Contribution of Evolutionary Selected Immune Gene Polymorphism to Immune-Related Disorders: The Case of Lymphocyte Scavenger Receptors CD5 and CD6
Abstract
1. Evolution and Selective Pressure on Immune Receptor Genes: Examples of Selection
1.1. Human Evolution and Pathogens
1.2. Detecting Local Adaptation in the Human Genome
1.3. Examples of Positive Selection at Immune Response Receptors
2. The Case of Lymphocyte Scavenger Receptors CD5 and CD6
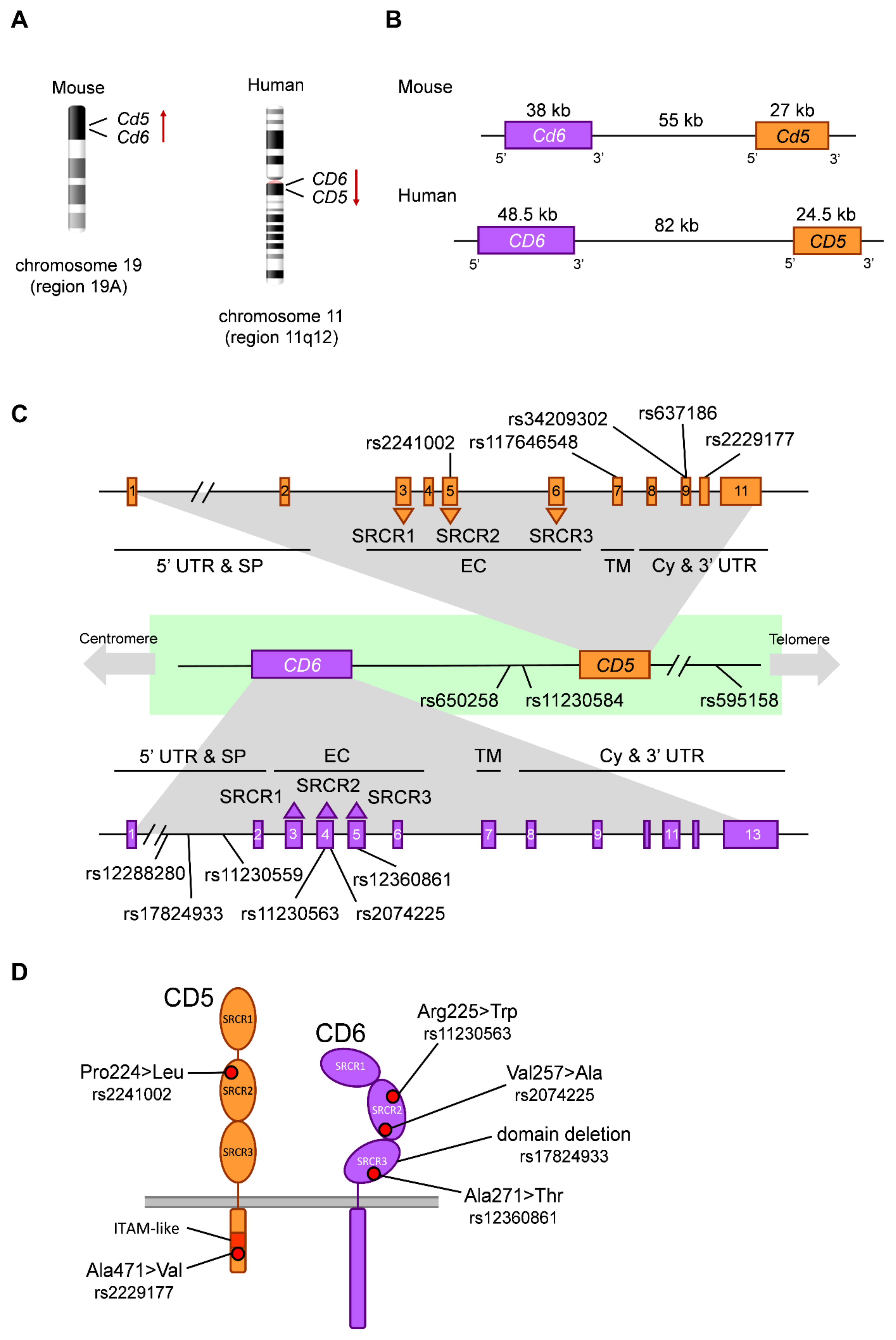
2.1. The CD5 and CD6 Protein Receptors: Structure and Function
2.2. The CD5 and CD6 Genes: Location, Exon/Intron Organization and Isoforms
2.3. Functionally Relevant CD5 and CD6 Polymorphisms
2.4. Discovery of the CD5 and CD6 Loci as Targets of Natural Selection
2.5. CD5 Polymorphism in Autoimmunity and Cancer
2.6. CD6 Polymorphism in Autoimmunity and Cancer
| Gene | SNP | Alleles * | Change | CADD | AFR | EUR | EAS | SAS | AMR | Functional/Clinical Relevance |
|---|---|---|---|---|---|---|---|---|---|---|
| CD5 | rs2241002 | C>T | Pro224>Leu | 11.06 | 0.31 | 0.15 | 0.06 | 0.15 | 0.14 | T allele associated to lower risk of lupus nephritis [66] and higher melanoma mortality [73]. Haplotypic combinations with rs2229177 associated to lupus nephritis [66] and survival in melanoma [73] and chronic lymphocytic leukemia (CLL) [75]. |
| rs117646548 | G>A | Ala377>Thr | 11.86 | 0.00 | 0.01 | 0.00 | 0.00 | 0.01 | ||
| rs34209302 | C>T | His461>Tyr | 0.092 | 0.08 | 0.01 | 0.00 | 0.08 | 0.01 | ||
| rs637186 | G>A | Arg461>His | 0.014 | 0.01 | 0.08 | 0.00 | 0.04 | 0.05 | ||
| rs2229177 | C>T | Ala471>Val | 25.2 | 0.51 | 0.55 | 0.99 | 0.80 | 0.66 | T allele associated to more signaling upon CD5 stimulation [65], stronger TCR inhibition [66], decreased lupus nephritis risk [66] and lower survival in melanoma [73] and CLL [75]. | |
| Inter-genic | rs650258 | T>C | 0.051 | 0.65 | 0.63 | 0.88 | 0.76 | 0.76 | C allele associated to increased multiple sclerosis (MS) risk [67,79]. | |
| rs11230584 | G>A | 1.789 | 0.25 | 0.15 | 0.13 | 0.18 | 0.11 | Modulation of CD5 and CD6 expression [69]. | ||
| rs595158 | C>A | 2.165 | 0.55 | 0.54 | 0.99 | 0.79 | 0.67 | Risk locus in rheumatoid arthritis [90]. | ||
| CD6 | rs12288280 | G>T | Intronic | 2.973 | 0.50 | 0.10 | 0.10 | 0.05 | 0.14 | T allele associated to decreased neuromyelitis optica risk in an Asian cohort [84]. |
| rs17824933 | C>G | Intronic | 7.58 | 0.01 | 0.23 | 0.03 | 0.07 | 0.12 | G allele associated to increased expression of CD6Δd3 [68], increased MS risk in European cohorts [76,77,78] and increased psoriasis severity [85]. | |
| rs11230559 | T>C | Intronic | 4.239 | 0.01 | 0.25 | 0.04 | 0.07 | 0.12 | In linkage disequilibrium with rs17824933 [67]. | |
| rs11230563 | C>T | Arg225>Trp | 22.4 | 0.61 | 0.36 | 0.17 | 0.21 | 0.30 | Haplotypic combinations with rs2074225 associated to differential CD6 expression [67]. T allele associated to decreased MS risk in an African American cohort [83], decreased psoriasis severity [85] and increased Behçet’s disease risk in a Han population [86]. Involvement in inflammatory bowel disease [87,88]. | |
| rs2074225 | T>C | Val257>Ala | 17.66 | 0.33 | 0.38 | 0.59 | 0.54 | 0.56 | Haplotypic combinations with rs11230563 associated to differential CD6 expression [67]. T allele associated to increased MS risk in a European cohort [67]. | |
| rs12360861 | G>A | Ala271>Thr | 0.001 | 0.04 | 0.19 | 0.00 | 0.05 | 0.12 | A allele associated to decreased MS risk in a European cohort [80] and increased psoriasis severity [85]. |
3. Concluding Remark
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cohen, M.N.; Armelagos, G. Paleopathology at the Origins of Agriculture; Cohen, M., Armelagos, G., Eds.; Academic Press: Orlando, FL, USA, 1984; ISBN 978-0-8130-4489-7. [Google Scholar]
- Morens, D.M.; Fauci, A.S. Emerging Pandemic Diseases: How We Got to COVID-19. Cell 2020, 182, 1077–1092. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Andrés, J.; Netea, M.G. Impact of Historic Migrations and Evolutionary Processes on Human Immunity. Trends Immunol. 2019, 40, 1105–1119. [Google Scholar] [CrossRef] [PubMed]
- Barreiro, L.B.; Quintana-Murci, L. From evolutionary genetics to human immunology: How selection shapes host defence genes. Nat. Rev. Genet. 2010, 11, 17–30. [Google Scholar] [CrossRef] [PubMed]
- Okada, H.; Kuhn, C.; Feillet, H.; Bach, J.F. The “hygiene hypothesis” for autoimmune and allergic diseases: An update. Clin. Exp. Immunol. 2010, 160, 1–9. [Google Scholar] [CrossRef]
- Akey, J.M. Constructing genomic maps of positive selection in humans: Where do we go from here? Genome Res. 2009, 19, 711–722. [Google Scholar] [CrossRef]
- Sabeti, P.C.; Schaffner, S.F.; Fry, B.; Lohmueller, J.; Varilly, P.; Shamovsky, O.; Palma, A.; Mikkelsen, T.S.; Altshuler, D.; Lander, E.S. Positive natural selection in the human lineage. Science 2006, 312, 1614–1620. [Google Scholar] [CrossRef]
- Williamson, S.H.; Hubisz, M.J.; Clark, A.G.; Payseur, B.A.; Bustamante, C.D.; Nielsen, R. Localizing recent adaptive evolution in the human genome. PLoS Genet. 2007, 3, 0030090. [Google Scholar] [CrossRef]
- Grossman, S.R.; Shlyakhter, I.; Shylakhter, I.; Karlsson, E.K.; Byrne, E.H.; Morales, S.; Frieden, G.; Hostetter, E.; Angelino, E.; Garber, M.; et al. A composite of multiple signals distinguishes causal variants in regions of positive selection. Science 2010, 327, 883–886. [Google Scholar] [CrossRef]
- Fan, S.; Hansen, M.E.B.; Lo, Y.; Tishkoff, S.A. Going global by adapting local: A review of recent human adaptation. Science 2016, 354, 54–59. [Google Scholar] [CrossRef]
- Aguirre-Gamboa, R.; Joosten, I.; Urbano, P.C.M.; van der Molen, R.G.; van Rijssen, E.; van Cranenbroek, B.; Oosting, M.; Smeekens, S.; Jaeger, M.; Zorro, M.; et al. Differential Effects of Environmental and Genetic Factors on T and B Cell Immune Traits. Cell Rep. 2016, 17, 2474–2487. [Google Scholar] [CrossRef]
- Li, Y.; Oosting, M.; Smeekens, S.P.; Jaeger, M.; Aguirre-Gamboa, R.; Le, K.T.T.; Deelen, P.; Ricaño-Ponce, I.; Schoffelen, T.; Jansen, A.F.M.; et al. A Functional Genomics Approach to Understand Variation in Cytokine Production in Humans. Cell 2016, 167, 1099–1110.e14. [Google Scholar] [CrossRef]
- Quach, H.; Rotival, M.; Pothlichet, J.; Loh, Y.H.E.; Dannemann, M.; Zidane, N.; Laval, G.; Patin, E.; Harmant, C.; Lopez, M.; et al. Genetic Adaptation and Neandertal Admixture Shaped the Immune System of Human Populations. Cell 2016, 167, 643–656.e17. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31, 3812–3814. [Google Scholar] [CrossRef]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef]
- Kircher, M.; Witten, D.M.; Jain, P.; O′roak, B.J.; Cooper, G.M.; Shendure, J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 2014, 46, 310–315. [Google Scholar] [CrossRef]
- Barreiro, L.B.; Ben-Ali, M.; Quach, H.; Laval, G.; Patin, E.; Pickrell, J.K.; Bouchier, C.; Tichit, M.; Neyrolles, O.; Gicquel, B.; et al. Evolutionary dynamics of human toll-like receptors and their different contributions to host defense. PLoS Genet. 2009, 5. [Google Scholar] [CrossRef]
- Laayouni, H.; Oosting, M.; Luisi, P.; Ioana, M.; Alonso, S.; Ricano-Ponce, I.; Trynka, G.; Zhernakova, A.; Plantinga, T.S.; Cheng, S.C.; et al. Convergent evolution in European and Rroma populations reveals pressure exerted by plague on Toll-like receptors. Proc. Natl. Acad. Sci. USA 2014, 111, 2668–2673. [Google Scholar] [CrossRef]
- Dannemann, M.; Andrés, A.M.; Kelso, J. Introgression of Neandertal-and Denisovan-like Haplotypes Contributes to Adaptive Variation in Human Toll-like Receptors. Am. J. Hum. Genet. 2016, 98, 22–33. [Google Scholar] [CrossRef]
- Fry, A.E.; Ghansa, A.; Small, K.S.; Palma, A.; Auburn, S.; Diakite, M.; Green, A.; Campino, S.; Teo, Y.Y.; Clark, T.G.; et al. Positive selection of a CD36 nonsense variant in sub-Saharan Africa, but no association with severe malaria phenotypes. Hum. Mol. Genet. 2009, 18, 2683–2692. [Google Scholar] [CrossRef][Green Version]
- Wang, J.; Li, Y. CD36 tango in cancer: Signaling pathways and functions. Theranostics 2019, 9, 4893–4908. [Google Scholar] [CrossRef]
- Park, Y.M. CD36, a scavenger receptor implicated in atherosclerosis. Exp. Mol. Med. 2014, 46, e99. [Google Scholar] [CrossRef]
- Zhao, L.; Varghese, Z.; Moorhead, J.F.; Chen, Y.; Ruan, X.Z. CD36 and lipid metabolism in the evolution of atherosclerosis. Br. Med. Bull. 2018, 126, 101–112. [Google Scholar] [CrossRef]
- Hughes, A.L.; Nei, M. Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 1988, 335, 167–170. [Google Scholar] [CrossRef]
- Hedrick, P.W. Pathogen resistance and genetic variation at MHC loci. Evolution 2002, 56, 1902–1908. [Google Scholar] [CrossRef]
- Prugnolle, F.; Manica, A.; Charpentier, M.; Guégan, J.F.; Guernier, V.; Balloux, F. Pathogen-driven selection and worldwide HLA class I diversity. Curr. Biol. 2005, 15, 1022–1027. [Google Scholar] [CrossRef] [PubMed]
- Walsh, E.C.; Mather, K.A.; Schaffner, S.F.; Farwell, L.; Daly, M.J.; Patterson, N.; Cullen, M.; Carrington, M.; Bugawan, T.L.; Erlich, H.; et al. An integrated haplotype map of the human major histocompatibility complex. Am. J. Hum. Genet. 2003, 73, 580–590. [Google Scholar] [CrossRef]
- De Bakker, P.I.W.; McVean, G.; Sabeti, P.C.; Miretti, M.M.; Green, T.; Marchini, J.; Ke, X.; Monsuur, A.J.; Whittaker, P.; Delgado, M.; et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 2006, 38, 1166–1172. [Google Scholar] [CrossRef]
- Santos, R.F.; Oliveira, L.; Carmo, A.M. Tuning T Cell Activation: The Function of CD6 At the Immunological Synapse and in T Cell Responses. Curr. Drug Targets 2016, 17, 630–639. [Google Scholar] [CrossRef]
- Burgueño-Bucio, E.; Mier-Aguilar, C.A.; Soldevila, G. The multiple faces of CD5. J. Leukoc. Biol. 2019, 105, 891–904. [Google Scholar] [CrossRef]
- Cho, J.-H.; Sprent, J. TCR tuning of T cell subsets. Immunol. Rev. 2018, 283, 129–137. [Google Scholar] [CrossRef]
- Chappell, P.E.; Garner, L.I.; Yan, J.; Metcalfe, C.; Hatherley, D.; Johnson, S.; Robinson, C.V.; Lea, S.M.; Brown, M.H. Structures of CD6 and Its Ligand CD166 Give Insight into Their Interaction. Structure 2015, 23, 1426–1436. [Google Scholar] [CrossRef] [PubMed]
- Bowen, M.A.; Patel, D.D.; Li, X.; Modrell, B.; Malacko, A.R.; Wang, W.C.; Marquardt, H.; Neubauer, M.; Pesando, J.M.; Francke, U.; et al. Cloning, mapping, and characterization of activated leukocyte-cell adhesion molecule (ALCAM), a CD6 ligand. J. Exp. Med. 1995, 181, 2213–2220. [Google Scholar] [CrossRef] [PubMed]
- Levin, T.G.; Powell, A.E.; Davies, P.S.; Silk, A.D.; Dismuke, A.D.; Anderson, E.C.; Swain, J.R.; Wong, M.H. Characterization of the intestinal cancer stem cell marker CD166 in the human and mouse gastrointestinal tract. Gastroenterology 2010, 139, 2072–2082.e5. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.D.; Wee, S.F.; Whichard, L.P.; Bowen, M.A.; Pesando, J.M.; Aruffo, A.; Haynes, B.F. Identification and characterization of a 100-kD ligand for CD6 on human thymic epithelial cells. J. Exp. Med. 1995, 181, 1563–1568. [Google Scholar] [CrossRef]
- Donizy, P.; Zietek, M.; Halon, A.; Leskiewicz, M.; Kozyra, C.; Matkowski, R. Prognostic significance of ALCAM (CD166/MEMD) expression in cutaneous melanoma patients. Diagn. Pathol. 2015, 10, 86. [Google Scholar] [CrossRef]
- Escoda-Ferran, C.; Carrasco, E.; Caballero-Baños, M.; Miró-Julià, C.; Martínez-Florensa, M.; Consuegra-Fernández, M.; Martínez, V.G.; Liu, F.-T.; Lozano, F. Modulation of CD6 function through interaction with Galectin-1 and -3. FEBS Lett. 2014, 588, 2805–2813. [Google Scholar] [CrossRef]
- Enyindah-Asonye, G.; Li, Y.; Ruth, J.H.; Spassov, D.S.; Hebron, K.E.; Zijlstra, A.; Moasser, M.M.; Wang, B.; Singer, N.G.; Cui, H.; et al. CD318 is a ligand for CD6. Proc. Natl. Acad. Sci. USA 2017, 114, E6912–E6921. [Google Scholar] [CrossRef]
- Cayrol, R.; Wosik, K.; Berard, J.L.; Dodelet-Devillers, A.; Ifergan, I.; Kebir, H.; Haqqani, A.S.; Kreymborg, K.; Krug, S.; Moumdjian, R.; et al. Activated leukocyte cell adhesion molecule promotes leukocyte trafficking into the central nervous system. Nat. Immunol. 2008, 9, 137–145. [Google Scholar] [CrossRef]
- Tabbekh, M.; Mokrani-Hammani, M.; Bismuth, G.; Mami-Chouaib, F. T-cell modulatory properties of CD5 and its role in antitumor immune responses. Oncoimmunology 2013, 2, e22841. [Google Scholar] [CrossRef]
- Consuegra-Fernández, M.; Aranda, F.; Simões, I.; Orta, M.; Sarukhan, A.; Lozano, F. CD5 as a target for immune-based therapies. Crit. Rev. Immunol. 2015, 35, 85–115. [Google Scholar] [CrossRef]
- Dennehy, K.M.; Broszeit, R.; Ferris, W.F.; Beyers, A.D. Thymocyte activation induces the association of the proto-oncoprotein c-cbl and ras GTPase-activating protein with CD5. Eur. J. Immunol. 1998, 28, 1617–1625. [Google Scholar] [CrossRef]
- Demydenko, D. c-Cbl mediated ubiquitylation and regulation of cell surface exposure of CD5. Biochem. Biophys. Res. Commun. 2010, 392, 500–504. [Google Scholar] [CrossRef]
- Axtell, R.C.; Xu, L.; Barnum, S.R.; Raman, C. CD5-CK2 Binding/Activation-Deficient Mice Are Resistant to Experimental Autoimmune Encephalomyelitis: Protection Is Associated with Diminished Populations of IL-17-Expressing T Cells in the Central Nervous System. J. Immunol. 2006, 177, 8542–8549. [Google Scholar] [CrossRef]
- Soldevila, G.; Raman, C.; Lozano, F. The immunomodulatory properties of the CD5 lymphocyte receptor in health and disease. Curr. Opin. Immunol. 2011, 23, 310–318. [Google Scholar] [CrossRef]
- Mori, D.; Grégoire, C.; Voisinne, G.; Celis-Gutierrez, J.; Aussel, R.; Girard, L.; Camus, M.; Marcellin, M.; Argenty, J.; Burlet-Schiltz, O.; et al. The T cell CD6 receptor operates a multitask signalosome with opposite functions in T cell activation. J. Exp. Med. 2021, 218, e20201011. [Google Scholar] [CrossRef]
- Gimferrer, I.; Ibáñez, A.; Farnós, M.; Sarrias, M.-R.; Fenutría, R.; Roselló, S.; Zimmermann, P.; David, G.; Vives, J.; Serra-Pagès, C.; et al. The Lymphocyte Receptor CD6 Interacts with Syntenin-1, a Scaffolding Protein Containing PDZ Domains. J. Immunol. 2005, 175, 1406–1414. [Google Scholar] [CrossRef]
- Hassan, N.J.; Simmonds, S.J.; Clarkson, N.G.; Hanrahan, S.; Puklavec, M.J.; Bomb, M.; Barclay, A.N.; Brown, M.H. CD6 Regulates T-Cell Responses through Activation-Dependent Recruitment of the Positive Regulator SLP-76. Mol. Cell. Biol. 2006, 26, 6727–6738. [Google Scholar] [CrossRef]
- Breuning, J.; Brown, M.H. T Cell Costimulation by CD6 Is Dependent on Bivalent Binding of a GADS/SLP-76 Complex. Mol. Cell. Biol. 2017, 37, 71–88. [Google Scholar] [CrossRef]
- Hem, C.D.; Ekornhol, M.; Granum, S.; Sundvold-Gjerstad, V.; Spurkland, A. CD6 and Linker of Activated T Cells are Potential Interaction Partners for T Cell-Specific Adaptor Protein. Scand. J. Immunol. 2017, 85, 104–112. [Google Scholar] [CrossRef]
- Vera, J.; Fenutria, R.; Canadas, O.; Figueras, M.; Mota, R.; Sarrias, M.-R.; Williams, D.L.; Casals, C.; Yelamos, J.; Lozano, F. The CD5 ectodomain interacts with conserved fungal cell wall components and protects from zymosan-induced septic shock-like syndrome. Proc. Natl. Acad. Sci. USA 2009, 106, 1506–1511. [Google Scholar] [CrossRef]
- Sarhan, M.A.; Pham, T.N.Q.; Chen, A.Y.; Michalak, T.I. Hepatitis C Virus Infection of Human T Lymphocytes Is Mediated by CD5. J. Virol. 2012, 86, 3723–3735. [Google Scholar] [CrossRef] [PubMed]
- Mourglia-Ettlin, G.; Miles, S.; Velasco-De-Andrés, M.; Armiger-Borràs, N.; Cucher, M.; Dematteis, S.; Lozano, F. The ectodomains of the lymphocyte scavenger receptors CD5 and CD6 interact with tegumental antigens from Echinococcus granulosus sensu lato and protect mice against secondary cystic echinococcosis. PLoS Negl. Trop. Dis. 2018, 12, e0006891. [Google Scholar] [CrossRef] [PubMed]
- Sarrias, M.-R.; Farnós, M.; Mota, R.; Sánchez-Barbero, F.; Ibáñez, A.; Gimferrer, I.; Vera, J.; Fenutría, R.; Casals, C.; Yélamos, J.; et al. CD6 binds to pathogen-associated molecular patterns and protects from LPS-induced septic shock. Proc. Natl. Acad. Sci. USA 2007, 104, 11724–11729. [Google Scholar] [CrossRef] [PubMed]
- Carrasco, E.; Escoda, C.; Alvarez-Fenrández, C.; Sanchez-Palomino, S.; Carreras, E.; Gatell, J.M.; Gallart, T.; García, F.; Climent, N.; Lozano, F. A role for scavenger-like lymphocyte receptor CD6 in HIV-1 viral infection. AIDS Res. Hum. Retrovir. 2014, 30, A49–A50. [Google Scholar] [CrossRef]
- Velasco-de-Andrés, M.; Català, C.; Casadó-Llombart, S.; Simões, I.; Zaragoza, O.; Carreras, E.; Lozano, F. The lymphocyte scavenger receptor CD5 plays a nonredundant role in fungal infection. Cell. Mol. Immunol. 2021, 18, 498–500. [Google Scholar] [CrossRef]
- Català, C.; Velasco-de Andrés, M.; Casadó-Llombart, S.; Martínez-Florensa, M.; García-Luna, J.; Leyton-Pereira, A.; Aranda, F.; Consuegra-Fernández, M.; Mourglia-Ettlin, G.; Lozano, F. CD6 deficiency confers susceptibility to bacterial sepsis. In Proceedings of the 42nd Congress of Sociedad Española de Inmunología, Madrid, Spain, 24–26 March 2021. [Google Scholar]
- Lecomte, O.; Bock, J.B.; Birren, B.W.; Vollrath, D.; Parnes, J.R. Molecular linkage of the mouse CD5 and CD6 genes. Immunogenetics 1996, 44, 385–390. [Google Scholar] [CrossRef]
- Padilla, O.; Calvo, J.; Vila, J.M.; Arman, M.; Gimferrer, I.; Places, L.; Arias, M.T.; Pujana, M.A.; Vives, J.; Lozano, F. Genomic organization of the human CD5 gene. Immunogenetics 2000, 51, 993–1001. [Google Scholar] [CrossRef]
- Jones, N.H.; Clabby, M.L.; Dialynas, D.P.; Huang, H.J.S.; Herzenberg, L.A.; Strominger, J.L. Isolation of complementary DNA clones encoding the human lymphocyte glycoprotein T1/Leu-1. Nature 1986, 323, 346–349. [Google Scholar] [CrossRef]
- Renaudineau, Y.; Hillion, S.; Saraux, A.; Mageed, R.A.; Youinou, P. An alternative exon 1 of the CD5 gene regulates CD5 expression in human B lymphocytes. Blood 2005, 106, 2781–2789. [Google Scholar] [CrossRef]
- Bowen, M.A.; Whitney, G.S.; Neubauer, M.; Starling, G.C.; Palmer, D.; Zhang, J.; Nowak, N.J.; Shows, T.B.; Aruffo, A. Structure and Chromosomal Location of the Human CD6 Gene: Detection of Five Human CD6 Isoforms. J. Immunol. 1997, 158, 1149–1156. [Google Scholar]
- Bonet, L.; Farnós, M.; Martínez-Florensa, M.; Martínez, V.G.; Lozano, F. Identification of functionally relevant phoshorylatable serine clusters in the cytoplasmic region of the human CD6 lymphocyte surface receptor. FEBS Lett. 2013, 587, 2205–2213. [Google Scholar] [CrossRef]
- Castro, M.A.A.; Oliveira, M.I.; Nunes, R.J.; Fabre, S.; Barbosa, R.; Peixoto, A.; Brown, M.H.; Parnes, J.R.; Bismuth, G.; Moreira, A.; et al. Extracellular Isoforms of CD6 Generated by Alternative Splicing Regulate Targeting of CD6 to the Immunological Synapse. J. Immunol. 2007, 178, 4351–4361. [Google Scholar] [CrossRef]
- Carnero-Montoro, E.; Bonet, L.; Engelken, J.; Bielig, T.; Martínez-Florensa, M.; Lozano, F.; Bosch, E. Evolutionary and functional evidence for positive selection at the human CD5 immune receptor gene. Mol. Biol. Evol. 2012, 29, 811–823. [Google Scholar] [CrossRef]
- Cenit, M.C.; Martínez-Florensa, M.; Consuegra, M.; Bonet, L.; Carnero-Montoro, E.; Armiger, N.; Caballero-Baños, M.; Arias, M.T.; Benitez, D.; Ortego-Centeno, N.; et al. Analysis of ancestral and functionally relevant CD5 variants in systemic lupus erythematosus patients. PLoS ONE 2014, 9, e113090. [Google Scholar] [CrossRef]
- Swaminathan, B.; Cuapio, A.; Alloza, I.; Matesanz, F.; Alcina, A.; García-Barcina, M.; Fedetz, M.; Fernández, Ó.; Lucas, M.; Órpez, T.; et al. Fine Mapping and Functional Analysis of the Multiple Sclerosis Risk Gene CD6. PLoS ONE 2013, 8, e62376. [Google Scholar] [CrossRef]
- Kofler, D.M.; Severson, C.A.; Mousissian, N.; De Jager, P.L.; Hafler, D.A. The CD6 Multiple Sclerosis Susceptibility Allele Is Associated with Alterations in CD4+ T Cell Proliferation. J. Immunol. 2011, 187, 3286–3291. [Google Scholar] [CrossRef]
- Peters, J.E.; Lyons, P.A.; Lee, J.C.; Richard, A.C.; Fortune, M.D.; Newcombe, P.J.; Richardson, S.; Smith, K.G.C. Insight into Genotype-Phenotype Associations through eQTL Mapping in Multiple Cell Types in Health and Immune-Mediated Disease. PLoS Genet. 2016, 12, e1005908. [Google Scholar] [CrossRef]
- Moreno-Estrada, A.; Tang, K.; Sikora, M.; Marqus-Bonet, T.; Casals, F.; Navarro, A.; Calafell, F.; Bertranpetit, J.; Stoneking, M.; Bosch, E. Interrogating 11 fast-evolving genes for signatures of recent positive selection in worldwide human populations. Mol. Biol. Evol. 2009, 26, 2285–2297. [Google Scholar] [CrossRef]
- Eastman, S.W.; Martin-Serrano, J.; Chung, W.; Zang, T.; Bieniasz, P.D. Identification of human VPS37C, a component of endosomal sorting complex required for transport-I important for viral budding. J. Biol. Chem. 2005, 280, 628–636. [Google Scholar] [CrossRef]
- Casadó-Llombart, S.; Velasco-de Andrés, M.; Català, C.; Leyton-Pereira, A.; Suárez, B.; Armiger, N.; Carreras, E.; Esteller, M.; Ricart, E.; Ordás, E.; et al. CD5 and CD6 lymphocyte co-receptor expression and variation impact experimental and clinical inflammatory bowel disease. 2021; submitted. [Google Scholar]
- Potrony, M.; Carreras, E.; Aranda, F.; Zimmer, L.; Puig-Butille, J.-A.; Tell-Martí, G.; Armiger, N.; Sucker, A.; Giménez-Xavier, P.; Martínez-Florensa, M.; et al. Inherited functional variants of the lymphocyte receptor CD5 influence melanoma survival. Int. J. Cancer 2016, 139, 1297–1302. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Sawada, H.; Izumi, Y.; Fukuda, T.; Utsunomiya, A.; Ikeda, S.; Uike, N.; Tsukada, J.; Kawano, F.; Shibuya, T.; et al. Chronic lymphocytic leukemia (CLL) is rare, but the proportion of T-CLL is high in Japan. Eur. J. Haematol. 2001, 67, 152–157. [Google Scholar] [CrossRef] [PubMed]
- Delgado, J.; Bielig, T.; Bonet, L.; Carnero-Montoro, E.; Puente, X.S.; Colomer, D.; Bosch, E.; Campo, E.; Lozano, F. Impact of the functional CD5 polymorphism A471V on the response of chronic lymphocytic leukaemia to conventional chemotherapy regimens. Br. J. Haematol. 2017, 177, 147–150. [Google Scholar] [CrossRef] [PubMed][Green Version]
- De Jager, P.L.; Jia, X.; Wang, J.; de Bakker, P.I.W.; Ottoboni, L.; Aggarwal, N.T.; Piccio, L.; Raychaudhuri, S.; Tran, D.; Aubin, C.; et al. Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat. Genet. 2009, 41, 776–782. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, B.; Matesanz, F.; Cavanillas, M.L.; Alloza, I.; Otaegui, D.; Olascoaga, J.; Cénit, M.C.; de las Heras, V.; Barcina, M.G.; Arroyo, R.; et al. Validation of the CD6 and TNFRSF1A loci as risk factors for multiple sclerosis in Spain. J. Neuroimmunol. 2010, 223, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Leppä, V.; Surakka, I.; Tienari, P.J.; Elovaara, I.; Compston, A.; Sawcer, S.; Robertson, N.; de Jager, P.L.; Aubin, C.; Hafler, D.A.; et al. The Genetic Association of Variants in CD6, TNFRSF1A and IRF8 to Multiple Sclerosis: A Multicenter Case-Control Study. PLoS ONE 2011, 6, e18813. [Google Scholar] [CrossRef]
- Sawcer, S.; Hellenthal, G.; Pirinen, M.; Spencer, C.C.A.; Patsopoulos, N.A.; Moutsianas, L.; Dilthey, A.; Su, Z.; Freeman, C.; Hunt, S.E.; et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 2011, 476, 214–219. [Google Scholar] [CrossRef]
- Wagner, M.; Bilinska, M.; Pokryszko-Dragan, A.; Sobczynski, M.; Cyrul, M.; Kusnierczyk, P.; Jasek, M. ALCAM and CD6—Multiple sclerosis risk factors. J. Neuroimmunol. 2014, 276, 98–103. [Google Scholar] [CrossRef]
- Wagner, M.; Wiśniewski, A.; Bilińska, M.; Pokryszko-Dragan, A.; Nowak, I.; Kuśnierczyk, P.; Jasek, M. ALCAM—Novel multiple sclerosis locus interfering with HLA-DRB1*1501. J. Neuroimmunol. 2013, 258, 71–76. [Google Scholar] [CrossRef]
- Zhou, P.; Du, L.F.; Lv, G.Q.; Yu, X.M.; Gu, Y.L.; Li, J.P.; Zhang, C. Functional polymorphisms in CD166/ALCAM gene associated with increased risk for breast cancer in a Chinese population. Breast Cancer Res. Treat. 2011, 128, 527–534. [Google Scholar] [CrossRef]
- Johnson, B.A.; Wang, J.; Taylor, E.M.; Caillier, S.J.; Herbert, J.; Khan, O.A.; Cross, A.H.; De Jager, P.L.; Gourraud, P.A.F.; Cree, B.C.A.; et al. Multiple sclerosis susceptibility alleles in African Americans. Genes Immun. 2010, 11, 343–350. [Google Scholar] [CrossRef]
- Park, T.J.; Kim, H.J.; Kim, J.H.; Bae, J.S.; Cheong, H.S.; Park, B.L.; Shin, H.D. Associations of CD6, TNFRSF1A and IRF8 polymorphisms with risk of inflammatory demyelinating diseases. Neuropathol. Appl. Neurobiol. 2013, 39, 519–530. [Google Scholar] [CrossRef]
- Consuegra-Fernández, M.; Julià, M.; Martínez-Florensa, M.; Aranda, F.; Català, C.; Armiger-Borràs, N.; Arias, M.T.; Santiago, F.; Guilabert, A.; Esteve, A.; et al. Genetic and experimental evidence for the involvement of the CD6 lymphocyte receptor in psoriasis. Cell. Mol. Immunol. 2018, 15, 898–906. [Google Scholar] [CrossRef]
- Zheng, M.; Zhang, L.; Yu, H.; Hu, J.; Cao, Q.; Huang, G.; Huang, Y.; Yuan, G.; Kijlstra, A.; Yang, P. Genetic polymorphisms of cell adhesion molecules in Behçet’s disease in a Chinese Han population. Sci. Rep. 2016, 6, 24974. [Google Scholar] [CrossRef]
- Jostins, L.; Ripke, S.; Weersma, R.K.; Duerr, R.H.; McGovern, D.P.; Hui, K.Y.; Lee, J.C.; Philip Schumm, L.; Sharma, Y.; Anderson, C.A.; et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 2012, 491, 119–124. [Google Scholar] [CrossRef]
- Ellinghaus, D.; Jostins, L.; Spain, S.L.; Cortes, A.; Bethune, J.; Han, B.; Park, Y.R.; Raychaudhuri, S.; Pouget, J.G.; Hübenthal, M.; et al. Analysis of five chronic inflammatory diseases identifies 27 new associations and highlights disease-specific patterns at shared loci. Nat. Genet. 2016, 48, 510–518. [Google Scholar] [CrossRef]
- Darvishi, B.; Boroumandieh, S.; Majidzadeh-A, K.; Salehi, M.; Jafari, F.; Farahmand, L. The role of activated leukocyte cell adhesion molecule (ALCAM) in cancer progression, invasion, metastasis and recurrence: A novel cancer stem cell marker and tumor-specific prognostic marker. Exp. Mol. Pathol. 2020, 115, 104443. [Google Scholar] [CrossRef]
- Eyre, S.; Bowes, J.; Diogo, D.; Lee, A.; Barton, A.; Martin, P.; Zhernakova, A.; Stahl, E.; Viatte, S.; McAllister, K.; et al. High-density genetic mapping identifies new susceptibility loci for rheumatoid arthritis. Nat. Genet. 2012, 44, 1336–1340. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Casadó-Llombart, S.; Velasco-de Andrés, M.; Català, C.; Leyton-Pereira, A.; Lozano, F.; Bosch, E. Contribution of Evolutionary Selected Immune Gene Polymorphism to Immune-Related Disorders: The Case of Lymphocyte Scavenger Receptors CD5 and CD6. Int. J. Mol. Sci. 2021, 22, 5315. https://doi.org/10.3390/ijms22105315
Casadó-Llombart S, Velasco-de Andrés M, Català C, Leyton-Pereira A, Lozano F, Bosch E. Contribution of Evolutionary Selected Immune Gene Polymorphism to Immune-Related Disorders: The Case of Lymphocyte Scavenger Receptors CD5 and CD6. International Journal of Molecular Sciences. 2021; 22(10):5315. https://doi.org/10.3390/ijms22105315
Chicago/Turabian StyleCasadó-Llombart, Sergi, María Velasco-de Andrés, Cristina Català, Alejandra Leyton-Pereira, Francisco Lozano, and Elena Bosch. 2021. "Contribution of Evolutionary Selected Immune Gene Polymorphism to Immune-Related Disorders: The Case of Lymphocyte Scavenger Receptors CD5 and CD6" International Journal of Molecular Sciences 22, no. 10: 5315. https://doi.org/10.3390/ijms22105315
APA StyleCasadó-Llombart, S., Velasco-de Andrés, M., Català, C., Leyton-Pereira, A., Lozano, F., & Bosch, E. (2021). Contribution of Evolutionary Selected Immune Gene Polymorphism to Immune-Related Disorders: The Case of Lymphocyte Scavenger Receptors CD5 and CD6. International Journal of Molecular Sciences, 22(10), 5315. https://doi.org/10.3390/ijms22105315






