Differential Regulation of NAPDH Oxidases in Salt-Tolerant Eutrema salsugineum and Salt-Sensitive Arabidopsis thaliana
Abstract
:1. Introduction
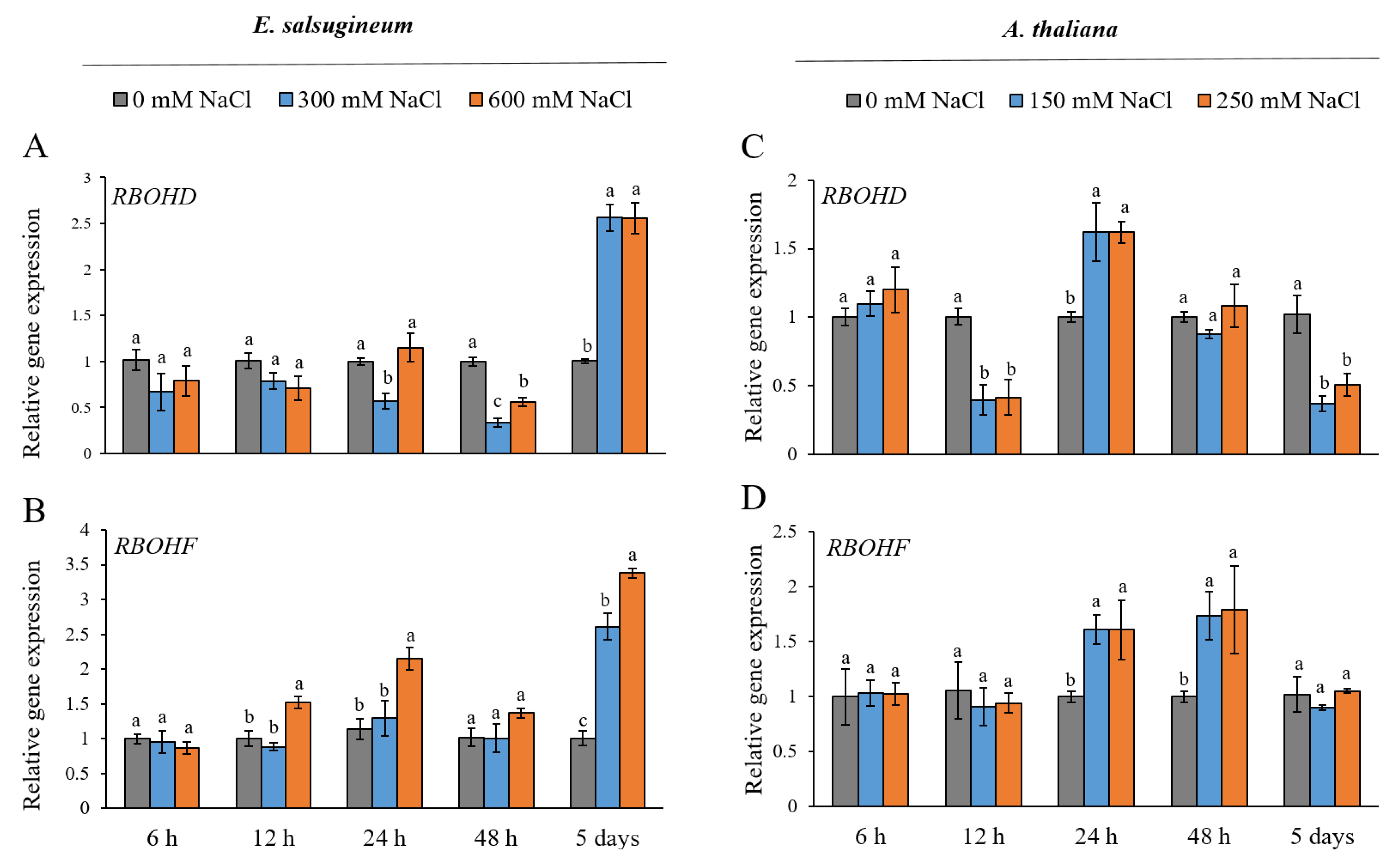
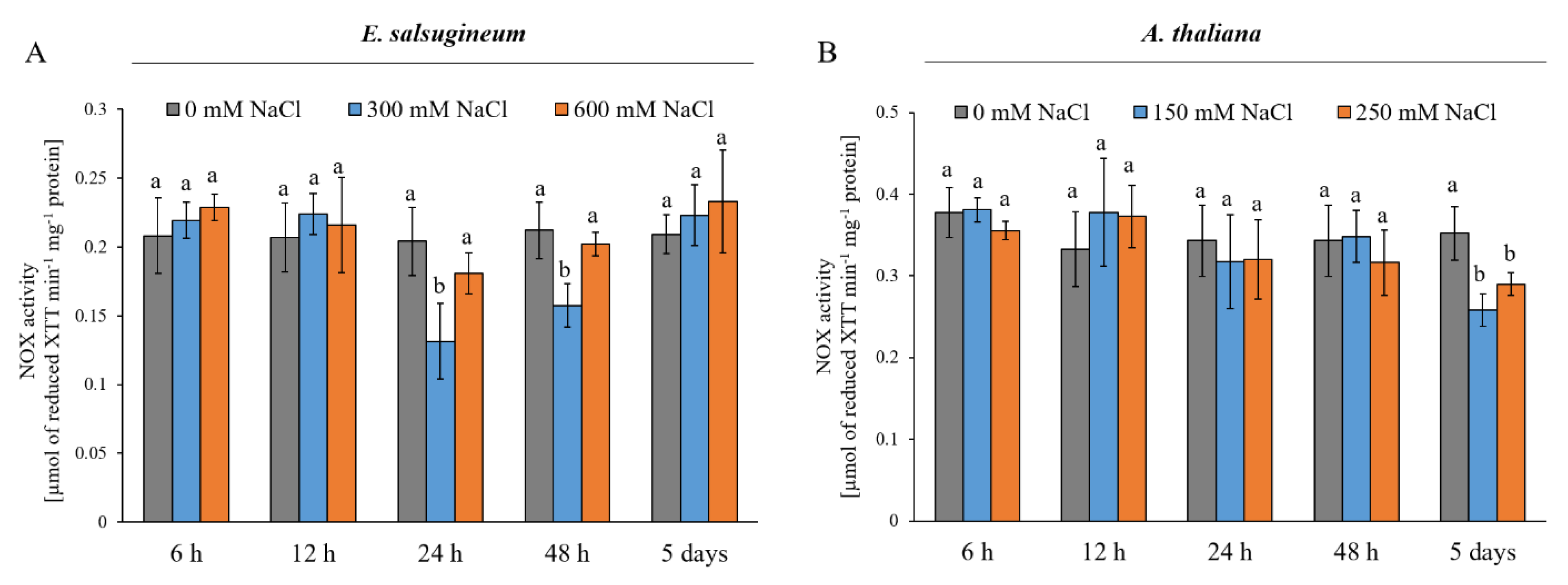
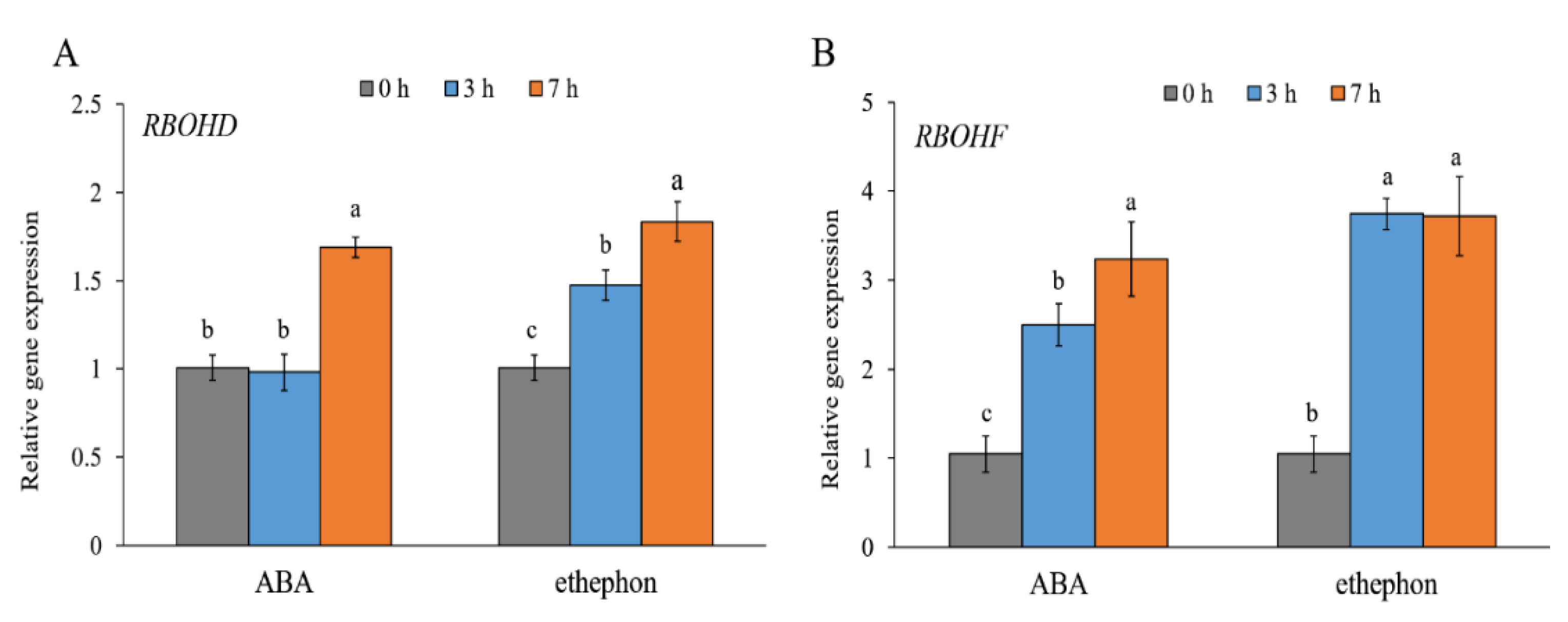
2. Results
3. Discussion
4. Methods
4.1. Plant Material, Growth Conditions, and Treatments
4.2. Database Search and Prediction of Cis-Acting Regulatory Elements
4.3. Gene Expression Analysis
4.4. Membrane Protein Extraction
4.5. NADPH Oxidase (NOX) Activity
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pan, T.; Liu, M.; Kreslavski, V.D.; Zharmukhamedov, S.K.; Nie, C.; Yu, M.; Kuznetsov, V.V.; Allakhverdiev, S.I.; Shabala, S. Non-stomatal limitation of photosynthesis by soil salinity. Crit. Rev. Environ. Sci. Technol. 2020, 51, 791–825. [Google Scholar] [CrossRef]
- Zhao, C.; Zhang, H.; Song, C.; Zhu, J.K.; Shabala, S. Mechanisms of plant responses and adaptation to soil salinity. Innovation 2020, 1, 100017. [Google Scholar] [CrossRef]
- Bose, J.; Rodrigo-Moreno, A.; Shabala, S. ROS homeostasis in halophytes in the context of salinity stress tolerance. J. Exp. Bot. 2014, 65, 1241–1257. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Khare, T.; Sharma, M.; Wani, S.H. ROS-induced signaling and gene expression in crops under salinity stress. In Reactive Oxygen Species and Antioxidant Systems in Plants: Role and Regulation under Abiotic Stress; Khan, M.I.R., Khan, N.A., Eds.; Springer: Singapore, 2017; pp. 159–184. [Google Scholar]
- Møller, I.M.; Jensen, P.E.; Hansson, A. Oxidative modifications to cellular components in plants. Annu. Rev. Plant Biol. 2007, 58, 459–481. [Google Scholar] [CrossRef] [Green Version]
- Foyer, C.H.; Ruban, A.V.; Noctor, G. Viewing oxidative stress through the lens of oxidative signaling rather than damage. Biochem. J. 2017, 474, 877–883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Czarnocka, W.; Karpiński, S. Friend or foe? Reactive oxygen species production, scavenging and signaling in plant response to environmental stresses. Free Radic. Biol. Med. 2018, 122, 4–20. [Google Scholar] [CrossRef] [PubMed]
- Fichman, Y.; Mittler, R. Rapid systemic signaling during abiotic and biotic stresses: Is the ROS wave master of all trades? Plant J. 2020, 102, 887–896. [Google Scholar] [CrossRef] [Green Version]
- Kaur, G.; Guruprasad, K.; Temple, B.R.; Shirvanyants, D.G.; Dokholyan, N.V.; Pati, P.K. Structural complexity and functional diversity of plant NADPH oxidases. Amino Acids. 2018, 50, 79–94. [Google Scholar] [CrossRef]
- Suzuki, N.; Miller, G.; Morales, J.; Shulaev, V.; Torres, M.A.; Mittler, R. Respiratory burst oxidases: The engines of ROS signaling. Curr. Opin. Plant Biol. 2011, 14, 691–699. [Google Scholar] [CrossRef]
- Sagi, M.; Fluhr, R. Production of reactive oxygen species by plant NADPH oxidases. Plant Physiol. 2006, 141, 336–340. [Google Scholar] [CrossRef] [Green Version]
- Kaur, G.; Pati, P.K. Analysis of cis-acting regulatory elements of respiratory burst oxidase homolog (Rboh) gene families in Arabidopsis and rice provides clues for their diverse functions. Comput Biol. Chem. 2016, 62, 104–118. [Google Scholar] [CrossRef]
- Müller, K.; Carstens, A.C.; Linkies, A.; Torres, M.A.; Leubner-Metzger, G. The NADPH-oxidase AtrbohB plays a role in Arabidopsis seed after-ripening. New Phytol. 2009, 184, 885–897. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwak, J.M.; Mori, I.C.; Pei, Z.M.; Leonhardt, N.; Torres, M.A.; Dangl, J.L.; Bloom, R.E.; Bodde, S.; Jones, J.D.; Schroeder, J.I. NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J. 2003, 22, 2623–2633. [Google Scholar] [CrossRef] [PubMed]
- Monshausen, G.B.; Bibikova, T.N.; Messerli, M.A.; Shi, C.; Gilroy, S. Oscillations in extracellular pH and reactive oxygen species modulate tip growth of Arabidopsis root hairs. Proc. Natl. Acad. Sci. USA 2007, 104, 20996–21001. [Google Scholar] [CrossRef] [Green Version]
- Potocký, M.; Jones, M.A.; Bezvoda, R.; Smirnoff, N.; Žárský, V. Reactive oxygen species produced by NADPH oxidase are involved in pollen tube growth. New Phytol. 2007, 174, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Larkindale, J.; Hall, J.D.; Knight, M.R.; Vierling, E. Heat stress phenotypes of Arabidopsis mutants implicate multiple signaling pathways in the acquisition of thermotolerance. Plant Physiol. 2005, 138, 882–897. [Google Scholar] [CrossRef] [Green Version]
- Miller, G.; Schlauch, K.; Tam, R.; Cortes, D.; Torres, M.A.; Shulaev, V.; Dangl, J.L.; Mittler, R. The plant NADPH oxidase RbohD mediates rapid systemic signaling in response to diverse stimuli. Sci. Signal. 2009, 2, ra45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Zhang, M.M.; Wang, Y.J.; Gao, Y.T.; Li, R.; Wang, G.F.; Li, V.Q.; Liu, W.T.; Chen, K.M. The plasma membrane NADPH oxidase OsRbohA plays a crucial role in developmental regulation and drought-stress response in rice. Physiol. Plant. 2016, 156, 421–443. [Google Scholar] [CrossRef]
- Ma, L.; Zhang, H.; Sun, L.; Jiao, Y.; Zhang, G.; Miao, C.; Hao, F. NADPH oxidase AtrbohD and AtrbohF function in ROS dependent regulation of Na+/K+ homeostasis in Arabidopsis under salt stress. J. Exp. Bot. 2012, 63, 305–317. [Google Scholar] [CrossRef]
- Rejeb, K.B.; Benzarti, M.; Debez, A.; Bailly, C.; Savouré, A.; Abdelly, C. NADPH oxidase-dependent H2O2 production is required for salt-induced antioxidant defense in Arabidopsis thaliana. J. Plant Physiol. 2015, 174, 5–15. [Google Scholar] [CrossRef]
- Ogasawara, Y.; Kaya, H.; Hiraoka, G.; Yumoto, F.; Kimura, S.; Kadota, Y.; Hishinuma, H.; Senzaki, E.; Yamagoe, S.; Nagata, K.; et al. Synergistic activation of the Arabidopsis NADPH oxidase AtrbohD by Ca2+ and phosphorylation. J. Biol. Chem. 2008, 283, 8885–8892. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Li, M.; Yu, L.; Zhou, Z.; Liang, X.; Liu, Z.; Cai, G.; Gao, L.; Zhang, X.; Wang, Y.; et al. The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe. 2014, 15, 329–338. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Tan, J.; Guo, Z.; Lu, S.; He, S.; Shu, W.; Zhou, B. Increased abscisic acid levels in transgenic tobacco over-expressing 9 cis-epoxycarotenoid dioxygenase influence H2O2 and NO production and antioxidant defences. Plant Cell Environ. 2009, 32, 509–519. [Google Scholar] [CrossRef]
- Evans, M.J.; Choi, W.G.; Gilroy, S.; Morris, R.J. A ROS-assisted calcium wave dependent on the AtRBOHD NADPH oxidase and TPC1 cation channel propagates the systemic response to salt stress. Plant Physiol. 2016, 171, 1771–1784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, J.S.; Zhu, J.K.; Bressan, R.A.; Hasegawa, P.M.; Shi, H. Reactive oxygen species mediate Na+-induced SOS1 mRNA stability in Arabidopsis. Plant J. 2008, 53, 554–565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ben Rejeb, K.B.; Vos, D.L.; Disquet, I.L.; Leprince, A.; Bordenave, M.; Maldiney, R.; Jdey, A.; Abdelly, C.; Savouré, A. Hydrogen peroxide produced by NADPH oxidases increases proline accumulation during salt or mannitol stress in Arabidopsis thaliana. New Phytol. 2015, 208, 138–1148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grigore, M.N. Defining halophytes: A conceptual and historical approach in an ecological frame. In Halophytes and Climate Change: Adaptive Mechanisms and Potential Uses; Hasanuzzaman, M., Shabala, S., Fujita, M., Eds.; CABI: Boston, MA, USA, 2019; pp. 3–18. [Google Scholar]
- Niewiadomska, E.; Wiciarz, M. Adaptations of chloroplastic metabolism in halophytic plants. In Progress in Botany; Lüttge, U., Beyschlag, W., Eds.; Springer: Cham, Switzerland, 2015; pp. 177–193. [Google Scholar]
- Bose, J.; Munns, R.; Shabala, S.; Gilliham, M.; Pogson, B.; Tyerman, S.D. Chloroplast function and ion regulation in plants growing on saline soils: Lessons from halophytes. J. Exp. Bot. 2017, 68, 3129–3143. [Google Scholar] [CrossRef] [PubMed]
- Kazachkova, Y.; Eshel, G.; Pantha, P.; Cheeseman, J.M.; Dassanayake, M.; Barak, S. Halophytism: What have we learnt from Arabidopsis thaliana relative model systems? Plant Physiol. 2018, 178, 972–988. [Google Scholar] [CrossRef] [Green Version]
- Ozgur, R.; Uzilday, B.; Sekmen, A.H.; Turkan, I. Reactive oxygen species regulation and antioxidant defence in halophytes. Funct. Plant Biol. 2013, 40, 832–847. [Google Scholar] [CrossRef]
- Surówka, E.; Latowski, D.; Libik-Konieczny, M.; Miszalski, Z. ROS Signalling, and Antioxidant Defence Network in Halophytes. In Halophytes and Climate Change: Adaptive Mechanisms and Potential Uses; Hasanuzzaman, M., Shabala, S., Fujita, M., Eds.; CABI: Boston, MA, USA, 2019; pp. 179–195. [Google Scholar]
- Wiciarz, M.; Gubernator, B.; Kruk, J.; Niewiadomska, E. Enhanced chloroplastic generation of H2O2 in stress-resistant Thellungiella salsuginea in comparison to Arabidopsis thaliana. Physiol. Plant. 2015, 153, 467–476. [Google Scholar] [CrossRef] [Green Version]
- Pilarska, M.; Wiciarz, M.; Jajić, I.; Kozieradzka-Kiszkurno, M.; Dobrev, P.; Vanková, R.; Niewiadomska, E. A different pattern of production and scavenging of reactive oxygen species in halophytic Eutrema salsugineum (Thellungiella salsuginea) plants in comparison to Arabidopsis thaliana and its relation to salt stress signaling. Front. Plant Sci. 2016, 7, 1179. [Google Scholar] [CrossRef] [PubMed]
- Sewelam, N.; Jaspert, N.; Van Der Kelen, K.; Tognetti, V.B.; Schmitz, J.; Frerigmann, H.; Stahl, E.; Zeier, J.; Van Breusegem, F.; Maurino, V.G. Spatial H2O2 signaling specificity: H2O2 from chloroplasts and peroxisomes modulates the plant transcriptome differentially. Mol. Plant 2014, 7, 1191–1210. [Google Scholar] [CrossRef] [Green Version]
- Torres, M.A.; Onouchi, H.; Hamada, S.; Machida, C.; Hammond-Kosack, K.E.; Jones, J.D. Six Arabidopsis thaliana homologues of the human respiratory burst oxidase (gp91phox). Plant J. 1998, 14, 365–370. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Wen, C.K. Preparation of ethylene gas and comparison of ethylene responses induced by ethylene, ACC, and ethephon. Plant Physiol. Bioch. 2010, 48, 45–53. [Google Scholar] [CrossRef]
- Cheng, C.; Xu, X.; Gao, M.; Li, J.; Guo, C.; Song, J.; Wang, X. Genome-wide analysis of respiratory burst oxidase homologs in grape (Vitis vinifera L.). Int. J. Mol. Sci. 2013, 14, 24169–24186. [Google Scholar] [CrossRef]
- Zhu, J.-K. Plant salt tolerance. Trends Plant Sci. 2001, 6, 66–71. [Google Scholar] [CrossRef]
- Hasan, M.; Rahman, A.; Skalicky, M.; Alabdallah, N.; Waseem, M.; Jahan, M.; Ahammed, G.; El-Mogy, M.; El-Yazied, A.; Ibrahim, M.; et al. Ozone Induced Stomatal Regulations, MAPK and Phytohormone Signaling in Plants. Int. J. Mol. Sci. 2021, 22, 6304. [Google Scholar] [CrossRef]
- Hou, Q.; Bartels, D. Comparative study of the aldehyde dehydrogenase (ALDH) gene superfamily in the glycophyte Arabidopsis thaliana and Eutrema halophytes. Ann. Bot. 2014, 115, 465–479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Y.J.; Xu, S.; Han, B.; Wu, M.Z.; Yuan, X.X.; Han, Y.; Gu, Q.; Xu, D.K.; Yang, Q.; Shen, W.B. Evidence of Arabidopsis salt acclimation induced by up-regulation of HY1 and the regulatory role of RbohD-derived reactive oxygen species synthesis. Plant J. 2011, 66, 280–292. [Google Scholar] [CrossRef]
- Srivastava, A.K.; Srivastava, S.; Lokhande, V.H.; D’souza, S.F.; Suprasanna, P. Salt stress reveales differential antioxidant and energetics responses in, glycophyte (Brassica juncea L.) and halophyte (Sesuvium portulacastrum L.). Front. Environ. Sci. 2015, 3, 19. [Google Scholar] [CrossRef] [Green Version]
- Jakubowska, D.; Janicka-Russak, M.; Kabała, K.; Migocka, M.; Reda, M. Modification of plasma membrane NADPH oxidase activity in cucumber seedling roots in response to cadmium stress. Plant Sci. 2015, 234, 50–59. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhu, H.; Zhang, Q.; Li, M.; Yan, M.; Wang, R.; Wang, L.; Welti, R.; Zhang, W.; Wang, X. Phospholipase Da1 and phosphatidic acid regulate NADPH oxidase activity and production of reactive oxygen species in ABA-mediated stomatal closure in Arabidopsis. Plant Cell 2009, 21, 2357–2377. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Smith, J.A.C.; Harberd, N.P.; Jiang, C. The regulatory roles of ethylene and reactive oxygen species (ROS) in plant salt stress responses. Plant Mol. Biol. 2016, 91, 651–659. [Google Scholar] [CrossRef]
- Yu, Z.; Duan, X.; Luo, L.; Dai, S.; Ding, Z.; Xia, G. How plant hormones mediate salt stress responses. Trends Plant Sci. 2020, 25, 1117–1130. [Google Scholar] [CrossRef]
- Sewelam, N.; Kazan, K.; Thomas-Hall, S.R.; Kidd, B.N.; Manners, J.M.; Schenk, P.M. Ethylene response factor 6 is a regulator of reactive oxygen species signaling in Arabidopsis. PLoS ONE 2013, 8, e70289. [Google Scholar] [CrossRef] [Green Version]
- Ellouzi, H.; Ben Hamed, K.; Hernández, I.; Cela, J.; Müller, M.; Magné, C.; Abdelly, C.; Munné-Bosch, S. A comparative study of the early osmotic, ionic, redox and hormonal signaling response in leaves and roots of two halophytes and a glycophyte to salinity. Planta 2014, 240, 1299–1317. [Google Scholar] [CrossRef] [PubMed]
- Arbona, V.; Argamasilla, R.; Gómez-Cadenas, A. Common and divergent physiological, hormonal and metabolic responses of Arabidopsis thaliana and Thellungiella halophila to water and salt stress. J. Plant. Physiol. 2010, 167, 1342–1350. [Google Scholar] [CrossRef] [PubMed]
- Prerostova, S.; Dobrev, P.I.; Gaudinova, A.; Hosek, P.; Soudek, P.; Knirsch, V.; Vankova, R. Hormonal dynamics during salt stress responses of salt-sensitive Arabidopsis thaliana and salt-tolerant Thellungiella salsuginea. Plant Sci. 2017, 264, 188–198. [Google Scholar] [CrossRef] [PubMed]
- Higo, K.; Ugawa, Y.; Iwamoto, M.; Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 1999, 27, 297–300. [Google Scholar] [CrossRef] [Green Version]
- Valenzuela-Avendaño, J.; Mota, I.E.; Uc, G.; Perera, R.; Valenzuela-Soto, E.; Aguilar, J.Z. Use of a simple method to isolate intact RNA from partially hydrated Selaginella lepidophylla plants. Plant Mol. Biol. Rep. 2005, 23, 199–200. [Google Scholar] [CrossRef]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Janeczko, A.; Budziszewska, B.; Skoczowski, A.; Dybała, M. Specific binding sites for progesterone and 17ß-estradiol in cells of Triticum aestivum L. Acta Biochem. Pol. 2008, 55, 708–711. [Google Scholar]
- Bradford, M.M. A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of proteindye- binding. Anal. Biochem. 1975, 72, 248–254. [Google Scholar] [CrossRef]
- Able, A.; Guest, D.; Sutherland, M. Use of a new tetrazolium-based assay to study the production of superoxide radicals by tobacco cell cultures challenged with avirulent zoospores of Phytophthora parasitica var nicotianae. Plant Physiol. 1998, 117, 491–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Potocký, M.; Pejchar, P.; Gutkowska, M.; Jiménez-Quesada, M.J.; Potocká, A.; de Dios, A.J.; Kost, B.; Žárský, V. NADPH oxidase activity in pollen tubes is affected by calcium ions, signaling phospholipids and Rac/Rop GTPases. J. Plant Physiol. 2012, 169, 1654–1663. [Google Scholar] [CrossRef] [PubMed]
| Cis-Acting Regulatory Elements | Core Sequence | Hormone | Number of Elements | |||
|---|---|---|---|---|---|---|
| EsRBOHD | EsRBOHF | AtRBOHD | AtRBOHF | |||
| ERELEE4 | AWTTCAAA | Ethylene | 1 (+) | 1 (−) | 1 (+) 2 (−) | 1 (−) |
| GT1CONSENSUS | GRWAAW | Salicylic acid | 5 (+) 10 (−) | 7 (+) 7 (−) | 7 (+) 6 (−) | 11 (+) 9 (−) |
| WBOXATNPR1 | TTGAC | Salicylic acid | 2 (+) 3 (−) | 2 (−) 2 (−) | 4 (+) 1 (−) | 1 (+) 2 (−) |
| ASF1MOTIFCAMV | TGACG | Salicylic acid, Auxins | - | 1 (−) | - | 3 (−) |
| ARFAT | TGTCTC | Auxins | 1 (+) | - | 1 (−) | - |
| CATATGGMSAUR | CATATG | Auxins | 1 (+) 1 (−) | - | - | - |
| CPBCSPOR | TATTAG | Cytokinins | 1 (+) | 1 (−) | 2 (−) | 1 (+) 3 (−) |
| WRKY71OS | TGAC | Gibberellins | 6 (+) 6 (−) | 4 (+) 4 (−) | 8 (+) 6 (−) | 2 (+) 5 (−) |
| MYBGAHV | TAACAAA | Gibberellins | 1 (−) | 1 (+) | 2 (−) | - |
| GAREAT | TAACAAR | Gibberellins | 1 (−) | 1 (+) | 3 (−) | 1 (−) |
| PYRIMIDINEBOXHVEPB1 | TTTTTTCC | Gibberellins, ABA | - | 1 (+) | - | 1 (−) |
| DPBFCOREDCDC3 | ACACNNG | ABA | 1 (+) | 2 (+) | 2 (+) 2 (−) | 1 (+) |
| RYREPEATVFLEB4 | CATGCATG | ABA | - | 1 (+) 1 (−) | - | - |
| MYB1AT | WAACCA | ABA | 1 (+) 1 (−) | 1 (+) | 1 (+) 2 (−) | 1 (+) |
| MYCATRD22 | CACATG | ABA | 1 (+) | 1 (+) | 1 (+) | - |
| LTRECOREATCOR15 | CCGAC | ABA | - | - | 1 (+) | 1 (+) 1 (−) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pilarska, M.; Bartels, D.; Niewiadomska, E. Differential Regulation of NAPDH Oxidases in Salt-Tolerant Eutrema salsugineum and Salt-Sensitive Arabidopsis thaliana. Int. J. Mol. Sci. 2021, 22, 10341. https://doi.org/10.3390/ijms221910341
Pilarska M, Bartels D, Niewiadomska E. Differential Regulation of NAPDH Oxidases in Salt-Tolerant Eutrema salsugineum and Salt-Sensitive Arabidopsis thaliana. International Journal of Molecular Sciences. 2021; 22(19):10341. https://doi.org/10.3390/ijms221910341
Chicago/Turabian StylePilarska, Maria, Dorothea Bartels, and Ewa Niewiadomska. 2021. "Differential Regulation of NAPDH Oxidases in Salt-Tolerant Eutrema salsugineum and Salt-Sensitive Arabidopsis thaliana" International Journal of Molecular Sciences 22, no. 19: 10341. https://doi.org/10.3390/ijms221910341
APA StylePilarska, M., Bartels, D., & Niewiadomska, E. (2021). Differential Regulation of NAPDH Oxidases in Salt-Tolerant Eutrema salsugineum and Salt-Sensitive Arabidopsis thaliana. International Journal of Molecular Sciences, 22(19), 10341. https://doi.org/10.3390/ijms221910341







