Application of Machine-Learning Methods to Recognize mitoBK Channels from Different Cell Types Based on the Experimental Patch-Clamp Results
Abstract
1. Introduction
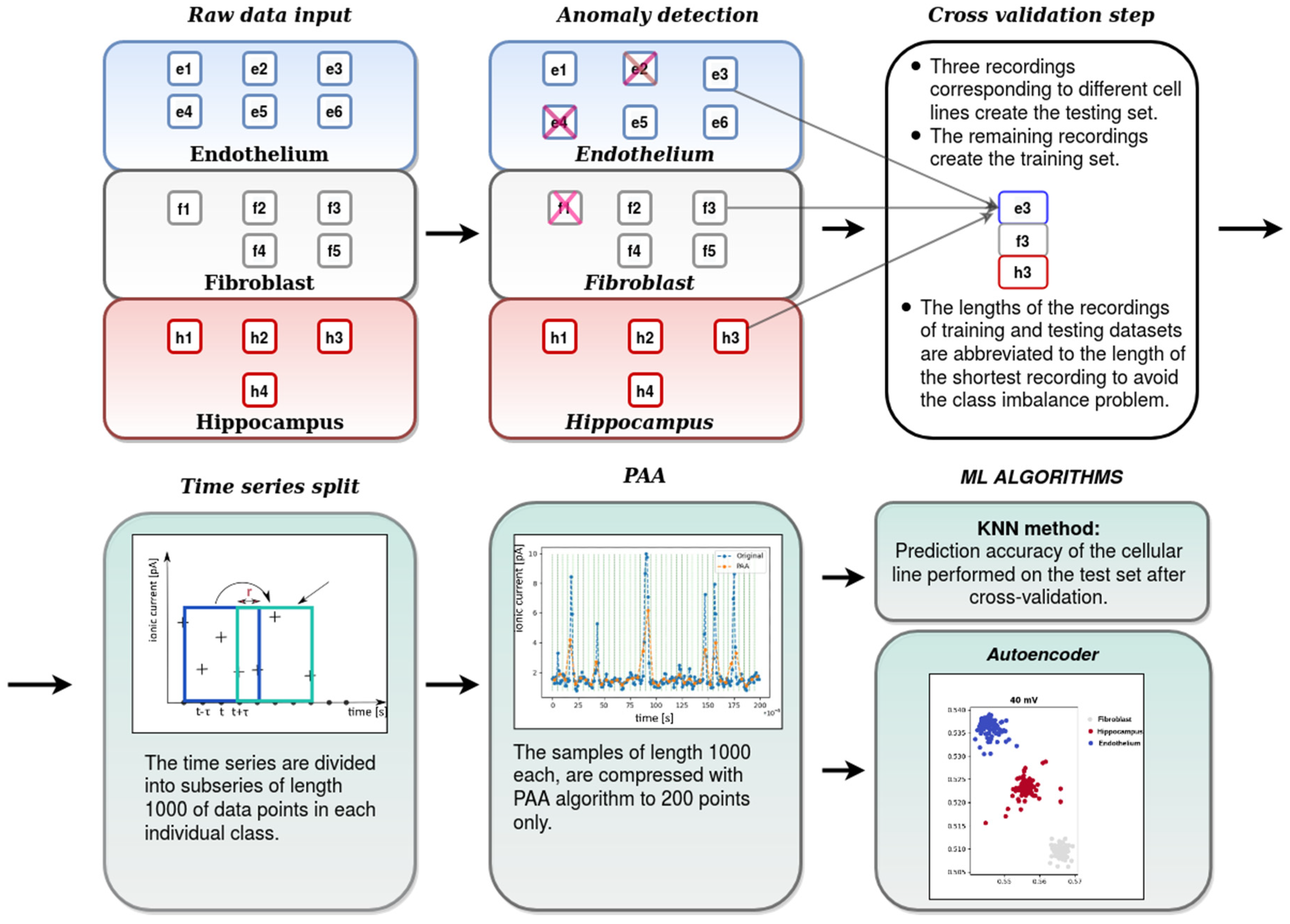
- the first data preprocessing stage-where the abnormal recordings were rejected from further analysis. This stage of analysis was performed by means of the autoencoder anomaly detection technique.
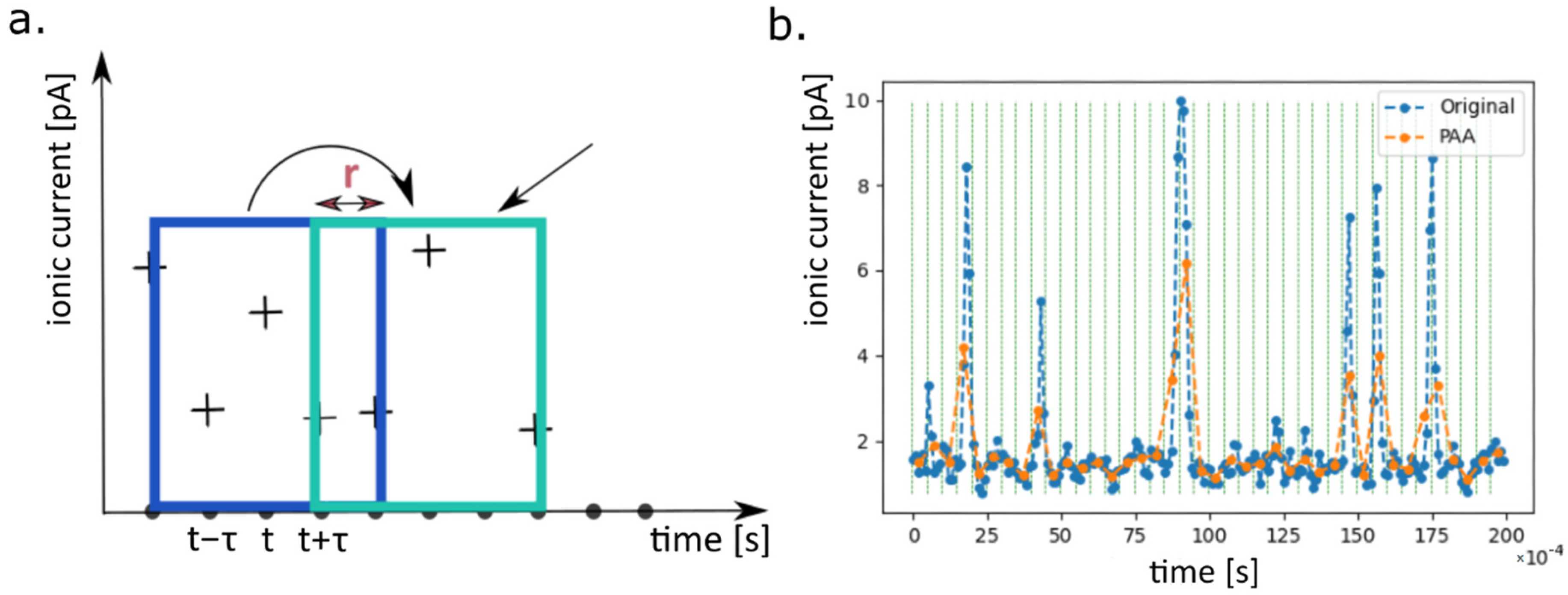
- the second data preprocessing stage including partition of the recordings into smaller subseries (samples), dimensionality reduction by Piecewise Aggregate Approximation (PAA) and scaling
- the cross-validation step: partition of the recordings into the training and test sets,
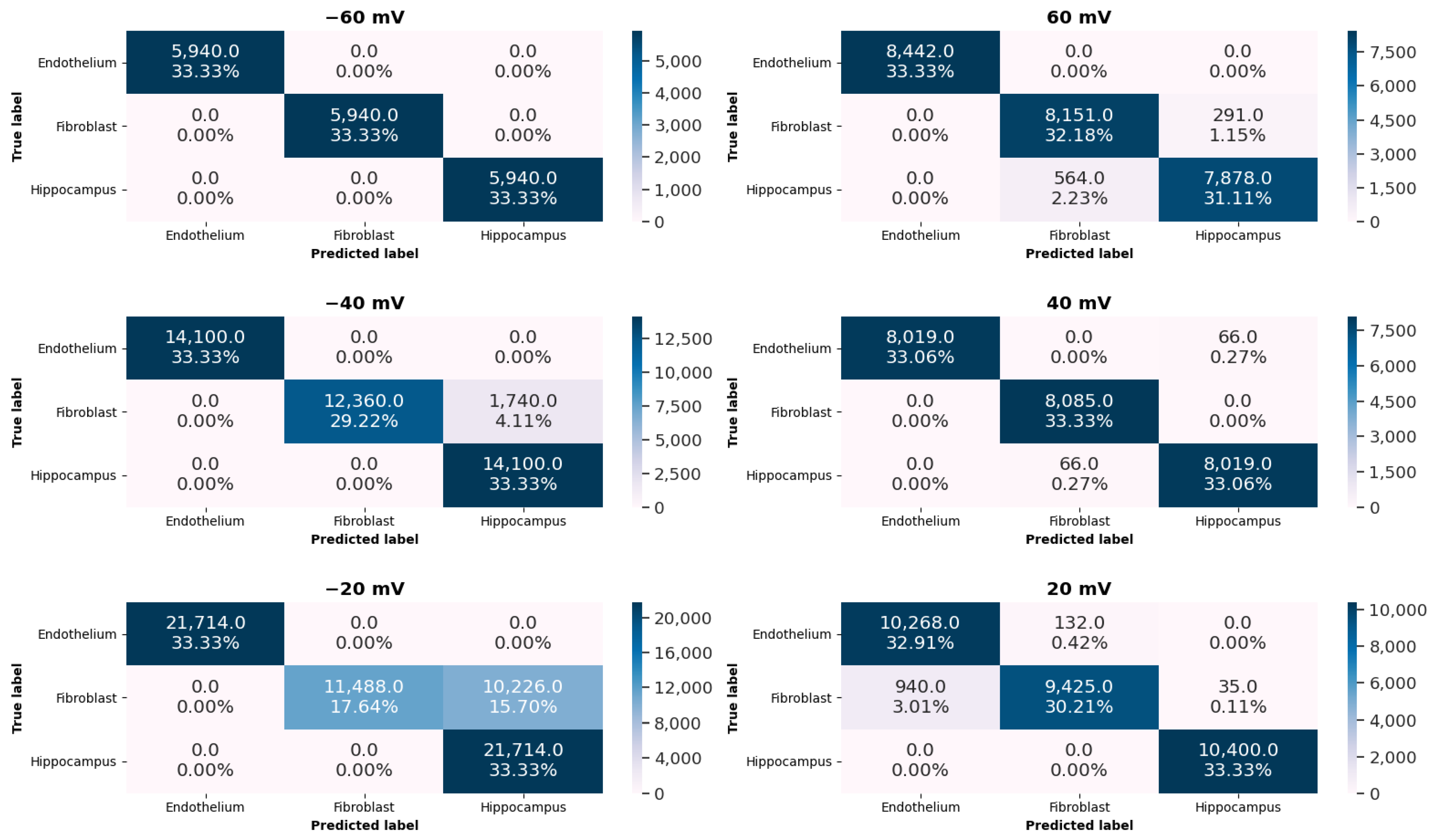
- classification of tissue-specific single-channel activity patterns by means of the K-Nearest Neighbors (KNN) algorithm and evaluation of its accuracy,
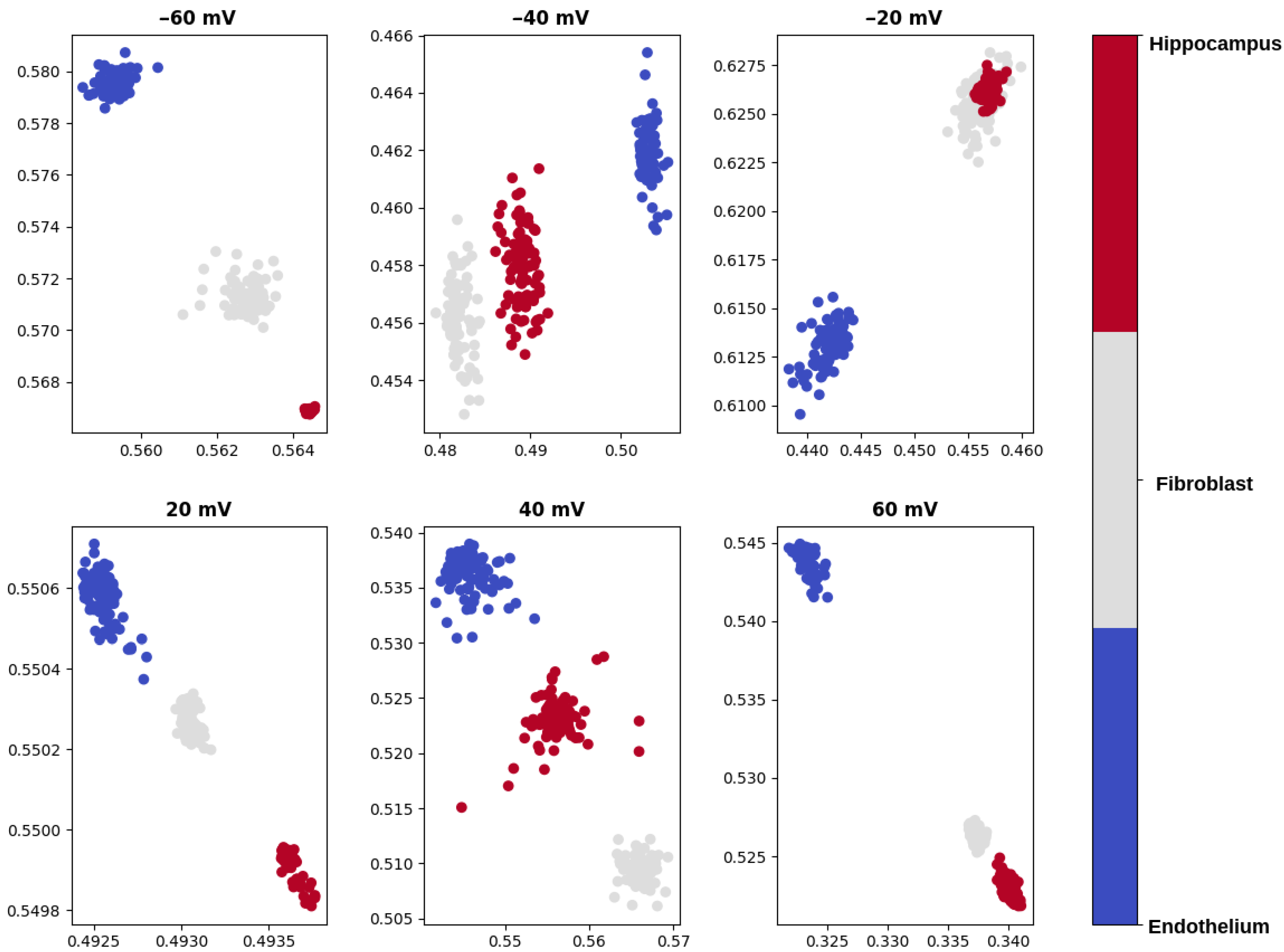
- visualization of the test data set compressed to a 2-dimensional space by means of the autoencoder,
- calculation of the average distances between the signal samples corresponding to various mitoBK channel groups in the 2-dimensional latent space.
2. Results
2.1. Kinetic Properties of Experimental Data
2.2. Machine-Learning Results
3. Discussion
4. Methods and Materials
4.1. Electrophysiology
4.1.1. Mitochondria and Mitoplast Preparation
4.1.2. Patch-Clamp Experiments
4.1.3. Basic Analysis of Experimental Results
4.2. Machine-Learning Approaches
4.2.1. Data Preprocessing
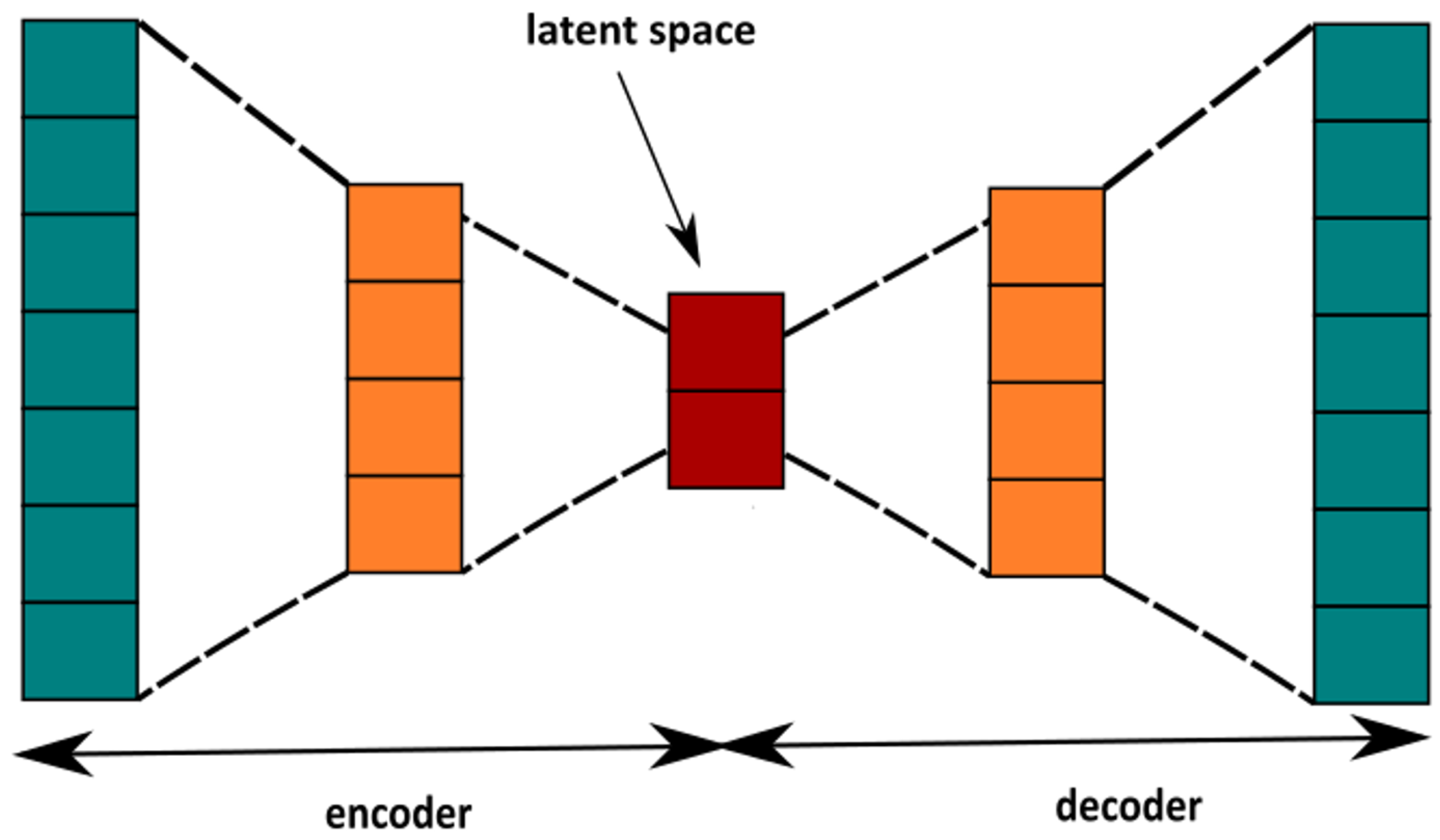
4.2.2. Autoencoder
4.2.3. Anomaly Detection with Autoencoder
4.2.4. PAA
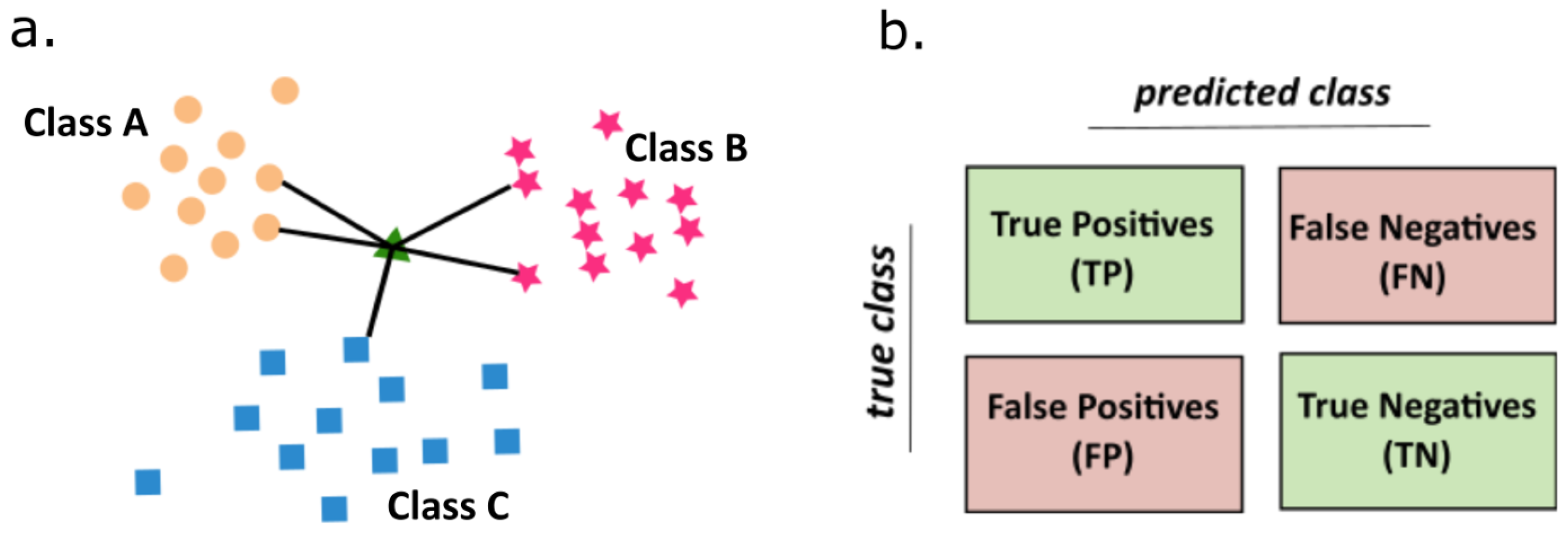
4.2.5. K-Nearest Neighbors Algorithm
- The values of the parameter K and the distance metric d are selected. This prior selection has a great impact on the obtained results. The values of K is a key choice that gives a balance between overtraining and the under-fitting. In our case K = 5 was the optimal selection. As a metric of distance a simple Euclidean relation was used (4).where N is a space dimensionality.
- For a sample subjected to the classification the number of K-nearest neighbors are determined by the calculation of distance to the nearest neighbors. At the final step, a class etiquette is assigned by majority of vote. Thus, the KNN algorithm classifies the object to this class, which is most frequently represented among K-nearest neighbors from the training set.
- To avoid the problems with the overfitting or the selection bias [56] the cross-validation technique is applied. One step in this technique is about partitioning the data into training and test sets. In our case, in every round, one record from each cellular line, serves as a test set, whereas the remaining data forms the training set. We make so many rounds, each involving a different collection of the testing records.
4.2.6. K-Means
- the number of clusters is determined. Their centroids are chosen randomly.
- the kind of metric is selected. Similar to the KNN algorithm, the Euclidean distance is the most popular choice.
- the distance from all points to centroids is determined. The new object is assigned to the cluster with the closest center.
- based on the mean of points assigned to a given cluster, the new centers of concentration are set down.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Amato, F.; López, A.; Peña-Méndez, E.M.; Vaňhara, P.; Hampl, A.; Havel, J. Artificial neural networks in medical diagnosis. J. Appl. Biomed. 2013, 11, 47–58. [Google Scholar] [CrossRef]
- Hessler, G.; Baringhaus, K.H. Artificial intelligence in drug design. Molecules 2018, 23, 2520. [Google Scholar] [CrossRef]
- Jordan, A.M. Artificial intelligence in drug design—The storm before the calm? ACS Med. Chem. Lett. 2018, 9, 1150–1152. [Google Scholar] [CrossRef] [PubMed]
- Mak, K.K.; Pichika, M.R. Artificial intelligence in drug development: Present status and future prospects. Drug Discov. Today 2019, 24, 773–780. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Sheridan, R.P.; Liaw, A.; Dahl, G.E.; Svetnik, V. Deep neural nets as a method for quantitative structure–activity relationships. J. Chem. Inf. Model. 2015, 55, 263–274. [Google Scholar] [CrossRef] [PubMed]
- Wulff, H.; Castle, N.A.; Pardo, L.A. Voltage-gated potassium channels as therapeutic targets. Nat. Rev. Drug Discov. 2009, 8, 982–1001. [Google Scholar] [CrossRef]
- Szewczyk, A.; Jarmuszkiewicz, W.; Kunz, W.S. Mitochondrial potassium channels. IUBMB Life 2009, 61, 134–143. [Google Scholar] [CrossRef]
- Laskowski, M.; Augustynek, B.; Kulawiak, B.; Koprowski, P.; Bednarczyk, P.; Jarmuszkiewicz, W.; Szewczyk, A. What do we not know about mitochondrial potassium channels? Biochim. Biophys. Acta BBA Bioenerg. 2016, 1857, 1247–1257. [Google Scholar] [CrossRef]
- Balderas, E.; Zhang, J.; Stefani, E.; Toro, L. Mitochondrial BKCa channel. Front. Physiol. 2015, 6, 104. [Google Scholar] [CrossRef]
- Marty, A. Ca-dependent K channels with large unitary conductance in chromaffin cell membranes. Nature 1981, 291, 497–500. [Google Scholar] [CrossRef]
- Pallotta, B.S.; Magleby, K.L.; Barrett, J.N. Single channel recordings of Ca2+-activated K+ currents in rat muscle cell culture. Nature 1981, 293, 471–474. [Google Scholar] [CrossRef] [PubMed]
- Latorre, R.; Vergara, C.; Hidalgo, C. Reconstitution in planar lipid bilayers of a Ca2+-dependent K+ channel from transverse tubule membranes isolated from rabbit skeletal muscle. Proc. Natl. Acad. Sci. USA 1982, 79, 805–809. [Google Scholar] [CrossRef]
- Szewczyk, A.; Skalska, J.; Głąb, M.; Kulawiak, B.; Malińska, D.; Koszela-Piotrowska, I.; Kunz, W.S. Mitochondrial potassium channels: From pharmacology to function. Biochim. Biophys. Acta BBA Bioenerg. 2006, 1757, 715–720. [Google Scholar] [CrossRef] [PubMed]
- Frankenreiter, S.; Bednarczyk, P.; Kniess, A.; Bork, N.I.; Straubinger, J.; Koprowski, P.; Wrzosek, A.; Mohr, E.; Logan, A.; Murphy, M.P.; et al. cGMP-elevating compounds and ischemic conditioning provide cardioprotection against ischemia and reperfusion injury via cardiomyocyte-specific BK channels. Circulation 2017, 136, 2337–2355. [Google Scholar] [CrossRef] [PubMed]
- Bednarczyk, P.; Wieckowski, M.R.; Broszkiewicz, M.; Skowronek, K.; Siemen, D.; Szewczyk, A. Putative structural and functional coupling of the mitochondrial BKCa channel to the respiratory chain. PLoS ONE 2013, 8, e68125. [Google Scholar] [CrossRef]
- Szewczyk, A.; Kajma, A.; Malinska, D.; Wrzosek, A.; Bednarczyk, P.; Zabłocka, B.; Dołowy, K. Pharmacology of mitochondrial potassium channels: Dark side of the field. FEBS Lett. 2010, 584, 2063–2069. [Google Scholar] [CrossRef]
- Tseng-Crank, J.; Foster, C.D.; Krause, J.D.; Mertz, R.; Godinot, N.; DiChiara, T.J.; Reinhart, P.H. Cloning, expression, and distribution of functionally distinct Ca2+-activated K+ channel isoforms from human brain. Neuron 1994, 13, 1315–1330. [Google Scholar] [CrossRef]
- Fury, M.; Marx, S.O.; Marks, A.R. Molecular BKology: The study of splicing and dicing. Sci. STKE 2002, 2002, pe12. [Google Scholar] [CrossRef]
- Schubert, R.; Nelson, M.T. Protein kinases: Tuners of the BKCa channel in smooth muscle. Trends Pharmacol. Sci. 2001, 22, 505–512. [Google Scholar] [CrossRef]
- Li, M.; Tanaka, Y.; Alioua, A.; Wu, Y.; Lu, R.; Kundu, P.; Sanchez-Pastor, E.; Marijic, J.; Stefani, E.; Toro, L. Thromboxane A2 receptor and MaxiK-channel intimate interaction supports channel trans-inhibition independent of G-protein activation. Proc. Natl. Acad. Sci. USA 2010, 107, 19096–19101. [Google Scholar] [CrossRef]
- Shipston, M.J. Alternative splicing of potassium channels: A dynamic switch of cellular excitability. Trends Cell Biol. 2001, 11, 353–358. [Google Scholar] [CrossRef]
- Singh, H.; Lu, R.; Bopassa, J.C.; Meredith, A.L.; Stefani, E.; Toro, L. mitoBKCa is encoded by the Kcnma1 gene, and a splicing sequence defines its mitochondrial location. Proc. Natl. Acad. Sci. USA 2013, 110, 10836–10841. [Google Scholar] [CrossRef] [PubMed]
- Contreras, G.F.; Neely, A.; Alvarez, O.; Gonzalez, C.; Latorre, R. Modulation of BK channel voltage gating by different auxiliary β subunits. Proc. Natl. Acad. Sci. USA 2012, 109, 18991–18996. [Google Scholar] [CrossRef] [PubMed]
- Latorre, R.; Castillo, K.; Carrasquel-Ursulaez, W.; Sepulveda, R.V.; Gonzalez-Nilo, F.; Gonzalez, C.; Alvarez, O. Molecular determinants of BK channel functional diversity and functioning. Physiol. Rev. 2017, 97, 39–87. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Yan, J. Modulation of BK channel function by auxiliary beta and gamma subunits. In International Review of Neurobiology; Elsevier: Amsterdam, The Netherlands, 2016; Volume 128, pp. 51–90. [Google Scholar]
- Gonzalez-Perez, V.; Lingle, C.J. Regulation of BK channels by beta and gamma subunits. Annu. Rev. Physiol. 2019, 81, 113–137. [Google Scholar] [CrossRef]
- Tao, X.; MacKinnon, R. Molecular structures of the human Slo1 K+ channel in complex with β4. eLife 2019, 8, e51409. [Google Scholar] [CrossRef]
- Tao, X.; Hite, R.K.; MacKinnon, R. Cryo-EM structure of the open high-conductance Ca 2+-activated K+ channel. Nature 2017, 541, 46–51. [Google Scholar] [CrossRef]
- Han, K.; Wang, M.; Zhang, L.; Wang, Y.; Guo, M.; Zhao, M.; Zhao, Q.; Zhang, Y.; Zeng, N.; Wang, C. Predicting Ion Channels Genes and Their Types With Machine Learning Techniques. Front. Genet. 2019, 10, 399. [Google Scholar] [CrossRef]
- Celik, N.; O’Brien, F.; Brennan, S.; Rainbow, R.D.; Dart, C.; Zheng, Y.; Coenen, F.; Barrett-Jolley, R. Deep-Channel uses deep neural networks to detect single-molecule events from patch-clamp data. Commun. Biol. 2020, 3, 1–10. [Google Scholar] [CrossRef]
- Sakmann, B. Single-Channel Recording; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Piwońska, M.; Szewczyk, A.; Schröder, U.H.; Reymann, K.G.; Bednarczyk, P. Effectors of large-conductance calcium-activated potassium channel modulate glutamate excitotoxicity in organotypic hippocampal slice cultures. Acta Neurobiol. Exp. 2016, 76, 20–31. [Google Scholar] [CrossRef]
- Bednarczyk, P.; Koziel, A.; Jarmuszkiewicz, W.; Szewczyk, A. Large-conductance Ca2+-activated potassium channel in mitochondria of endothelial EA. hy926 cells. Am. J. Physiol. Heart Circ. Physiol. 2013, 304, H1415–H1427. [Google Scholar] [CrossRef] [PubMed]
- Kicinska, A.; Augustynek, B.; Kulawiak, B.; Jarmuszkiewicz, W.; Szewczyk, A.; Bednarczyk, P. A large-conductance calcium-regulated K+ channel in human dermal fibroblast mitochondria. Biochem. J. 2016, 473, 4457–4471. [Google Scholar] [CrossRef] [PubMed]
- Chaovalitwongse, W.A.; Fan, Y.J.; Sachdeo, R.C. On the time series k-nearest neighbor classification of abnormal brain activity. IEEE Trans. Syst. Man Cybern. Part A Syst. Hum. 2007, 37, 1005–1016. [Google Scholar] [CrossRef]
- Murugappan, M. Electromyogram signal based human emotion classification using KNN and LDA. In Proceedings of the 2011 IEEE International Conference on System Engineering and Technology, Shah Alam, Malaysia, 27–28 June 2011; pp. 106–110. [Google Scholar]
- Saini, I.; Singh, D.; Khosla, A. QRS detection using K-Nearest Neighbor algorithm (KNN) and evaluation on standard ECG databases. J. Adv. Res. 2013, 4, 331–344. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.M.; Walsh, J.R.; Long, J.; Davis, C.B.; Henstock, P.; Hodge, M.R.; Maciejewski, M.; Mu, X.J.; Ra, S.; Zhang, S.; et al. Deep learning of representations for transcriptomics-based phenotype prediction. BioRxiv 2019, 574723. [Google Scholar] [CrossRef]
- Deng, Y.; Bao, F.; Dai, Q.; Wu, L.F.; Altschuler, S.J. Scalable analysis of cell-type composition from single-cell transcriptomics using deep recurrent learning. Nat. Methods 2019, 16, 311–314. [Google Scholar] [CrossRef]
- Abdolhosseini, F.; Azarkhalili, B.; Maazallahi, A.; Kamal, A.; Motahari, S.A.; Sharifi-Zarchi, A.; Chitsaz, H. Cell identity codes: Understanding cell identity from gene expression profiles using deep neural networks. Sci. Rep. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Danaee, P.; Ghaeini, R.; Hendrix, D.A. A deep learning approach for cancer detection and relevant gene identification. In Pacific Symposium on Biocomputing 2017; World Scientific: Singapore, 2017; pp. 219–229. [Google Scholar]
- Wen, T.; Zhang, Z. Deep convolution neural network and autoencoders-based unsupervised feature learning of EEG signals. IEEE Access 2018, 6, 25399–25410. [Google Scholar] [CrossRef]
- Kieu, T.; Yang, B.; Guo, C.; Jensen, C.S. Outlier Detection for Time Series with Recurrent Autoencoder Ensembles. In Proceedings of the IJCAI, Macao, China, 10–16 August 2019; pp. 2725–2732. [Google Scholar]
- Edgell, C.J.; McDonald, C.C.; Graham, J.B. Permanent cell line expressing human factor VIII-related antigen established by hybridization. Proc. Natl. Acad. Sci. USA 1983, 80, 3734–3737. [Google Scholar] [CrossRef]
- Hunt, S.E.; McLaren, W.; Gil, L.; Thormann, A.; Schuilenburg, H.; Sheppard, D.; Parton, A.; Armean, I.M.; Trevanion, S.J.; Flicek, P.; et al. Ensembl variation resources. Database 2018. [Google Scholar] [CrossRef]
- Benzinger, G.R.; Xia, X.M.; Lingle, C.J. Direct observation of a preinactivated, open state in BK channels with β2 subunits. J. Gen. Physiol. 2006, 127, 119–131. [Google Scholar] [CrossRef] [PubMed]
- Wallner, M.; Meera, P.; Toro, L. Molecular basis of fast inactivation in voltage and Ca2+-activated K+ channels: A transmembrane β-subunit homolog. Proc. Natl. Acad. Sci. USA 1999, 96, 4137–4142. [Google Scholar] [CrossRef] [PubMed]
- Mercik, S.; Weron, K.; Siwy, Z. Statistical analysis of ionic current fluctuations in membrane channels. Phys. Rev. E 1999, 60, 7343. [Google Scholar] [CrossRef] [PubMed]
- Longadge, R.; Dongre, S. Class Imbalance Problem in Data Mining Review. Int. J. Comput. Sci. Netw. 2013, 2, 83–87. [Google Scholar]
- Lin, J.; Keogh, E.; Wei, L.; Lonardi, S. Experiencing SAX: A Novel Symbolic Representation of Time Series. Data Min. Knowl. Discov. 2007, 15, 107–144. [Google Scholar] [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, USA, 2016; Available online: http://www.deeplearningbook.org (accessed on 1 June 2020).
- Kramer, M. Nonlinear principal component analysis using autoassosiative neural networks. AIChE J. 1991, 37, 233–243. [Google Scholar] [CrossRef]
- Hinton, G.E.; Salakhutdinov, R.R. Reducing the dimensionality of data with neural networks. Science 2006, 313, 504–507. [Google Scholar] [CrossRef]
- Xia, Y.; Cao, X.; Wen, F.; Hua, G.; Sun, J. Learning discriminative reconstructions for unsupervised outlier removal. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 11–18 September 2015; pp. 1511–1519. [Google Scholar]
- Yang, K.; Shahabi, C. An efficient k nearest neighbor search for multivariate time series. Inf. Comput. 2007, 205, 65–98. [Google Scholar] [CrossRef]
- Cawley, G.; Talbot, N. On Over-fitting in Model Selection and Subsequent Selection Bias in Performance Evaluation. J. Mach. Learn. Res. 2010, 11, 2079–2107. [Google Scholar]
- MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proceedings of the Fifth Berkeley Symposium on Mathematical Statistics and Probability, Oakland, CA, USA, 18–21 June 1967; Volume 1, pp. 281–297. [Google Scholar]


| Pipette Potential [mV] | Acc without AD | Acc with AD | Dist FH | Dist FE | Dist HE |
|---|---|---|---|---|---|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Richter-Laskowska, M.; Trybek, P.; Bednarczyk, P.; Wawrzkiewicz-Jałowiecka, A. Application of Machine-Learning Methods to Recognize mitoBK Channels from Different Cell Types Based on the Experimental Patch-Clamp Results. Int. J. Mol. Sci. 2021, 22, 840. https://doi.org/10.3390/ijms22020840
Richter-Laskowska M, Trybek P, Bednarczyk P, Wawrzkiewicz-Jałowiecka A. Application of Machine-Learning Methods to Recognize mitoBK Channels from Different Cell Types Based on the Experimental Patch-Clamp Results. International Journal of Molecular Sciences. 2021; 22(2):840. https://doi.org/10.3390/ijms22020840
Chicago/Turabian StyleRichter-Laskowska, Monika, Paulina Trybek, Piotr Bednarczyk, and Agata Wawrzkiewicz-Jałowiecka. 2021. "Application of Machine-Learning Methods to Recognize mitoBK Channels from Different Cell Types Based on the Experimental Patch-Clamp Results" International Journal of Molecular Sciences 22, no. 2: 840. https://doi.org/10.3390/ijms22020840
APA StyleRichter-Laskowska, M., Trybek, P., Bednarczyk, P., & Wawrzkiewicz-Jałowiecka, A. (2021). Application of Machine-Learning Methods to Recognize mitoBK Channels from Different Cell Types Based on the Experimental Patch-Clamp Results. International Journal of Molecular Sciences, 22(2), 840. https://doi.org/10.3390/ijms22020840






