A Confocal Microscopic Study of Gene Transfer into the Mesencephalic Tegmentum of Juvenile Chum Salmon, Oncorhynchus keta, Using Mouse Adeno-Associated Viral Vectors
Abstract
1. Introduction
2. Results
2.1. Anterior Hypothalamic Ventricle
2.2. Posterior Tuberculum Area
2.3. Dorsal Thalamus
2.4. Postcommissural Area
2.5. Dorso-Medial Tegmentum
2.6. Dorso-Lateral Tegmentum
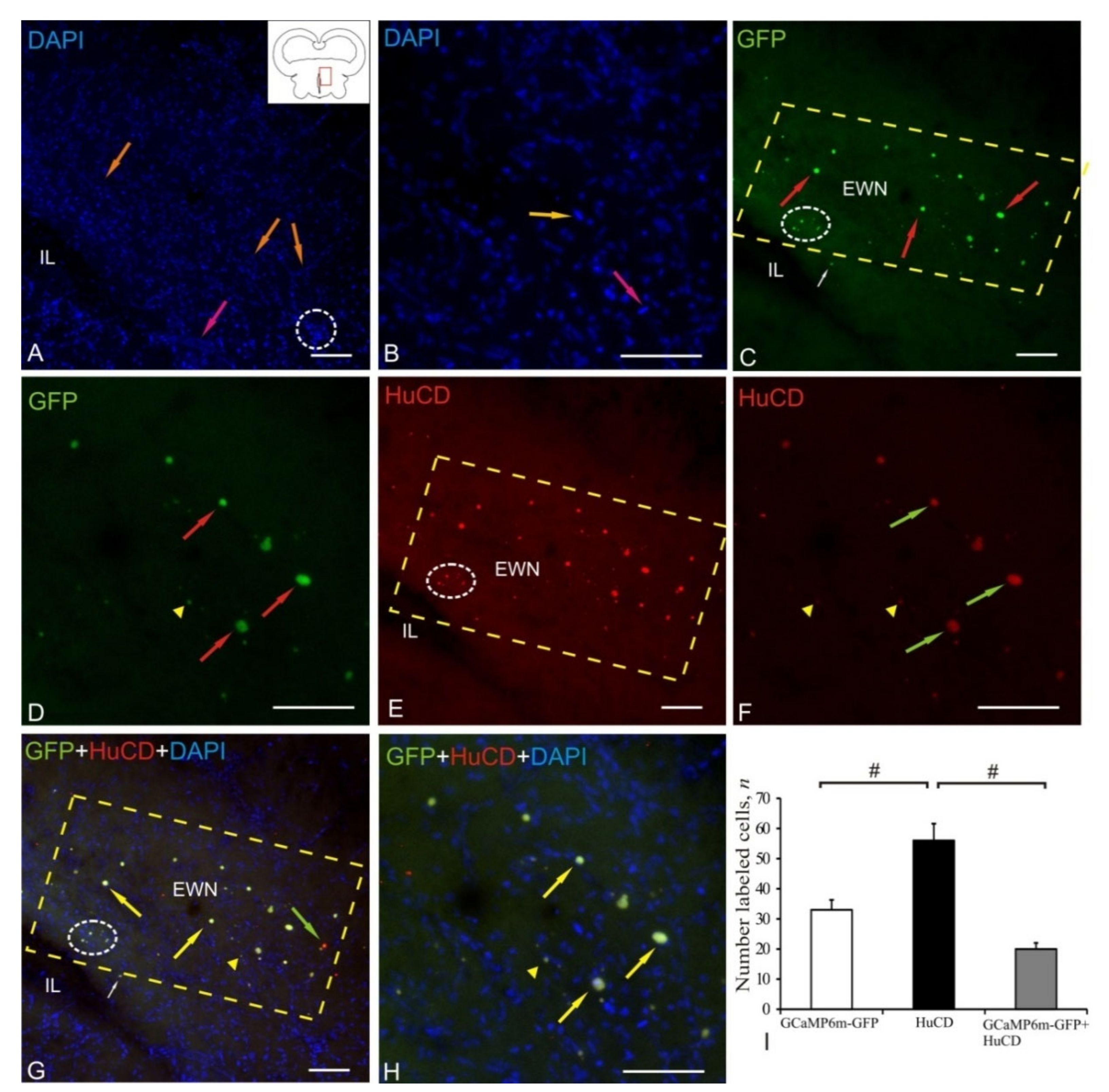
2.7. Edinger–Westphal Nucleus
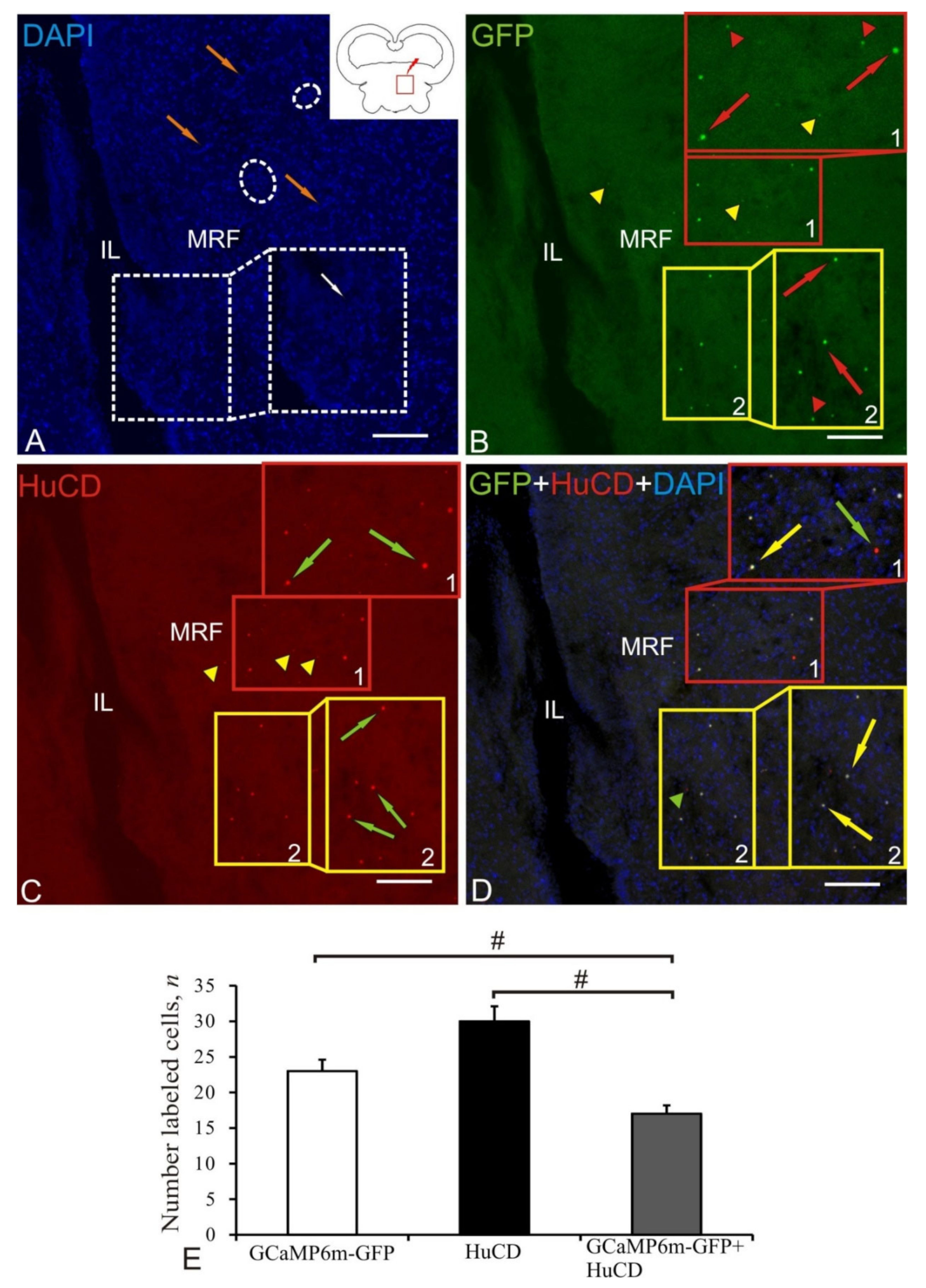
2.8. Mesencephalic Reticular Formation
3. Discussion
3.1. Cytotoxicity
3.2. Receptor Specificity
3.3. Transduction of AdV into Fish Cells
3.4. Efficiency of Infection the Recombinant Mouse Hippocampal AAV on the Juvenile Chum Salmon Brain Cells
3.5. The Use of AdV and AAV in a Fish Model for Gene Therapy
4. Material and Methods
4.1. Experimental Animals
4.2. Injection of Recombinant Adeno-Associated Viruses
4.3. Sample Preparation
4.4. Immunofluorescence Labeling
4.5. Microscopy
4.6. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAV | adeno-associated virus |
| AdV | adenovirus |
| Ad5Luc1 | AdV serotype 5 lucefirase |
| Ad5LucRGD | AdV serotype 5 lucefirase |
| Ad-EGFP | adenovirus enchased green fluorescent protein |
| BBB | blood–brain barrier |
| CAR | Coxsackie adenovirus receptor |
| CHSE-214 | rainbow trout cell line |
| CLSM | confocal laser scanning microscopy |
| CMV | cytomegalovirus |
| DAPI | 4′,6-Diamidino-2-phenylindole dihydrochloride |
| DLT | dorso-lateral tegmentum |
| DMT | dorso-medial tegmentum |
| Dth | dorsal thalamus |
| EWN | Edinger-Westphal nucleus |
| FrAdV-1 | frog adenovirus type 1 |
| GCaMPs | genetically encoded indicators of calcium |
| GFP | green fluorescent protein |
| HC-AdV | high-capacity adenoviral vectors |
| HCA-EGFP | high-capacity adenovirus enchased green fluorescent protein |
| hCAR | human Coxsackie adenovirus receptor |
| HuCD | RNA-binding proteins which induce neuronal differentiation activity |
| IF | immunofluorescence |
| IL | injection lumen |
| ITRs | inverted terminal repeats |
| LH | lateral hypothalamus |
| mCAR | mouse Coxsackie adenovirus receptor |
| MH | medial hypothalamus |
| MRF | mesencephalic reticular formation |
| PCR | Polymerase chain reaction |
| PRT | pretectal nucleus |
| PTA | posterior tuberculum area |
| PVZ | periventricular zone |
| PZ | parenchimal zone |
| rAAV | recombinant adeno-associated viral vectors |
| RBP | RNA binding protein |
| RGD motif | argininine-glycine-aspartic acid |
| rRF | rostral part of the reticular formation |
| SnAdV-1 | snake adenovirus type 1 |
| SVZ | subventricular zone |
| TAdV-3 | turkey adenovirus type 3 |
| VSVG | vesicular stomatitis glycoprotein |
| Vth | ventral thalamus |
| WSAdV-1 | white sturgeon adenovirus type 1 |
References
- Haery, L.; Deverman, B.E.; Matho, K.S.; Cetin, A.; Woodard, K.; Cepko, C.; Guerin, K.I.; Rego, M.A.; Ersing, I.; Bachle, S.M.; et al. Adeno-associated virus technologies and methods for targeted neuronal manipulation. Front. Neuroanat. 2019, 13, 93. [Google Scholar] [CrossRef]
- Muramatsu, S.; Fujimoto, K.; Kato, S.; Mizukami, H.; Asari, S.; Ikeguchi, K. A phase I study of aromatic L-amino acid decarboxylase gene therapy for Parkinson’s disease. Mol. Ther. 2010, 18, 1731–1735. [Google Scholar] [CrossRef] [PubMed]
- Gessler, D.J.; Tai, P.W.L.; Li, J.; Gao, G. Intravenous infusion of AAV for widespread gene delivery to the nervous system. Methods Mol. Biol. 2019, 1950, 143–163. [Google Scholar] [CrossRef] [PubMed]
- Bence, G.; Zachary, F.; Matheus, H.; Crommentuijn, W.; Dakai, M.; Maguire, C.A. Naturally enveloped AAV vectors for shielding neutralizing antibodies and robust gene delivery in vivo. Biomaterials 2014, 35, 7598–7609. [Google Scholar]
- Shinohara, Y.; Konno, A.; Nitta, K.; Matsuzaki, Y.; Yasui, H.; Suwa, J.; Hiromura, K.; Hirai, H. Effects of neutralizing antibody production on AAV-PHP.B-mediated ransduction of the mouse central nervous system. Mol. Neurobiol. 2019, 56, 4203–4214. [Google Scholar] [CrossRef] [PubMed]
- Rincon, M.V.; de Vin, F.; Duqué, S.I.; Fripont, S.; Castaldo, S.A.; Bouhuijzen-Wenger, J.; Holt, M.J. Widespread transduction of astrocytes and neurons in the mouse central nervous system after systemic delivery of a self-complementary AAV-PHP.B vector. Gene Ther. 2018, 25, 83–92. [Google Scholar] [CrossRef]
- Vite, C.H.; Passini, M.A.; Haskins, M.E.; Wolfe, J.H. Adeno-associated virus vector-mediated transduction in the cat brain. Gene Ther. 2003, 10, 1874–1881. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Wan, J.; Wu, Y.; Tian, Y.; Yao, Y.; Yao, S.; Ji, X.; Wang, S.; Su, Z.; Xu, H. Challenges in adeno-associated virus-based treatment of central nervous system diseases through systemic injection. Life Sci. 2021, 270, 119142. [Google Scholar] [CrossRef]
- Liguore, W.A.; Domire, J.S.; Button, D.; Wang, Y.; Dufour, B.D.; Srinivasan, S.; McBride, J.L. AAV-PHP.B administration results in a differential pattern of CNS biodistribution in non-human primates compared with mice. Mol. Ther. 2019, 27, 2018–2037. [Google Scholar] [CrossRef]
- Russell, W.C.; Benko, M. Animal adenoviruses. In Encyclopedia of Virology; Granoff, A., Webster, R.G., Eds.; Academic Press: New York, NY, USA, 1999; pp. 14–21. [Google Scholar]
- Wickham, T.J. Targeting adenovirus. Gene Ther. 2000, 7, 110–114. [Google Scholar] [CrossRef]
- Krasnykh, V.N.; Douglas, J.T.; van Beusechem, B.W. Genetic targeting of adenoviral vectors. Mol. Ther. 2000, 1, 391–405. [Google Scholar] [CrossRef] [PubMed]
- Essbauer, S.; Ahne, W. Viruses of lower vertebrates. J. Vet. Med. B Infect. Dis. Vet. Public Health 2001, 48, 403–475. [Google Scholar] [CrossRef]
- Davison, A.J.; Wright, K.M.; Harrach, B. DNA sequence of frog adenovirus. J. Gen. Virol. 2000, 81, 2431–2439. [Google Scholar] [CrossRef] [PubMed]
- Pitcovski, J.; Mualem, M.; Rei-Koren, Z.; Krispel, S.; Shmueli, E.; Peretz, Y.; Gutter, B.; Gallili, G.E.; Michael, A.; Goldberg, D. The complete DNA sequence and genome organization of the avian adenovirus, haemorrhagic enteritis virus. Virology 1998, 249, 307–315. [Google Scholar] [CrossRef]
- Farkas, L.S.; Benko, M.; Élo, P.; Ursu, K.; Dán, Á.; Ahne, W.; Harrach, B. Genomic and phylogenetic analyses of an adenovirus isolated from a corn snake (Elaphe guttata) imply common origin with the members of the proposed new genus Atadenovirus. J. Gen. Virol. 2002, 83, 2403–2410. [Google Scholar] [CrossRef] [PubMed]
- Kovács, G.M.; LaPatra, S.E.; D’Halluin, J.C.; Benko, M. Phylogenetic analysis of the hexon and protease genes of a fish adenovirus isolated from white sturgeon (Acipenser transmontanus) supports the proposal for a new adenovirus genus. Virus Res. 2003, 98, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Benko, M.; Élo, P.; Ursu, K.; Ahne, W.; LaPatra, S.E.; Thomson, D.; Harrach, B. First molecular evidence for the existence of distinct fish and snake adenoviruses. J. Virol. 2002, 76, 10056–10059. [Google Scholar] [CrossRef][Green Version]
- Gulías, P.; Guerra-Varela, J.; Gonzalez-Aparicio, M.; Ricobaraza, A.; Vales, A.; Gonzalez-Aseguinolaza, G.; Hernandez-Alcoceba, R.; Sánchez, L. Danio rerio as model organism for adenoviral vector evaluation. Genes 2019, 10, 1053. [Google Scholar] [CrossRef]
- Pushchina, E.V.; Alexandrova, Y.N.; Varaksin, A.A.; Obukhov, D.K. The efficacy of gene delivery into fish brain cells using the recombinant adeno-associated mouse hippocampal viruses. Rus. J. Physiol. 2020, 106, 1571–1584. [Google Scholar] [CrossRef]
- Stukaneva, M.E.; Pushchina, E.V.; Varaksin, A.A. Transduction of the recombinant adeno-associated mammalian hippocampal virus vector in cells of the cerebellum of juvenile Oncorhynchus keta. Rus. J. Physiol. 2020, 106, 1412–1420. [Google Scholar]
- Pushchina, E.V.; Kapustyanov, I.A.; Shamshurina, E.V.; Varaksin, A.A. Labeling of mesencephalic tegmental neurons in a juvenile pacific chum salmon Oncorhynchus keta with mouse hippocampal adeno-associated viral vectors. J. Evol. Biochem. Physiol. 2021, 57, 88–94. [Google Scholar] [CrossRef]
- Lee, J.Y.; Hirono, I.I.; Aoki, T. Stable expression of a foreign gene, delivered by gene gun, in the muscle of rainbow trout (Oncorhynchus mykiss). Mar. Biotechnol. 2000, 2, 254–258. [Google Scholar] [CrossRef] [PubMed]
- Rainbow, A.J.; Zacal, N.J. Expression of an adenovirus encoded reporter gene and its reactivation following UVC and oxidative damage in cultured fish cells. Int. J. Radiat. Biol. 2008, 84, 455–466. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Mikoshiba, K. Neuronal birthdate-specific gene transfer with adenoviral vectors. J. Neurosci. 2004, 24, 286–296. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Mikoshiba, K. Mediolateral compartmentalization of the cerebellum is determined on the ‘‘birth date” of Purkinje cells. J. Neurosci. 2003, 23, 11342–11351. [Google Scholar] [CrossRef]
- Zhao, H.Q.; Ivic, L.; Otaki, J.M.; Hashimoto, M.; Mikoshiba, K.; Firestein, S. Functional expression of a mammalian odorant receptor. Science 1998, 279, 237–242. [Google Scholar] [CrossRef]
- Albain, J.; Zon, L.I. Of fish and men: Using zebrafish to fight human diseases. Trends. Cell Biol. 2013, 23, 584–586. [Google Scholar]
- Broussard, G.J.; Liang, R.; Tian, L. Monitoring activity in neural circuits with genetically encoded indicators. Front. Mol. Neurosci. 2014, 7, 97. [Google Scholar] [CrossRef]
- Cearley, C.N.; Wolfe, J.H. Transduction characteristics of adeno-associated virus vectors expressing cap serotypes 7, 8, 9, and Rh10 in the mouse brain. Mol. Ther. 2006, 13, 528–537. [Google Scholar] [CrossRef]
- Cearley, C.N.; Wolfe, J.H. A single Injection of an adeno-associated virus vector into nuclei with divergent connections results in widespread vector distribution in the brain and global correction of a neurogenetic disease. J. Neurosci. 2007, 27, 9928–9940. [Google Scholar] [CrossRef]
- Lorenzen, N.; Lorenzen, E.; Einer-Jensen, K.; LaPatra, S.E. Immunity induced shortly after DNA vaccination of rainbow trout against rhabdoviruses protects against heterologous virus but not against bacterial pathogens. Dev. Comp. Immunol. 2002, 2, 173–179. [Google Scholar] [CrossRef]
- Zelenin, A.V.; Alimov, A.A.; Barmintzev, V.A.; Beniumov, A.O.; Zelenina, I.A.; Krasnov, A.M.; Kolesnikov, V.A. The delivery of foreign genes into fertilized fish eggs using high-velocity microprojectiles. FEBS Lett. 1991, 287, 118–120. [Google Scholar] [CrossRef]
- Sussman, R. Direct DNA delivery into zebrafish embryos employing tissue culture techniques. Genesis 2001, 31, 1–5. [Google Scholar] [CrossRef]
- Muller, F.; Lele, Z.; Varadi, L.; Menczel, L.; Orban, L. Efficient transient expression system based on square pulse electroporation and in vivo luciferase assay of fertilized fish eggs. FEBS Lett. 1993, 324, 27–32. [Google Scholar] [CrossRef]
- Burns, J.C.; Friedmann, T.; Driever, W.; Burrascano, M.; Yee, J.K. Vesicular stomatitis virus G glycoprotein pseudotyped retroviral vectors: Concentration to very high titer and efficient gene transfer into mammalian and non-mammalian cells. Proc. Natl. Acad. Sci. USA 1993, 90, 8033–8037. [Google Scholar] [CrossRef] [PubMed]
- Phenix, K.V.; McKenna, B.; Fitzpatrick, R.; Vaughan, L.; Atkins, G.; Liljestrom, P.; Todd, D. Cell culture evaluation of the Semliki Forest Virus expression system as a novel approach for antigen delivery and expression in fish. Mar. Biotechnol. 2000, 2, 27–37. [Google Scholar] [CrossRef]
- Overturf, K.; LaPatra, S.; Renolds, P.N. The effectiveness of adenoviral vectors to deliver and express genes in rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2003, 26, 91–101. [Google Scholar] [CrossRef]
- Knöpfel, T.; Lin, M.Z.; Levskaya, A.; Tian, L.; Lin, J.Y.; Boyden, E.S. Toward the second generation of optogenetic tools. J. Neurosci. 2010, 30, 14998–15004. [Google Scholar] [CrossRef]
- Pérez Koldenkova, V.; Nagai, T. Genetically encoded Ca(2+) indicators: Properties and evaluation. Biochim. Biophys. Acta 2013, 1833, 1787–1797. [Google Scholar] [CrossRef]
- Zhu, P.; Narita, Y.; Bundschuh, S.T.; Fajardo, O.; Scharer, Y.P.; Chattopadhyaya, B. Optogenetic dissection of neuronal circuits in zebrafish using viral gene transfer and the tet system. Front. Neural Circuits 2009, 3, 21. [Google Scholar] [CrossRef]
- Pushchina, E.V.; Stukaneva, M.E.; Varaksin, A.A. Hydrogen sulfide modulates adult and reparative neurogenesis in the cerebellum of juvenile masu salmon, Oncorhynchus masou. Int. J. Mol. Sci. 2020, 21, 9638. [Google Scholar] [CrossRef] [PubMed]
- Pushchina, E.V.; Zharikova, E.I.; Varaksin, A.A. Mechanical brain injury increases cells’ production of cystathionine β-synthase and glutamine synthetase, but feduces Pax2 expression in the telencephalon of juvenile chum salmon, Oncorhynchus keta. Int. J. Mol. Sci. 2021, 22, 1279. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, F.; Dang, L.; Liang, C.; He, B.; Liu, J. In vivo delivery systems for therapeutic genome editing. Int. J. Mol. Sci. 2016, 17, 626. [Google Scholar] [CrossRef] [PubMed]
- Lundstrom, K. Viral vectors in gene therapy. Diseases 2018, 6, 42. [Google Scholar] [CrossRef]
- Crustal, R. Adenovirus: The first effective in vivo gene delivery vector. Hum. Gene. Ther. 2014, 25, 3–11. [Google Scholar] [CrossRef]
- Barcia, C.; Jimenez-Dalmaroni, M.; Kroeger, K.; Puntel, M.; Rapaport, A.; Larocque, D.; King, G. One-year expression from high-capacity adenoviral vectors in the brains of animals with pre-existing anti-adenoviral immunity: Clinical implications. Mol. Gene Ther. 2007, 15, 2154–2163. [Google Scholar] [CrossRef] [PubMed]
- Brunetti-Pierri, N.; Ng, T.; Iannitti, D.; Cioffi, W.; Stapleton, G.; Law, M.; Breinholt, J. Transgene expression up to 7 years in nonhuman primates following hepatic transduction with helper-dependent adenoviral vectors. Hum. Gene Ther. 2013, 24, 761–765. [Google Scholar] [CrossRef]
- Zhang, B.; Shimada, Y.; Hirota, T.; Ariyoshi, M.; Kuroyanagi, J.; Nishimura, Y.; Tanaka, T. Novel immunologic tolerance of human cancer cell xenotransplants in zebrafish. Transl. Res. 2016, 170, 89–98. [Google Scholar] [CrossRef] [PubMed]
- Lam, S.H.; Chua, H.L.; Gong, Z.; Lam, T.J.; Sin, Y.M. Development and maturation of the immune system in zebrafish, Danio rerio: A gene expression profiling, in situ hybridization and immunological study. Dev. Comp. Immunol. 2004, 28, 9–28. [Google Scholar] [CrossRef]
- Zirger, J.M.; Puntel, M.; Bergeron, J.; Wibowo, M.; Moridzadeh, R.; Bondale, N.; Barcia, C.; Kroeger, K.M.; Liu, C.; Castro, M.G. Immune-mediated loss of transgene expression from virally transduced brain cells is irreversible, mediated by IFNγ, perforin, and TNFα, and due to the elimination of transduced cells. Mol. Ther. 2012, 20, 808–819. [Google Scholar] [CrossRef]
- Tomko, R.P.; Xu, R.; Philipson, L. HCAR and MCAR: The human and mouse cellular receptors for subgroup C adenoviruses and group B coxsackieviruses. Proc. Natl. Acad. Sci. USA 1997, 94, 3352–3356. [Google Scholar] [CrossRef] [PubMed]
- Okegawa, T.; Li, Y.; Pong, R.C.; Bergelson, J.M.; Zhou, J.; Hsieh, J.T. The dual impact of coxsackie and adenovirus receptor expression on human prostate cancer gene therapy. Cancer Res. 2000, 60, 5031–5036. [Google Scholar]
- Wickham, T.J.; Mathias, P.; Cheresh, D.A.; Nemerow, G.R. Integrins alpha v beta 3 and alpha v beta 5 promote adenovirus internalization but not virus attachment. Cell 1993, 73, 309–319. [Google Scholar] [CrossRef]
- Einfeld, D.A.; Schroeder, R.; Roelvink, P.W.; Lizonova, A.; King, C.R.; Kovesdi, I.; Wickham, T.J. Reducing the native tropism of adenovirus vectors requires removal of both CAR and integrin interactions. J. Virol. 2001, 75, 11284–11291. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, P.N.; Curiel, D.T. New generation adenoviral vectors improve gene transfer by CAR independent cell entry. Kidney Int. 2001, 61 (Suppl. S1), 24–31. [Google Scholar]
- Dmitriev, I.; Krasnykh, K.; Miller, C.R.; Wang, M.; Kashentseva, E.; Mikheeva, G.; Belousova, N.; Curiel, D.T. An adenovirus vector with genetically modified fibers demonstrates expanded tropism via utilization of a coxsackievirus and adenovirus receptor-independent cell entry mechanism. J. Virol. 1998, 72, 9706–9713. [Google Scholar] [CrossRef] [PubMed]
- Corbeil, S.; Kurath, G.; LaPatra, S.E. Fish DNA vaccine against infectious hematopoietic necrosis virus: Efficacy of various routes of immunization. Fish Shellfish Immunol. 2000, 10, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, T.; Saito, K.; Mitsui, K.; Ikawa, M.; Yamashita, M.; Taniguchi, Y.; Takeda, S.; Mitani, K.; Sakai, N. Introduction of a foreign gene into zebrafish and medaka cells using adenoviral vectors. Zebrafish 2009, 6, 3. [Google Scholar] [CrossRef]
- Jensen, N.J.; Bloch, B. Adenovirus-like particles associated with epidermal hyperplasia in cod (Gadus morhua). Nord. Vet. Med. 1980, 32, 173–175. [Google Scholar]
- Bloch, B.; Mellergaard, S.; Nielsen, E. Adenovirus-like particles associated with epithelial hyperplasias in dab, Limanda limandra (L.). J. Fish Dis. 1986, 9, 281–285. [Google Scholar] [CrossRef]
- Miyazaki, T.; Asai, Y.; Kobayashi, T.; Miyata, M. Lympholeukemia in madai Pagrus major in Japan. Dis. Aquat. Org. 2000, 40, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Hedrick, R.P.; Speas, J.; Kent, M.L.; McDowell, T. Adenovirus-like particles associated with a disease of cultured white sturgeon, Acipenser Transmontanus. Can. J. Fish. Aquat. Sci. 1985, 42, 1321–1325. [Google Scholar] [CrossRef]
- Benko, M.; Harrach, B. Molecular evolution of adenoviruses. Cur. Top. Microbiol. Immunol. 2003, 272, 4–35. [Google Scholar]
- Furutani-Seiki, M.; Wittbrodt, J. Medaka and zebrafish, an evolutionary twin study. Mech. Dev. 2004, 121, 629–637. [Google Scholar] [CrossRef] [PubMed]
- Mathivanan, S.; Fahner, C.J.; Gavin, E.; Reid, G.E.; Simpson, R.J. ExoCarta 2012: Database of exosomal proteins, RNA and lipids. Nucl. Acids Res. 2012, 40, D1241–D1244. [Google Scholar] [CrossRef]
- Hudry, E.; Martin, C.; Gandhi, S.; György, B.; Scheffer, D.I.; Mu, D.; Merkel, S.F.; Mingozzi, F.; Fitzpatrick, Z.; Dimant, H.; et al. Exosome-associated AAV vector as a robust and convenient neuroscience tool. Gene Ther. 2016, 23, 380–392. [Google Scholar] [CrossRef]
- Andaloussi, S.E.L.; Mäger, I.; Breakefield, X.O.; Wood, M.J.A. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug. Discov. 2013, 12, 47–57. [Google Scholar] [CrossRef]
- Feng, Z.; Hensley, L.; McKnight, K.L.; Hu, F.; Madden, V.; Ping, L.; Jeong, S.-H.; Walker, C.; Lanford, R.E.; Lemon, S.M. A pathogenic picornavirus acquires an envelope by hijacking cellular membranes. Nature 2013, 496, 367–371. [Google Scholar] [CrossRef]
- Ramakrishnaiah, V.; Thumann, C.; Fofana, I.; Habersetzer, F.; Pan, Q.; de Ruiter, P.E.; Willemsen, R.; Demmers, J.A.A.; Raj, V.S.; Jenster, G.; et al. WExosome-mediated transmission of hepatitis C virus between human hepatoma Huh7.5 cells. Proc. Natl. Acad. Sci. USA 2013, 110, 13109–13113. [Google Scholar] [CrossRef]
- Zhang, B.; Yin, Y.; Lai, R.C.; Lim, S.K. Immunotherapeutic potential of extracellular vesicles. Front. Immunol. 2014, 5, 518. [Google Scholar] [CrossRef] [PubMed]
- Wakamatsu, Y.; Weston, J.A. Sequential expression and role of Hu RNA-binding proteins during neurogenesis. Development 1997, 124, 3449–3460. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.D.; Morin, M.A.; Beckel-Mitchener, A.; Mobarak, C.D.; Neve, R.L.; Furneaux, H.M.; Burry, R.; Perrone-Bizzozero, N.I. Overexpression of HuD, but not of its truncated form HuD I+II, promotes GAP-43 gene expression and neurite outgrowth in PC12 cells in the absence of nerve growth factor. J. Neurochem. 2000, 75, 1103–1114. [Google Scholar] [CrossRef] [PubMed]
- Akamatsu, W. The RNA-binding protein HuD regulates neuronal cell identity and maturation. Proc. Natl. Acad. Sci. USA 2005, 102, 4625–4630. [Google Scholar] [CrossRef] [PubMed]
- Akamatsu, W.; Okano, H.J.; Osumi, N.; Inoue, T.; Nakamura, S.; Sakakibara, S.; Miura, M.; Matsuo, N.; Darnell, R.B.; Okano, H. Mammalian ELAV-like neuronal RNA-binding proteins HuB and HuC promote neuronal development in both the central and the peripheral nervous systems. Proc. Natl. Acad. Sci. USA 1999, 96, 9885–9890. [Google Scholar] [CrossRef] [PubMed]
- Hinman, M.N.; Lou, H. Diverse molecular functions of Hu proteins. Cell. Mol. Life Sci. 2008, 65, 3168–3181. [Google Scholar] [CrossRef]
- Zhu, H.; Hasman, R.A.; Barron, V.A.; Luo, G.; Lou, H. A nuclear function of Hu proteins as neuron-specific alternative RNA processing regulators. Mol. Biol. Cell 2006, 17, 5105–5114. [Google Scholar] [CrossRef] [PubMed]
- Rogers, G.L.; Martino, A.T.; Aslanidi, G.V.; Jayandharan, G.R.; Srivastava, A.; Herzog, R.W. Innate immune responses to AAV vectors. Front. Microbiol. 2011, 2, 194. [Google Scholar] [CrossRef]
- Mingozzi, F.; High, K.A. Immune responses to AAV in clinical trials. Curr. Gene Ther. 2011, 11, 321–330. [Google Scholar] [CrossRef]
- Xiong, W.; Wu, D.M.; Xue, Y.; Wang, S.K.; Chung, M.J.; Ji, X. AAV cis-regulatory sequences are correlated with ocular toxicity. Proc. Natl. Acad. Sci. USA 2019, 116, 5785–5794. [Google Scholar] [CrossRef]
- Touahri, Y.; Dixit, R.; Kofoed, R.H.; Miloska, K.; Park, E.; Raeisossadati, R.; Markham-Coultes, K.; David, L.A.; Rijal, H.; Zhao, J. Focused ultrasound as a novel strategy for noninvasive gene delivery to retinal Müller glia. Theranostics 2020, 10, 2982–2999. [Google Scholar] [CrossRef]
- Wagle, M.; Grunewald, B.; Subburaju, S.; Barzaghi, C.; Le Guyader, S.; Chan, J.; Jesuthasan, S. EphrinB2a in the zebrafish retinotectal system. J. Neurobiol. 2004, 59, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Rothenaigner, I.; Krecsmarik, M.; Hayes, J.A.; Bahn, B.; Lepier, A.; Fortin, G. Clonal analysis by distinct viral vectors identifies bona fide neural stem cells in the adult zebrafish telencephalon and characterizes their division properties and fate. Development 2011, 138, 1459–1469. [Google Scholar] [CrossRef]
- Luo, L.; Callaway, E.M.; Svoboda, K. Genetic dissection of neural circuits. Neuron 2008, 57, 634–660. [Google Scholar] [CrossRef]
- Burgos, J.S.; Ripoll-Gomez, J.; Alfaro, J.M.; Sastre, I.; Valdivieso, F. Zebrafish as a new model for herpes simplex virus type 1 infection. Zebrafish 2008, 5, 323–333. [Google Scholar] [CrossRef] [PubMed]
- De Vry, J.; Martínez-Martínez, P.; Losen, M.; Temel, Y.; Steckler, T.; Steinbusch, H.W.M. In vivo electroporation of the central nervous system: A non-viral approach for targeted gene delivery. Prog. Neurobiol. 2010, 92, 227–244. [Google Scholar] [CrossRef] [PubMed]
- Barnabé-Heider, F.; Meletis, K.; Eriksson, M.; Bergmann, O.; Sabelström, H.; Harvey, M.A.; Mikkers, H.; Frisén, J. Genetic manipulation of adult mouse neurogenic niches by in vivo electroporation. Nat. Methods 2008, 5, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Tabata, H.; Nakajima, K. Efficient in utero gene transfer system to the developing mouse brain using electroporation: Visualization of neuronal migration in the developing cortex. Neuroscience 2001, 103, 865–872. [Google Scholar] [CrossRef]
- Bianco, I.H.; Carl, M.; Russell, C.; Clarke, J.D.; Wilson, S.W. Brain asymmetry is encoded at the level of axon terminal morphology. Neural Dev. 2008, 3, 9. [Google Scholar] [CrossRef]
- Cerda, G.A.; Thomas, J.E.; Allende, M.L.; Karlstrom, R.O.; Palma, V. Electroporation of DNA, RNA, and morpholinos into zebrafish embryos. Methods 2006, 39, 207–211. [Google Scholar] [CrossRef]
- Rambabu, K.M.; Rao, S.H.N.; Rao, N.M. Efficient expression of transgenes in adult zebrafish by electroporation. BMC Biotechnol. 2005, 5, 29. [Google Scholar] [CrossRef]
- Gao, G.; Qu, G.; Burnham, M.S.; Huang, J.; Chirmule, N.; Joshi, B.; Yu, Q.C.; Marsh, J.A.; Conceicao, C.M.; Wilson, J.M. Purification of recombinant adeno-associated virus vectors by column chromatography and its performance in vivo. Hum. Gene Ther. 2000, 11, 2079–2091. [Google Scholar] [CrossRef] [PubMed]

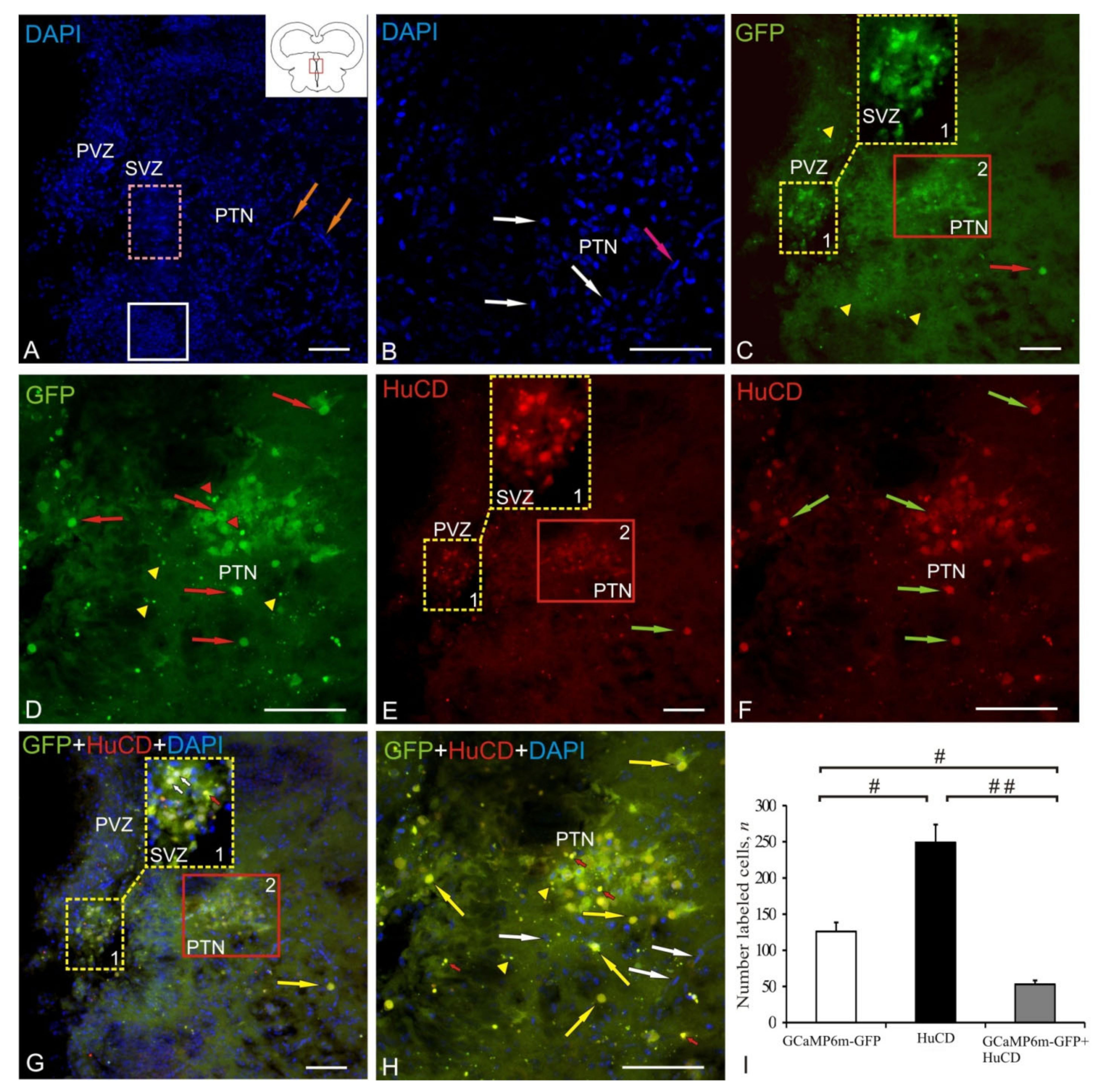
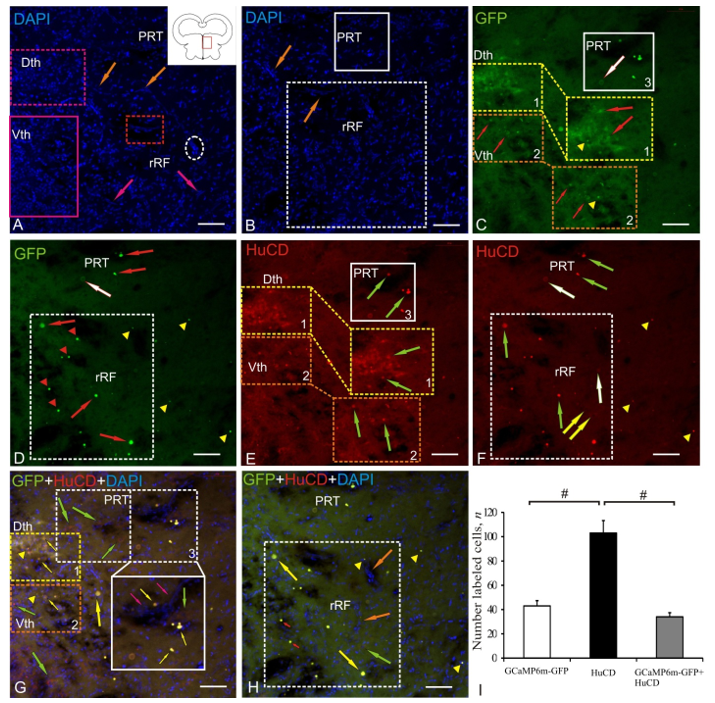
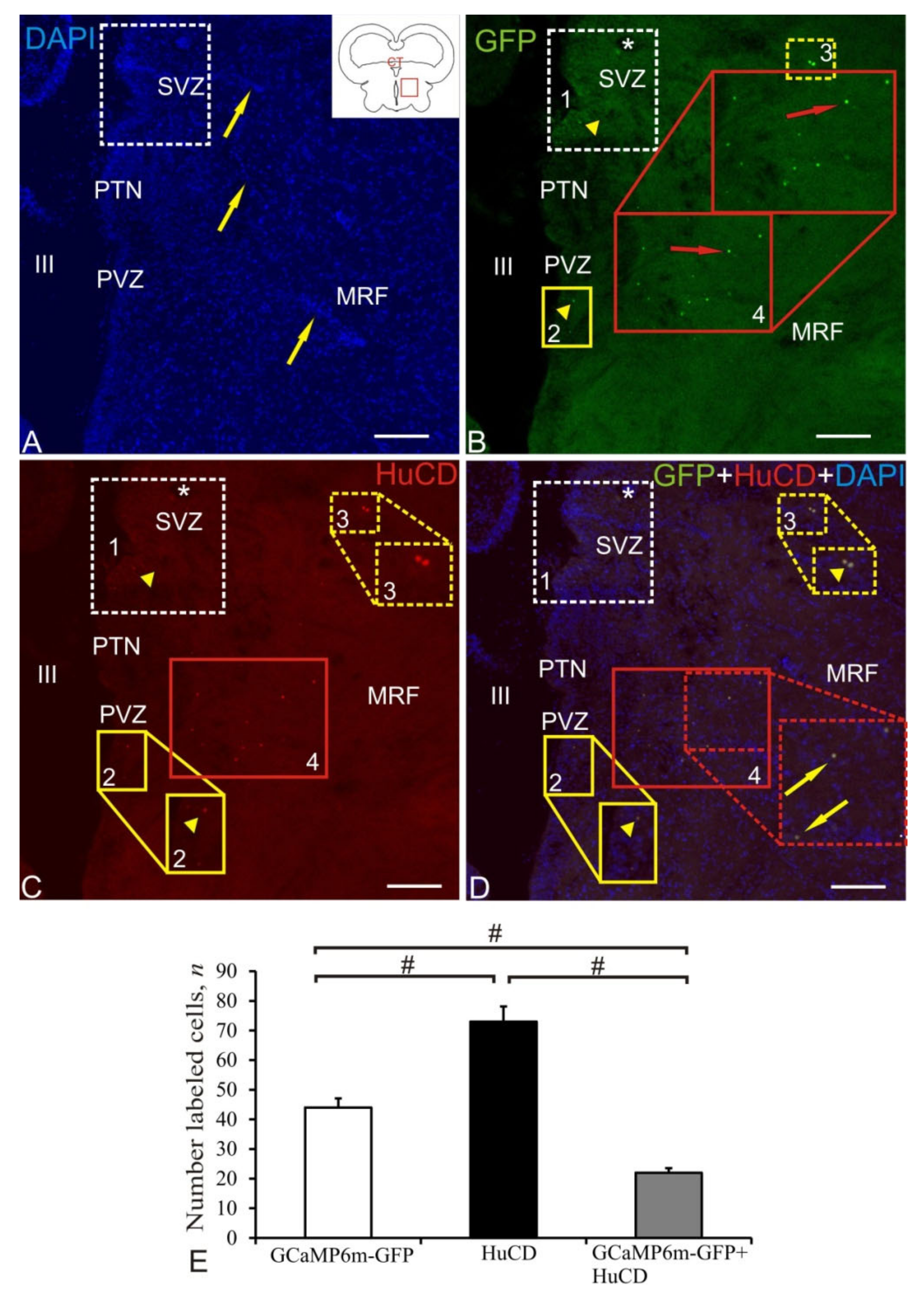
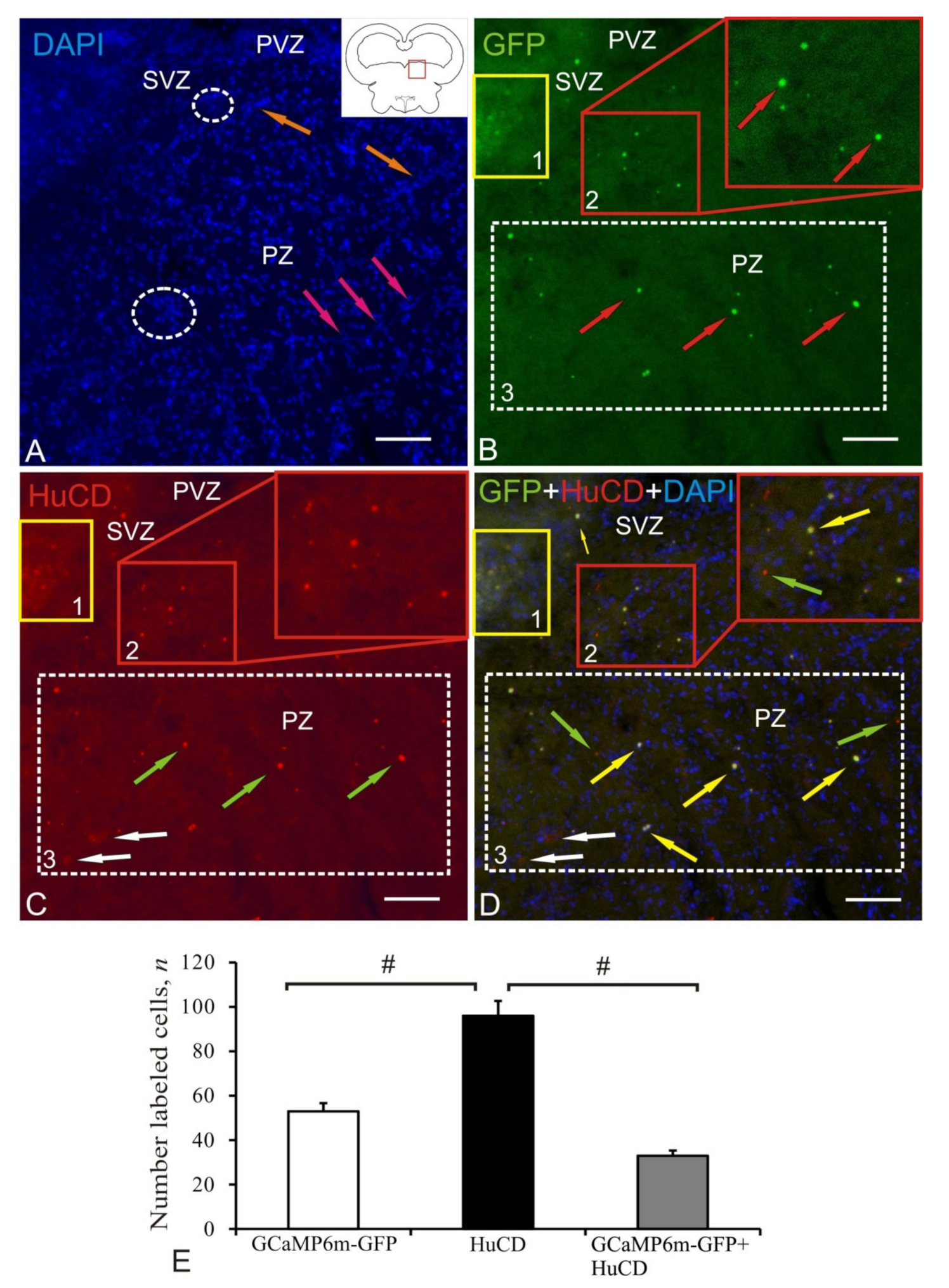

| Brain Area | DAPI Size of Nuclei/ Cells, μm | AAV Size of Cells, Nuclei, Granules, μm | Optical Density, UOD | HuCD Cell Size, μm | Optical Density, UOD | AAV + HuCD Size of Cells, Nuclei, Granules, μm | Optical Density, UOD |
|---|---|---|---|---|---|---|---|
| Anterior hypothalamic ventricle | 4.9 ± 0.3/3.4 ± 0.4 4.2 ± 0.1/3.1 ± 0.3 3.8 ± 0.1/2.9 ± 0.3 3.1 ± 0.4/2.8 ± 0.2 | 6.2 ± 0.6/6.2 ± 0.5 4.9 ± 0.3/4.6 ± 0.4 4.5 ± 0.1/3.6 ± 1.1 3.6 ± 0.2/3.1 ± 0.3 2.6 ± 0.4/2.3 ± 0.4 | +++ +++ +++/++ ++ +/++ | 8.7 ± 0.4/6.0 ± 0.8 7.4 ± 0.4/5.8 ± 1.9 6.1 ± 0.3/5.1 ± 0.6 4.9 ± 0.3/4.4 ± 0.1 3.9 ± 0.3/3.4 ± 0.4 | ++ ++ +++ +++ ++/+++ | 6.5 ± 0.2/5.6 ± 0.4 4.8 ± 0.5/4.6 ± 0.6 3.8 ± 0.2/3.2 ± 0.5 3.2 ± 0.2/2.7 ± 0.3 | ++ +++ ++/+++ ++ |
| Posterior tuberculum area | 8.5 ± 0.7/6.0 ± 1.8 6.2 ± 0.3/4.3 ± 0.7 4.8 ± 0.4/3.9 ± 0.5 | 8.4 ± 0.9/6.0 ± 0.8 5.8 ± 1.1/4.5 ± 0.8 3.4 ± 0.4/2.7 ± 0.7 2.4 ± 0.3/2.0 ± 0.4 | +++ ++ ++ +/++ | 8.2 ± 0.8/6.9 ± 0.8 6.5 ± 0.4/5.2 ± 0.4 3.9 ± 0.2/2.9 ± 0.4 | ++ +++ ++ | 8.1 ± 1.0/5.7 ± 1.4 6.3 ± 0.3/4.7 ± 0.7 4.4 ± 0.7/3.5 ± 0.5 | ++/+++ ++ +++ |
| Dorsal thalamus | 4.3 ± 0.2/3.2 ± 0.6 3.6 ± 0.2/2.8 ± 0.4 3.2 ± 0.2/2.6 ± 0.2 | 6.3 ± 0.3/5.4 ± 1.0 5.1 ± 0.4/4.0 ± 0.7 3.8 ± 0.4/3.4 ± 0.4 | ++/+++ +++ ++ | 7.2 ± 0.5/5.8 ± 0.9 6.0 ± 0.3/4.8 ± 0.6 4.8 ± 0.4/3.8 ± 0.7 3.0 ± 0.4/3.0 ± 0.3 | ++ ++/+++ +++ ++ | 8.1 ± 0.7/6.7 ± 1.4 6.9 ± 0.2/5.4 ± 0.9 5.8 ± 0.3/4.9 ± 0.4 4.9 ± 0.4/4.5 ± 0.4 | ++/+++ ++ +++/++ +++ |
| Postcommissural area | 5.3 ± 0.2/4.0 ± 0.5 4.5 ± 0.2/3.8 ± 0.4 3.7 ± 0.3/3.4 ± 0.4 | 8.8 ± 0.4/6.6 ± 0.2 5.2 ± 0.3/3.3 ± 0.8 3.0 ± 0.3/2.3 ± 0.3 2.4 ± 0.2/2.1 ± 0.3 | +/++ +++ ++ +/++ | 7.8 ± 2.1/5.9 ± 2.6 4.6 ± 0.2/3.7 ± 0.4 3.7 ± 0.3/2.8 ± 0.5 | +/++ +++ +++ | 5.7 ± 0.4/3.5 ± 0.5 4.6 ± 0.3/3.4 ± 0.4 3.8 ± 0.1/3.2 ± 0.5 2.3 ± 0.4/2.2 ± 0.3 | ++ + ++ + |
| Dorso-medial tegmentum | 7.8 ± 0.4/5.0 ± 0.6 5.9 ± 0.6/4.4 ± 0.5 4.5 ± 0.5/3.6 ± 0.2 | 10.8 ± 0.3/8.0 ± 0.7 8.7 ± 0.1/6.6 ± 0.5 7.2 ± 0.4/5.5 ± 0.6 5.9 ± 0.5/4.7 ± 0.7 4.5 ± 0.4/3.9 ± 0.6 2.4 ± 0.4/2.2 ± 0.2 | ++ +++ +++ + +/++ +++ | 26.1 ± 1.3/13.4 ± 1.5 11.0 ± 0.7/7.3 ± 2.0 8.3 ± 0.6/6.4 ± 1.2 6.5 ± 0.6/5.2 ± 0.9 5.0 ± 0.1/4.6 ± 0.3 | + ++/+++ +++ + +/++ | 8.3 ± 0.7/6.4 ± 0.8 6.3 ± 0.8/5.1 ± 0.6 4.5 ± 0.2/3.8 ± 0.6 | ++ +++ +/++ |
| Dorso-lateral tegmentum | 4.8 ± 0.3/3.2 ± 0.5 4.2 ± 0.2/3.1 ± 0.3 3.7 ± 0.1/2.8 ± 0.3 3.2 ± 0.2/2.7 ± 0.5 | 5.9 ± 0.6/4.7 ± 0.2 4.9 ± 0.1/3.6 ± 0.4 4.4 ± 0.1/3.6 ± 0.4 3.6 ± 0.2/2.8 ± 0.3 2.9 ± 0.4/2.6 ± 0.3 | +++ ++/+++ ++ ++ +/++ | 5.6 ± 0.3/4.2 ± 0.5 4.7 ± 0.2/3.6 ± 0.4 4.3 ± 0.2/3.3 ± 0.4 3.5 ± 0.2/2.8 ± 0.3 | +++ +++ ++/+++ ++ | 5.5 ± 0.6/4.3 ± 0.8 4.5 ± 0.3/3.8 ± 0.6 3.9 ± 0.1/3.5 ± 0.2 2.9 ± 0.4/2.6 ± 0.3 | +++ ++/+++ ++ + |
| Edinger–Westphal nucleus | 4.0 ± 0.3/2.8 ± 0.8 3.4 ± 0.2/2.9 ± 0.3 2.8 ± 0.2/2.5 ± 0.3 | 10.2 ± 1/7.6 ± 0.6 6.4 ± 0.5/5.3 ± 0.6 4.5 ± 0.3/3.7 ± 0.4 3.5 ± 0.5/2.6 ± 0.5 | +++ +++ ++/+++ ++ | 10.6 ± 1.2/7.7 ± 0.6 7.6 ± 0.4/6.4 ± 1.1 5.8 ± 0.7/4.5 ± 0.8 4.1 ± 0.3/3.4 ± 0.5 | +++ +++/++ ++ ++ | 7.4 ± 0.3/6.2 ± 0.6 5.3 ± 0.5/4.2 ± 0.4 3.8 ± 0.3/2.8 ± 0.8 | +++ +++/++ ++ |
| Mesencephalic reticular formation | 5.4 ± 0.3/3.8 ± 0.3 4.2 ± 0.3/3.3 ± 0.4 3.4 ± 0.3/2.7 ± 0.1 | 5.8 ± 0.3/4.4 ± 0.7 4.8 ± 0.3/3.9 ± 0.8 2.3 ± 0.1/2.2 ± 0.3 | +++ ++/+++ +/++ | 5.7 ± 0.4/4.3 ± 0.7 4.7 ± 0.2/3.7 ± 0.7 4.0 ± 0.3/3.5 ± 0.4 | +++ +++ ++/++ | 5.7 ± 0.5/4.5 ± 0.2 4.4 ± 0.3/4.1 ± 0.4 | +++ ++ |
| Brain Area | AAV, % | HuCD, % | AAV + HuCD, % |
|---|---|---|---|
| Anterior hypothalamic ventricle | 26.4 | 57 | 16.5 |
| Posterior tuberculum area | 29.4 | 58.2 | 12.4 |
| Dorsal thalamus | 23.9 | 57.2 | 18.9 |
| Postcommissural area | 31.6 | 52.5 | 15.8 |
| Dorso-medial tegmentum | 29.1 | 52.7 | 18.1 |
| Dorso-lateral tegmentum | 24.7 | 62.1 | 13.1 |
| Edinger–Westphal nucleus | 30.3 | 51.2 | 18.3 |
| Mesencephalic reticular formation | 32.8 | 42.8 | 24.3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pushchina, E.V.; Kapustyanov, I.A.; Shamshurina, E.V.; Varaksin, A.A. A Confocal Microscopic Study of Gene Transfer into the Mesencephalic Tegmentum of Juvenile Chum Salmon, Oncorhynchus keta, Using Mouse Adeno-Associated Viral Vectors. Int. J. Mol. Sci. 2021, 22, 5661. https://doi.org/10.3390/ijms22115661
Pushchina EV, Kapustyanov IA, Shamshurina EV, Varaksin AA. A Confocal Microscopic Study of Gene Transfer into the Mesencephalic Tegmentum of Juvenile Chum Salmon, Oncorhynchus keta, Using Mouse Adeno-Associated Viral Vectors. International Journal of Molecular Sciences. 2021; 22(11):5661. https://doi.org/10.3390/ijms22115661
Chicago/Turabian StylePushchina, Evgeniya V., Ilya A. Kapustyanov, Ekaterina V. Shamshurina, and Anatoly A. Varaksin. 2021. "A Confocal Microscopic Study of Gene Transfer into the Mesencephalic Tegmentum of Juvenile Chum Salmon, Oncorhynchus keta, Using Mouse Adeno-Associated Viral Vectors" International Journal of Molecular Sciences 22, no. 11: 5661. https://doi.org/10.3390/ijms22115661
APA StylePushchina, E. V., Kapustyanov, I. A., Shamshurina, E. V., & Varaksin, A. A. (2021). A Confocal Microscopic Study of Gene Transfer into the Mesencephalic Tegmentum of Juvenile Chum Salmon, Oncorhynchus keta, Using Mouse Adeno-Associated Viral Vectors. International Journal of Molecular Sciences, 22(11), 5661. https://doi.org/10.3390/ijms22115661






