Global Changes of 5-mC/5h-mC Ratio and Methylation of Adiponectin and Leptin Gene in Placenta Depending on Mode of Delivery
Abstract
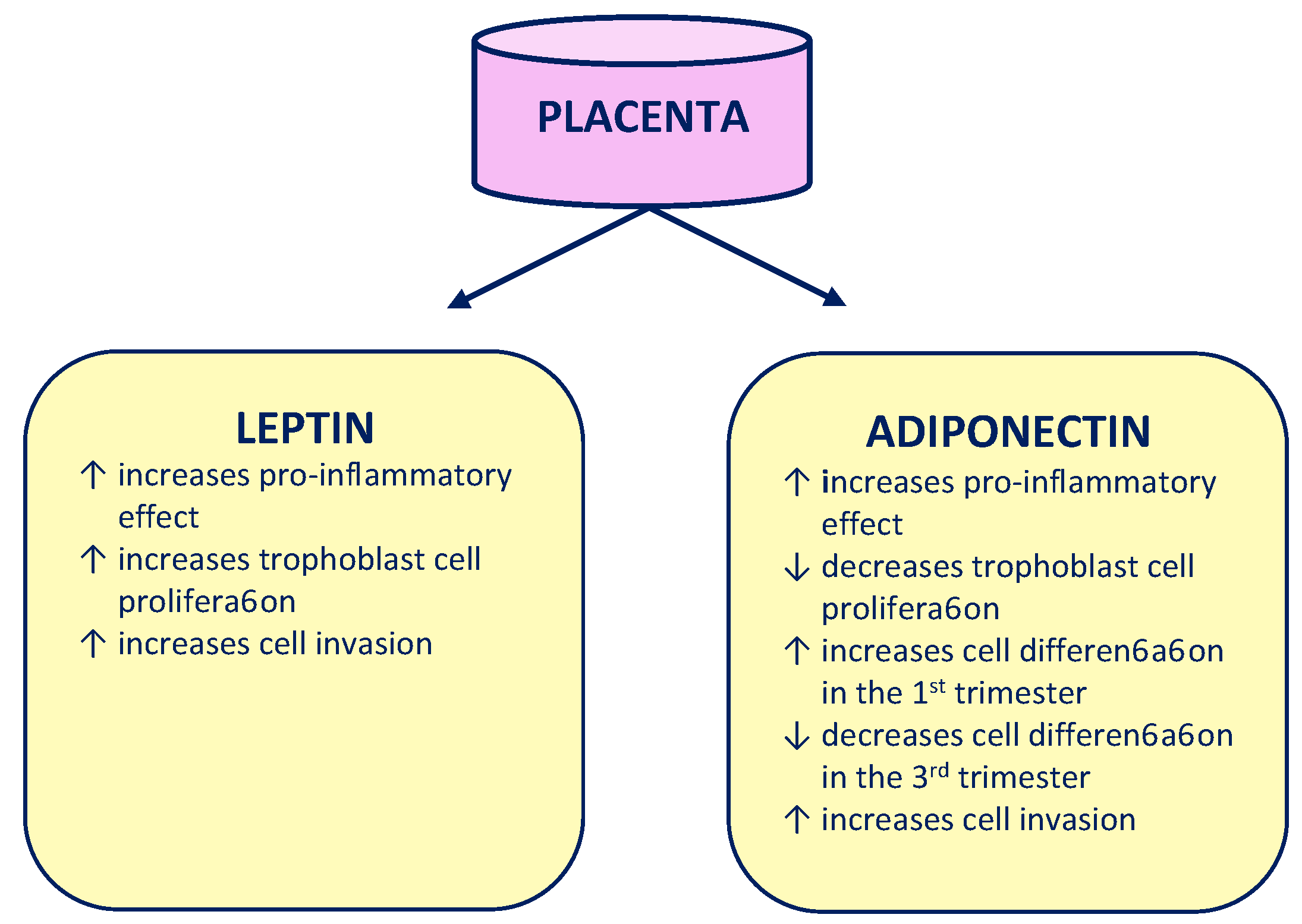
1. Introduction
2. Results
2.1. Characteristics of the Study Group
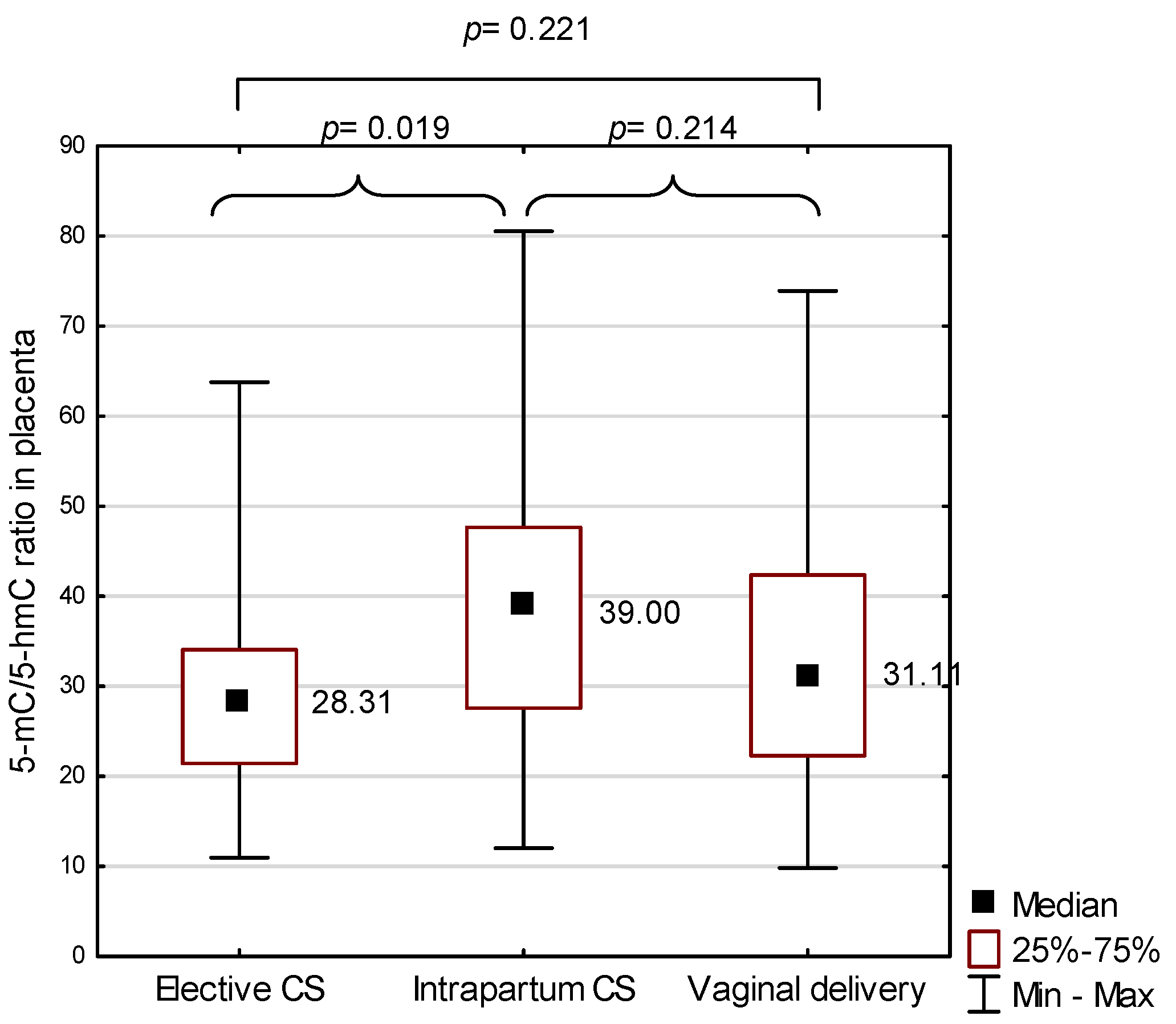
2.2. 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine (5-hmC) Ratio in the Placenta in the Study Group
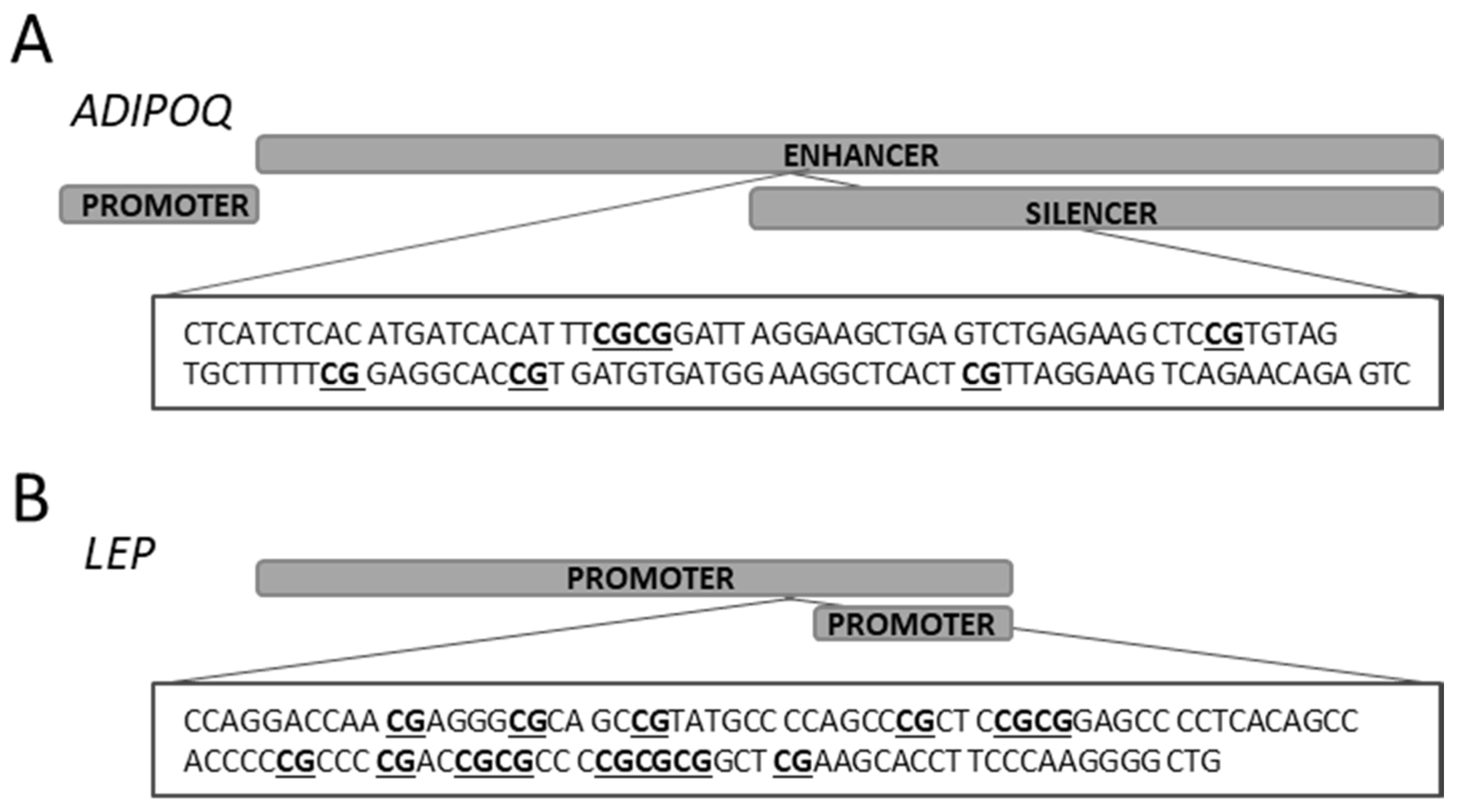
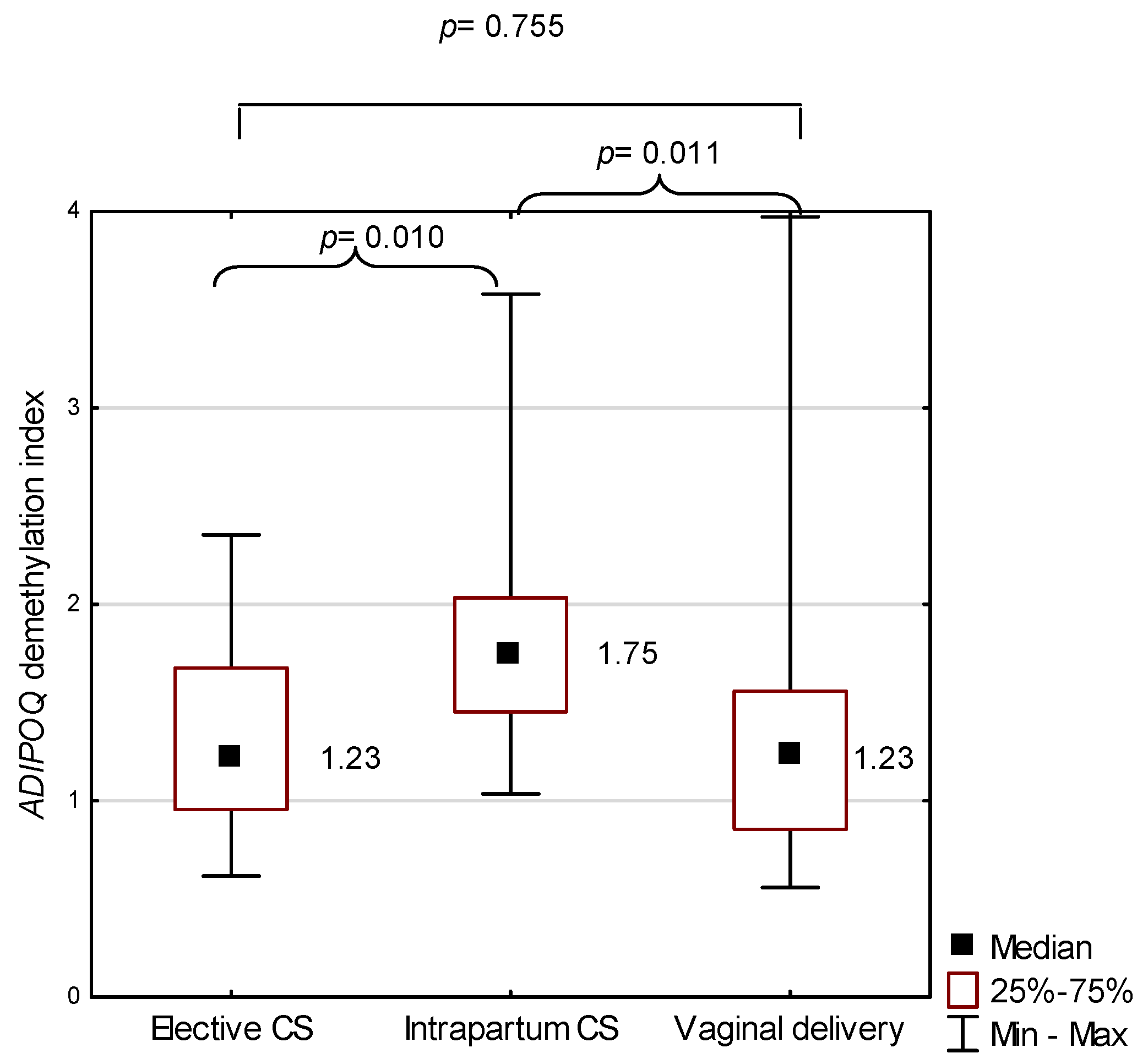
2.3. ADIPOQ in the Study Group
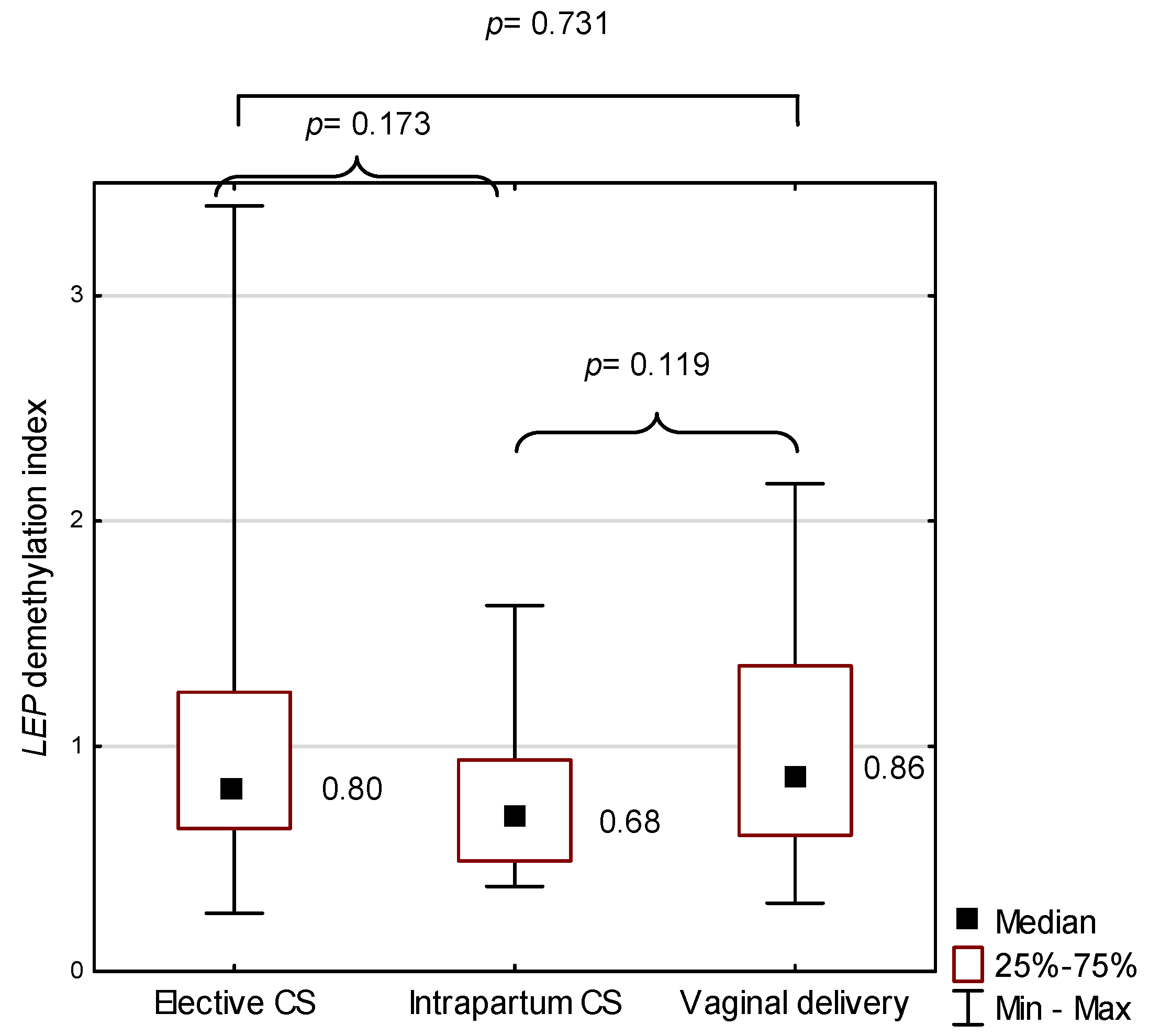
2.4. LEP in the Study Group
2.5. Mutual Correlations between Global Locus-Specific Methylation Status for ADIPOQ and LEP Demethylation Index in the Study Group
3. Discussion
4. Materials and Methods
4.1. Study Group
4.2. DNA Extraction and Sodium Bisulfite Conversion
4.3. Methylation-Specific Real-Time qPCR
4.4. Global DNA Methylation Status
4.5. Statistical Analysis
- The Kruskal–Wallis H test for the comparison of continuous variables among the three modes of birth;
- Fisher’s exact test for the comparison of categorical variables among the three modes of birth;
- The Mann–Whitney U test for the comparison of 5-mC/5-hmC ratio in the placenta and ADIPOQ and LEP between two modes of birth in pairs, between genders, or for epidural analgesia versus none;
- Spearman’s correlation coefficient r for the correlation of 5-mC/5-hmC ratio in the placenta, ADIPOQ vs. LEP, 5-mC/5-hmC ratio in the placenta, and ADIPOQ/LEP with numerical maternal or offspring outcomes.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Betrán, A.P.; Ye, J.; Moller, A.B.; Zhang, J.; Gülmezoglu, A.M.; Torloni, M.R. The increasing trend in caesarean section rates: Global, regional and national estimates: 1990–2014. PLoS ONE 2016, 11, e0148343. [Google Scholar] [CrossRef]
- Kuhle, S.; Tong, O.S.; Woolcott, C.G. Association between caesarean section and childhood obesity: A systematic review and meta-analysis. Obes. Rev. 2015, 16, 295–303. [Google Scholar] [CrossRef]
- Keag, O.E.; Norman, J.E.; Stock, S.J. Long-term risks and benefits associated with cesarean delivery for mother, baby, and subsequent pregnancies: Systematic review and meta-analysis. PLoS Med. 2018, 15, e1002494. [Google Scholar] [CrossRef] [PubMed]
- Ouchi, N.; Parker, J.L.; Lugus, J.J.; Walsh, K. Adipokines in inflammation and metabolic disease. Nat. Rev. Immunol. 2011, 11, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.V.; Scherer, P.E. Adiponectin, the past two decades. J. Mol. Cell Biol. 2016, 8, 93–100. [Google Scholar] [CrossRef]
- Sagawa, N.; Yura, S.; Itoh, H.; Kakui, K.; Takemura, M.; Nuamah, M.A.; Ogawa, Y.; Masuzaki, H.; Nakao, K.; Fujii, S. Possible role of placental leptin in pregnancy: A review. Endocrine 2002, 19, 65–71. [Google Scholar] [CrossRef]
- Señarís, R.; Garcia-Caballero, T.; Casabiell, X.; Gallego, R.; Castro, R.; Considine, R.V.; Dieguez, C.; Casanueva, F.F. Synthesis of leptin in human placenta. Endocrinology 1997, 138, 4501–4504. [Google Scholar] [CrossRef]
- Ashworth, C.J.; Hoggard, N.; Thomas, L.; Mercer, J.G.; Wallace, J.M.; Lea, R.G. Placental leptin. Rev. Reprod. 2000, 5, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Castellucci, M.; De Matteis, R.; Meisser, A.; Cancello, R.; Monsurrò, V.; Islami, D.; Sarzani, R.; Marzioni, D.; Cinti, S.; Bischof, P. Leptin modulates extracellular matrix molecules and metalloproteinases: Possible implications for trophoblast invasion. Mol. Hum. Reprod. 2000, 6, 951–958. [Google Scholar] [CrossRef]
- Chardonnens, D.; Cameo, P.; Aubert, M.L.; Pralong, F.P.; Islami, D.; Campana, A.; Gaillard, R.C.; Bischof, P. Modulation of human cytotrophoblastic leptin secretion by interleukin-1alpha and 17beta-oestradiol and its effect on HCG secretion. Mol. Hum. Reprod. 1999, 5, 1077–1082. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.M. Leptin and the endocrine control of energy balance. Nat. Metab. 2019, 1, 754–764. [Google Scholar] [CrossRef] [PubMed]
- Farley, D.M.; Choi, J.; Dudley, D.J.; Li, C.; Jenkins, S.L.; Myatt, L.; Nathanielsz, P.W. Placental amino acid transport and placental leptin resistance in pregnancies complicated by maternal obesity. Placenta 2010, 31, 718–724. [Google Scholar] [CrossRef] [PubMed]
- McDonald, E.A.; Wolfe, M.W. The pro-inflammatory role of adiponectin at the maternal-fetal interface. Am. J. Reprod. Immunol. 2011, 66, 128–136. [Google Scholar] [CrossRef] [PubMed]
- Benaitreau, D.; Dos Santos, E.; Leneveu, M.-C.; Alfaidy, N.; Feige, J.-J.; de Mazancourt, P.; Pecquery, R.; Dieudonné, M.-N. Effects of adiponectin on human trophoblast invasion. J. Endocrinol. 2010, 207, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, E.; Duval, F.; Vialard, F.; Dieudonné, M.-N. The roles of leptin and adiponectin at the fetal-maternal interface in humans. Horm. Mol. Biol. Clin. Investig. 2015, 24, 47–63. [Google Scholar] [CrossRef]
- Bianco-Miotto, T.; Mayne, B.T.; Buckberry, S.; Breen, J.; Rodriguez Lopez, C.M.; Roberts, C.T. Recent progress towards understanding the role of DNA methylation in human placental development. Reproduction 2016, 152, R23–R30. [Google Scholar] [CrossRef]
- Nugent, B.M.; Bale, T.L. The omniscient placenta: Metabolic and epigenetic regulation of fetal programming. Front. Neuroendocrinol. 2015, 39, 28–37. [Google Scholar] [CrossRef]
- Vlahos, A.; Mansell, T.; Saffery, R.; Novakovic, B. Human placental methylome in the interplay of adverse placental health, environmental exposure, and pregnancy outcome. PLoS Genet. 2019, 15, e1008236. [Google Scholar] [CrossRef]
- Nelissen, E.C.M.; van Montfoort, A.P.A.; Dumoulin, J.C.M.; Evers, J.L.H. Epigenetics and the placenta. Hum. Reprod. Update 2011, 17, 397–417. [Google Scholar] [CrossRef] [PubMed]
- Robinson, W.P.; Price, E.M. The human placental methylome. Cold Spring Harb. Perspect. Med. 2015, 5, a023044. [Google Scholar] [CrossRef]
- Virani, S.; Dolinoy, D.C.; Halubai, S.; Jones, T.R.; Domino, S.E.; Rozek, L.S.; Nahar, M.S.; Padmanabhan, V. Delivery type not associated with global methylation at birth. Clin. Epigenetics 2012, 4, 8. [Google Scholar] [CrossRef]
- Li, L.-C.; Dahiya, R. MethPrimer: Designing primers for methylation PCRs. Bioinformatics 2002, 18, 1427–1431. [Google Scholar] [CrossRef] [PubMed]
- Novakovic, B.; Saffery, R. DNA methylation profiling highlights the unique nature of the human placental epigenome. Epigenomics 2010, 2, 627–638. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, D.I.; Blair, J.D.; Lott, P.; Yu, H.O.K.; Hong, D.; Crary, F.; Ashwood, P.; Walker, C.; Korf, I.; Robinson, W.P.; et al. The human placenta methylome. Proc. Natl. Acad. Sci. USA 2013, 110, 6037–6042. [Google Scholar] [CrossRef] [PubMed]
- Dahlen, H.G.; Kennedy, H.P.; Anderson, C.M.; Bell, A.F.; Clark, A.; Foureur, M.; Ohm, J.E.; Shearman, A.M.; Taylor, J.Y.; Wright, M.L.; et al. The EPIIC hypothesis: Intrapartum effects on the neonatal epigenome and consequent health outcomes. Med. Hypotheses 2013, 80, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Słabuszewska-Jóźwiak, A.; Włodarczyk, M.; Ciebiera, M.; Zwolińska, J.; Wojtyła, C.; Nowicka, G.; Jakiel, G.; Raczkiewicz, D. Placental DNA methylation in caesarean sections—A pilot study. Arch. Med. Sci. 2020. [Google Scholar] [CrossRef]
- Bachman, M.; Uribe-Lewis, S.; Yang, X.; Williams, M.; Murrell, A.; Balasubramanian, S. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat. Chem. 2014, 6, 1049–1055. [Google Scholar] [CrossRef]
- Sun, M.; Song, M.M.; Wei, B.; Gao, Q.; Li, L.; Yao, B.; Chen, L.; Lin, L.; Dai, Q.; Zhou, X.; et al. 5-Hydroxymethylcytosine-mediated alteration of transposon activity associated with the exposure to adverse in utero environments in human. Hum. Mol. Genet. 2016, 25, 2208–2219. [Google Scholar] [CrossRef]
- Schlinzig, T.; Johansson, S.; Gunnar, A.; Ekström, T.J.; Norman, M. Epigenetic modulation at birth—Altered DNA-methylation in white blood cells after Caesarean section. Acta Paediatr. 2009, 98, 1096–1099. [Google Scholar] [CrossRef]
- Franz, M.B.; Poterauer, M.; Elhenicky, M.; Stary, S.; Birner, P.; Vinatzer, U.; Husslein, P.; Streubel, B.; Husslein, H. Global and single gene DNA methylation in umbilical cord blood cells after elective caesarean: A pilot study. Eur. J. Obstet. Gynecol. Reprod. Biol. 2014, 179, 121–124. [Google Scholar] [CrossRef]
- Bouchard, L.; Hivert, M.-F.; Guay, S.-P.; St-Pierre, J.; Perron, P.; Brisson, D. Placental adiponectin gene DNA methylation levels are associated with mothers’ blood glucose concentration. Diabetes 2012, 61, 1272–1280. [Google Scholar] [CrossRef]
- Nogues, P.; Dos Santos, E.; Jammes, H.; Berveiller, P.; Arnould, L.; Vialard, F.; Dieudonné, M.-N. Maternal obesity influences expression and DNA methylation of the adiponectin and leptin systems in human third-trimester placenta. Clin. Epigenetics 2019, 11, 20. [Google Scholar] [CrossRef]
- Mitsuya, K.; Parker, A.N.; Liu, L.; Ruan, J.; Vissers, M.C.M.; Myatt, L. Alterations in the placental methylome with maternal obesity and evidence for metabolic regulation. PLoS ONE 2017, 12, e0186115. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Yamada, T.; Abe, K.; Okamura, K.; Kamura, H.; Akaishi, R.; Minakami, H.; Nakabayashi, K.; Hata, K. Increased epigenetic alterations at the promoters of transcriptional regulators following inadequate maternal gestational weight gain. Sci. Rep. 2015, 5, 14224. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, D.; Ouidir, M.; Workalemahu, T.; Zeng, X.; Tekola-Ayele, F. Placental DNA methylation changes associated with maternal prepregnancy BMI and gestational weight gain. Int. J. Obes. 2020, 44, 1406–1416. [Google Scholar] [CrossRef]
- Kim, A.Y.; Park, Y.J.; Pan, X.; Shin, K.C.; Kwak, S.-H.; Bassas, A.F.; Sallam, R.M.; Park, K.S.; Alfadda, A.A.; Xu, A.; et al. Obesity-induced DNA hypermethylation of the adiponectin gene mediates insulin resistance. Nat. Commun. 2015, 6, 7585. [Google Scholar] [CrossRef] [PubMed]
- Ott, R.; Stupin, J.H.; Melchior, K.; Schellong, K.; Ziska, T.; Dudenhausen, J.W.; Henrich, W.; Rancourt, R.C.; Plagemann, A. Alterations of adiponectin gene expression and DNA methylation in adipose tissues and blood cells are associated with gestational diabetes and neonatal outcome. Clin. Epigenetics 2018, 10, 131. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 2012, 13, 484–492. [Google Scholar] [CrossRef]
- Jara, A.; Dreher, M.; Porter, K.; Christian, L.M. The association of maternal obesity and race with serum adipokines in pregnancy and postpartum: Implications for gestational weight gain and infant birth weight. Brain Behav. Immun. Health 2020, 3, 100053. [Google Scholar] [CrossRef]
- Gambino, Y.P.; Pérez Pérez, A.; Dueñas, J.L.; Calvo, J.C.; Sánchez-Margalet, V.; Varone, C.L. Regulation of leptin expression by 17beta-estradiol in human placental cells involves membrane associated estrogen receptor alpha. Biochim. Biophys. Acta 2012, 1823, 900–910. [Google Scholar] [CrossRef] [PubMed]
- Hauguel-de Mouzon, S.; Lepercq, J. Placental leptin and pregnancy pathologies. Gynecol. Obstet. Fertil. 2001, 29, 534–537. [Google Scholar] [CrossRef]
- Mansell, T.; Ponsonby, A.-L.; Collier, F.; Burgner, D.; Pezic, A.; Vuillermin, P.; Ryan, J.; Saffery, R. Methylation of the LEP gene promoter in blood at 12 months and BMI at 4 years of age-a population-based cohort study. Int. J. Obes. 2020, 44, 842–847. [Google Scholar] [CrossRef] [PubMed]
- Hauguel-de Mouzon, S.; Lepercq, J.; Catalano, P. The known and unknown of leptin in pregnancy. Am. J. Obstet. Gynecol. 2006, 194, 1537–1545. [Google Scholar] [CrossRef] [PubMed]
- Aye, I.L.M.H.; Powell, T.L.; Jansson, T. Review: Adiponectin--the missing link between maternal adiposity, placental transport and fetal growth? Placenta 2013, 34, S40–S45. [Google Scholar] [CrossRef] [PubMed]
- Akirav, E.M.; Lebastchi, J.; Galvan, E.M.; Henegariu, O.; Akirav, M.; Ablamunits, V.; Lizardi, P.M.; Herold, K.C. Detection of cell death in diabetes using differentially methylated circulating DNA. Proc. Natl. Acad. Sci. USA 2011, 108, 19018–19023. [Google Scholar] [CrossRef] [PubMed]
| Variable, Parameter | Unit or Category | Total Group (N = 70) | Elective CS (N = 30) | Intrapartum CS (N = 13) | Vaginal Delivery (N = 27) | p1 |
|---|---|---|---|---|---|---|
| Maternal age, Me (IQR) | years | 32 (28–35) | 34 (31–37) | 29 (27–34) | 31 (28–33) | 0.036 |
| Gestational age (weeks), n (%) | 37 | 6 (8.57) | 2 (6.67) | 1 (7.69) | 3 (11.11) | 0.026 |
| 38 | 11 (15.71) | 5 (16.67) | 2 (15.38) | 4 (14.81) | ||
| 39 | 32 (45.71) | 20 (66.67) | 4 (30.77) | 8 (29.63) | ||
| 40 | 9 (12.86) | 1 (3.33) | 1 (7.69) | 7 (25.93) | ||
| 41 | 12 (17.14) | 2 (6.67) | 5 (38.46) | 5 (18.52) | ||
| Parity, n (%) | 1 | 38 (54.29) | 13 (43.33) | 11 (84.62) | 14 (51.85) | 0.092 |
| 2 | 29 (41.43) | 16 (53.33) | 2 (15.38) | 11 (40.74) | ||
| 3 | 3 (4.29) | 1 (3.33) | 0 (0.00) | 2 (7.41) | ||
| Pre-gestational BMI, Me (IQR) | kg/m2 | 22.89 (20.82–25.01) | 23.58 (20.90–25.46) | 21.48 (19.47–22.31) | 22.86 (21.26–24.91) | 0.127 |
| Intragestational BMI, Me (IQR) | kg/m2 | 28.49 (25.95–30.80) | 29.10 (26.93–30.85) | 25.95 (25.71–29.30) | 28.26 (26.64–30.45) | 0.476 |
| Intragestational weight gain, Me (IQR) | kg | 16 (11–19) | 16 (12–20) | 17 (14–19) | 15 (11–18) | 0.551 |
| % | 25.00 (18.33–31.75) | 24.23 (19.05–28.17) | 32.08 (25.00–33.93) | 22.22 (16.92–29.23) | 0.129 | |
| Newborn weight, Me (IQR) | kg | 3.58 (3.30–3.85) | 3.53 (3.32–3.90) | 3.60 (3.45–3.80) | 3.65 (3.30–3.90) | 0.962 |
| Newborn gender, n (%) | male | 32 (45.71) | 13 (43.33) | 8 (61.54) | 11 (40.74) | 0.470 |
| female | 38 (54.29) | 17 (56.67) | 5 (38.46) | 16 (59.26) | ||
| 1 min Apgar score, n (%) | 8 | 2 (2.86) | 1 (3.33) | 1 (7.69) | 0 (0.00) | 0.066 |
| 9 | 10 (14.29) | 7 (23.33) | 2 (15.38) | 1 (3.70) | ||
| 10 | 58 (82.86) | 22 (73.33) | 10 (76.92) | 26 (96.30) | ||
| 5 min Apgar score, n (%) | 9 | 3 (4.29) | 2 (6.67) | 0 (0.00) | 1 (3.70) | 0.999 |
| 10 | 67 (95.71) | 28 (93.33) | 13 (100.00) | 26 (96.30) | ||
| Duration of uterine contraction 2, Me (IQR) | hours | - | - | - | 5.17 (3.42–7.67) | - |
| Analgesia, n (%) | spinal for cesarean section or epidural for vaginal delivery | 59 (84.29) | 30 (100.00) | 13 (100.00) | 16 (59.26) | - |
| Variable | Method | Total Group (N = 70) | Elective CS (N = 30) | Intrapartum CS (N = 13) | Vaginal Delivery (N = 27) | ||||
|---|---|---|---|---|---|---|---|---|---|
| Test | p | Test | p | Test | p | Test | p | ||
| Maternal age (years) | r | −0.008 | 0.946 | 0.347 | 0.060 | −0.434 | 0.139 | 0.172 | 0.390 |
| Gestational age (weeks) | r | 0.105 | 0.388 | 0.028 | 0.88 | 0.020 | 0.948 | 0.096 | 0.634 |
| Parity | r | −0.133 | 0.273 | 0.196 | 0.299 | −0.114 | 0.711 | −0.308 | 0.118 |
| Pregestational BMI (kg/m2) | r | −0.084 | 0.491 | 0.206 | 0.275 | −0.121 | 0.694 | −0.233 | 0.243 |
| Intragestational BMI (kg/m) | r | 0.047 | 0.701 | 0.317 | 0.088 | 0.143 | 0.641 | −0.162 | 0.421 |
| Intragestational weight gain (kg) | r | 0.238 | 0.047 | 0.122 | 0.522 | 0.520 | 0.069 | 0.096 | 0.632 |
| Intragestational weight gain (%) | r | 0.212 | 0.078 | −0.034 | 0.858 | 0.512 | 0.074 | 0.150 | 0.455 |
| Newborn weight (kg) | r | 0.093 | 0.444 | 0.355 | 0.054 | −0.174 | 0.571 | 0.015 | 0.940 |
| Newborn gender | Z | −1.438 | 0.150 | −0.942 | 0.346 | −2.196 | 0.028 | 0.444 | 0.657 |
| 1 min Apgar score | r | −0.112 | 0.357 | −0.271 | 0.147 | 0.056 | 0.856 | −0.050 | 0.803 |
| 5 min Apgar score | r | −0.072 | 0.556 | −0.216 | 0.251 | na | - | −0.050 | 0.803 |
| Epidural analgesia 1 | Z | - | - | - | - | - | - | −0.493 | 0.622 |
| Uterine contraction (duration in hours) 1 | r | - | - | - | - | - | - | 0.160 | 0.426 |
| Target Gene | Target CpG Status | Primer Name | Primer Sequence | Product Size (bp) | Ta (°C) | Number of CpG Sites | Reference: Amplified Region |
|---|---|---|---|---|---|---|---|
| ADIPOQ | Methylated | ADIPOQ_M3 ADIPOQ_M4 | 5′–TATTTTATATGATTATATTTCGCGG 5′–AACTCTATTCTAACTTCCTAACGAA | 121 | 57 | 6 | NG_021140.1: 8205..8327 nt |
| Unmethylated | ADIPOQ_U3 ADIPOQ_U4 | 5′–TTTATTTTATATGATTATATTTTGTGG 5′–AACTCTATTCTAACTTCCTAACAAA | 123 | 57 | |||
| LEP | Methylated | LEP_M1 LEP_M2 | 5′–TAGGATTAACGAGGGCGTAGTC 5′–AACCCCTTAAAAAAATACTTCGAA | 111 | 60 | 14 | NG_007450.1: 4586..4698 nt |
| Unmethylated | LEP_U1 LEP_U2 | 5′–TTAGGATTAATGAGGGTGTAGTTGT 5′–CAACCCCTTAAAAAAATACTTCAAA | 113 | 60 |
| Variable | Method | Total Group (N = 70) | Elective CS (N = 30) | Intrapartum CS (N = 13) | Vaginal Delivery (N = 27) | ||||
|---|---|---|---|---|---|---|---|---|---|
| Test | p | Test | p | Test | p | Test | p | ||
| Maternal age (years) | r | −0.115 | 0.340 | −0.216 | 0.251 | −0.041 | 0.893 | 0.309 | 0.117 |
| Gestational age (weeks) | r | 0.002 | 0.988 | −0.145 | 0.445 | 0.316 | 0.293 | −0.113 | 0.575 |
| Parity | r | −0.240 | 0.044 | −0.212 | 0.262 | 0.342 | 0.253 | −0.249 | 0.211 |
| Pregestational BMI (kg/m2) | r | −0.080 | 0.516 | −0.270 | 0.149 | 0.385 | 0.194 | 0.085 | 0.672 |
| Intragestational BMI (kg/m) | r | 0.021 | 0.855 | −0.195 | 0.303 | 0.523 | 0.067 | 0.192 | 0.336 |
| Intragestational weight gain (kg) | r | 0.200 | 0.097 | 0.221 | 0.240 | 0.190 | 0.534 | 0.172 | 0.390 |
| Intragestational weight gain (%) | r | 0.177 | 0.142 | 0.193 | 0.308 | 0.066 | 0.830 | 0.162 | 0.421 |
| Newborn weight (kg) | r | −0.009 | 0.944 | 0.186 | 0.324 | 0.135 | 0.660 | −0.114 | 0.571 |
| Newborn gender | Z | 1.002 | 0.316 | 0.146 | 0.884 | 0.001 | 0.999 | 2.245 | 0.025 |
| 1 min Apgar score | r | 0.033 | 0.087 | 0.242 | 0.198 | −0.268 | 0.376 | 0.101 | 0.617 |
| 5 min Apgar score | r | 0.182 | 0.133 | 0.208 | 0.269 | na | - | 0.101 | 0.617 |
| Epidural analgesia 1 | Z | - | - | - | - | - | - | 1.900 | 0.057 |
| Uterine contraction (duration in hours) 1 | r | - | - | - | - | - | - | 0.307 | 0.119 |
| Variable | Method | Total Group (N = 70) | Elective CS (N = 30) | Intrapartum CS (N = 13) | Vaginal Delivery (N = 27) | ||||
|---|---|---|---|---|---|---|---|---|---|
| Test | p | Test | p | Test | p | Test | p | ||
| Maternal age (years) | r | −0.059 | 0.625 | −0.090 | 0.635 | −0.036 | 0.907 | 0.022 | 0.913 |
| Gestational age (weeks) | r | −0.056 | 0.644 | 0.035 | 0.855 | −0.270 | 0.372 | 0.026 | 0.899 |
| Parity | r | 0.294 | 0.013 | −0.024 | 0.899 | −0.228 | 0.454 | 0.606 | 0.001 |
| Pregestational BMI (kg/m2) | r | −0.103 | 0.397 | −0.347 | 0.060 | 0.267 | 0.378 | −0.306 | 0.120 |
| Intragestational BMI (kg/m) | r | −0.104 | 0.392 | −0.307 | 0.099 | 0.294 | 0.329 | −0.183 | 0.360 |
| Intragestational weight gain (kg) | r | 0.020 | 0.867 | 0.264 | 0.158 | −0.218 | 0.474 | −0.013 | 0.948 |
| Intragestational weight gain (%) | r | 0.022 | 0.854 | 0.276 | 0.140 | −0.305 | 0.310 | 0.143 | 0.476 |
| Newborn weight (kg) | r | 0.087 | 0.474 | 0.022 | 0.910 | 0.171 | 0.577 | 0.140 | 0.485 |
| Newborn gender | Z | 0.318 | 0.750 | 0.167 | 0.867 | 0.000 | 1.000 | −0.419 | 0.675 |
| 1 min Apgar score | r | 0.047 | 0.702 | 0.056 | 0.768 | 0.179 | 0.559 | −0.252 | 0.205 |
| 5 min Apgar score | r | 0.063 | 0.605 | 0.309 | 0.097 | na | - | −0.252 | 0.205 |
| Epidural analgesia 1 | Z | - | - | - | - | - | - | −2.245 | 0.025 |
| Uterine contraction (duration in hours) 1 | r | - | - | - | - | - | - | −0.395 | 0.041 |
| Variable | Total Group (N = 70) | Elective CS (N = 30) | Intrapartum CS (N = 13) | Vaginal Delivery (N = 27) | ||||
|---|---|---|---|---|---|---|---|---|
| r | p | r | p | r | p | r | p | |
| 5-mC/5-hmC ratio in the placenta and ADIPOQ | 0.144 | 0.235 | 0.147 | 0.437 | −0.236 | 0.437 | 0.190 | 0.343 |
| 5-mC/5-hmC ratio in the placenta and LEP | −0.024 | 0.841 | 0.072 | 0.706 | −0.077 | 0.803 | 0.004 | 0.983 |
| ADIPOQ and LEP | −0.156 | 0.198 | 0.226 | 0.230 | −0.027 | 0.929 | −0.456 | 0.017 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Słabuszewska-Jóźwiak, A.; Malinowska, M.; Kloska, A.; Jakóbkiewicz-Banecka, J.; Gujski, M.; Bojar, I.; Raczkiewicz, D.; Jakiel, G. Global Changes of 5-mC/5h-mC Ratio and Methylation of Adiponectin and Leptin Gene in Placenta Depending on Mode of Delivery. Int. J. Mol. Sci. 2021, 22, 3195. https://doi.org/10.3390/ijms22063195
Słabuszewska-Jóźwiak A, Malinowska M, Kloska A, Jakóbkiewicz-Banecka J, Gujski M, Bojar I, Raczkiewicz D, Jakiel G. Global Changes of 5-mC/5h-mC Ratio and Methylation of Adiponectin and Leptin Gene in Placenta Depending on Mode of Delivery. International Journal of Molecular Sciences. 2021; 22(6):3195. https://doi.org/10.3390/ijms22063195
Chicago/Turabian StyleSłabuszewska-Jóźwiak, Aneta, Marcelina Malinowska, Anna Kloska, Joanna Jakóbkiewicz-Banecka, Mariusz Gujski, Iwona Bojar, Dorota Raczkiewicz, and Grzegorz Jakiel. 2021. "Global Changes of 5-mC/5h-mC Ratio and Methylation of Adiponectin and Leptin Gene in Placenta Depending on Mode of Delivery" International Journal of Molecular Sciences 22, no. 6: 3195. https://doi.org/10.3390/ijms22063195
APA StyleSłabuszewska-Jóźwiak, A., Malinowska, M., Kloska, A., Jakóbkiewicz-Banecka, J., Gujski, M., Bojar, I., Raczkiewicz, D., & Jakiel, G. (2021). Global Changes of 5-mC/5h-mC Ratio and Methylation of Adiponectin and Leptin Gene in Placenta Depending on Mode of Delivery. International Journal of Molecular Sciences, 22(6), 3195. https://doi.org/10.3390/ijms22063195








