Identification of DNA Damage Repair-Associated Prognostic Biomarkers for Prostate Cancer Using Transcriptomic Data Analysis
Abstract
:1. Introduction
2. Results
2.1. Identification of DDR Genes Associated with Overall Survival in Prostate Cancer
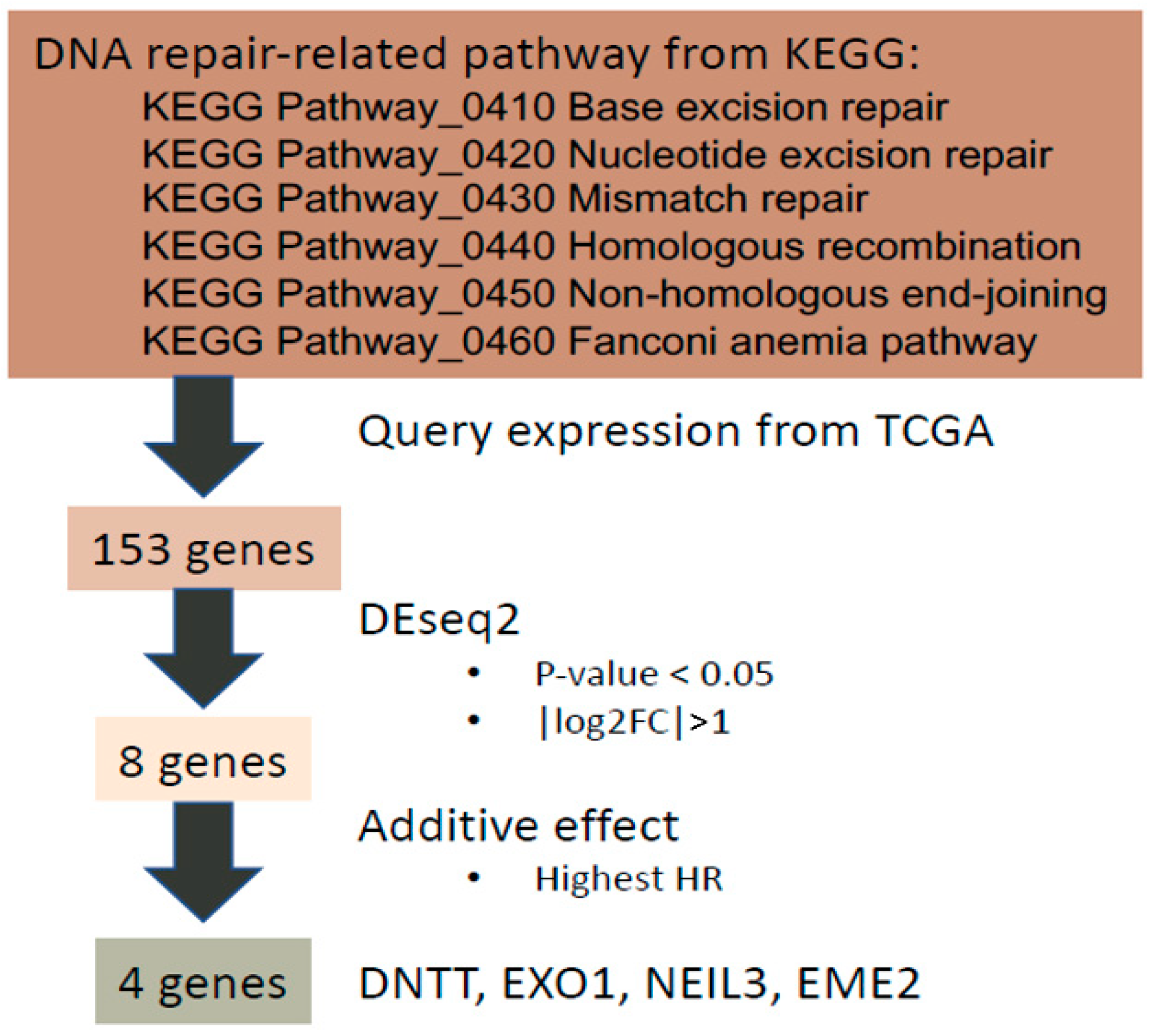
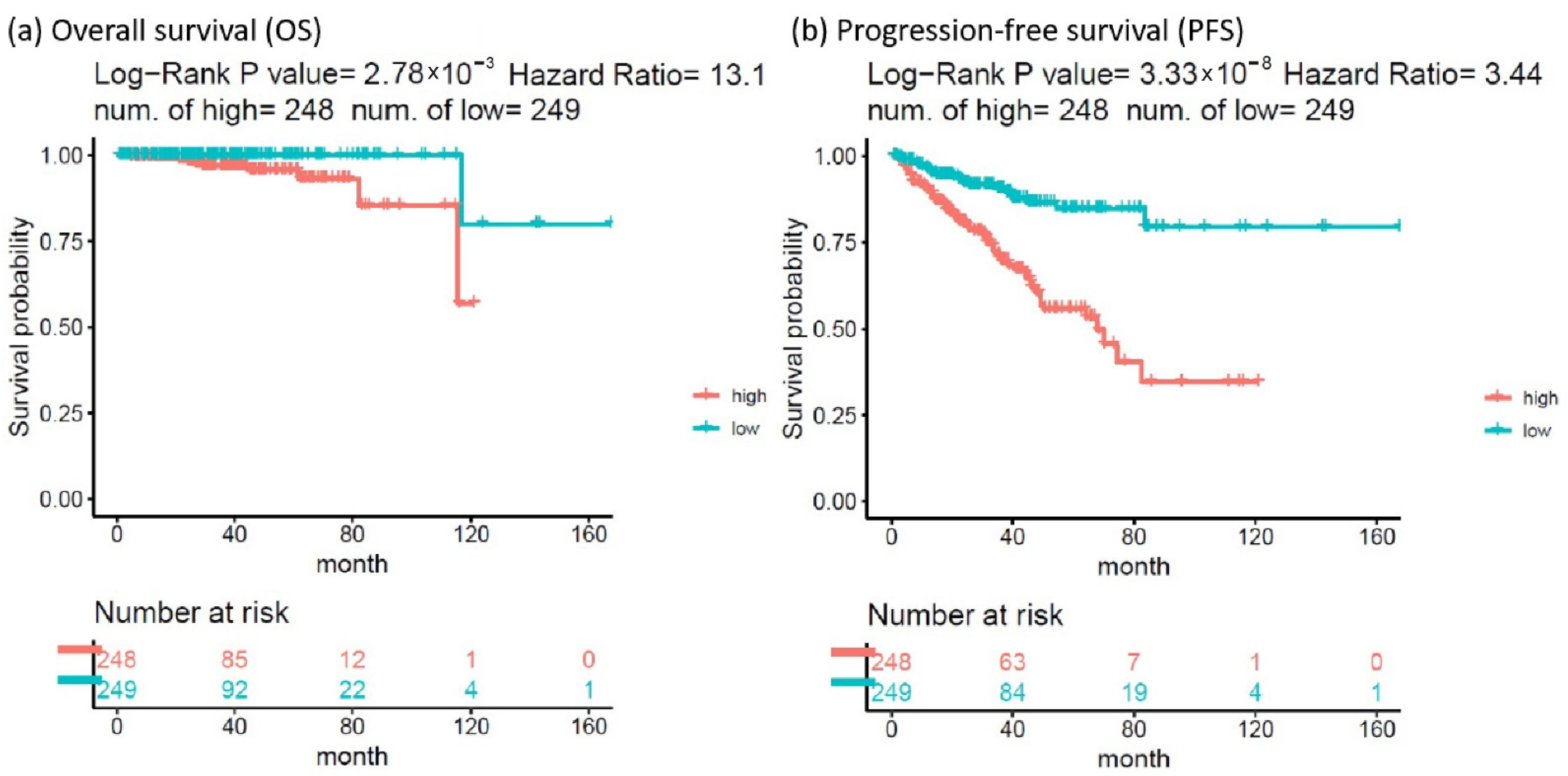
2.2. Identification of a Four-Gene Panel as Prognostic Markers for PC Survival
2.3. Multivariate Cox Regression Analysis
2.4. Expression of the Four-Gene Panel Based on Luminal and Basal Subtypes of PC
3. Discussion
4. Materials and Methods
4.1. Patient Clinical Information and Gene Expression Data Source
4.2. DNA-Repair Related Gene Selection
4.3. Differential Expression Analysis
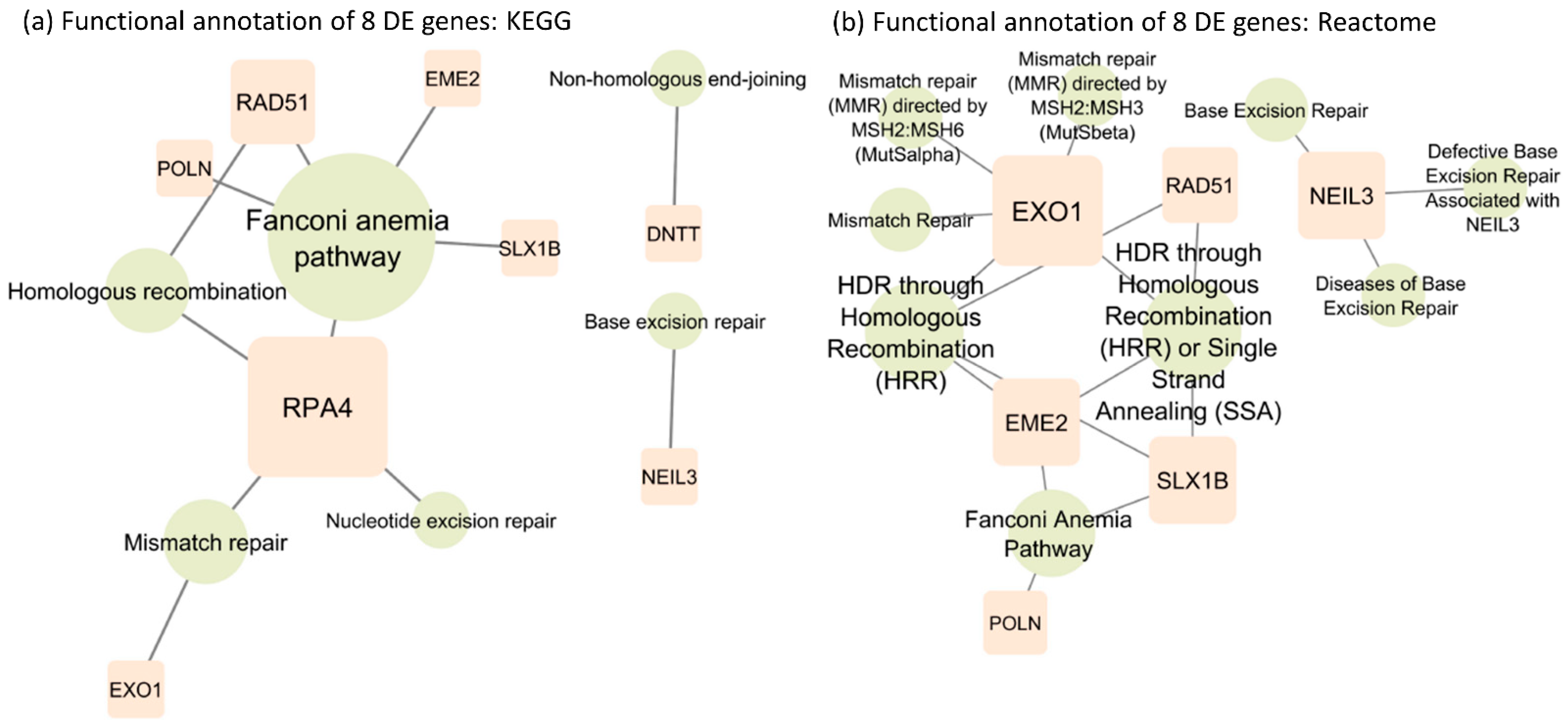
4.4. Functional Annotation
4.5. Additive Effect Analysis
4.6. Survival Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- De Bono, J.S.; Logothetis, C.J.; Molina, A.; Fizazi, K.; North, S.; Chu, L.; Chi, K.N.; Jones, R.J.; Goodman, O.B., Jr.; Saad, F.; et al. Abiraterone and increased survival in metastatic prostate cancer. N. Engl. J. Med. 2011, 364, 1995–2005. [Google Scholar] [CrossRef]
- Scher, H.I.; Fizazi, K.; Saad, F.; Taplin, M.E.; Sternberg, C.N.; Miller, K.; de Wit, R.; Mulders, P.; Chi, K.N.; Shore, N.D.; et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N. Engl. J. Med. 2012, 367, 1187–1197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, M.R.; Saad, F.; Chowdhury, S.; Oudard, S.; Hadaschik, B.A.; Graff, J.N.; Olmos, D.; Mainwaring, P.N.; Lee, J.Y.; Uemura, H.; et al. Apalutamide Treatment and Metastasis-free Survival in Prostate Cancer. N. Engl. J. Med. 2018, 378, 1408–1418. [Google Scholar] [CrossRef]
- Fizazi, K.; Shore, N.; Tammela, T.L.; Ulys, A.; Vjaters, E.; Polyakov, S.; Jievaltas, M.; Luz, M.; Alekseev, B.; Kuss, I.; et al. Darolutamide in Nonmetastatic, Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2019, 380, 1235–1246. [Google Scholar] [CrossRef]
- Nuhn, P.; De Bono, J.S.; Fizazi, K.; Freedland, S.J.; Grilli, M.; Kantoff, P.W.; Sonpavde, G.; Sternberg, C.N.; Yegnasubramanian, S.; Antonarakis, E.S. Update on Systemic Prostate Cancer Therapies: Management of Metastatic Castration-resistant Prostate Cancer in the Era of Precision Oncology. Eur. Urol. 2018, 75, 88–99. [Google Scholar] [CrossRef]
- Hussain, M.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Survival with Olaparib in Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 383, 2345–2357. [Google Scholar] [CrossRef]
- De Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Cancer Genome Atlas Research Network. The Molecular Taxonomy of Primary Prostate Cancer. Cell 2015, 163, 1011–1025. [Google Scholar] [CrossRef] [Green Version]
- Lang, S.H.; Swift, S.L.; White, H.; Misso, K.; Kleijnen, J.; Quek, R.G. A systematic review of the prevalence of DNA damage response gene mutations in prostate cancer. Int. J. Oncol. 2019, 55, 597–616. [Google Scholar] [CrossRef] [Green Version]
- Robinson, D.; Van Allen, E.M.; Wu, Y.M.; Schultz, N.; Lonigro, R.J.; Mosquera, J.M.; Montgomery, B.; Taplin, M.E.; Pritchard, C.C.; Attard, G.; et al. Integrative clinical genomics of advanced prostate cancer. Cell 2015, 161, 1215–1228. [Google Scholar] [CrossRef] [Green Version]
- Jin, M.H.; Oh, D.Y. ATM in DNA repair in cancer. Pharmacol. Ther. 2019, 203, 107391. [Google Scholar] [CrossRef]
- Li, X.; Heyer, W.D. Homologous recombination in DNA repair and DNA damage tolerance. Cell Res. 2008, 18, 99–113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mateo, J.; Boysen, G.; Barbieri, C.E.; Bryant, H.E.; Castro, E.; Nelson, P.S.; Olmos, D.; Pritchard, C.C.; Rubin, M.A.; de Bono, J.S. DNA Repair in Prostate Cancer: Biology and Clinical Implications. Eur. Urol. 2017, 71, 417–425. [Google Scholar] [CrossRef] [Green Version]
- Bryant, H.E.; Schultz, N.; Thomas, H.D.; Parker, K.M.; Flower, D.; Lopez, E.; Kyle, S.; Meuth, M.; Curtin, N.J.; Helleday, T. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 2005, 434, 913–917. [Google Scholar] [CrossRef] [PubMed]
- Mateo, J.; Carreira, S.; Sandhu, S.; Miranda, S.; Mossop, H.; Perez-Lopez, R.; Nava Rodrigues, D.; Robinson, D.; Omlin, A.; Tunariu, N.; et al. DNA-Repair Defects and Olaparib in Metastatic Prostate Cancer. N. Engl. J. Med. 2015, 373, 1697–1708. [Google Scholar] [CrossRef]
- Chatterjee, N.; Walker, G.C. Mechanisms of DNA damage, repair, and mutagenesis. Env. Mol. Mutagen 2017, 58, 235–263. [Google Scholar] [CrossRef] [Green Version]
- Gong, J.; Posadas, E.; Bhowmick, N.; Kim, H.; Daskivich, T.; Gupta, A.; Sandler, H.; Kamrava, M.; Zumsteg, Z.; Freedland, S.; et al. Integrating PARP Inhibitors Into Advanced Prostate Cancer Therapeutics. Oncology 2021, 35, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Walden, H.; Deans, A.J. The Fanconi anemia DNA repair pathway: Structural and functional insights into a complex disorder. Annu. Rev. Biophys. 2014, 43, 257–278. [Google Scholar] [CrossRef]
- Liu, S.H.; Shen, P.C.; Chen, C.Y.; Hsu, A.N.; Cho, Y.C.; Lai, Y.L.; Chen, F.H.; Li, C.Y.; Wang, S.C.; Chen, M.; et al. DriverDBv3: A multi-omics database for cancer driver gene research. Nucleic Acids Res. 2020, 48, 863–870. [Google Scholar] [CrossRef] [Green Version]
- Zhao, S.G.; Chang, S.L.; Erho, N.; Yu, M.; Lehrer, J.; Alshalalfa, M.; Speers, C.; Cooperberg, M.R.; Kim, W.; Ryan, C.J. Associations of Luminal and Basal Subtyping of Prostate Cancer With Prognosis and Response to Androgen Deprivation Therapy. JAMA Oncol. 2017, 3, 1663–1672. [Google Scholar] [CrossRef]
- Yoon, J.; Kim, M.; Posadas, E.M.; Freedland, S.J.; Liu, Y.; Davicioni, E.; Den, R.B.; Trock, B.J.; Karnes, R.J.; Klein, E.A. A comparative study of PCS and PAM50 prostate cancer classification schemes. Prostate Cancer Prostatic Dis. 2021, 24, 733–742. [Google Scholar] [CrossRef] [PubMed]
- You, S.; Knudsen, B.S.; Erho, N.; Alshalalfa, M.; Takhar, M.; Al-Deen Ashab, H.; Davicioni, E.; Karnes, R.J.; Klein, E.A.; Den, R.B. Integrated Classification of Prostate Cancer Reveals a Novel Luminal Subtype with Poor Outcome. Cancer Res. 2016, 76, 4948–4958. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, F.Y.; Thomas, S.; Saad, F.; Gormley, M.; Yu, M.K.; Ricci, D.S.; Rooney, B.; Brookman-May, S.; McCarthy, S.; Olmos, D.; et al. Association of Molecular Subtypes With Differential Outcome to Apalutamide Treatment in Nonmetastatic Castration-Resistant Prostate Cancer. JAMA Oncol. 2021, 7, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Jan, Y.J.; Yoon, J.; Chen, J.F.; Teng, P.C.; Yao, N.; Cheng, S.; Lozano, A.; Chu, G.C.Y.; Chung, H.; Lu, Y.T.; et al. A Circulating Tumor Cell-RNA Assay for Assessment of Androgen Receptor Signaling Inhibitor Sensitivity in Metastatic Castration-Resistant Prostate Cancer. Theranostics 2019, 9, 2812–2826. [Google Scholar] [CrossRef]
- Ogata, H.; Goto, S.; Fujibuchi, W.; Kanehisa, M. Computation with the KEGG pathway database. Biosystems 1998, 47, 119–128. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, 457–462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jassal, B.; Matthews, L.; Viteri, G.; Gong, C.; Lorente, P.; Fabregat, A.; Sidiropoulos, K.; Cook, J.; Gillespie, M.; Haw, R.; et al. The reactome pathway knowledgebase. Nucleic Acids Res. 2020, 48, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Christensen, E. Multivariate survival analysis using Cox’s regression model. Hepatology 1987, 7, 1346–1358. [Google Scholar] [CrossRef] [PubMed]
- Teng, P.-C.; Jan, Y.J.; Kim, M.; Chen, J.-F.; Yoon, J.; Wang, J.J.; Chen, P.-J.; Yao, N.; Lee, Y.-T.; Lozano, A.; et al. Development of a circulating tumor cell-based RNA classifier for patients with castration-resistant prostate cancer: CTC-PCS/PAM50. J. Clin. Oncol. 2020, 38, e17509. [Google Scholar] [CrossRef]
- Hamid, A.; Wang, X.V.; Chen, Y.-H.; Feng, F.Y.; Den, R.B.; Attard, G.; Allen, E.M.V.; Huang, H.-C.; Karns, A.; Dittamore, R.; et al. Luminal B subtype as a predictive biomarker of docetaxel benefit for newly diagnosed metastatic hormone sensitive prostate cancer (mHSPC): A correlative study of E3805 CHAARTED. J. Clin. Oncol. 2020, 38, 162. [Google Scholar] [CrossRef]
- Feng, F.Y.; Thomas, S.; Aguilar-Bonavides, C.; Gormley, M.; Agarwal, N.; Attard, G.; Wyatt, A.W.; Davicioni, E.; Ricci, D.S.; Lopez-Gitlitz, A.; et al. Molecular determinants of outcome for metastatic castration-sensitive prostate cancer (mCSPC) with addition of apalutamide (APA) or placebo (PBO) to androgen deprivation therapy (ADT) in TITAN. J. Clin. Oncol. 2020, 38, 5535. [Google Scholar] [CrossRef]
- Kim, W.; Small, E.J.; Aggarwal, R.R.; Den, R.B.; Lehrer, J.; Zhang, L.; Youngren, J.; Goldstein, T.C.; Alumkal, J.J.; Gleave, M.; et al. Luminal and basal subtyping of metastatic castration-resistant prostate cancer (mCRPC) and its clinical implications. J. Clin. Oncol. 2018, 36, 197. [Google Scholar] [CrossRef]
- Cooperberg, M.R.; Erho, N.; Chan, J.M.; Feng, F.Y.; Fishbane, N.; Zhao, S.G.; Simko, J.P.; Cowan, J.E.; Lehrer, J.; Alshalalfa, M.; et al. The Diverse Genomic Landscape of Clinically Low-risk Prostate Cancer. Eur. Urol. 2018, 74, 444–452. [Google Scholar] [CrossRef]
- Goodwin, J.F.; Schiewer, M.J.; Dean, J.L.; Schrecengost, R.S.; de Leeuw, R.; Han, S.; Ma, T.; Den, R.B.; Dicker, A.P.; Feng, F.Y. A hormone-DNA repair circuit governs the response to genotoxic insult. Cancer Discov. 2013, 3, 1254–1271. [Google Scholar] [CrossRef] [Green Version]
- Mehra, R.; Tomlins, S.A.; Shen, R.; Nadeem, O.; Wang, L.; Wei, J.T.; Pienta, K.J.; Ghosh, D.; Rubin, M.A.; Chinnaiyan, A.M.; et al. Comprehensive assessment of TMPRSS2 and ETS family gene aberrations in clinically localized prostate cancer. Mod. Pathol. 2007, 20, 538–544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, C.; Wyatt, A.W.; McPherson, A.; Lin, D.; McConeghy, B.J.; Mo, F.; Shukin, R.; Lapuk, A.V.; Jones, S.J.; Zhao, Y.; et al. Poly-gene fusion transcripts and chromothripsis in prostate cancer. Genes Chromosomes Cancer 2012, 51, 1144–1153. [Google Scholar] [CrossRef]
- Mani, R.S.; Tomlins, S.A.; Callahan, K.; Ghosh, A.; Nyati, M.K.; Varambally, S.; Palanisamy, N.; Chinnaiyan, A.M. Induced chromosomal proximity and gene fusions in prostate cancer. Science 2009, 326, 1230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ju, B.G.; Lunyak, V.V.; Perissi, V.; Garcia-Bassets, I.; Rose, D.W.; Glass, C.K.; Rosenfeld, M.G. A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription. Science 2006, 312, 1798–1802. [Google Scholar] [CrossRef] [PubMed]
- Tomlins, S.A.; Rhodes, D.R.; Perner, S.; Dhanasekaran, S.M.; Mehra, R.; Sun, X.W.; Varambally, S.; Cao, X.; Tchinda, J.; Kuefer, R.; et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 2005, 310, 644–648. [Google Scholar] [CrossRef]
- Miller, E.T.; You, S.; Cadaneanu, R.M.; Kim, M.; Yoon, J.; Liu, S.T.; Li, X.; Kwan, L.; Hodge, J.; Quist, M.J.; et al. Chromosomal instability in untreated primary prostate cancer as an indicator of metastatic potential. BMC Cancer 2020, 20, 398. [Google Scholar] [CrossRef]
- Cooperberg, M.R.; Broering, J.M.; Carroll, P.R. Time trends and local variation in primary treatment of localized prostate cancer. J. Clin. Oncol. 2010, 28, 1117–1123. [Google Scholar] [CrossRef] [Green Version]
- Erho, N.; Crisan, A.; Vergara, I.A.; Mitra, A.P.; Ghadessi, M.; Buerki, C.; Bergstralh, E.J.; Kollmeyer, T.; Fink, S.; Haddad, Z.; et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS ONE 2013, 8, e66855. [Google Scholar] [CrossRef] [Green Version]
- Cuzick, J.; Swanson, G.P.; Fisher, G.; Brothman, A.R.; Berney, D.M.; Reid, J.E.; Mesher, D.; Speights, V.O.; Stankiewicz, E.; Foster, C.S.; et al. Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: A retrospective study. Lancet Oncol 2011, 12, 245–255. [Google Scholar] [CrossRef] [Green Version]
- Casanova-Salas, I.; Athie, A.; Boutros, P.C.; Del Re, M.; Miyamoto, D.T.; Pienta, K.J.; Posadas, E.M.; Sowalsky, A.G.; Stenzl, A.; Wyatt, A.W.; et al. Quantitative and Qualitative Analysis of Blood-based Liquid Biopsies to Inform Clinical Decision-making in Prostate Cancer. Eur. Urol. 2021, 79, 762–771. [Google Scholar] [CrossRef] [PubMed]
- Duijvesz, D.; Versluis, C.Y.; van der Fels, C.A.; Vredenbregt-van den Berg, M.S.; Leivo, J.; Peltola, M.T.; Bangma, C.H.; Pettersson, K.S.; Jenster, G. Immuno-based detection of extracellular vesicles in urine as diagnostic marker for prostate cancer. Int. J. Cancer 2015, 137, 2869–2878. [Google Scholar] [CrossRef]
- Scher, H.I.; Morris, M.J.; Stadler, W.M.; Higano, C.; Basch, E.; Fizazi, K.; Antonarakis, E.S.; Beer, T.M.; Carducci, M.A.; Chi, K.N.; et al. Trial Design and Objectives for Castration-Resistant Prostate Cancer: Updated Recommendations From the Prostate Cancer Clinical Trials Working Group 3. J. Clin. Oncol. 2016, 34, 1402–1418. [Google Scholar] [CrossRef] [Green Version]
- Armstrong, A.J.; Halabi, S.; Luo, J.; Nanus, D.M.; Giannakakou, P.; Szmulewitz, R.Z.; Danila, D.C.; Healy, P.; Anand, M.; Rothwell, C.J.; et al. Prospective Multicenter Validation of Androgen Receptor Splice Variant 7 and Hormone Therapy Resistance in High-Risk Castration-Resistant Prostate Cancer: The PROPHECY Study. J. Clin. Oncol. 2019, 37, 1120–1129. [Google Scholar] [CrossRef] [PubMed]
- Del Re, M.; Biasco, E.; Crucitta, S.; Derosa, L.; Rofi, E.; Orlandini, C.; Miccoli, M.; Galli, L.; Falcone, A.; Jenster, G.W.; et al. The Detection of Androgen Receptor Splice Variant 7 in Plasma-derived Exosomal RNA Strongly Predicts Resistance to Hormonal Therapy in Metastatic Prostate Cancer Patients. Eur. Urol. 2017, 71, 680–687. [Google Scholar] [CrossRef] [PubMed]
- Bolderson, E.; Tomimatsu, N.; Richard, D.J.; Boucher, D.; Kumar, R.; Pandita, T.K.; Burma, S.; Khanna, K.K. Phosphorylation of Exo1 modulates homologous recombination repair of DNA double-strand breaks. Nucleic Acids Res. 2010, 38, 1821–1831. [Google Scholar] [CrossRef] [Green Version]
- Keijzers, G.; Bakula, D.; Petr, M.A.; Madsen, N.G.K.; Teklu, A.; Mkrtchyan, G.; Osborne, B.; Scheibye-Knudsen, M. Human Exonuclease 1 (EXO1) Regulatory Functions in DNA Replication with Putative Roles in Cancer. Int. J. Mol. Sci. 2018, 20, 74. [Google Scholar] [CrossRef] [Green Version]
- Luo, F.; Wang, Y.Z.; Lin, D.; Li, J.; Yang, K. Exonuclease 1 expression is associated with clinical progression, metastasis, and survival prognosis of prostate cancer. J. Cell Biochem. 2019, 120, 11383–11389. [Google Scholar] [CrossRef] [PubMed]
- Hua, X.; Ge, S.; Chen, J.; Zhang, L.; Tai, S.; Liang, C. Effects of RNA Binding Proteins on the Prognosis and Malignant Progression in Prostate Cancer. Front. Genet. 2020, 11, 591667. [Google Scholar] [CrossRef] [PubMed]
- Imtiaz, H.; Afroz, S.; Hossain, M.A.; Bellah, S.F.; Rahman, M.M.; Kadir, M.S.; Sultana, R.; Mazid, M.A.; Rahman, M.M. Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population. Tumour Biol. 2019, 41, 1010428319830837. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Berends, M.J.; Post, J.G.; Mensink, R.G.; Verlind, E.; Van Der Sluis, T.; Kempinga, C.; Sijmons, R.H.; van der Zee, A.G.; Hollema, H.; et al. Germline mutations of EXO1 gene in patients with hereditary nonpolyposis colorectal cancer (HNPCC) and atypical HNPCC forms. Gastroenterology 2001, 120, 1580–1587. [Google Scholar] [CrossRef]
- He, D.; Li, T.; Sheng, M.; Yang, B. Exonuclease 1 (Exo1) Participates in Mammalian Non-Homologous End Joining and Contributes to Drug Resistance in Ovarian Cancer. Med. Sci. Monit. 2020, 26, e918751. [Google Scholar] [CrossRef]
- Venditti, A.; Del Poeta, G.; Buccisano, F.; Tamburini, A.; Cox-Froncillo, M.C.; Aronica, G.; Bruno, A.; Del Moro, B.; Epiceno, A.M.; Battaglia, A.; et al. Prognostic relevance of the expression of Tdt and CD7 in 335 cases of acute myeloid leukemia. Leukemia 1998, 12, 1056–1063. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ok, C.Y.; Medeiros, L.J.; Thakral, B.; Tang, G.; Jain, N.; Jabbour, E.; Pierce, S.A.; Konoplev, S. High-grade B-cell lymphomas with TdT expression: A diagnostic and classification dilemma. Mod. Pathol. 2019, 32, 48–58. [Google Scholar] [CrossRef]
- Sidiropoulos, M.; Hanna, W.; Raphael, S.J.; Ghorab, Z. Expression of TdT in Merkel cell carcinoma and small cell lung carcinoma. Am. J. Clin. Pathol. 2011, 135, 831–838. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Chan, J.; Lambele, M.; Yusufzai, T.; Stumpff, J.; Opresko, P.L.; Thali, M.; Wallace, S.S. NEIL3 Repairs Telomere Damage during S Phase to Secure Chromosome Segregation at Mitosis. Cell Rep. 2017, 20, 2044–2056. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Xu, L.; Shi, S.; Wu, S.; Meng, R.; Chen, H.; Jiang, Z. Deficiency of NEIL3 Enhances the Chemotherapy Resistance of Prostate Cancer. Int. J. Mol. Sci. 2021, 22, 4098. [Google Scholar] [CrossRef]
- Tran, O.T.; Tadesse, S.; Chu, C.; Kidane, D. Overexpression of NEIL3 associated with altered genome and poor survival in selected types of human cancer. Tumor Biol. 2020, 42, 1010428320918404. [Google Scholar] [CrossRef] [PubMed]
- Shinmura, K.; Kato, H.; Kawanishi, Y.; Igarashi, H.; Goto, M.; Tao, H.; Inoue, Y.; Nakamura, S.; Misawa, K.; Mineta, H.; et al. Abnormal Expressions of DNA Glycosylase Genes NEIL1, NEIL2, and NEIL3 Are Associated with Somatic Mutation Loads in Human Cancer. Oxid. Med. Cell Longev. 2016, 2016, 1546392. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, W.; Yin, Q.; Guo, S.; Wang, J. NEIL3 contributes toward the carcinogenesis of liver cancer and regulates PI3K/Akt/mTOR signaling. Exp. Ther. Med. 2021, 22, 1053. [Google Scholar] [CrossRef] [PubMed]
- Techer, H.; Koundrioukoff, S.; Carignon, S.; Wilhelm, T.; Millot, G.A.; Lopez, B.S.; Brison, O.; Debatisse, M. Signaling from Mus81-Eme2-Dependent DNA Damage Elicited by Chk1 Deficiency Modulates Replication Fork Speed and Origin Usage. Cell Rep. 2016, 14, 1114–1127. [Google Scholar] [CrossRef] [Green Version]
- Pepe, A.; West, S.C. MUS81-EME2 promotes replication fork restart. Cell Rep. 2014, 7, 1048–1055. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Wang, Y.; Zhang, W.; Wang, X.; Chen, L.; Wang, S. BKM120 sensitizes BRCA-proficient triple negative breast cancer cells to olaparib through regulating FOXM1 and Exo1 expression. Sci. Rep. 2021, 11, 4774. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Shi, J.; Chen, S.H.; Bian, C.; Yu, X. The PIN domain of EXO1 recognizes poly(ADP-ribose) in DNA damage response. Nucleic Acids Res. 2015, 43, 10782–10794. [Google Scholar] [CrossRef] [Green Version]
- Cheruiyot, A.; Paudyal, S.C.; Kim, I.K.; Sparks, M.; Ellenberger, T.; Piwnica-Worms, H.; You, Z. Poly(ADP-ribose)-binding promotes Exo1 damage recruitment and suppresses its nuclease activities. DNA Repair 2015, 35, 106–115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prensner, J.R.; Rubin, M.A.; Wei, J.T.; Chinnaiyan, A.M. Beyond PSA: The next generation of prostate cancer biomarkers. Sci. Transl. Med. 2012, 4, 127rv3. [Google Scholar] [CrossRef] [Green Version]
- Mohler, J.L.; Antonarakis, E.S.; Armstrong, A.J.; D’Amico, A.V.; Davis, B.J.; Dorff, T.; Eastham, J.A.; Enke, C.A.; Farrington, T.A.; Higano, C.S.; et al. Prostate Cancer, Version 2.2019, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. 2019, 17, 479–505. [Google Scholar] [CrossRef] [Green Version]
- Meacham, C.E.; Morrison, S.J. Tumour heterogeneity and cancer cell plasticity. Nature 2013, 501, 328–337. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scher, H.I.; Solo, K.; Valant, J.; Todd, M.B.; Mehra, M. Prevalence of Prostate Cancer Clinical States and Mortality in the United States: Estimates Using a Dynamic Progression Model. PLoS ONE 2015, 10, e0139440. [Google Scholar] [CrossRef] [Green Version]
- Halabi, S.; Kelly, W.K.; Ma, H.; Zhou, H.; Solomon, N.C.; Fizazi, K.; Tangen, C.M.; Rosenthal, M.; Petrylak, D.P.; Hussain, M.; et al. Meta-Analysis Evaluating the Impact of Site of Metastasis on Overall Survival in Men With Castration-Resistant Prostate Cancer. J. Clin. Oncol. 2016, 34, 1652–1659. [Google Scholar] [CrossRef]
- Phillips, R.; Shi, W.Y.; Deek, M.; Radwan, N.; Lim, S.J.; Antonarakis, E.S.; Rowe, S.P.; Ross, A.E.; Gorin, M.A.; Deville, C.; et al. Outcomes of Observation vs Stereotactic Ablative Radiation for Oligometastatic Prostate Cancer: The ORIOLE Phase 2 Randomized Clinical Trial. JAMA Oncol. 2020, 6, 650–659. [Google Scholar] [CrossRef] [Green Version]
- Scher, H.I.; Lu, D.; Schreiber, N.A.; Louw, J.; Graf, R.P.; Vargas, H.A.; Johnson, A.; Jendrisak, A.; Bambury, R.; Danila, D.; et al. Association of AR-V7 on Circulating Tumor Cells as a Treatment-Specific Biomarker With Outcomes and Survival in Castration-Resistant Prostate Cancer. JAMA Oncol. 2016, 2, 1441–1449. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef] [Green Version]
- Perou, C.M.; Sorlie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Zhao, S.G.; Chen, W.S.; Das, R.; Chang, S.L.; Tomlins, S.A.; Chou, J.; Quigley, D.A.; Dang, H.X.; Barnard, T.J.; Mahal, B.A.; et al. Clinical and Genomic Implications of Luminal and Basal Subtypes Across Carcinomas. Clin. Cancer Res. 2019, 25, 2450–2457. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]


| SDE Gene | Log2 Fold Change | Adjusted p Value of SDE | HR of OS | p Value of OS |
|---|---|---|---|---|
| RAD51 | 1.02 | 5.92 × 10−11 | 1.88 | 0.36 |
| RPA4 | 1.18 | 1.83 × 10−6 | 2.15 | 0.26 |
| SLX1B | 1.02 | 2.2 × 10−3 | 1.25 | 0.73 |
| POLN | 1.00 | 7.16 × 10−19 | 1.99 | 0.30 |
| EME2 | 1.29 | 1.16 × 10−18 | 5.67 | 0.07 |
| EXO1 | 1.27 | 2.82 × 10−11 | 1.75 | 0.51 |
| DNTT | −1.20 | 1.1 × 10−2 | 1.22 | 0.85 |
| NEIL3 | 1.93 | 1.21 × 10−18 | 2.01 | 0.31 |
| Variables for OS | HR | Lower 95% CI | Higher 95% CI | p Value |
| 4-gene expression | ||||
| Lower | ref | ref | ref | ref |
| Higher | 13.8 | 1.21 | 158 | 0.0348 |
| Age (year) | 1.05 | 0.95 | 1.15 | 0.3667 |
| PSA (ng/mL) | 0.99 | 0.99 | 1.00 | 0.2950 |
| TNM stages | ||||
| T1N0M0 or T2N0M0 | ref | ref | ref | ref |
| Others | 7.27 | 0.92 | 57.39 | 0.0597 |
| Gleason scores | ||||
| ≤7 | ref | ref | ref | ref |
| >7 | 0.72 | 0.14 | 3.62 | 0.6897 |
| Variables for PFS | HR | Lower 95% CI | Higher 95% CI | p Value |
| 4-gene expression | ||||
| Lower | ref | ref | ref | ref |
| Higher | 2.32 | 1.39 | 3.95 | 0.0020 |
| Age (year) | 1.00 | 0.98 | 1.04 | 0.6517 |
| PSA (ng/mL) | 0.99 | 0.99 | 1.00 | 0.2380 |
| TNM stages | ||||
| T1N0M0 or T2N0M0 | ref | ref | ref | ref |
| Others | 1.33 | 0.86 | 2.08 | 0.2034 |
| Gleason scores | ||||
| ≤7 | ref | ref | ref | ref |
| >7 | 2.70 | 1.61 | 4.56 | 0.0002 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Teng, P.-C.; Huang, S.-P.; Liu, C.-H.; Lin, T.-Y.; Cho, Y.-C.; Lai, Y.-L.; Wang, S.-C.; Yeh, H.-C.; Chuu, C.-P.; Chen, D.-N.; et al. Identification of DNA Damage Repair-Associated Prognostic Biomarkers for Prostate Cancer Using Transcriptomic Data Analysis. Int. J. Mol. Sci. 2021, 22, 11771. https://doi.org/10.3390/ijms222111771
Teng P-C, Huang S-P, Liu C-H, Lin T-Y, Cho Y-C, Lai Y-L, Wang S-C, Yeh H-C, Chuu C-P, Chen D-N, et al. Identification of DNA Damage Repair-Associated Prognostic Biomarkers for Prostate Cancer Using Transcriptomic Data Analysis. International Journal of Molecular Sciences. 2021; 22(21):11771. https://doi.org/10.3390/ijms222111771
Chicago/Turabian StyleTeng, Pai-Chi, Shu-Pin Huang, Chia-Hsin Liu, Ting-Yi Lin, Yi-Chun Cho, Yo-Liang Lai, Shu-Chi Wang, Hsin-Chih Yeh, Chih-Pin Chuu, Deng-Neng Chen, and et al. 2021. "Identification of DNA Damage Repair-Associated Prognostic Biomarkers for Prostate Cancer Using Transcriptomic Data Analysis" International Journal of Molecular Sciences 22, no. 21: 11771. https://doi.org/10.3390/ijms222111771
APA StyleTeng, P.-C., Huang, S.-P., Liu, C.-H., Lin, T.-Y., Cho, Y.-C., Lai, Y.-L., Wang, S.-C., Yeh, H.-C., Chuu, C.-P., Chen, D.-N., Cheng, W.-C., & Li, C.-Y. (2021). Identification of DNA Damage Repair-Associated Prognostic Biomarkers for Prostate Cancer Using Transcriptomic Data Analysis. International Journal of Molecular Sciences, 22(21), 11771. https://doi.org/10.3390/ijms222111771







