Multi-Omics Perspective Reveals the Different Patterns of Tumor Immune Microenvironment Based on Programmed Death Ligand 1 (PD-L1) Expression and Predictor of Responses to Immune Checkpoint Blockade across Pan-Cancer
Abstract
1. Introduction
2. Results
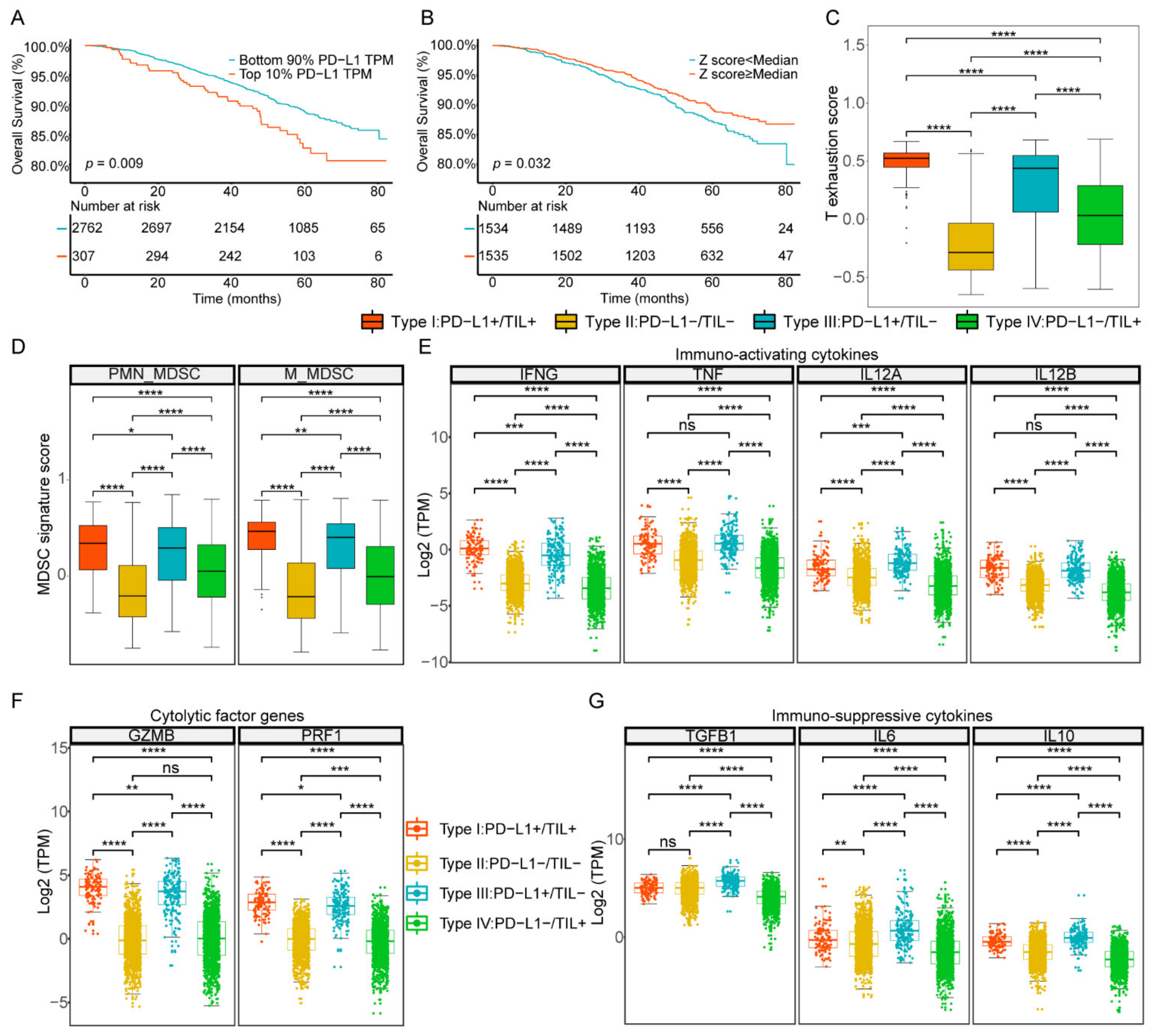
2.1. Prognostic Significance of TIL Z Score/PD-L1 to ICI Response Prediction and Stratification of Four TIME Subtypes across Pan-Cancer Types
2.2. The Composition and Abundance of Lymphocyte among Four Subtypes
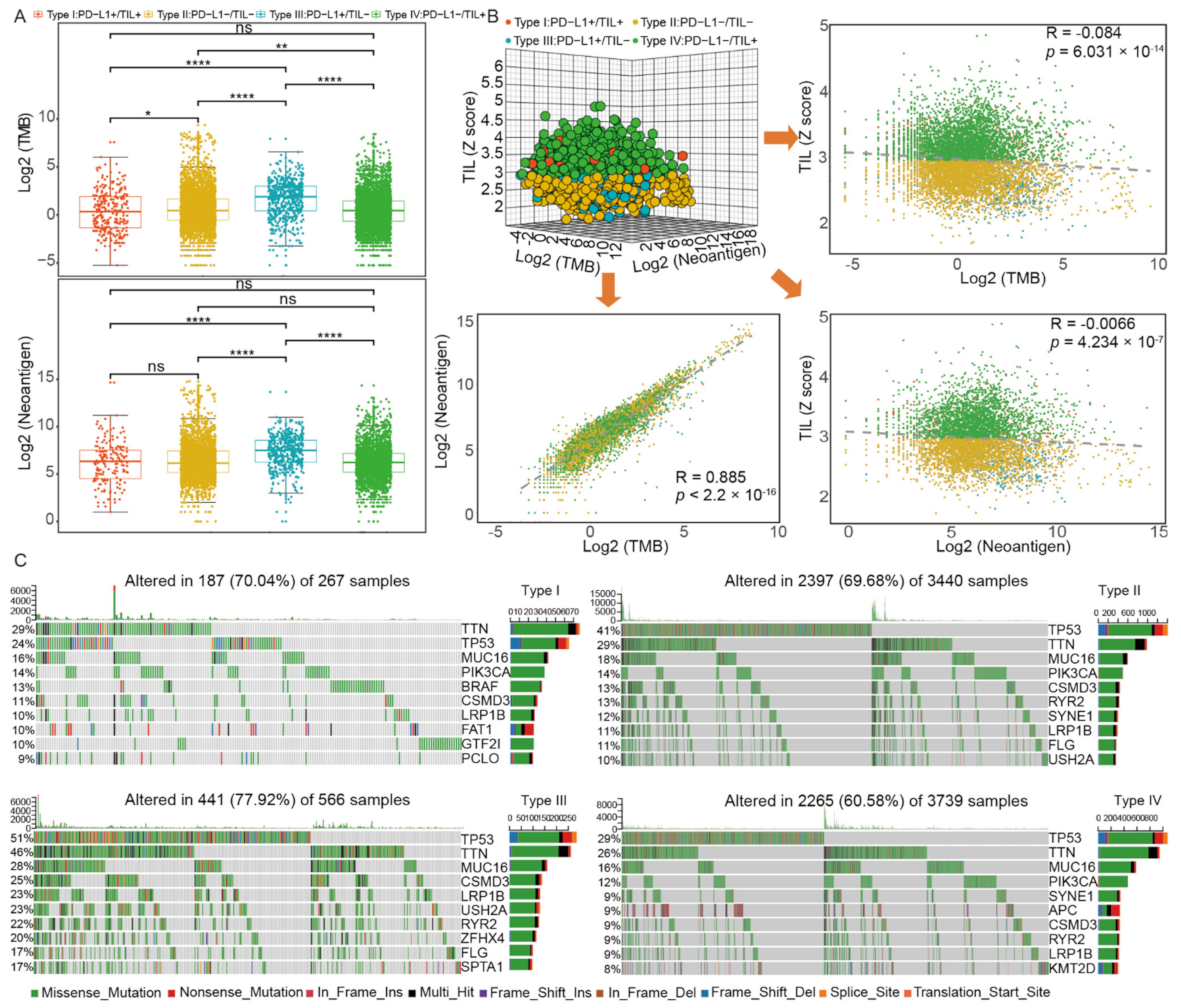
2.3. Genomics Pattern Discrepancy in Four TIME Subtypes
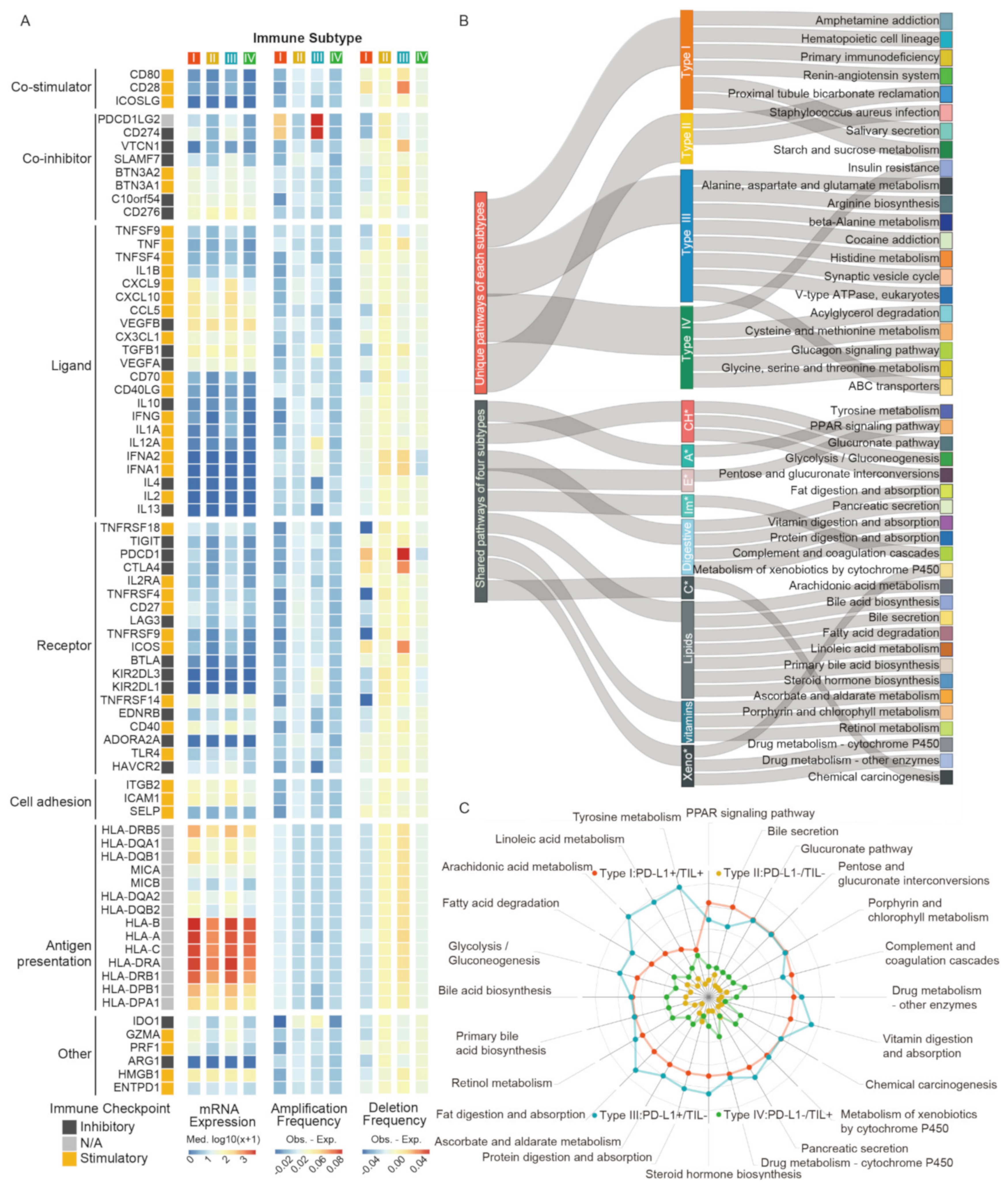
2.4. Transcriptomics Pattern Discrepancy in Four TIME Subtypes
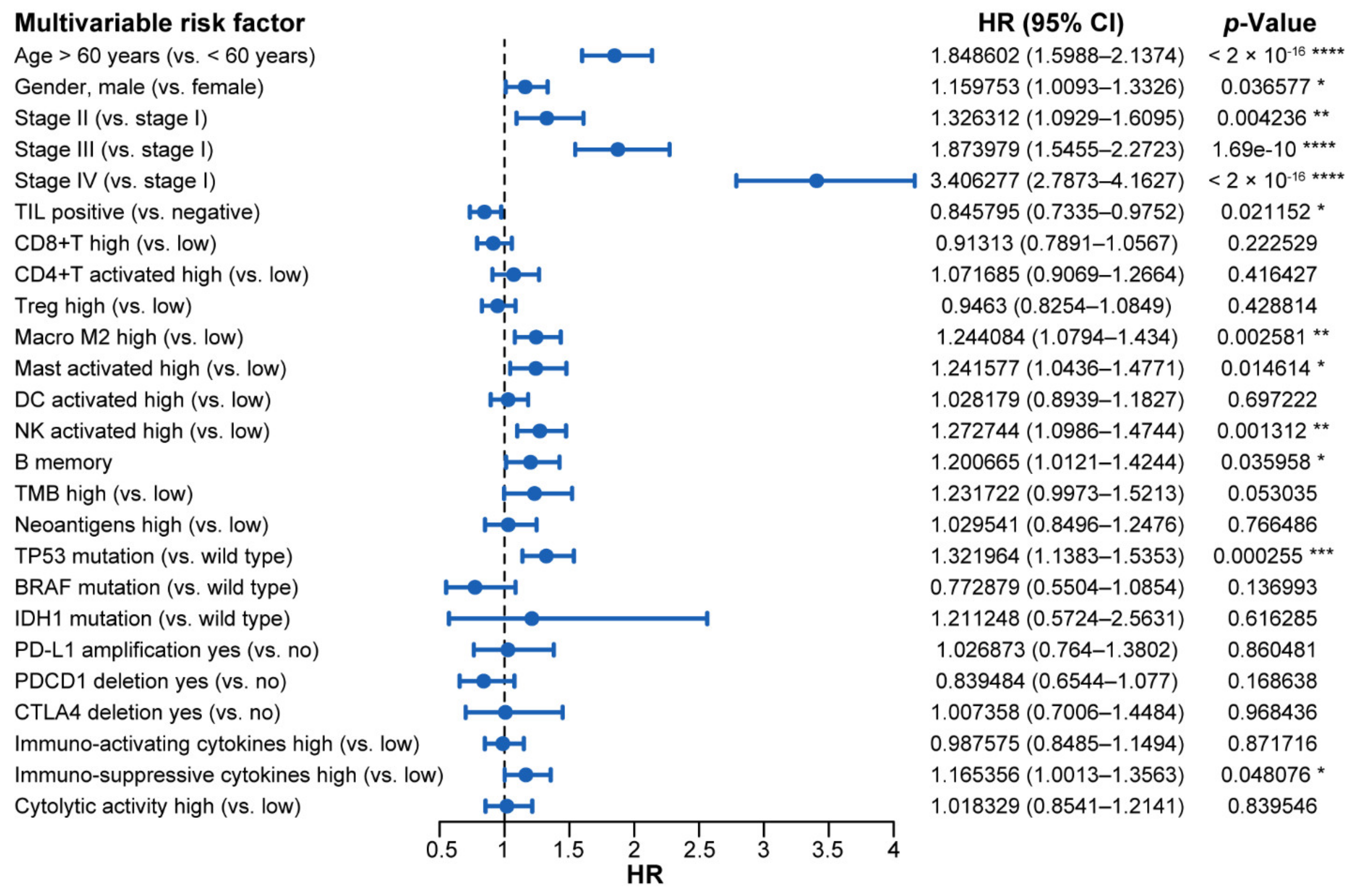
2.5. Hazard Analysis for Multiple Omics Factors across Four TIME Subtypes
2.6. Validation in GEO Dataset
3. Discussion
4. Materials and Methods
4.1. Data Collection and Preprocessing
4.2. Tumor-Infiltrating Lymphocyte Z Score
4.3. TIME Subtypes and Immune Cells Proportion
4.4. Genomic Analysis
4.5. Differential Gene Analysis and Pathway Score Analysis
4.6. Gene Set Variation Analysis (GSVA) Score of Gene Expression Signature
4.7. Survival Analysis
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ICI | Immune checkpoint inhibitors; |
| TIME | Tumor immune microenvironment; |
| PD-1 | Programmed death1; |
| PD-L1 | Programmed death ligand 1; |
| TCGA | The Cancer Genome Atlas; |
| TIL | Tumor-infiltrating lymphocyte; |
| TMB | Tumor mutation burden; |
| IHC | Immunohistochemistry; |
| THYM | Thymoma; |
| UCS | Uterine carcinosarcoma; |
| LUSC | Lung squamous cell carcinoma; |
| LIHC | Liver hepatocellular carcinoma; |
| TP53 | Tumor protein 53; |
| TTN | Titin; |
| LRP1B | LDL receptor related protein 1B; |
| CSMD3 | CUB and Sushi Multiple Domains 3; |
| BRAF | B-Raf Proto-Oncogene; |
| FAT1 | FAT Atypical Cadherin 1; |
| GTF2I | General Transcription Factor Iii; |
| PCLO | Piccolo; |
| ZFHX4 | Zinc Finger Homeobox 4; |
| SPTA1 | Spectrin alpha, erythrocytic 1; |
| APC | Adenomatous polyposis coli; |
| KMT2D | Lysine methyltransferase 2D; |
| PIK3CA | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; |
| PDCD1LG2 | Programmed Cell Death 1 Ligand 2; |
| VTCN1 | V-set domain containing T cell activation inhibitor 1; |
| PDCD1 | Programmed Cell Death 1; |
| CTLA4 | Cytotoxic T-lymphocyte-associated protein 4; |
| DEG | Different expressed gene; |
| KEGG | Kyoto Encyclopedia of Genes and Genomes; |
| IFNG | Interferon gamma; |
| TNF | Tumor necrosis factor; |
| TNFA | Tumor Necrosis Factor Alpha; |
| IL6 | Interleukin 6; |
| IL12 | Interleukin 12; |
| IL12A | Interleukin 12A; |
| IL12B | Interleukin 12B; |
| IL10 | Interleukin 10; |
| GZMB | Granzyme B; |
| PRF1 | Perforin-1; |
| KRAS | Kirsten ras; |
| VEGFA | Vascular endothelial growth factor A; |
| TGFB1 | Transforming growth factor beta 1; |
| HRAS | HRas proto-oncogene; |
| IDH1 | Isocitrate dehydrogenase (NADP(+)) 1; |
| POLE | DNA polymerase epsilon; |
| POLD1 | DNA polymerase delta 1; |
| MUC16 | Mucin 16; |
| RYR2 | Ryanodine receptor 2; |
| SYNE1 | Spectrin repeat containing nuclear envelope protein 1; |
| FLG | Filaggrin; |
| USH2A | Usherin; |
| CDKN2A | Cyclin dependent kinase inhibitor 2A; |
| MB21D2 | Mab-21 domain containing 2; |
| NDUFA13 | NADH:ubiquinone oxidoreductase subunit A13; |
| DGCR6L | DiGeorge syndrome critical region gene 6 like; |
| S100A1 | S100 calcium binding protein A1; |
| IAPP | Islet amyloid polypeptide; |
| SLC3A2 | Solute carrier family 3 member 2; |
| KLF3 | Kruppel like factor 3; |
| GNG12 | G protein subunit gamma 12; |
| NRAS | NRAS proto-oncogene; |
| RAB9B | RAB9B, member RAS oncogene family; |
| SH3BGRL2 | SH3 domain binding glutamate rich protein like 2; |
| TNP1 | Transition protein 1; |
| RPL22 | Ribosomal protein L22; |
| MRPL22 | Mitochondrial ribosomal protein L22; |
| CBLN3 | Cerebellin 3 precursor; |
| PAIP2 | Poly(A) binding protein interacting protein 2; |
| SEC61B | SEC61 translocon subunit beta; |
| DBI | Diazepam binding inhibitor; |
| GNA11 | G protein subunit alpha 11; |
| ARHGAP1 | Rho GTPase activating protein 1. |
References
- Havel, J.J.; Chowell, D.; Chan, T.A. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat. Rev. Cancer 2019, 19, 133–150. [Google Scholar] [CrossRef]
- Sunshine, J.; Taube, J.M. PD-1/PD-L1 inhibitors. Curr. Opin. Pharmacol. 2015, 23, 32–38. [Google Scholar] [CrossRef]
- Smyth, M.J.; Ngiow, S.F.; Ribas, A.; Teng, M.W. Combination cancer immunotherapies tailored to the tumour microenvironment. Nat. Rev. Clin. Oncol. 2016, 13, 143–158. [Google Scholar] [CrossRef] [PubMed]
- Braun, D.A.; Burke, K.P.; Van Allen, E.M. Genomic Approaches to Understanding Response and Resistance to Immunotherapy. Clin. Cancer Res. 2016, 22, 5642–5650. [Google Scholar] [CrossRef]
- Balar, A.V.; Weber, J.S. PD-1 and PD-L1 antibodies in cancer: Current status and future directions. Cancer Immunol. Immunother. 2017, 66, 551–564. [Google Scholar] [CrossRef]
- Chen, D.S.; Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 2017, 541, 321–330. [Google Scholar] [CrossRef]
- Gasser, S.; Lim, L.H.K.; Cheung, F.S.G. The role of the tumour microenvironment in immunotherapy. Endocr. Relat. Cancer 2017, 24, T283–T295. [Google Scholar] [CrossRef]
- Chalmers, Z.R.; Connelly, C.F.; Fabrizio, D.; Gay, L.; Ali, S.M.; Ennis, R.; Schrock, A.; Campbell, B.; Shlien, A.; Chmielecki, J.; et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Rizvi, N.A.; Hellmann, M.D.; Snyder, A.; Kvistborg, P.; Makarov, V.; Havel, J.J.; Lee, W.; Yuan, J.; Wong, P.; Ho, T.S.; et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015, 348, 124–128. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, T.N.; Schreiber, R.D. Neoantigens in cancer immunotherapy. Science 2015, 348, 69–74. [Google Scholar] [CrossRef]
- Fridman, W.H.; Pages, F.; Sautes-Fridman, C.; Galon, J. The immune contexture in human tumours: Impact on clinical outcome. Nat. Rev. Cancer 2012, 12, 298–306. [Google Scholar] [CrossRef]
- Bremnes, R.M.; Al-Shibli, K.; Donnem, T.; Sirera, R.; Al-Saad, S.; Andersen, S.; Stenvold, H.; Camps, C.; Busund, L.T. The role of tumor-infiltrating immune cells and chronic inflammation at the tumor site on cancer development, progression, and prognosis: Emphasis on non-small cell lung cancer. J. Thorac. Oncol. 2011, 6, 824–833. [Google Scholar] [CrossRef]
- Gibney, G.T.; Weiner, L.M.; Atkins, M.B. Predictive biomarkers for checkpoint inhibitor-based immunotherapy. Lancet Oncol. 2016, 17, e542–e551. [Google Scholar] [CrossRef]
- Jin, Z.; Yoon, H.H. The promise of PD-1 inhibitors in gastro-esophageal cancers: Microsatellite instability vs. PD-L1. J. Gastrointest. Oncol. 2016, 7, 771–788. [Google Scholar] [CrossRef] [PubMed]
- Teng, M.W.; Ngiow, S.F.; Ribas, A.; Smyth, M.J. Classifying Cancers Based on T-cell Infiltration and PD-L1. Cancer Res. 2015, 75, 2139–2145. [Google Scholar] [CrossRef]
- Taube, J.M.; Anders, R.A.; Young, G.D.; Xu, H.; Sharma, R.; McMiller, T.L.; Chen, S.; Klein, A.P.; Pardoll, D.M.; Topalian, S.L.; et al. Colocalization of inflammatory response with B7-h1 expression in human melanocytic lesions supports an adaptive resistance mechanism of immune escape. Sci. Transl. Med. 2012, 4, 127ra37. [Google Scholar] [CrossRef]
- Sznol, M.; Chen, L. Antagonist antibodies to PD-1 and B7-H1 (PD-L1) in the treatment of advanced human cancer. Clin. Cancer Res. 2013, 19, 1021–1034. [Google Scholar] [CrossRef] [PubMed]
- Sanmamed, M.F.; Chen, L. A Paradigm Shift in Cancer Immunotherapy: From Enhancement to Normalization. Cell 2018, 175, 313–326. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, L. Classification of Advanced Human Cancers Based on Tumor Immunity in the MicroEnvironment (TIME) for Cancer Immunotherapy. JAMA Oncol. 2016, 2, 1403–1404. [Google Scholar] [CrossRef] [PubMed]
- Yagi, T.; Baba, Y.; Ishimoto, T.; Iwatsuki, M.; Miyamoto, Y.; Yoshida, N.; Watanabe, M.; Baba, H. PD-L1 Expression, Tumor-infiltrating Lymphocytes, and Clinical Outcome in Patients With Surgically Resected Esophageal Cancer. Ann. Surg. 2019, 269, 471–478. [Google Scholar] [CrossRef]
- Lin, Z.; Gu, J.; Cui, X.; Huang, L.; Li, S.; Feng, J.; Liu, B.; Zhou, Y. Deciphering Microenvironment of NSCLC based on CD8 + TIL Density and PD-1/PD-L1 Expression. J. Cancer 2019, 10, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Ricklefs, F.L.; Alayo, Q.; Krenzlin, H.; Mahmoud, A.B.; Speranza, M.C.; Nakashima, H.; Hayes, J.L.; Lee, K.; Balaj, L.; Passaro, C.; et al. Immune evasion mediated by PD-L1 on glioblastoma-derived extracellular vesicles. Sci. Adv. 2018, 4, eaar2766. [Google Scholar] [CrossRef] [PubMed]
- Poggio, M.; Hu, T.; Pai, C.C.; Chu, B.; Belair, C.D.; Chang, A.; Montabana, E.; Lang, U.E.; Fu, Q.; Fong, L.; et al. Suppression of Exosomal PD-L1 Induces Systemic Anti-tumor Immunity and Memory. Cell 2019, 177, 414–427.e13. [Google Scholar] [CrossRef] [PubMed]
- Ock, C.Y.; Keam, B.; Kim, S.; Lee, J.S.; Kim, M.; Kim, T.M.; Jeon, Y.K.; Kim, D.W.; Chung, D.H.; Heo, D.S. Pan-Cancer Immunogenomic Perspective on the Tumor Microenvironment Based on PD-L1 and CD8 T-Cell Infiltration. Clin. Cancer Res. 2016, 22, 2261–2270. [Google Scholar] [CrossRef]
- Chen, S.; Zhang, N.; Shao, J.; Wang, T.; Wang, X. Multi-omics Perspective on the Tumor Microenvironment based on PD-L1 and CD8 T-Cell Infiltration in Urothelial Cancer. J. Cancer 2019, 10, 697–707. [Google Scholar] [CrossRef]
- Kondou, R.; Iizuka, A.; Nonomura, C.; Miyata, H.; Ashizawa, T.; Nagashima, T.; Ohshima, K.; Urakami, K.; Kusuhara, M.; Yamaguchi, K.; et al. Classification of tumor microenvironment immune types based on immune response-associated gene expression. Int. J. Oncol. 2019, 54, 219–228. [Google Scholar] [CrossRef]
- Wang, Y.; Zhu, C.; Song, W.; Li, J.; Zhao, G.; Cao, H. PD-L1 Expression and CD8(+) T Cell Infiltration Predict a Favorable Prognosis in Advanced Gastric Cancer. J. Immunol. Res. 2018, 2018, 4180517. [Google Scholar] [CrossRef]
- Hugo, W.; Zaretsky, J.M.; Sun, L.; Song, C.; Moreno, B.H.; Hu-Lieskovan, S.; Berent-Maoz, B.; Pang, J.; Chmielowski, B.; Cherry, G.; et al. Genomic and Transcriptomic Features of Response to Anti-PD-1 Therapy in Metastatic Melanoma. Cell 2016, 165, 35–44. [Google Scholar] [CrossRef]
- Snyder, A.; Nathanson, T.; Funt, S.A.; Ahuja, A.; Buros Novik, J.; Hellmann, M.D.; Chang, E.; Aksoy, B.A.; Al-Ahmadie, H.; Yusko, E.; et al. Contribution of systemic and somatic factors to clinical response and resistance to PD-L1 blockade in urothelial cancer: An exploratory multi-omic analysis. PLoS Med. 2017, 14, e1002309. [Google Scholar] [CrossRef]
- Miao, D.; Margolis, C.A.; Gao, W.H.; Voss, M.H.; Li, W.; Martini, D.J.; Norton, C.; Bosse, D.; Wankowicz, S.M.; Cullen, D.; et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 2018, 359, 801–805. [Google Scholar] [CrossRef]
- Riaz, N.; Havel, J.J.; Makarov, V.; Desrichard, A.; Urba, W.J.; Sims, J.S.; Hodi, F.S.; Martin-Algarra, S.; Mandal, R.; Sharfman, W.H.; et al. Tumor and Microenvironment Evolution during Immunotherapy with Nivolumab. Cell 2017, 171, 934–949. [Google Scholar] [CrossRef] [PubMed]
- Mariathasan, S.; Turley, S.J.; Nickles, D.; Castiglioni, A.; Yuen, K.; Wang, Y.; Kadel, E.E., III; Koeppen, H.; Astarita, J.L.; Cubas, R.; et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 2018, 554, 544–548. [Google Scholar] [CrossRef]
- Dong, H.; Zhu, G.; Tamada, K.; Chen, L. B7-H1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion. Nat. Med. 1999, 5, 1365–1369. [Google Scholar] [CrossRef] [PubMed]
- Butte, M.J.; Keir, M.E.; Phamduy, T.B.; Sharpe, A.H.; Freeman, G.J. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity 2007, 27, 111–122. [Google Scholar] [CrossRef]
- Lee, J.S.; Ruppin, E. Multiomics Prediction of Response Rates to Therapies to Inhibit Programmed Cell Death 1 and Programmed Cell Death 1 Ligand 1. JAMA Oncol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Topalian, S.L.; Drake, C.G.; Pardoll, D.M. Immune checkpoint blockade: A common denominator approach to cancer therapy. Cancer Cell 2015, 27, 450–461. [Google Scholar] [CrossRef]
- Nowicki, T.S.; Akiyama, R.; Huang, R.R.; Shintaku, I.P.; Wang, X.; Tumeh, P.C.; Singh, A.; Chmielowski, B.; Denny, C.; Federman, N.; et al. Infiltration of CD8 T Cells and Expression of PD-1 and PD-L1 in Synovial Sarcoma. Cancer Immunol. Res. 2017, 5, 118–126. [Google Scholar] [CrossRef]
- Zhang, X.C.; Wang, J.; Shao, G.G.; Wang, Q.; Qu, X.; Wang, B.; Moy, C.; Fan, Y.; Albertyn, Z.; Huang, X.; et al. Comprehensive genomic and immunological characterization of Chinese non-small cell lung cancer patients. Nat. Commun. 2019, 10, 1772. [Google Scholar] [CrossRef]
- Binnewies, M.; Roberts, E.W.; Kersten, K.; Chan, V.; Fearon, D.F.; Merad, M.; Coussens, L.M.; Gabrilovich, D.I.; Ostrand-Rosenberg, S.; Hedrick, C.C.; et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 2018, 24, 541–550. [Google Scholar] [CrossRef]
- Li, B.; Li, T.; Pignon, J.C.; Wang, B.; Wang, J.; Shukla, S.A.; Dou, R.; Chen, Q.; Hodi, F.S.; Choueiri, T.K.; et al. Landscape of tumor-infiltrating T cell repertoire of human cancers. Nat. Genet. 2016, 48, 725–732. [Google Scholar] [CrossRef]
- Gentles, A.J.; Newman, A.M.; Liu, C.L.; Bratman, S.V.; Feng, W.; Kim, D.; Nair, V.S.; Xu, Y.; Khuong, A.; Hoang, C.D.; et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med. 2015, 21, 938–945. [Google Scholar] [CrossRef] [PubMed]
- Charoentong, P.; Finotello, F.; Angelova, M.; Mayer, C.; Efremova, M.; Rieder, D.; Hackl, H.; Trajanoski, Z. Pan-cancer Immunogenomic Analyses Reveal Genotype-Immunophenotype Relationships and Predictors of Response to Checkpoint Blockade. Cell Rep. 2017, 18, 248–262. [Google Scholar] [CrossRef]
- Baert, T.; Vankerckhoven, A.; Riva, M.; Van Hoylandt, A.; Thirion, G.; Holger, G.; Mathivet, T.; Vergote, I.; Coosemans, A. Myeloid Derived Suppressor Cells: Key Drivers of Immunosuppression in Ovarian Cancer. Front. Immunol. 2019, 10, 1273. [Google Scholar] [CrossRef]
- Samstein, R.M.; Lee, C.H.; Shoushtari, A.N.; Hellmann, M.D.; Shen, R.; Janjigian, Y.Y.; Barron, D.A.; Zehir, A.; Jordan, E.J.; Omuro, A.; et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 2019, 51, 202–206. [Google Scholar] [CrossRef]
- Yarchoan, M.; Hopkins, A.; Jaffee, E.M. Tumor Mutational Burden and Response Rate to PD-1 Inhibition. N. Engl. J. Med. 2017, 377, 2500–2501. [Google Scholar] [CrossRef]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef] [PubMed]
- Wasylecka-Juszczynska, M.M.; Gorniak, P.; Szydlowski, M.; Polak, A.; Juszczynski, P. BRAF V600E mutation in melanoma sustains IFN-gamma inducible PD-L1 expression by coactivating STAT1 and increasing protein translation. Ann. Oncol. 2018, 29, viii463–viii464. [Google Scholar] [CrossRef]
- Dudnik, E.; Peled, N.; Nechushtan, H.; Wollner, M.; Onn, A.; Agbarya, A.; Moskovitz, M.; Keren, S.; Popovits-Hadari, N.; Urban, D.; et al. BRAF Mutant Lung Cancer: Programmed Death Ligand 1 Expression, Tumor Mutational Burden, Microsatellite Instability Status, and Response to Immune Check-Point Inhibitors. J. Thorac. Oncol. 2018, 13, 1128–1137. [Google Scholar] [CrossRef]
- Feng, D.; Qin, B.; Pal, K.; Sun, L.; Dutta, S.; Dong, H.; Liu, X.; Mukhopadhyay, D.; Huang, S.; Sinicrope, F.A. BRAF(V600E)-induced, tumor intrinsic PD-L1 can regulate chemotherapy-induced apoptosis in human colon cancer cells and in tumor xenografts. Oncogene 2019, 38, 6752–6766. [Google Scholar] [CrossRef] [PubMed]
- Gibney, G.T.; Tang, S.J.; Poorman, K.; Olszanski, A.J.; Eisenberg, B.L.; Mehmi, I.; Farma, J.M.; In, G.K.; Amin, A.; Rapisuwon, S.; et al. Associations of age, PD-L1 status, BRAF mutation and tumor mutational burden (TMB) in advanced melanoma. J. Clin. Oncol. 2018, 36. [Google Scholar] [CrossRef]
- Aoki, K.; Taketo, M.M. Adenomatous polyposis coli (APC): A multi-functional tumor suppressor gene. J. Cell Sci. 2007, 120, 3327–3335. [Google Scholar] [CrossRef] [PubMed]
- Scheel, A.H.; Ansen, S.; Schultheis, A.M.; Scheffler, M.; Fischer, R.N.; Michels, S.; Hellmich, M.; George, J.; Zander, T.; Brockmann, M.; et al. PD-L1 expression in non-small cell lung cancer: Correlations with genetic alterations. Oncoimmunology 2016, 5, e1131379. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.Y.; Zhong, W.Z.; Zhang, X.C.; Su, J.; Xie, Z.; Liu, S.Y.; Tu, H.Y.; Chen, H.J.; Sun, Y.L.; Zhou, Q.; et al. Potential Predictive Value of TP53 and KRAS Mutation Status for Response to PD-1 Blockade Immunotherapy in Lung Adenocarcinoma. Clin. Cancer Res. 2017, 23, 3012–3024. [Google Scholar] [CrossRef] [PubMed]
- Riether, C.; Schurch, C.M.; Ochsenbein, A.F. Regulation of hematopoietic and leukemic stem cells by the immune system. Cell Death Differ. 2015, 22, 187–198. [Google Scholar] [CrossRef]
- Geiger, R.; Rieckmann, J.C.; Wolf, T.; Basso, C.; Feng, Y.; Fuhrer, T.; Kogadeeva, M.; Picotti, P.; Meissner, F.; Mann, M.; et al. L-Arginine Modulates T Cell Metabolism and Enhances Survival and Anti-tumor Activity. Cell 2016, 167, 829–842.e13. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.H.; Qiu, J.; O’Sullivan, D.; Buck, M.D.; Noguchi, T.; Curtis, J.D.; Chen, Q.; Gindin, M.; Gubin, M.M.; van der Windt, G.J.; et al. Metabolic Competition in the Tumor Microenvironment Is a Driver of Cancer Progression. Cell 2015, 162, 1229–1241. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Locasale, J.W.; Bielas, J.H.; O’Sullivan, J.; Sheahan, K.; Cantley, L.C.; Vander Heiden, M.G.; Vitkup, D. Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nat. Biotechnol. 2013, 31, 522–529. [Google Scholar] [CrossRef]
- Wang, G.Z.; Zhang, L.; Zhao, X.C.; Gao, S.H.; Qu, L.W.; Yu, H.; Fang, W.F.; Zhou, Y.C.; Liang, F.; Zhang, C.; et al. The Aryl hydrocarbon receptor mediates tobacco-induced PD-L1 expression and is associated with response to immunotherapy. Nat. Commun. 2019, 10, 1125. [Google Scholar] [CrossRef]
- Badur, M.G.; Metallo, C.M. Reverse engineering the cancer metabolic network using flux analysis to understand drivers of human disease. Metab. Eng. 2018, 45, 95–108. [Google Scholar] [CrossRef]
- O’Neill, L.A.; Pearce, E.J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 2016, 213, 15–23. [Google Scholar] [CrossRef]
- Motz, G.T.; Santoro, S.P.; Wang, L.P.; Garrabrant, T.; Lastra, R.R.; Hagemann, I.S.; Lal, P.; Feldman, M.D.; Benencia, F.; Coukos, G. Tumor endothelium FasL establishes a selective immune barrier promoting tolerance in tumors. Nat. Med. 2014, 20, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef]
- Mayakonda, A.; Lin, D.C.; Assenov, Y.; Plass, C.; Koeffler, H.P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018, 28, 1747–1756. [Google Scholar] [CrossRef]
- Mermel, C.H.; Schumacher, S.E.; Hill, B.; Meyerson, M.L.; Beroukhim, R.; Getz, G. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011, 12, R41. [Google Scholar] [CrossRef]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Ou Yang, T.H.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2018, 48, 812–830.e14. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- McCarthy, D.J.; Chen, Y.; Smyth, G.K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucl. Acids Res. 2012, 40, 4288–4297. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Veglia, F.; Sanseviero, E.; Gabrilovich, D.I. Myeloid-derived suppressor cells in the era of increasing myeloid cell diversity. Nat. Rev. Immunol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.Y.; Zhang, Y.Y.; Zheng, L.T.; Zheng, C.H.; Song, J.T.; Zhang, Q.M.; Kang, B.X.; Liu, Z.Z.R.; Jin, L.; Xing, R.; et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 2018, 24, 978–985. [Google Scholar] [CrossRef]
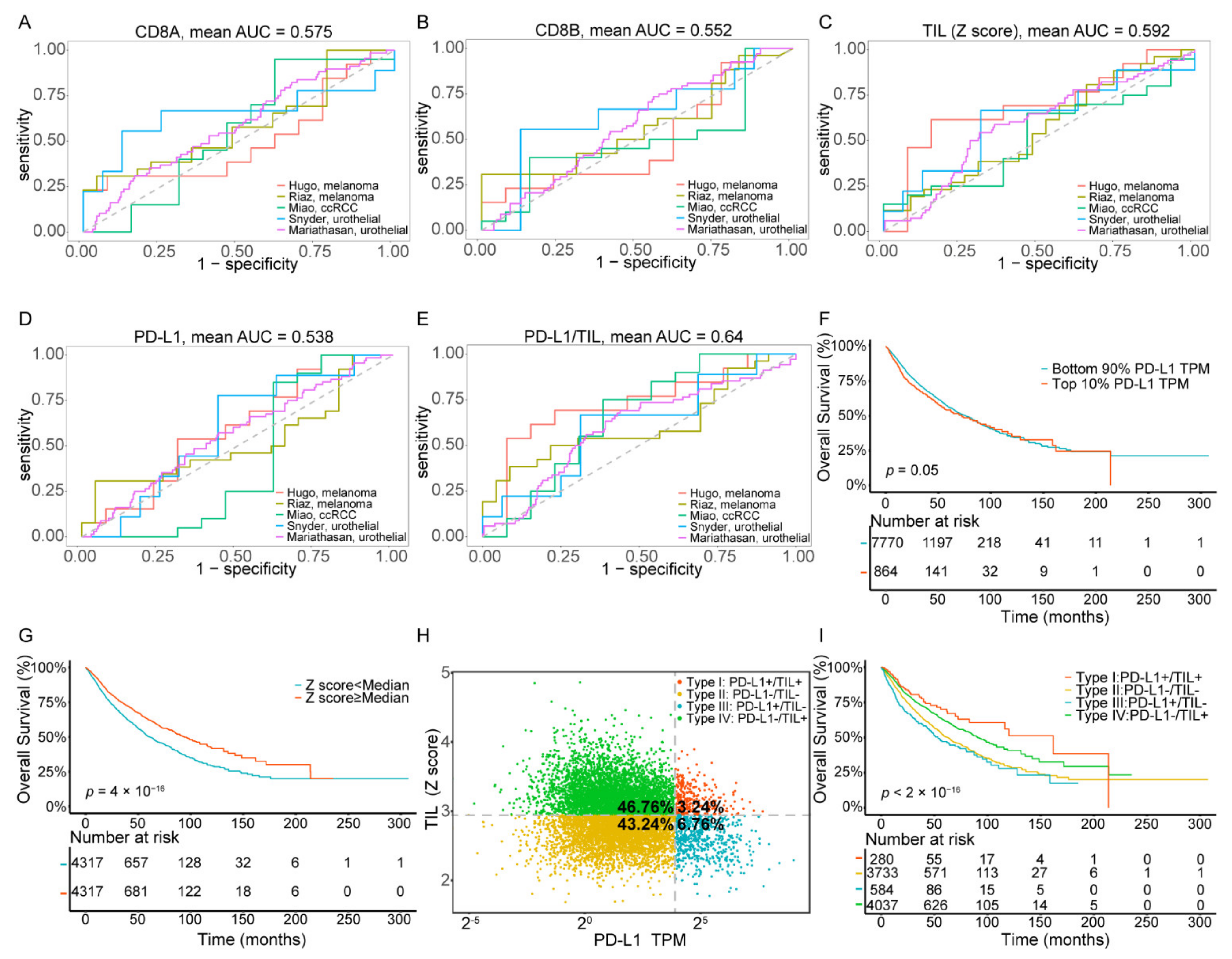

| Cohorts | Cancer Type | Drug | No. of Patients | No. of Responders | No. of Non-Responders | AUC Value | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| CD8A | CD8B | TIL (Z Score) | PD-L1 | PD-L1/TIL | ||||||
| Hugo [28] | melanoma | anti-PD-1 (pembrolizumab and nivolumab) | 26 | 13 | 13 | 0.503 | 0.497 | 0.686 | 0.598 | 0.722 |
| Riaz [31] | melanoma | anti-PD-1 (nivolumab) | 49 | 26 | 23 | 0.587 | 0.566 | 0.557 | 0.523 | 0.609 |
| Miao [30] | ccRCC | anti-PD-1 (nivolumab) | 33 | 20 | 13 | 0.554 | 0.488 | 0.515 | 0.415 | 0.658 |
| Snyder [29] | urothelial cancer | anti-PD-L1 (atezolizumab) | 25 | 9 | 16 | 0.646 | 0.632 | 0.611 | 0.59 | 0.611 |
| Mariat- Hasan [32] | urothelial cancer | anti-PD-L1 (atezolizumab) | 298 | 68 | 230 | 0.585 | 0.578 | 0.589 | 0.564 | 0.6 |
| Type I | Type II | Type III | Type IV | p Value | |
|---|---|---|---|---|---|
| No. | 280 | 3733 | 584 | 4037 | |
| Age | 56.22 ± 15.01 | 57.86 ± 14.87 | 61.84 ± 13.70 | 58.94 ± 13.72 | 9 × 10−11 |
| Gender | 0.0004998 | ||||
| Male | 133 (47.50%) | 1562 (41.84%) | 303 (51.88%) | 2077 (51.45%) | |
| Female | 147 (52.50%) | 2171 (58.16%) | 281 (48.12%) | 1960 (48.55%) | |
| Stage | 0.0004998 | ||||
| I | 40 (14.29%) | 630 (16.88%) | 157 (26.88%) | 933 (23.11%) | |
| II | 36 (12.86%) | 749 (20.06%) | 122 (20.89%) | 815 (20.19%) | |
| III | 41 (14.64%) | 489 (13.10%) | 107 (18.32%) | 649 (16.08%) | |
| IV | 34 (12.14%) | 231 (6.19%) | 61 (10.45%) | 334 (8.27%) | |
| T cells | 0.47 ± 0.18 | 0.28 ± 0.13 | 0.36 ± 0.14 | 0.36 ± 0.13 | <2.2 × 10−16 |
| B cells | 0.08 ± 0.08 | 0.09 ± 0.09 | 0.10 ± 0.09 | 0.09 ± 0.10 | 0.0086 |
| Macrophages | 0.31 ± 0.17 | 0.46 ± 0.17 | 0.41 ± 0.14 | 0.37 ± 0.15 | <2.2 × 10−16 |
| DC cells | 0.06 ± 0.06 | 0.04 ± 0.06 | 0.05 ±0.06 | 0.05 ± 0.06 | <2.2 × 10−16 |
| NK cells | 0.04 ± 0.04 | 0.04 ± 0.04 | 0.04 ± 0.03 | 0.05 ± 0.04 | <2.2 × 10−16 |
| Mast cells | 0.04 ± 0.04 | 0.08 ± 0.07 | 0.05 ± 0.04 | 0.07 ± 0.07 | <2.2 × 10−16 |
| Eosinophils | 0.00 ± 0.00 | 0.00 ± 0.02 | 0.00 ± 0.01 | 0.00 ± 0.01 | 4.2 × 10−11 |
| Neutrophils | 0.00 ± 0.01 | 0.01 ± 0.02 | 0.01 ± 0.02 | 0.01 ± 0.02 | 2.1 × 10−13 |
| TMB | 4.22 ± 13.22 | 6.76 ± 30.72 | 6.85 ± 13.61 | 3.65 ± 12.33 | 1.8 × 10−8 |
| Neoantigens | 333.62 ± 1972.69 | 353.96 ± 1625.59 | 313.25 ± 677.87 | 187.79 ± 619.51 | 1.4 × 10−5 |
| TP53-mut | 65 (23.21%) | 1409 (37.74%) | 286 (48.97%) | 1074 (26.60%) | <2.2 × 10−16 |
| BRAF-mut | 35 (12.50%) | 151 (4.05%) | 30 (5.14%) | 297 (7.36%) | <2.2 × 10−16 |
| HRAS-mut | 13 (4.64%) | 33 (0.88%) | 20 (3.42%) | 49 (1.21%) | 8.734 × 10−6 |
| IDH1-mut | 6 (2.14%) | 346 (9.27%) | 11 (1.88%) | 85 (2.11%) | <2.2 × 10−16 |
| POLE-mut | 4 (1.43%) | 120 (3.21%) | 27 (4.62%) | 92 (2.28%) | <2.2 × 10−16 |
| POLD1-mut | 5 (1.79%) | 65 (1.74%) | 6 (1.03%) | 38 (0.94%) | <2.2 × 10−16 |
| PDCD1LG2 CNA | <2.2 × 10−16 | ||||
| Amplification | 28 (10.00%) | 114 (3.05%) | 85 (14.55%) | 88 (2.18%) | |
| Deletion | 1 (0.36%) | 166 (4.45%) | 13 (2.23%) | 101 (2.50%) | |
| PD-L1 CNA | <2.2 × 10−16 | ||||
| Amplification | 28 (10.00%) | 114 (3.05%) | 84 (14.38%) | 87 (2.16%) | |
| Deletion | 1 (0.36%) | 166 (4.45%) | 13 (2.23%) | 100 (2.48%) | |
| PDCD1 CNA | 8.064 × 10−5 | ||||
| Amplification | 0 (0.00%) | 101 (2.71%) | 9 (1.54%) | 47 (1.16%) | |
| Deletion | 34 (12.14%) | 382 (10.23%) | 89 (15.24%) | 294 (7.28%) | |
| CTLA4 CNA | 0.001178 | ||||
| Amplification | 2 (0.71%) | 136 (3.64%) | 19 (3.25%) | 88 (2.18%) | |
| Deletion | 16 (5.71%) | 149 (3.99%) | 46 (7.88%) | 125 (3.10%) | |
| Immuno-activating cytokines | 2.81 ± 3.76 | 2.19 ± 3.49 | 4.52 ± 6.75 | 1.37 ± 2.37 | <2.2 × 10−16 |
| Immuno-suppressive cytokines | 39.38 ± 33.96 | 39.24 ± 39.31 | 50.47±29.03 | 34.76 ± 37.71 | <2.2 × 10−16 |
| Cytolytic activity | 34.68 ± 36.73 | 11.46 ± 19.96 | 47.71±72.69 | 12.15 ± 30.09 | <2.2 × 10−16 |
| Variable | Univariate Prognostic Analysis | Multivariate Prognostic Analysis | ||||
|---|---|---|---|---|---|---|
| HR | 95% CI | p-Value | HR | 95% CI | p-Value | |
| Age > 60 years (vs. < 60 years) | 1.87137 | 1.712–2.046 | < 2 × 10−16 | 1.848602 | 1.5988–2.1374 | <2 × 10−16 |
| Gender, male (vs. female) | 1.14972 | 1.054–1.255 | 0.002 | 1.159753 | 1.0093–1.3326 | 0.036577 |
| Stage II (vs. stage I) | 1.44218 | 1.219–1.706 | 1.89 × 10−5 | 1.326312 | 1.0929–1.6095 | 0.004236 |
| Stage III (vs. stage I) | 2.27638 | 1.934–2.679 | <2 × 10−16 | 1.873979 | 1.5455–2.2723 | 1.69e-10 |
| Stage IV (vs. stage I) | 4.66921 | 3.957–5.509 | <2 × 10−16 | 3.406277 | 2.7873–4.1627 | <2 × 10−16 |
| PD-L1 positive (vs. negative) | 1.1452 | 0.9999–1.312 | 0.0501 | ———— | ————— | ———— |
| TIL positive (vs. negative) | 0.69328 | 0.6345–0.7575 | 4 × 10−16 | 0.845795 | 0.7335–0.9752 | 0.021152 |
| CD8+T high (vs. low) | 0.7363 | 0.6744–0.8039 | 8.31 × 10−12 | 0.91313 | 0.7891–1.0567 | 0.222529 |
| CD4+T activated high (vs. low) | 1.1385 | 1.043–1.242 | 0.00355 | 1.071685 | 0.9069–1.2664 | 0.416427 |
| Treg high (vs. low) | 0.8552 | 0.7836–0.9333 | 0.000453 | 0.9463 | 0.8254–1.0849 | 0.428814 |
| Macro M2 high (vs. low) | 1.15472 | 1.058–1.26 | 0.00128 | 1.244084 | 1.0794–1.434 | 0.002581 |
| Mast activated high (vs. low) | 1.56816 | 1.422–1.73 | <2 × 10−16 | 1.241577 | 1.0436–1.4771 | 0.014614 |
| DC activated high (vs. low) | 1.18640 | 1.086–1.296 | 0.000148 | 1.028179 | 0.8939–1.1827 | 0.697222 |
| NK activated high (vs. low) | 0.81109 | 0.7432–0.8852 | 2.66 × 10−6 | 1.272744 | 1.0986–1.4744 | 0.001312 |
| B memory | 1.20193 | 1.087–1.329 | 0.000331 | 1.200665 | 1.0121–1.4244 | 0.035958 |
| TMB high (vs. low) | 1.71388 | 1.559–1.884 | <2 × 10−16 | 1.231722 | 0.9973–1.5213 | 0.053035 |
| Neoantigens high (vs. low) | 1.5202 | 1.361–1.698 | 1.01 × 10−13 | 1.029541 | 0.8496–1.2476 | 0.766486 |
| TP53 mutation (vs. wild type) | 1.72522 | 1.58–1.884 | <2 × 10−16 | 1.321964 | 1.1383–1.5353 | 0.000255 |
| BRAF mutation (vs. wild type) | 0.4703 | 0.3531–0.6263 | 2.44 × 10−7 | 0.772879 | 0.5504–1.0854 | 0.136993 |
| IDH1 mutation (vs. wild type) | 0.6939 | 0.5493–0.8765 | 0.00218 | 1.211248 | 0.5724–2.5631 | 0.616285 |
| POLE mutation (vs. wild type) | 0.96719 | 0.7449–1.256 | 0.802 | ———— | ————— | ———— |
| POLD1 mutation (vs. wild type) | 0.7212 | 0.4474–1.163 | 0.18 | ———— | —————— | ———— |
| PD-L1 amplification yes (vs. no) | 1.4735 | 1.208–1.797 | 0.000128 | 1.026873 | 0.764–1.3802 | 0.860481 |
| PDCD1 deletion yes (vs. no) | 1.24219 | 1.084–1.424 | 0.00182 | 0.839484 | 0.6544–1.077 | 0.168638 |
| CTLA4 deletion yes (vs. no) | 1.44534 | 1.193–1.752 | 0.000173 | 1.007358 | 0.7006–1.4484 | 0.968436 |
| Immuno-activating cytokines high (vs. low) | 1.33658 | 1.224–1.46 | 1.1 × 10−10 | 0.987575 | 0.8485–1.1494 | 0.871716 |
| Immuno-suppressive cytokines high (vs. low) | 1.69775 | 1.552–1.857 | <2 × 10−16 | 1.165356 | 1.0013–1.3563 | 0.048076 |
| Cytolytic activity high (vs. low) | 1.1153 | 1.022–1.217 | 0.0144 | 1.018329 | 0.8541–1.2141 | 0.839546 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, K.; Hu, M.; Chen, J.; Wei, J.; Qin, J.; Lin, S.; Du, H. Multi-Omics Perspective Reveals the Different Patterns of Tumor Immune Microenvironment Based on Programmed Death Ligand 1 (PD-L1) Expression and Predictor of Responses to Immune Checkpoint Blockade across Pan-Cancer. Int. J. Mol. Sci. 2021, 22, 5158. https://doi.org/10.3390/ijms22105158
Huang K, Hu M, Chen J, Wei J, Qin J, Lin S, Du H. Multi-Omics Perspective Reveals the Different Patterns of Tumor Immune Microenvironment Based on Programmed Death Ligand 1 (PD-L1) Expression and Predictor of Responses to Immune Checkpoint Blockade across Pan-Cancer. International Journal of Molecular Sciences. 2021; 22(10):5158. https://doi.org/10.3390/ijms22105158
Chicago/Turabian StyleHuang, Kaitang, Meiling Hu, Jiayun Chen, Jinfen Wei, Jingxin Qin, Shudai Lin, and Hongli Du. 2021. "Multi-Omics Perspective Reveals the Different Patterns of Tumor Immune Microenvironment Based on Programmed Death Ligand 1 (PD-L1) Expression and Predictor of Responses to Immune Checkpoint Blockade across Pan-Cancer" International Journal of Molecular Sciences 22, no. 10: 5158. https://doi.org/10.3390/ijms22105158
APA StyleHuang, K., Hu, M., Chen, J., Wei, J., Qin, J., Lin, S., & Du, H. (2021). Multi-Omics Perspective Reveals the Different Patterns of Tumor Immune Microenvironment Based on Programmed Death Ligand 1 (PD-L1) Expression and Predictor of Responses to Immune Checkpoint Blockade across Pan-Cancer. International Journal of Molecular Sciences, 22(10), 5158. https://doi.org/10.3390/ijms22105158






