? Int. J. Mol. Sci. 2022, 23, 6293. https://doi.org/10.3390/ijms23116293
Ludwig-Müller J. What Can We Learn from -Omics Approaches to Understand Clubroot Disease? International Journal of Molecular Sciences. 2022; 23(11):6293. https://doi.org/10.3390/ijms23116293
Chicago/Turabian StyleLudwig-Müller, Jutta. 2022. "What Can We Learn from -Omics Approaches to Understand Clubroot Disease?" International Journal of Molecular Sciences 23, no. 11: 6293. https://doi.org/10.3390/ijms23116293
APA StyleLudwig-Müller, J. (2022). What Can We Learn from -Omics Approaches to Understand Clubroot Disease? International Journal of Molecular Sciences, 23(11), 6293. https://doi.org/10.3390/ijms23116293