Synthesis of Dolichols in Candida albicans Is Co-Regulated with Elongation of Fatty Acids
Abstract
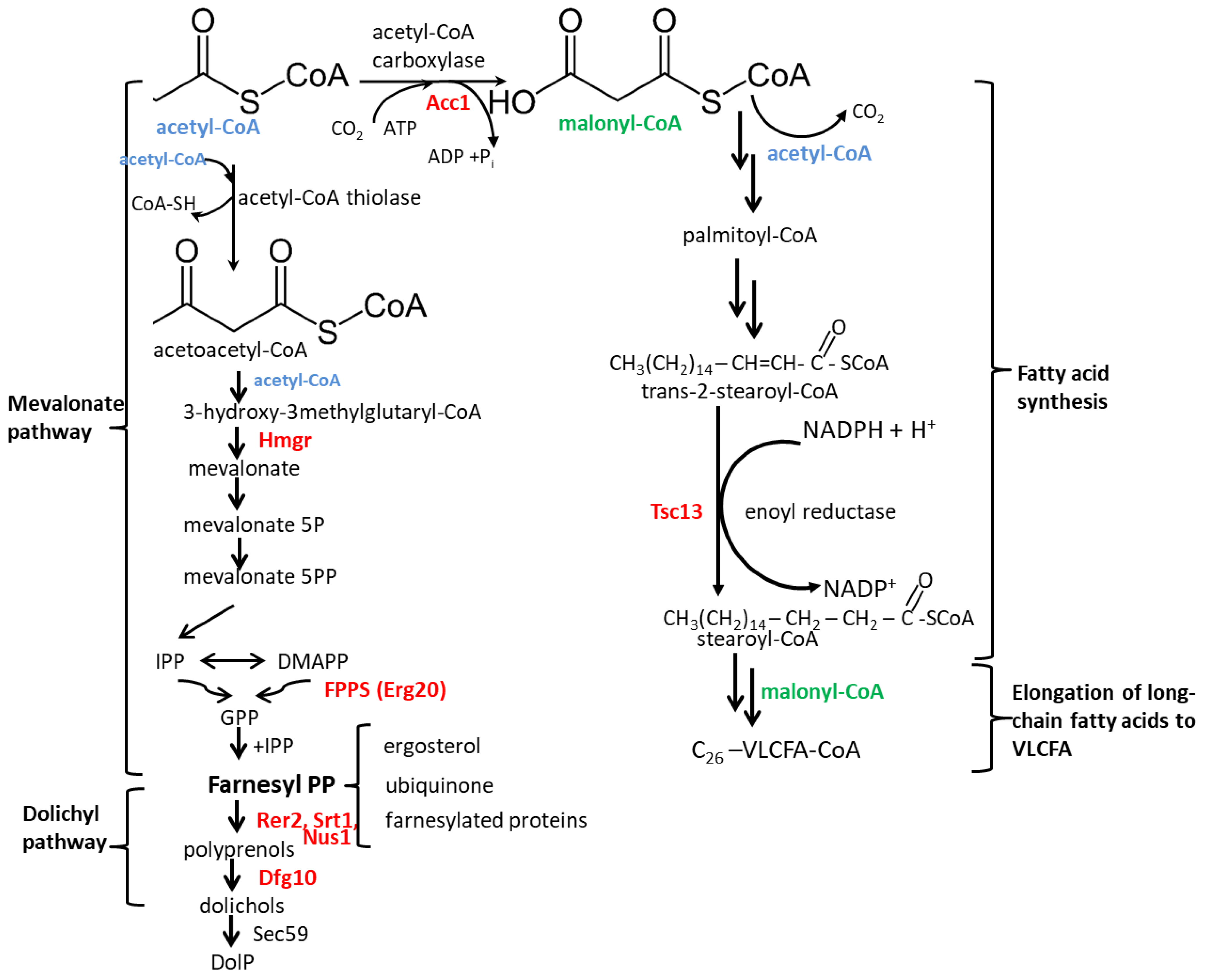
:1. Introduction
2. Results
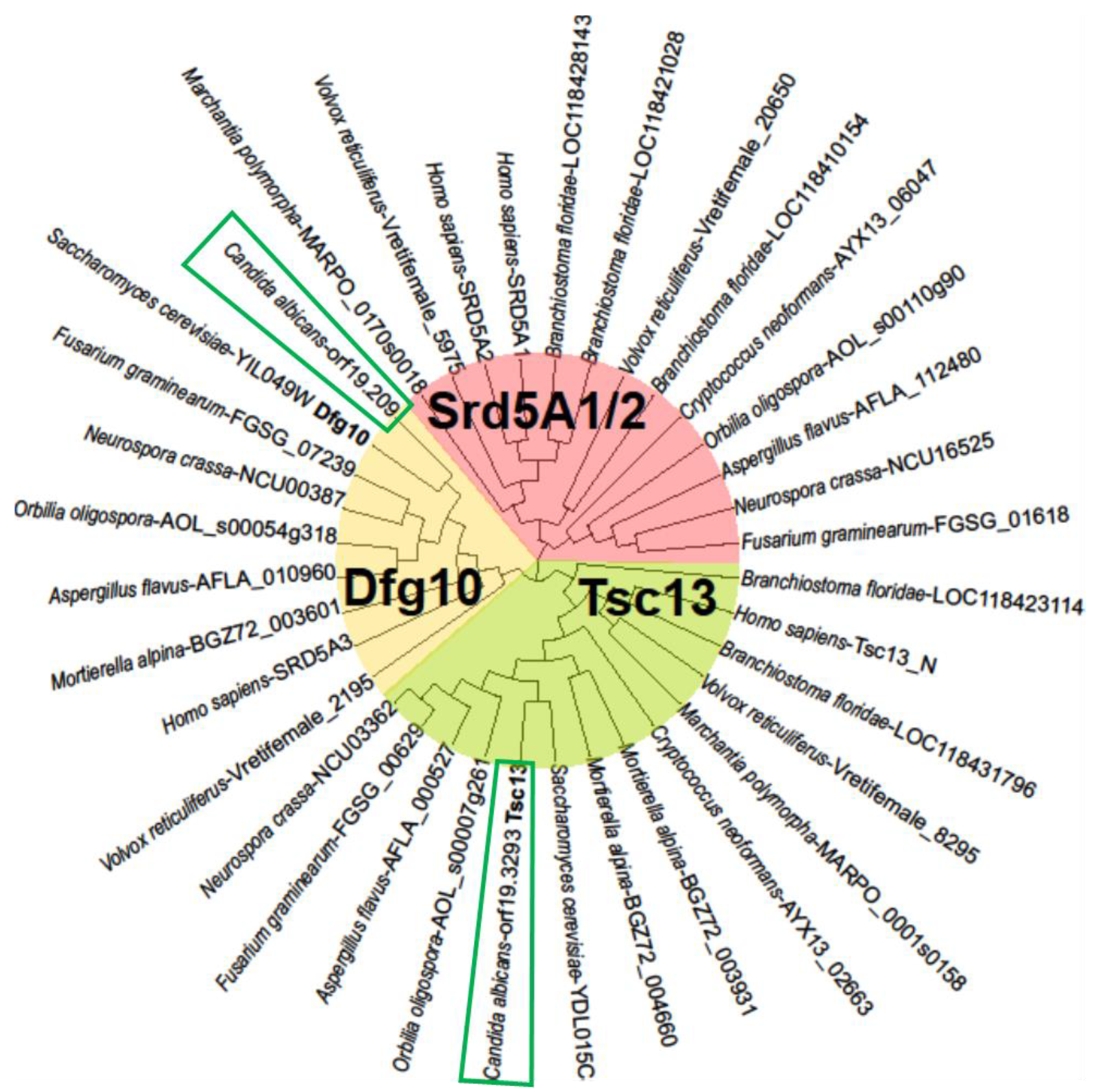
2.1. C. albicans Orthologues of the S. cerevisiae DFG10 and TSC13 Genes
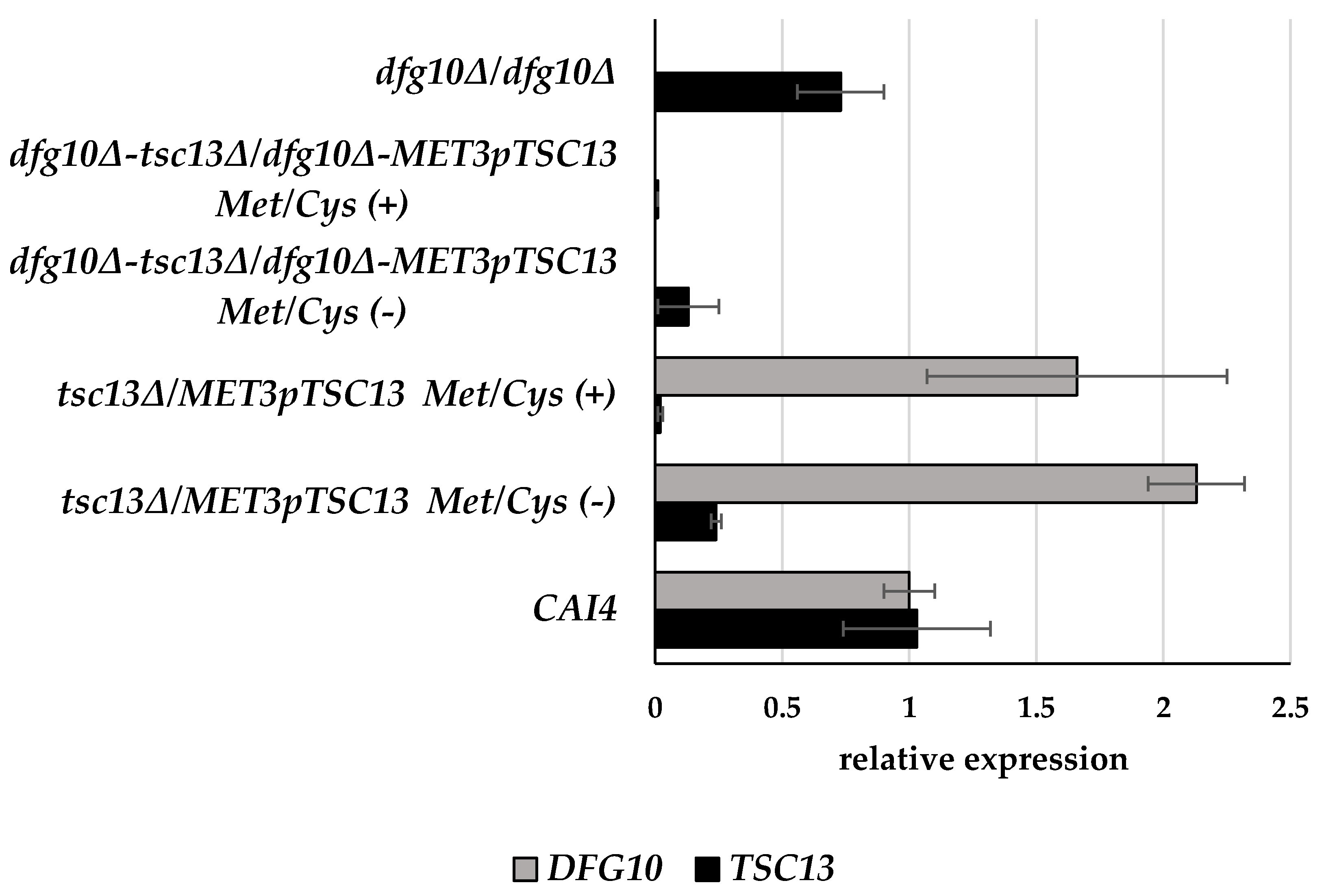
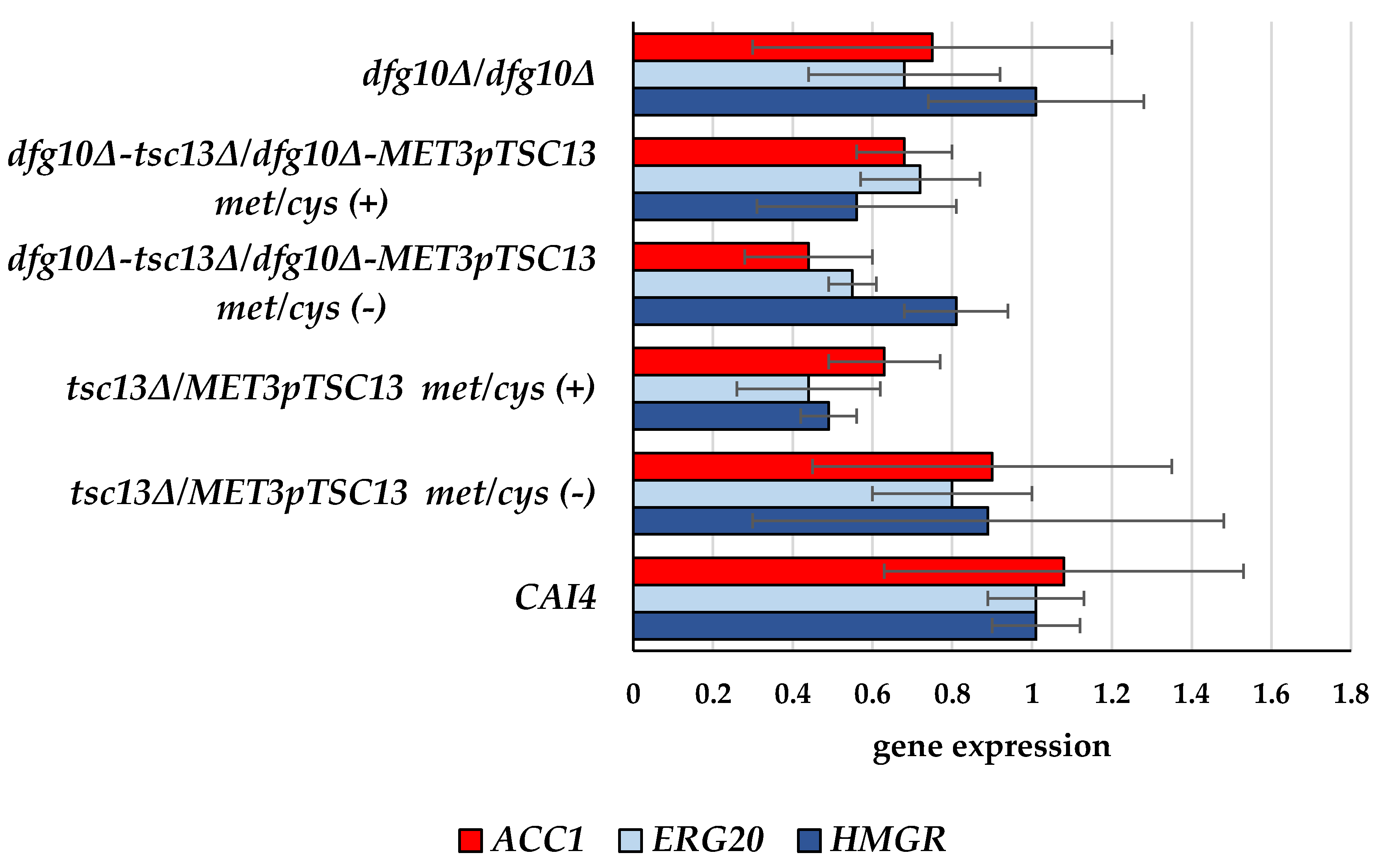
2.2. Gene Expression in DFG10 and TSC13 Mutants
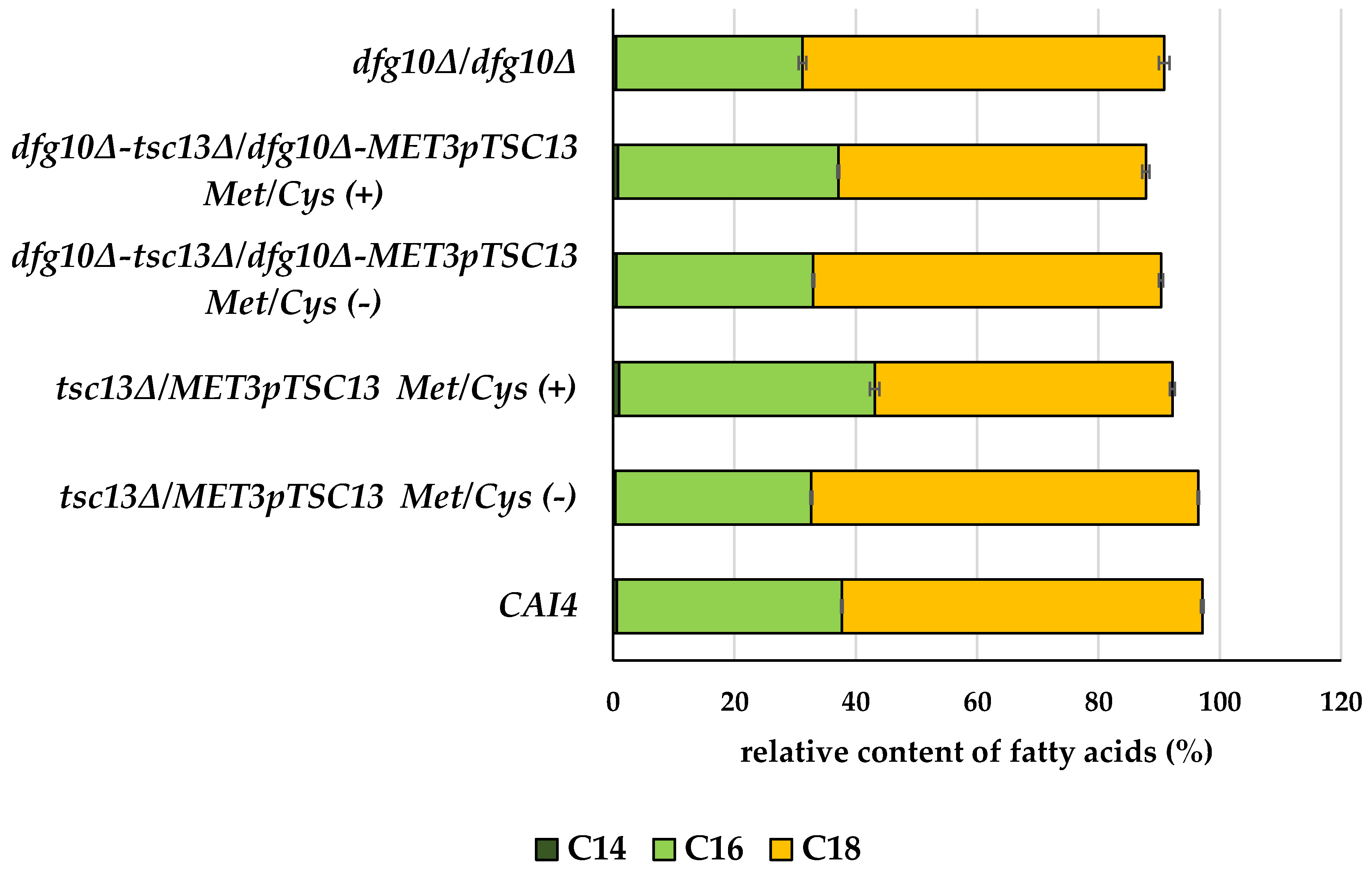
2.3. Fatty Acid Content in Membranes of DFG10 and TSC13 Mutants
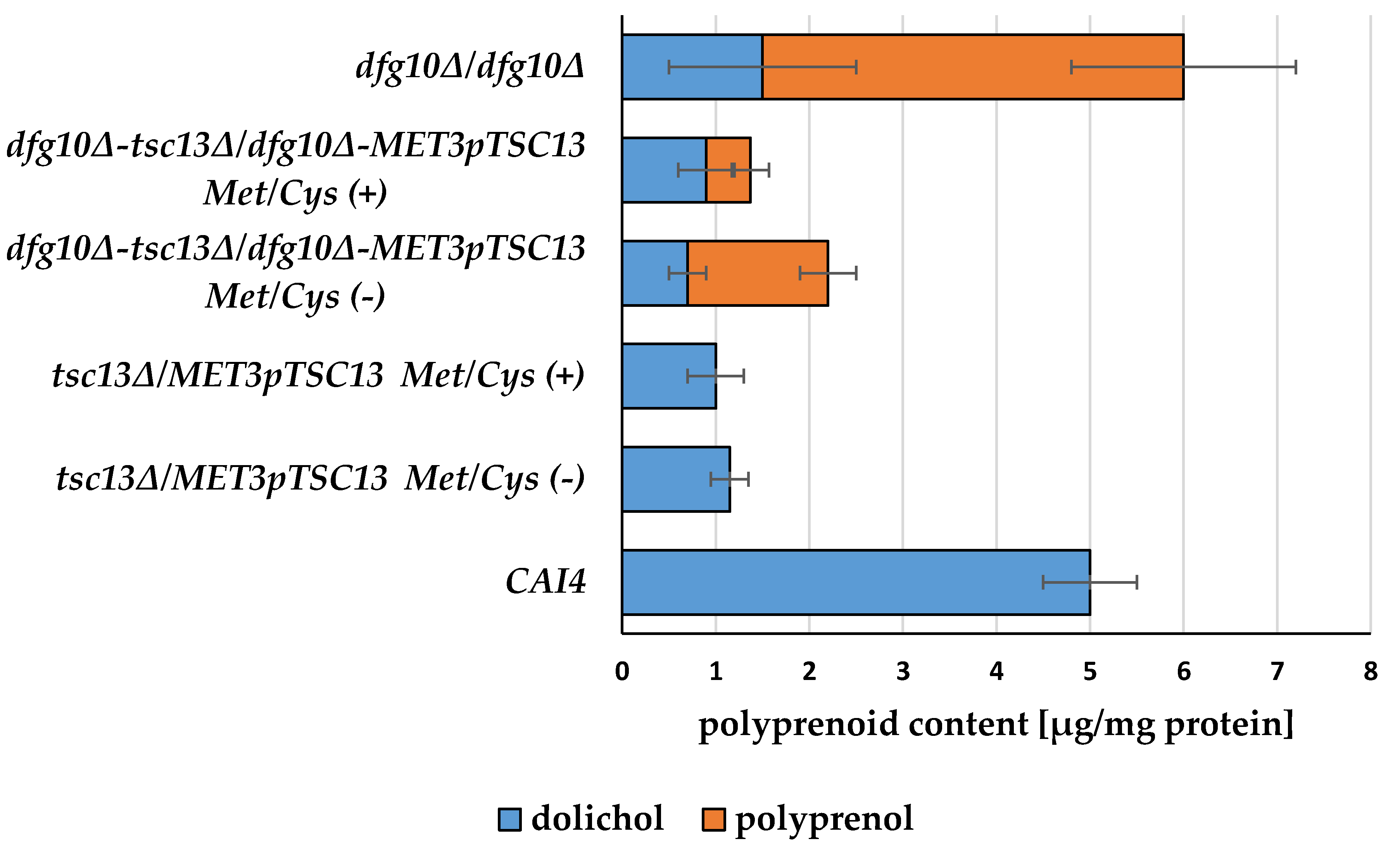
2.4. Defective Saturation of Polyprenols and Fatty Acid Elongation Alter Morphology of C. albicans
3. Discussion
4. Materials and Methods
4.1. Strains and Growth Conditions
4.2. Construction of Candida albicans Strains
4.3. RNA Isolation, RT-qPCR
4.4. Extraction and Purification of Polyisoprenoids
4.5. Lipid Analysis
4.6. Phylogenetic Analysi
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sagami, H.; Kurisaki, A.; Ogura, K. Formation of dolichol from dehydrodolichol is catalyzed by NADPH-dependent reductase localized in microsomes of rat liver. J. Biol. Chem. 1993, 268, 10109–10113. [Google Scholar] [CrossRef]
- Adair, W.L.J.; Cafmeyer, N. Characterization of the Saccharomyces cerevisiae cis-prenyltransferase required for dolichyl phosphate biosynthesis. Arch. Biochem. Biophys. 1987, 259, 589–596. [Google Scholar] [CrossRef]
- Sato, M.; Fujisaki, S.; Sato, K.; Nishimura, Y.; Nakano, A. Yeast Saccharomyces cerevisiae has two cis-prenyltransferases with different properties and localizations. Implication for their distinct physiological roles in dolichol synthesis. Genes Cells 2001, 6, 495–506. [Google Scholar] [CrossRef]
- Juchimiuk, M.; Orłowski, J.; Gawarecka, K.; Swiezewska, E.; Ernst, J.F.; Palamarczyk, G. Candida albicans cis-prenyltransferase Rer2 is required for protein glycosylation, cell wall integrity and hyphae formation. Fungal Genet. Biol. 2014, 69, 1–12. [Google Scholar] [CrossRef]
- Park, E.J.; Grabinska, K.A.; Guan, Z.; Stránecky, V.; Hartmannová, H.; Hodanova, K.; Baresova, V.; Sovova, J.; Levente, J.; Ondruskova, N.; et al. Mutation of Nogo-B receptor, a subunit of cis-prenyltransferase, causes a congenital disorder of glycosylation. Cell Metab. 2014, 20, 448–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Burton, W.A.; Scher, M.G.; Waechter, C.J. Enzymatic phosphorylation of dolichol in central nervous tissue. J. Biol. Chem. 1979, 254, 7129–7136. [Google Scholar] [CrossRef]
- Hemming, F.W. Glycosyl phosphopolyprenols. In Glycolipids; Wiegandt, L., Ed.; Elsevier Science Publishers: Amsterdam, The Netherlands, 1985; pp. 261–305. [Google Scholar]
- Schneiter, R.; Brügger, B.; Amann, C.M.; Prestwich, G.D.; Epand, R.F.; Zellnig, G.; Wieland, F.T.; Epand, R.M. Identification and biophysical characterization of a very-long-chain-fatty acid-substituted phosphatidylinositol in yeast subcellular membranes. Biochem. J. 2004, 381, 941–949. [Google Scholar] [CrossRef] [Green Version]
- Hunkeler, M.; Stuttfeld, E.; Hagmann, A.; Imseng, S.; Maier, T. The dynamic organization of fungal acetyl-CoA carboxylase. Nat. Commun. 2016, 7, 11196. [Google Scholar] [CrossRef] [Green Version]
- Kohlwein, S.D.; Eder, S.; Oh, C.-S.; Martin, C.E.; Gable, K.; Bacikova, D.; Dunn, T. Tsc13 is required for fatty acod elongation and localizes to a novel structure at the nuclear-vacuolar interface in Saccharomyces cerevisiae. Mol. Cell. Biol. 2001, 21, 109–125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tehlivests, O.; Scheuringer, K.; Kohlwein, S.D. Fatty acid synthesis and elongation in yeast. Biochim. Biophys. Acta 2007, 1771, 255–270. [Google Scholar] [CrossRef] [PubMed]
- Oh, C.-S.; Toke, D.A.; Mandala, S.; Martin, C.E. ELO2 and ELO3, homologues of the Saccharomyces cerevisiae ELO1 gene, function in fatty acid elongation and are required for sphingolipid formation. J. Biol. Chem. 1997, 272, 17376–17384. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rössler, H.; Rieck, C.; Delong, T.; Hoja, U.; Schweizer, E. Functional differentiation and selective inactivation of multiple Saccharomyces cerevisiae genes involved in very-long-chain fatty acid synthesis. Mol. Gen. Genomics 2003, 269, 290–298. [Google Scholar] [CrossRef]
- Erdbrügger, P.; Fröhlich, F. The role of very long chain fatty acids in yeast physiology and human diseases. Biol. Chem. 2021, 402, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2020, 49. [Google Scholar] [CrossRef] [PubMed]
- Stiles, A.R.; Russell, D.W. SRD5A3: A surprising role in glycosylation. Cell 2010, 142, 196–198. [Google Scholar] [CrossRef] [Green Version]
- Cantagrel, V.; Lefeber, D.J.; Ng, B.G.; Guan, Z.; Silhavy, J.L.; Bielas, S.L.; Lehle, L.; Hombauer, H.; Adamowicz, M.; Swiezewska, E.; et al. SRD5A3 is required for the conversion of polyprenol to dolichol, essential for N-linked protein glycosylation. Cell 2010, 142, 203–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mösch, H.-U.; Fink, G.R. Dissection of filamentous growth by transposon mutagenesis in Saccharomyces cerevisiae. Genetics 1997, 145, 671–684. [Google Scholar] [CrossRef]
- Behrens, N.H.; Leloir, L.F. Dolichol monophosphate glucose: An intermediate in glucose transfer in liver. Proc. Natl. Acad. Sci. USA 1970, 66, 153–159. [Google Scholar] [CrossRef] [Green Version]
- Hoja, U.; Wellein, C.; Greiner, E.; Schweizer, E. Pleiotropic phenotype of acetyl-CoA-carboxylase-defective yeast cells—Viability of a BPL1- amber mutation depending on its read through by normal tRNA(Gln) (CAG). Eur. J. Biochem. 1998, 254, 520–526. [Google Scholar] [CrossRef]
- Murthy, S.; Tong, H.; Hohl, R.J. Regulation of fatty acid synthesis by farnesyl pyrophosphate. J. Biol. Chem. 2005, 280, 41793–41804. [Google Scholar] [CrossRef] [Green Version]
- Care, R.S.; Trevethick, J.; Binley, K.M.; Sudbery, P.E. The MET3 promoter: A new tool for Candida albicans molecular genetics. Mol. Microbiol. 1999, 34, 792–798. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Kohler, J.; Fink, G.R. Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 1994, 266, 1723–1726. [Google Scholar] [CrossRef]
- Fonzi, W.A.; Irwin, M.Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 1993, 134, 717–728. [Google Scholar] [CrossRef]
- Chomczynski, P.; Sacchi, N. Single-step method of RNA isolation by acid guanidinum thiocyanate-phenol-chloroform extraction. Anal. Biochem. 1987, 162, 156–159. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Andersen, C.L.; Jensen, J.L.; Orntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [Green Version]
- Augustyniak, J.; Lenart, J.; Lipka, G.; Stepien, P.P.; Buzanska, L. Reference Gene Validation via RT-qPCR for human iPSC-derived neural stem cells and neural progenitors. Mol. Neurobiol. 2019, 56, 6820–6832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [Green Version]
- Stothard, P. The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques 2000, 28, 1102–1104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gawarecka, K.; Swiezewska, E. Analysis of plant polyisoprenoids. Methods Mol. Biol. 2014, 1153, 135–147. [Google Scholar]
- Bernat, P.; Nykiel-Szymańska, J.; Stolarek, P.; Słaba, M.; Szewczyk, R.; Różalska, S. 2,4-dichlorophenoxyacetic acid-induced oxidative stress: Metabolome and membrane modifications in Umbelopsis isabellina, a herbicide degrader. PLoS ONE 2018, 13, e01996772018. [Google Scholar] [CrossRef] [PubMed]
- Ichihara, K.; Fukabayashi, Y. Preparation of fatty acid methyl esters for gas-liquid chromatography. J. Lipid. Res. 2010, 51, 635–640. [Google Scholar] [CrossRef] [Green Version]
- Altschul, S.F.; Lipman, D.J. Protein database searches for multiple alignments. Proc. Natl. Acad. Sci. USA 1990, 87, 5509–5513. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Lefort, V.; Longueville, J.E.; Gascuel, O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef] [PubMed] [Green Version]

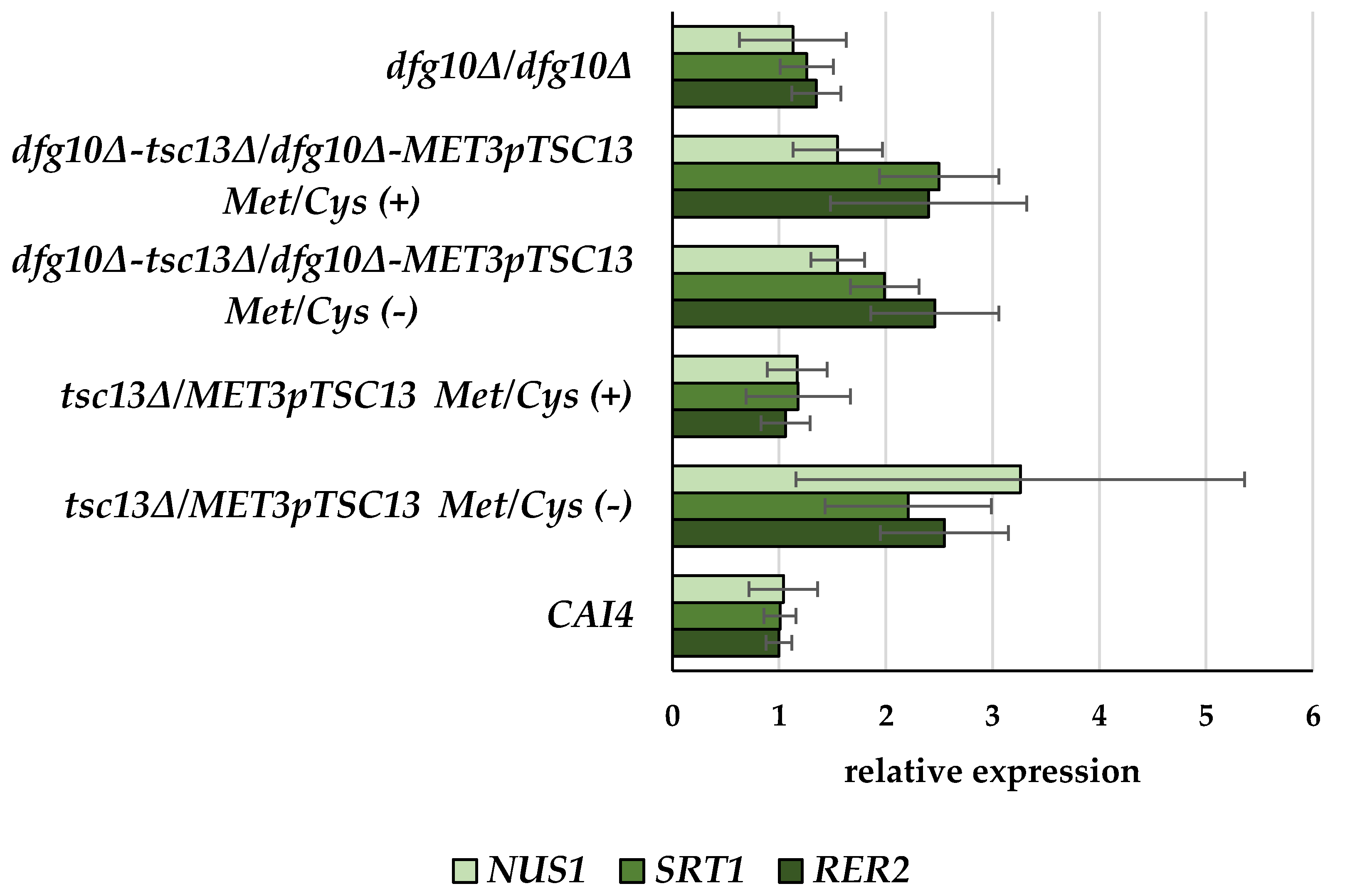

| Strain | Genotype | Source |
|---|---|---|
| CAI-4 | ura3Δ::im434/ura3Δ::imm434 | [26] |
| dfg10(URA3)/DFG10 | CAI4; dfg10Δ::hisG-URA3-hisG/DFG10 | This study |
| dfg10∆/DFG10 | CAI4; dfg10Δ::hisG/DFG10 | This study |
| dfg10(URA3)/dfg10∆ | CAI4 but dfg10Δ::hisG/dfg10Δ:hisG-URA3-hisG | This study |
| dfg10∆/dfg10∆ | CAI4; dfg10Δ:hisG/dfg10Δ::hisG | This study |
| tsc13(URA3)/TSC13 | CAI4; tsc13Δ::hisG-URA3-hisG/TSC13 | This study |
| tsc13∆/MET3pTSC13 | CAI4; tsc13Δ::hisG/MET3pTSC13 | This study |
| dfg10∆-tsc13∆/dfg10∆-MET3pTSC13 | CAI4; dfg10Δ:hisG/dfg10Δ::hisG- tsc13Δ::hisG/MET3pTSC13 | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Janik, A.; Perlińska-Lenart, U.; Gawarecka, K.; Augustyniak, J.; Bratek-Gerej, E.; Bernat, P.; Piłsyk, S.; Skalmowska, P.; Palamarczyk, G.; Swiezewska, E.; et al. Synthesis of Dolichols in Candida albicans Is Co-Regulated with Elongation of Fatty Acids. Int. J. Mol. Sci. 2022, 23, 409. https://doi.org/10.3390/ijms23010409
Janik A, Perlińska-Lenart U, Gawarecka K, Augustyniak J, Bratek-Gerej E, Bernat P, Piłsyk S, Skalmowska P, Palamarczyk G, Swiezewska E, et al. Synthesis of Dolichols in Candida albicans Is Co-Regulated with Elongation of Fatty Acids. International Journal of Molecular Sciences. 2022; 23(1):409. https://doi.org/10.3390/ijms23010409
Chicago/Turabian StyleJanik, Anna, Urszula Perlińska-Lenart, Katarzyna Gawarecka, Justyna Augustyniak, Ewelina Bratek-Gerej, Przemysław Bernat, Sebastian Piłsyk, Patrycja Skalmowska, Grażyna Palamarczyk, Ewa Swiezewska, and et al. 2022. "Synthesis of Dolichols in Candida albicans Is Co-Regulated with Elongation of Fatty Acids" International Journal of Molecular Sciences 23, no. 1: 409. https://doi.org/10.3390/ijms23010409
APA StyleJanik, A., Perlińska-Lenart, U., Gawarecka, K., Augustyniak, J., Bratek-Gerej, E., Bernat, P., Piłsyk, S., Skalmowska, P., Palamarczyk, G., Swiezewska, E., & Kruszewska, J. S. (2022). Synthesis of Dolichols in Candida albicans Is Co-Regulated with Elongation of Fatty Acids. International Journal of Molecular Sciences, 23(1), 409. https://doi.org/10.3390/ijms23010409








