Potent Activity of Hybrid Arthropod Antimicrobial Peptides Linked by Glycine Spacers
Abstract
1. Introduction
2. Results
2.1. Peptide Sequences and Properties
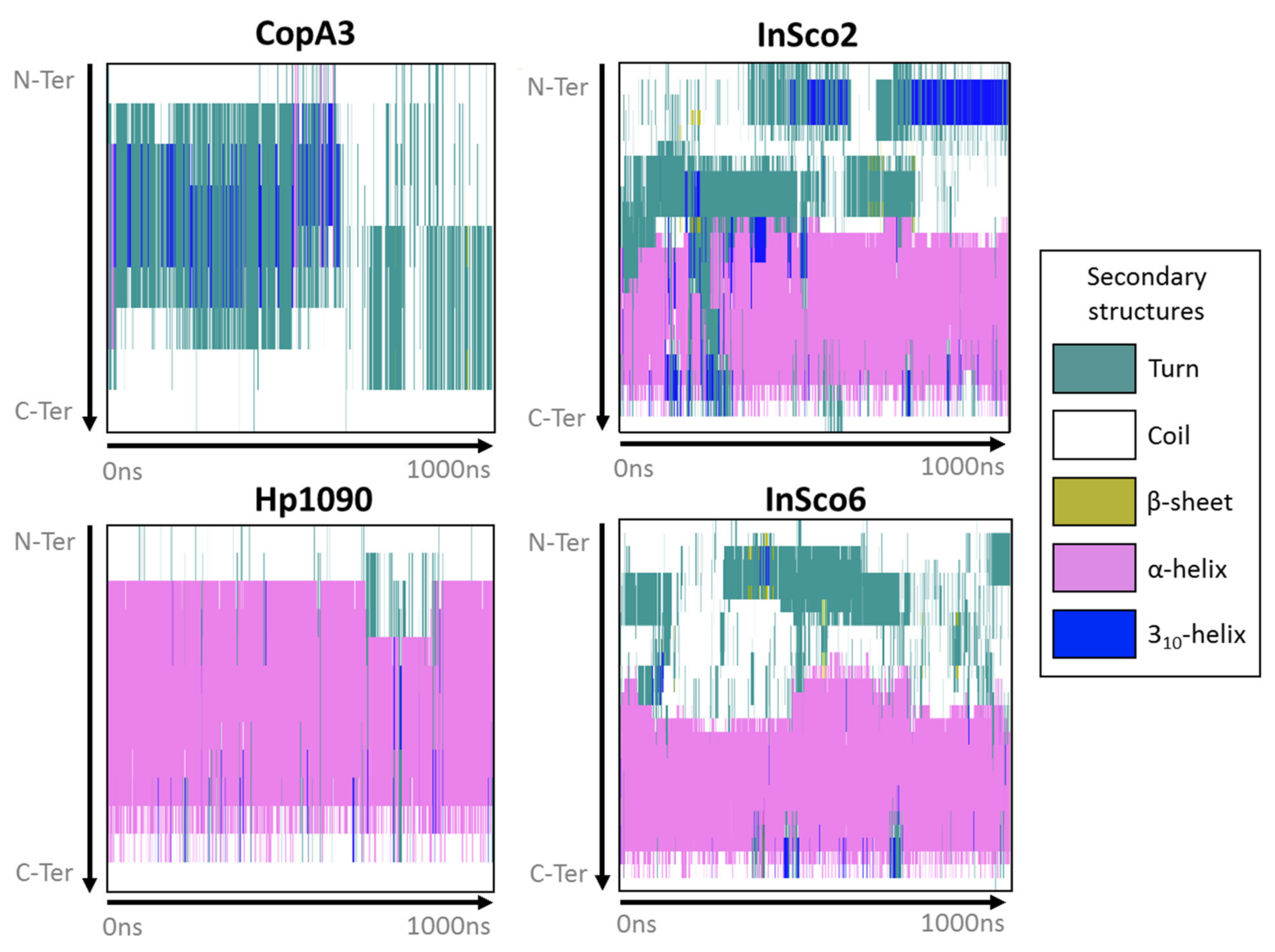
2.2. Predicted 3D Structure and Membrane Polarity
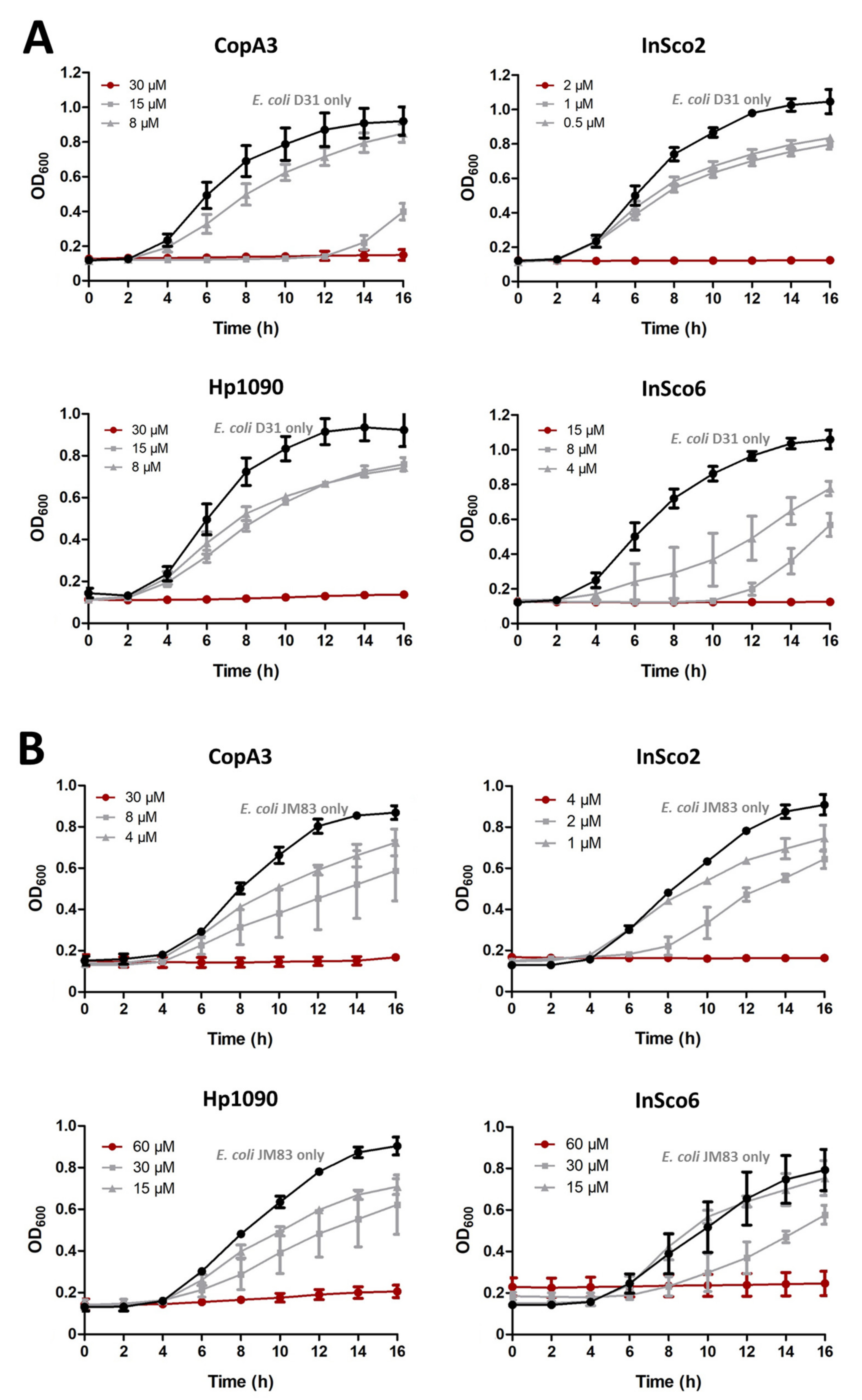
2.3. Antibacterial Activity Assays
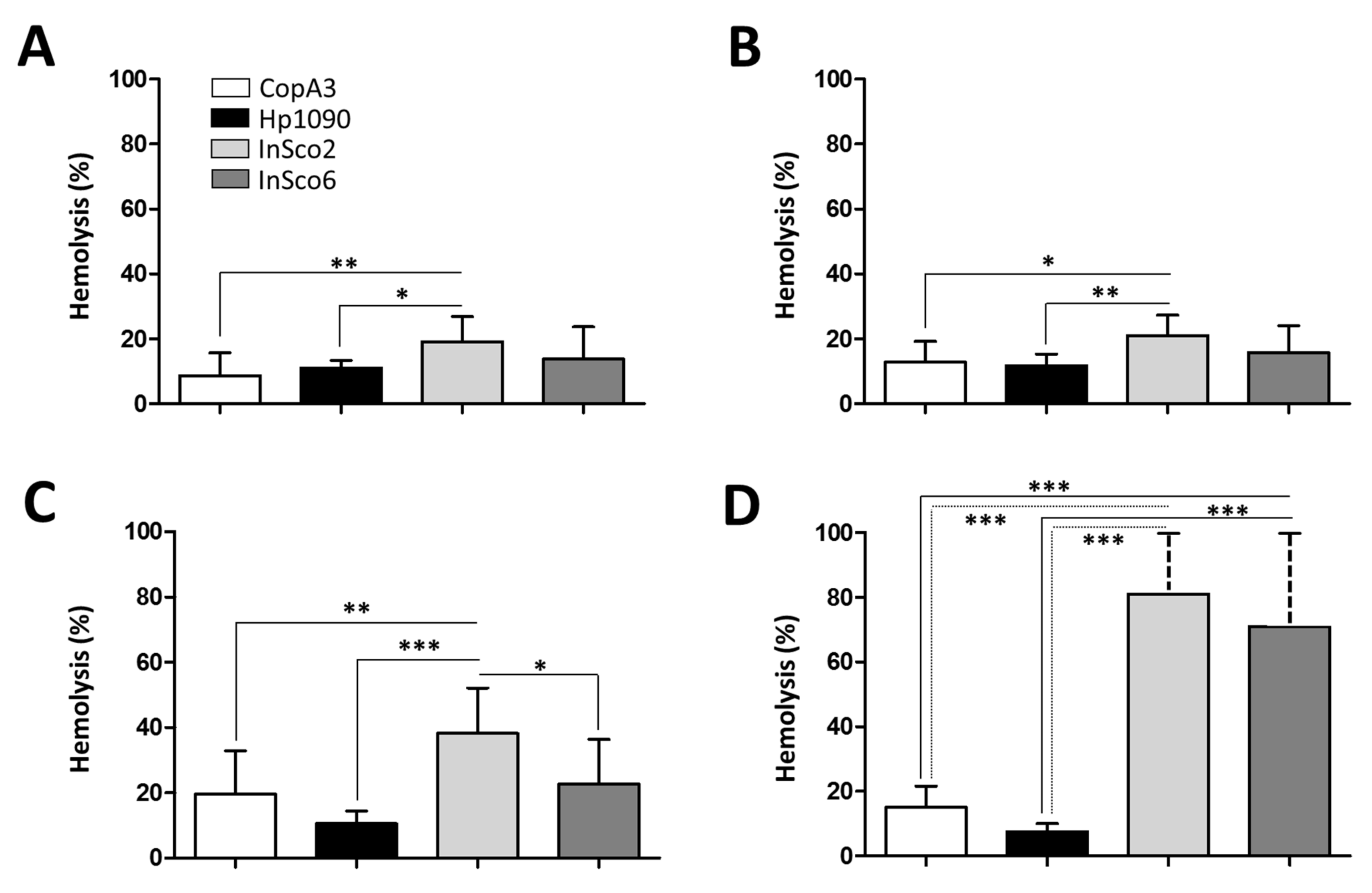
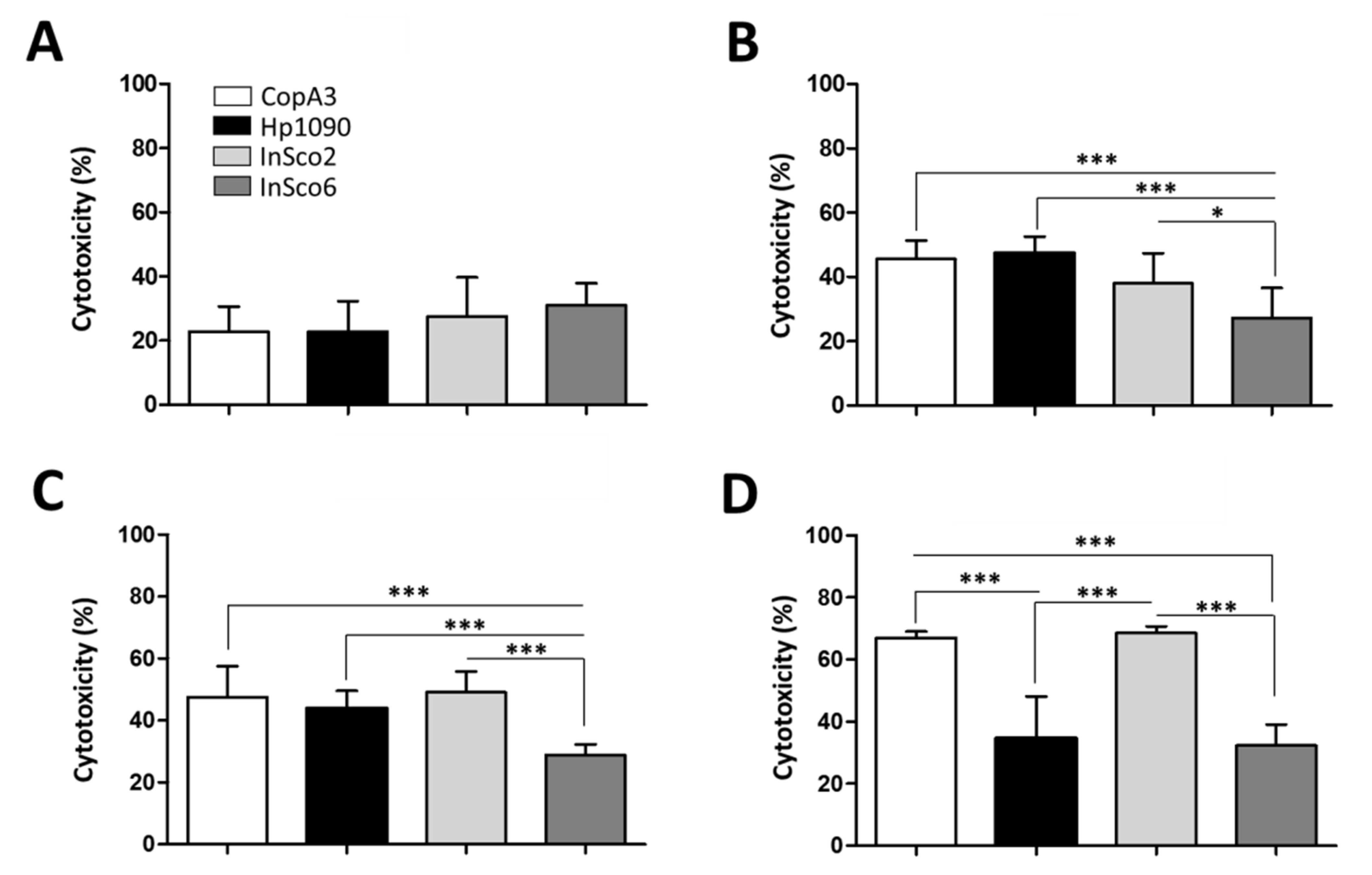
2.4. Screening AMPs for Hemolytic and Cytotoxic Activity
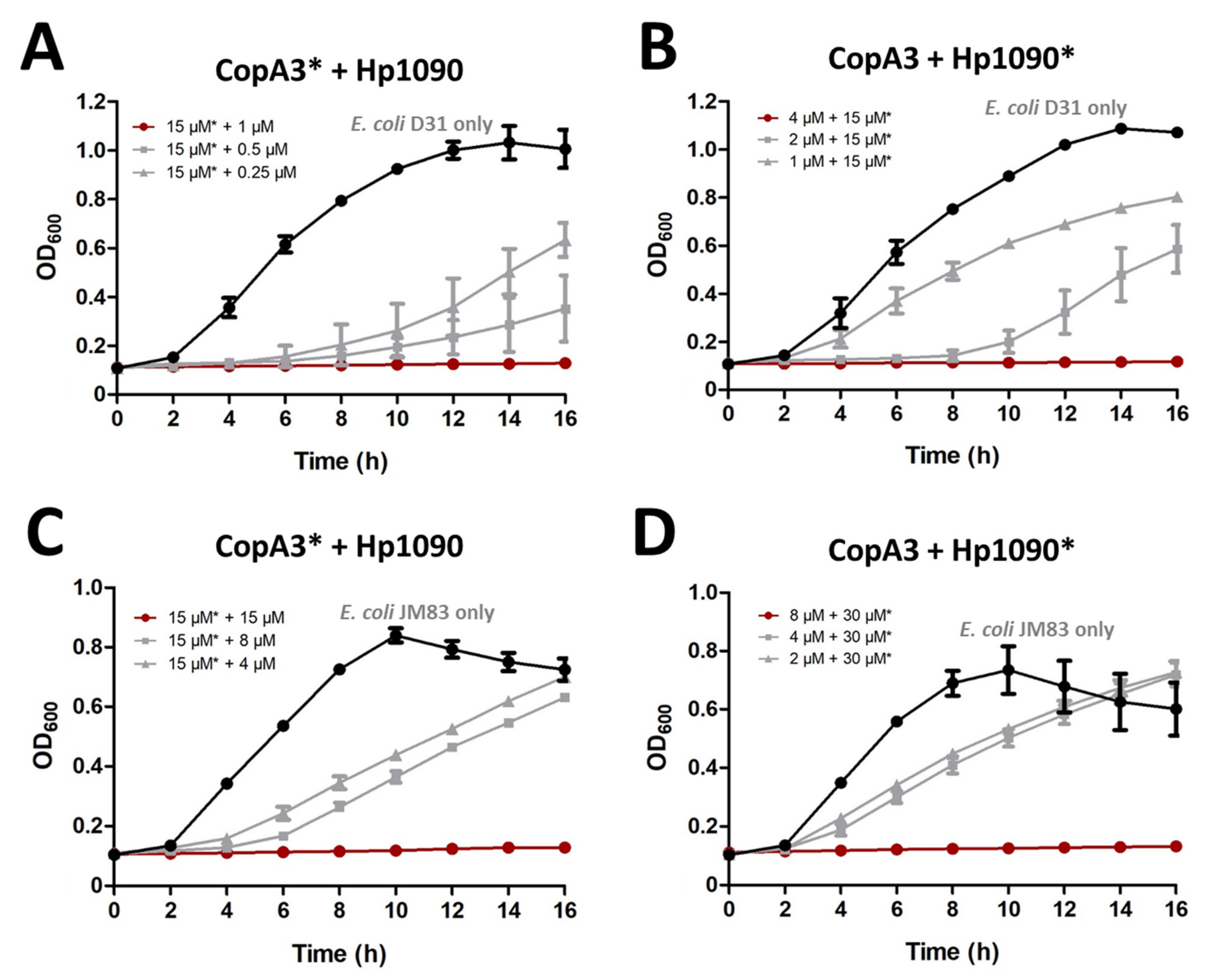
2.5. Synergistic Activity of CopA3 and Hp1090
3. Discussion
4. Materials and Methods
4.1. Peptide Sequences and Bioinformatics
4.2. In silico Peptide Structure Prediction
4.3. Peptide-Membrane Molecular Dynamics
4.4. Minimum Inhibitory Concentration
4.5. Measurement of Hemolytic Activity and Cytotoxicity
4.6. Synergistic Effect of CopA3 and Hp1090
4.7. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stern, S.; Chorzelski, S.; Franken, L.; Völler, S.; Rentmeister, H.; Grosch, B. Breaking through the Wall: A Call for Concerted Action on Antibiotics Research and Development; Global Union for Antibiotics Research and Development (GUARD) Initiative: Berlin, Germany, 2017. [Google Scholar]
- Taneja, N.; Kaur, H. Insights into Newer Antimicrobial Agents Against Gram-Negative Bacteria. Microbiol. Insights 2016, 9, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Kudryashova, E.; Seveau, S.M.; Kudryashov, D.S. Targeting and Inactivation of Bacterial Toxins by Human Defensins. Biol. Chem. 2017, 398, 1069–1085. [Google Scholar] [CrossRef] [PubMed]
- Jenssen, H.; Hamill, P.; Hancock, R.E.W. Peptide Antimicrobial Agents. Clin. Microbiol. Rev. 2006, 19, 491–511. [Google Scholar] [CrossRef] [PubMed]
- Lei, J.; Sun, L.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q. The Antimicrobial Peptides and Their Potential Clinical Applications. Am. J. Transl. Res. 2019, 11, 3919–3931. [Google Scholar] [PubMed]
- Mylonakis, E.; Podsiadlowski, L.; Muhammed, M.; Vilcinskas, A. Diversity, Evolution and Medical Applications of Insect Antimicrobial Peptides. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2016, 371. [Google Scholar] [CrossRef]
- Tonk, M.; Vilcinskas, A. The Medical Potential of Antimicrobial Peptides from Insects. Curr. Top. Med. Chem. 2017, 17, 554–575. [Google Scholar] [CrossRef] [PubMed]
- Vilcinskas, A.; Mukherjee, K.; Vogel, H. Expansion of the Antimicrobial Peptide Repertoire in the Invasive Ladybird Harmonia Axyridis. Proc. R. Soc. B Biol. Sci. 2013, 280, 20122113. [Google Scholar] [CrossRef] [PubMed]
- Gerardo, N.M.; Altincicek, B.; Anselme, C.; Atamian, H.; Barribeau, S.M.; de Vos, M.; Duncan, E.J.; Evans, J.D.; Gabaldón, T.; Ghanim, M.; et al. Immunity and Other Defenses in Pea Aphids, Acyrthosiphon Pisum. Genome Biol. 2010, 11, R21. [Google Scholar] [CrossRef]
- Choi, J.H.; Jang, A.Y.; Lin, S.; Lim, S.; Kim, D.; Park, K.; Han, S.-M.; Yeo, J.-H.; Seo, H.S. Melittin, a Honeybee Venom-Derived Antimicrobial Peptide, May Target Methicillin-Resistant Staphylococcus Aureus. Mol. Med. Rep. 2015, 12, 6483–6490. [Google Scholar] [CrossRef]
- Lu, X.; Shen, J.; Jin, X.; Ma, Y.; Huang, Y.; Mei, H.; Chu, F.; Zhu, J. Bactericidal Activity of Musca Domestica Cecropin (Mdc) on Multidrug-Resistant Clinical Isolate of Escherichia Coli. Appl. Microbiol. Biotechnol. 2012, 95, 939–945. [Google Scholar] [CrossRef]
- Oñate-Garzón, J.; Manrique-Moreno, M.; Trier, S.; Leidy, C.; Torres, R.; Patiño, E. Antimicrobial Activity and Interactions of Cationic Peptides Derived from Galleria Mellonella Cecropin D-like Peptide with Model Membranes. J. Antibiot. (Tokyo) 2017, 70, 238–245. [Google Scholar] [CrossRef]
- Ulvatne, H.; Karoliussen, S.; Stiberg, T.; Rekdal, O.; Svendsen, J.S. Short Antibacterial Peptides and Erythromycin Act Synergically against Escherichia Coli. J. Antimicrob. Chemother. 2001, 48, 203–208. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zerweck, J.; Strandberg, E.; Kukharenko, O.; Reichert, J.; Bürck, J.; Wadhwani, P.; Ulrich, A.S. Molecular Mechanism of Synergy between the Antimicrobial Peptides PGLa and Magainin 2. Sci. Rep. 2017, 7, 13153. [Google Scholar] [CrossRef] [PubMed]
- Bagheri, M.; Beyermann, M.; Dathe, M. Immobilization Reduces the Activity of Surface-Bound Cationic Antimicrobial Peptides with No Influence upon the Activity Spectrum. Antimicrob. Agents Chemother. 2009, 53, 1132–1141. [Google Scholar] [CrossRef]
- Liu, S.P.; Zhou, L.; Lakshminarayanan, R.; Beuerman, R.W. Multivalent Antimicrobial Peptides as Therapeutics: Design Principles and Structural Diversities. Int. J. Pept. Res. Ther. 2010, 16, 199–213. [Google Scholar] [CrossRef] [PubMed]
- McKenna, M. Antibiotic Resistance: The Last Resort. Nature 2013, 499, 394–396. [Google Scholar] [CrossRef] [PubMed]
- Mojsoska, B.; Zuckermann, R.N.; Jenssen, H. Structure-Activity Relationship Study of Novel Peptoids That Mimic the Structure of Antimicrobial Peptides. Antimicrob. Agents Chemother. 2015, 59, 4112–4120. [Google Scholar] [CrossRef]
- Klubthawee, N.; Adisakwattana, P.; Hanpithakpong, W.; Somsri, S.; Aunpad, R. A Novel, Rationally Designed, Hybrid Antimicrobial Peptide, Inspired by Cathelicidin and Aurein, Exhibits Membrane-Active Mechanisms against Pseudomonas Aeruginosa. Sci. Rep. 2020, 10, 9117. [Google Scholar] [CrossRef]
- Hwang, J.-S.; Lee, J.; Kim, Y.-J.; Bang, H.-S.; Yun, E.-Y.; Kim, S.-R.; Suh, H.-J.; Kang, B.-R.; Nam, S.-H.; Jeon, J.-P.; et al. Isolation and Characterization of a Defensin-Like Peptide (Coprisin) from the Dung Beetle, Copris Tripartitus. Int. J. Pept. 2009, 2009, 136284. [Google Scholar] [CrossRef]
- Kim, I.-W.; Kim, S.-J.; Kwon, Y.-N.; Yun, E.-Y.; Ahn, M.-Y.; Kang, D.-C.; Hwang, J.-S. Effects of the Synthetic Coprisin Analog Peptide, CopA3 in Pathogenic Microorganisms and Mammalian Cancer Cells. J. Microbiol. Biotechnol. 2012, 22, 156–158. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Kim, I.-W.; Kim, S.-H.; Yun, E.-Y.; Nam, S.-H.; Ahn, M.-Y.; Kang, D.-C.; Hwang, J.S. Anticancer Activity of CopA3 Dimer Peptide in Human Gastric Cancer Cells. BMB Rep. 2015, 48, 324–329. [Google Scholar] [CrossRef]
- Yan, R.; Zhao, Z.; He, Y.; Wu, L.; Cai, D.; Hong, W.; Wu, Y.; Cao, Z.; Zheng, C.; Li, W. A New Natural α-Helical Peptide from the Venom of the Scorpion Heterometrus Petersii Kills HCV. Peptides 2011, 32, 11–19. [Google Scholar] [CrossRef]
- Lee, E.; Kim, J.-K.; Shin, S.; Jeong, K.-W.; Shin, A.; Lee, J.; Lee, D.G.; Hwang, J.-S.; Kim, Y. Insight into the Antimicrobial Activities of Coprisin Isolated from the Dung Beetle, Copris Tripartitus, Revealed by Structure–Activity Relationships. Biochim. Biophys. Acta BBA-Biomembr. 2013, 1828, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Meletiadis, J.; Pournaras, S.; Roilides, E.; Walsh, T.J. Defining Fractional Inhibitory Concentration Index Cutoffs for Additive Interactions Based on Self-Drug Additive Combinations, Monte Carlo Simulation Analysis, and In Vitro-In Vivo Correlation Data for Antifungal Drug Combinations against Aspergillus Fumigatus. Antimicrob. Agents Chemother. 2010, 54, 602–609. [Google Scholar] [CrossRef] [PubMed]
- Rios, A.C.; Moutinho, C.G.; Pinto, F.C.; Del Fiol, F.S.; Jozala, A.; Chaud, M.V.; Vila, M.M.D.C.; Teixeira, J.A.; Balcão, V.M. Alternatives to Overcoming Bacterial Resistances: State-of-the-Art. Microbiol. Res. 2016, 191, 51–80. [Google Scholar] [CrossRef]
- Li, Y.; Xiang, Q.; Zhang, Q.; Huang, Y.; Su, Z. Overview on the Recent Study of Antimicrobial Peptides: Origins, Functions, Relative Mechanisms and Application. Peptides 2012, 37, 207–215. [Google Scholar] [CrossRef]
- Aoki, W.; Ueda, M. Characterization of Antimicrobial Peptides toward the Development of Novel Antibiotics. Pharmaceuticals 2013, 6, 1055–1081. [Google Scholar] [CrossRef]
- Kim, J.-S.; Jeong, J.-H.; Kim, Y. Design and Engineering of Antimicrobial Peptides Based on LPcin-YK3, an Antimicrobial Peptide Derivative from Bovine Milk. J. Microbiol. Biotechnol. 2018, 28, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhao, C.; Liang, G.; Zhang, M.; Zheng, J. Engineering Antimicrobial Peptides with Improved Antimicrobial and Hemolytic Activities. J. Chem. Inf. Model. 2013, 53, 3280–3296. [Google Scholar] [CrossRef] [PubMed]
- Sani, M.-A.; Saenger, C.; Juretic, D.; Separovic, F. Glycine Substitution Reduces Antimicrobial Activity and Helical Stretch of DiPGLa-H in Lipid Micelles. J. Phys. Chem. B 2017, 121, 4817–4822. [Google Scholar] [CrossRef] [PubMed]
- Taraballi, F.; Natalello, A.; Campione, M.; Villa, O.; Doglia, S.M.; Paleari, A.; Gelain, F. Glycine-Spacers Influence Functional Motifs Exposure and Self-Assembling Propensity of Functionalized Substrates Tailored for Neural Stem Cell Cultures. Front. Neuroeng. 2010, 3. [Google Scholar] [CrossRef]
- Ishida, A.; Watanabe, G.; Oshikawa, M.; Ajioka, I.; Muraoka, T. Glycine Substitution Effects on the Supramolecular Morphology and Rigidity of Cell-Adhesive Amphiphilic Peptides. Chem. Weinh. Bergstr. Ger. 2019, 25, 13523–13530. [Google Scholar] [CrossRef]
- Ruiz, N.; Kahne, D.; Silhavy, T.J. Advances in Understanding Bacterial Outer-Membrane Biogenesis. Nat. Rev. Microbiol. 2006, 4, 57–66. [Google Scholar] [CrossRef]
- Nikaido, H. Molecular Basis of Bacterial Outer Membrane Permeability Revisited. Microbiol. Mol. Biol. Rev. 2003, 67, 593–656. [Google Scholar] [CrossRef] [PubMed]
- Carnicelli, V.; Lizzi, A.R.; Ponzi, A.; Amicosante, G.; Bozzi, A.; Giulio, A. Interaction between Antimicrobial Peptides ( AMPs ) and Their Primary Target, the Biomembranes. Available online: https://www.semanticscholar.org/paper/Interaction-between-antimicrobial-peptides-(-AMPs-)-Carnicelli-Lizzi/cabb4ef72c702f54978fe84b3963f0e05bcf02f4 (accessed on 12 February 2021).
- Li, J.; Koh, J.-J.; Liu, S.; Lakshminarayanan, R.; Verma, C.S.; Beuerman, R.W. Membrane Active Antimicrobial Peptides: Translating Mechanistic Insights to Design. Front. Neurosci. 2017, 11. [Google Scholar] [CrossRef]
- Brogden, K.A. Antimicrobial Peptides: Pore Formers or Metabolic Inhibitors in Bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef]
- Hollmann, A.; Martinez, M.; Maturana, P.; Semorile, L.C.; Maffia, P.C. Antimicrobial Peptides: Interaction With Model and Biological Membranes and Synergism With Chemical Antibiotics. Front. Chem. 2018, 6. [Google Scholar] [CrossRef] [PubMed]
- Melo, M.N.; Sousa, F.J.R.; Carneiro, F.A.; Castanho, M.A.R.B.; Valente, A.P.; Almeida, F.C.L.; Da Poian, A.T.; Mohana-Borges, R. Interaction of the Dengue Virus Fusion Peptide with Membranes Assessed by NMR: The Essential Role of the Envelope Protein Trp101 for Membrane Fusion. J. Mol. Biol. 2009, 392, 736–746. [Google Scholar] [CrossRef]
- Kumar, P.; Kizhakkedathu, J.N.; Straus, S.K. Antimicrobial Peptides: Diversity, Mechanism of Action and Strategies to Improve the Activity and Biocompatibility In Vivo. Biomolecules 2018, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Yeaman, M.R.; Yount, N.Y. Mechanisms of Antimicrobial Peptide Action and Resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef]
- Sitaram, N.; Nagaraj, R. Interaction of Antimicrobial Peptides with Biological and Model Membranes: Structural and Charge Requirements for Activity. Biochim. Biophys. Acta 1999, 1462, 29–54. [Google Scholar] [CrossRef]
- Wimley, W.C. Describing the Mechanism of Antimicrobial Peptide Action with the Interfacial Activity Model. ACS Chem. Biol. 2010, 5, 905–917. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Patočka, J.; Kuča, K. Insect Antimicrobial Peptides, a Mini Review. Toxins 2018, 10, 461. [Google Scholar] [CrossRef]
- Lee, T.-H.; Hall, K.N.; Aguilar, M.-I. Antimicrobial Peptide Structure and Mechanism of Action: A Focus on the Role of Membrane Structure. Curr. Top. Med. Chem. 2016, 16, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.T.J.; Hale, J.D.; Elliott, M.; Hancock, R.E.W.; Straus, S.K. The Importance of Bacterial Membrane Composition in the Structure and Function of Aurein 2.2 and Selected Variants. Biochim. Biophys. Acta 2011, 1808, 622–633. [Google Scholar] [CrossRef]
- Sparr, E.; Ash, W.L.; Nazarov, P.V.; Rijkers, D.T.S.; Hemminga, M.A.; Tieleman, D.P.; Killian, J.A. Self-Association of Transmembrane Alpha-Helices in Model Membranes: Importance of Helix Orientation and Role of Hydrophobic Mismatch. J. Biol. Chem. 2005, 280, 39324–39331. [Google Scholar] [CrossRef]
- Hu, J.; Chen, C.; Zhang, S.; Zhao, X.; Xu, H.; Zhao, X.; Lu, J.R. Designed Antimicrobial and Antitumor Peptides with High Selectivity. Biomacromolecules 2011, 12, 3839–3843. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, A.; Vázquez, A. Bioactive Peptides: A Review. Food Qual. Saf. 2017, 1, 29–46. [Google Scholar] [CrossRef]
- Sd, S. Net charge, hydrophobicity and specific amino acids contribute to the activity of antimicrobial peptides. J. Health Transl. Med. 2014, 17, 1–7. [Google Scholar] [CrossRef]
- Friedrich, C.L.; Moyles, D.; Beveridge, T.J.; Hancock, R.E.W. Antibacterial Action of Structurally Diverse Cationic Peptides on Gram-Positive Bacteria. Antimicrob. Agents Chemother. 2000, 44, 2086–2092. [Google Scholar] [CrossRef]
- Toniolo, C.; Crisma, M.; Bonora, G.M.; Benedetti, E.; Blasio, B.D.; Pavone, V.; Pedone, C.; Santini, A. Preferred Conformation of the Terminally Blocked (Aib)10 Homo-Oligopeptide: A Long, Regular 310-Helix. Biopolymers 1991, 31, 129–138. [Google Scholar] [CrossRef]
- Conlon, J.M.; Al-Kharrge, R.; Ahmed, E.; Raza, H.; Galadari, S.; Condamine, E. Effect of Aminoisobutyric Acid (Aib) Substitutions on the Antimicrobial and Cytolytic Activities of the Frog Skin Peptide, Temporin-1DRa. Peptides 2007, 28, 2075–2080. [Google Scholar] [CrossRef] [PubMed]
- Dathe, M.; Nikolenko, H.; Meyer, J.; Beyermann, M.; Bienert, M. Optimization of the Antimicrobial Activity of Magainin Peptides by Modification of Charge. FEBS Lett. 2001, 501, 146–150. [Google Scholar] [CrossRef]
- Matsuzaki, K.; Sugishita, K.; Harada, M.; Fujii, N.; Miyajima, K. Interactions of an Antimicrobial Peptide, Magainin 2, with Outer and Inner Membranes of Gram-Negative Bacteria. Biochim. Biophys. Acta 1997, 1327, 119–130. [Google Scholar] [CrossRef]
- Andrä, J.; Monreal, D.; de Tejada, G.M.; Olak, C.; Brezesinski, G.; Gomez, S.S.; Goldmann, T.; Bartels, R.; Brandenburg, K.; Moriyon, I. Rationale for the Design of Shortened Derivatives of the NK-Lysin-Derived Antimicrobial Peptide NK-2 with Improved Activity against Gram-Negative Pathogens. J. Biol. Chem. 2007, 282, 14719–14728. [Google Scholar] [CrossRef]
- Jiang, Z.; Kullberg, B.J.; van der Lee, H.; Vasil, A.I.; Hale, J.D.; Mant, C.T.; Hancock, R.E.W.; Vasil, M.L.; Netea, M.G.; Hodges, R.S. Effects of Hydrophobicity on the Antifungal Activity of Alpha-Helical Antimicrobial Peptides. Chem. Biol. Drug Des. 2008, 72, 483–495. [Google Scholar] [CrossRef]
- Takahashi, D.; Shukla, S.K.; Prakash, O.; Zhang, G. Structural Determinants of Host Defense Peptides for Antimicrobial Activity and Target Cell Selectivity. Biochimie 2010, 92, 1236–1241. [Google Scholar] [CrossRef]
- Thaker, H.D.; Cankaya, A.; Scott, R.W.; Tew, G.N. Role of Amphiphilicity in the Design of Synthetic Mimics of Antimicrobial Peptides with Gram-Negative Activity. ACS Med. Chem. Lett. 2013, 4, 481–485. [Google Scholar] [CrossRef]
- Zhang, S.-K.; Song, J.; Gong, F.; Li, S.-B.; Chang, H.-Y.; Xie, H.-M.; Gao, H.-W.; Tan, Y.-X.; Ji, S.-P. Design of an α-Helical Antimicrobial Peptide with Improved Cell-Selective and Potent Anti-Biofilm Activity. Sci. Rep. 2016, 6, 27394. [Google Scholar] [CrossRef]
- Rodríguez, A.; Villegas, E.; Montoya-Rosales, A.; Rivas-Santiago, B.; Corzo, G. Characterization of Antibacterial and Hemolytic Activity of Synthetic Pandinin 2 Variants and Their Inhibition against Mycobacterium Tuberculosis. PLoS ONE 2014, 9, e101742. [Google Scholar] [CrossRef]
- Yin, L.M.; Edwards, M.A.; Li, J.; Yip, C.M.; Deber, C.M. Roles of Hydrophobicity and Charge Distribution of Cationic Antimicrobial Peptides in Peptide-Membrane Interactions. J. Biol. Chem. 2012, 287, 7738–7745. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Guarnieri, M.T.; Vasil, A.I.; Vasil, M.L.; Mant, C.T.; Hodges, R.S. Role of Peptide Hydrophobicity in the Mechanism of Action of Alpha-Helical Antimicrobial Peptides. Antimicrob. Agents Chemother. 2007, 51, 1398–1406. [Google Scholar] [CrossRef] [PubMed]
- Bjellqvist, B.; Hughes, G.J.; Pasquali, C.; Paquet, N.; Ravier, F.; Sanchez, J.C.; Frutiger, S.; Hochstrasser, D. The Focusing Positions of Polypeptides in Immobilized PH Gradients Can Be Predicted from Their Amino Acid Sequences. Electrophoresis 1993, 14, 1023–1031. [Google Scholar] [CrossRef] [PubMed]
- Bjellqvist, B.; Basse, B.; Olsen, E.; Celis, J.E. Reference Points for Comparisons of Two-Dimensional Maps of Proteins from Different Human Cell Types Defined in a PH Scale Where Isoelectric Points Correlate with Polypeptide Compositions. Electrophoresis 1994, 15, 529–539. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Springer Protocols Handbooks; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. ISBN 978-1-59259-890-8. [Google Scholar]
- Yang, J.; Zhang, Y. I-TASSER Server: New Development for Protein Structure and Function Predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef] [PubMed]
- Sastry, G.M.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and Ligand Preparation: Parameters, Protocols, and Influence on Virtual Screening Enrichments. J. Comput. Aided Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein Identification and Analysis Tools in the ExPASy Server. Methods Mol. Biol. Clifton NJ 1999, 112, 531–552. [Google Scholar] [CrossRef]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM Database and PPM Web Server: Resources for Positioning of Proteins in Membranes. Nucleic Acids Res. 2012, 40, D370–D376. [Google Scholar] [CrossRef]
- Bowers, K.J.; Chow, E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing, Association for Computing Machinery, Tampa, FL, USA, 11 November 2006; pp. 84-es. [Google Scholar]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of Simple Potential Functions for Simulating Liquid Water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Hornak, V.; Abel, R.; Okur, A.; Strockbine, B.; Roitberg, A.; Simmerling, C. Comparison of Multiple Amber Force Fields and Development of Improved Protein Backbone Parameters. Proteins 2006, 65, 712–725. [Google Scholar] [CrossRef]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved Side-Chain Torsion Potentials for the Amber Ff99SB Protein Force Field. Proteins 2010, 78, 1950–1958. [Google Scholar] [CrossRef]
- Wang, J.; Cieplak, P.; Kollman, P.A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 2000, 21, 1049–1074. [Google Scholar] [CrossRef]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D.; Pastor, R.W. Update of the CHARMM All-Atom Additive Force Field for Lipids: Validation on Six Lipid Types. J. Phys. Chem. B 2010, 114, 7830–7843. [Google Scholar] [CrossRef] [PubMed]
- Hoover, W.G. Canonical Dynamics: Equilibrium Phase-Space Distributions. Phys. Rev. A 1985, 31, 1695–1697. [Google Scholar] [CrossRef] [PubMed]
- Martyna, G.J.; Tobias, D.J.; Klein, M.L. Constant Pressure Molecular Dynamics Algorithms. J. Chem. Phys. 1994, 101, 4177–4189. [Google Scholar] [CrossRef]
- Tuckerman, M.; Berne, B.J.; Martyna, G.J. Reversible Multiple Time Scale Molecular Dynamics. J. Chem. Phys. 1992, 97, 1990–2001. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Tonk, M.; Cabezas-Cruz, A.; Valdés, J.J.; Rego, R.O.M.; Chrudimská, T.; Strnad, M.; Šíma, R.; Bell-Sakyi, L.; Franta, Z.; Vilcinskas, A.; et al. Defensins from the Tick Ixodes Scapularis Are Effective against Phytopathogenic Fungi and the Human Bacterial Pathogen Listeria Grayi. Parasit. Vectors 2014, 7, 554. [Google Scholar] [CrossRef] [PubMed]
- Tonk, M.; Pierrot, C.; Cabezas-Cruz, A.; Rahnamaeian, M.; Khalife, J.; Vilcinskas, A. The Drosophila Melanogaster Antimicrobial Peptides Mtk-1 and Mtk-2 Are Active against the Malarial Parasite Plasmodium Falciparum. Parasitol. Res. 2019, 118, 1993–1998. [Google Scholar] [CrossRef]
- Bolouri Moghaddam, M.R.; Tonk, M.; Schreiber, C.; Salzig, D.; Czermak, P.; Vilcinskas, A.; Rahnamaeian, M. The Potential of the Galleria Mellonella Innate Immune System Is Maximized by the Co-Presentation of Diverse Antimicrobial Peptides. Biol. Chem. 2016, 397, 939–945. [Google Scholar] [CrossRef]

| Peptides | Sequence | MW | pI | H | Net Charge * |
|---|---|---|---|---|---|
| Parental | |||||
| CopA3 | LLCIALRKK | 1.05 | 10.06 | 26.41 | +3 |
| Hp1090 | IFKAIWSGIKSLF | 1.5 | 10.00 | 44.75 | +2 |
| Hybrid | |||||
| InSco2 | LLCIALRKKGGIFKAIWSGIKSLF | 2.66 | 10.48 | 54.64 | +5 |
| InSco6 | LLCIALRKKGGGGGGIFKAIWSGIKSLF | 2.89 | 10.48 | 50.18 | +5 |
| Bacteria | Peptides | Calone (µM) | MICc (µM) | FICCopA3 + FICHp1090 | Combination Effect |
|---|---|---|---|---|---|
| E. coli D31 | CopA3 | 30 | 4 | 0.17 | Synergy |
| Hp1090 | 30 | 1 | |||
| E. coli JM83 | CopA3 | 30 | 8 | 0.52 | Partial synergy |
| Hp1090 | 60 | 15 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tonk, M.; Valdés, J.J.; Cabezas-Cruz, A.; Vilcinskas, A. Potent Activity of Hybrid Arthropod Antimicrobial Peptides Linked by Glycine Spacers. Int. J. Mol. Sci. 2021, 22, 8919. https://doi.org/10.3390/ijms22168919
Tonk M, Valdés JJ, Cabezas-Cruz A, Vilcinskas A. Potent Activity of Hybrid Arthropod Antimicrobial Peptides Linked by Glycine Spacers. International Journal of Molecular Sciences. 2021; 22(16):8919. https://doi.org/10.3390/ijms22168919
Chicago/Turabian StyleTonk, Miray, James J. Valdés, Alejandro Cabezas-Cruz, and Andreas Vilcinskas. 2021. "Potent Activity of Hybrid Arthropod Antimicrobial Peptides Linked by Glycine Spacers" International Journal of Molecular Sciences 22, no. 16: 8919. https://doi.org/10.3390/ijms22168919
APA StyleTonk, M., Valdés, J. J., Cabezas-Cruz, A., & Vilcinskas, A. (2021). Potent Activity of Hybrid Arthropod Antimicrobial Peptides Linked by Glycine Spacers. International Journal of Molecular Sciences, 22(16), 8919. https://doi.org/10.3390/ijms22168919









