Genome-Wide Identification and Expression Profiling of Wnt Family Genes in the Silkworm, Bombyx mori
Abstract
1. Introduction
2. Results
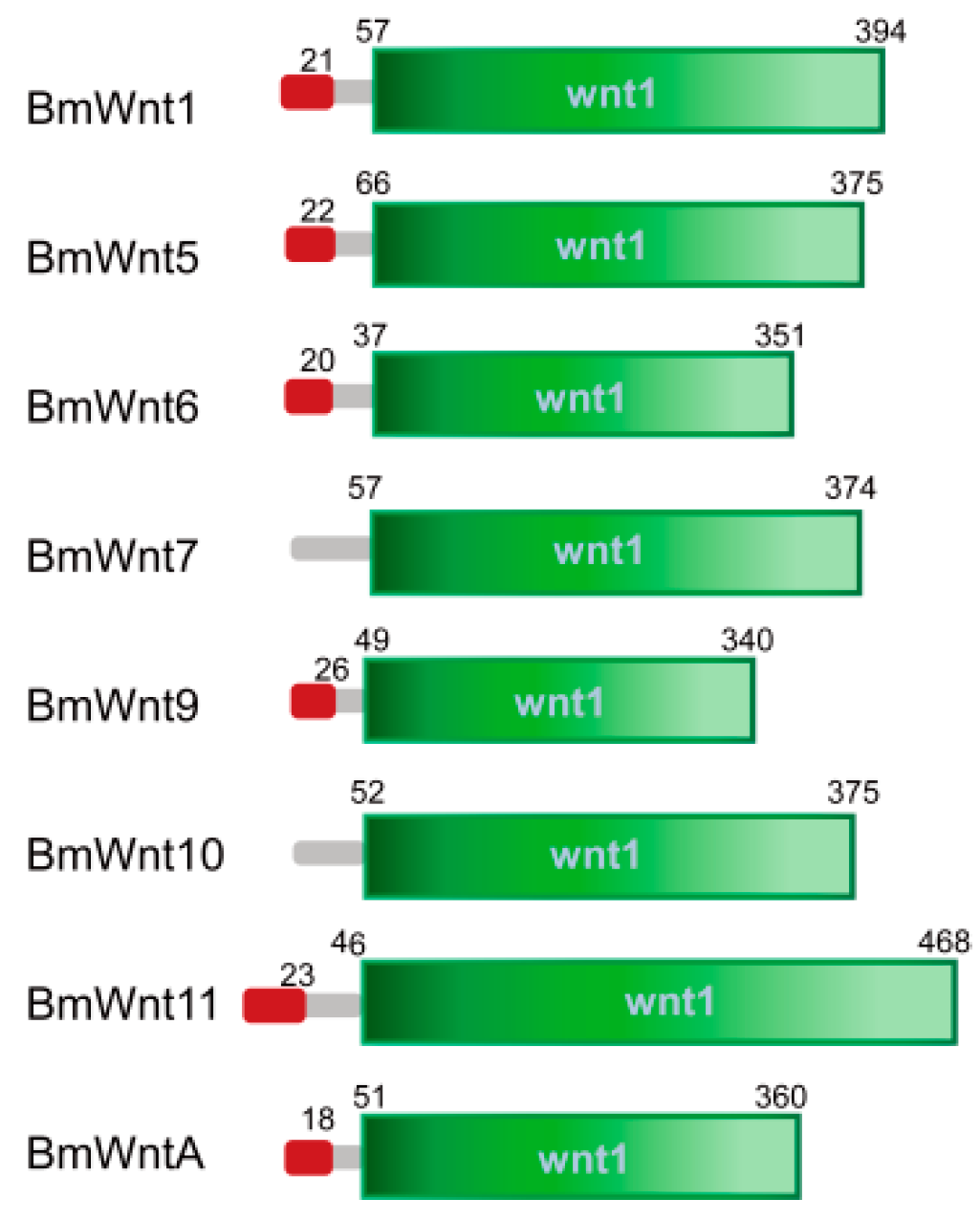
2.1. The Wnt Gene Repertoire in the Silkworm
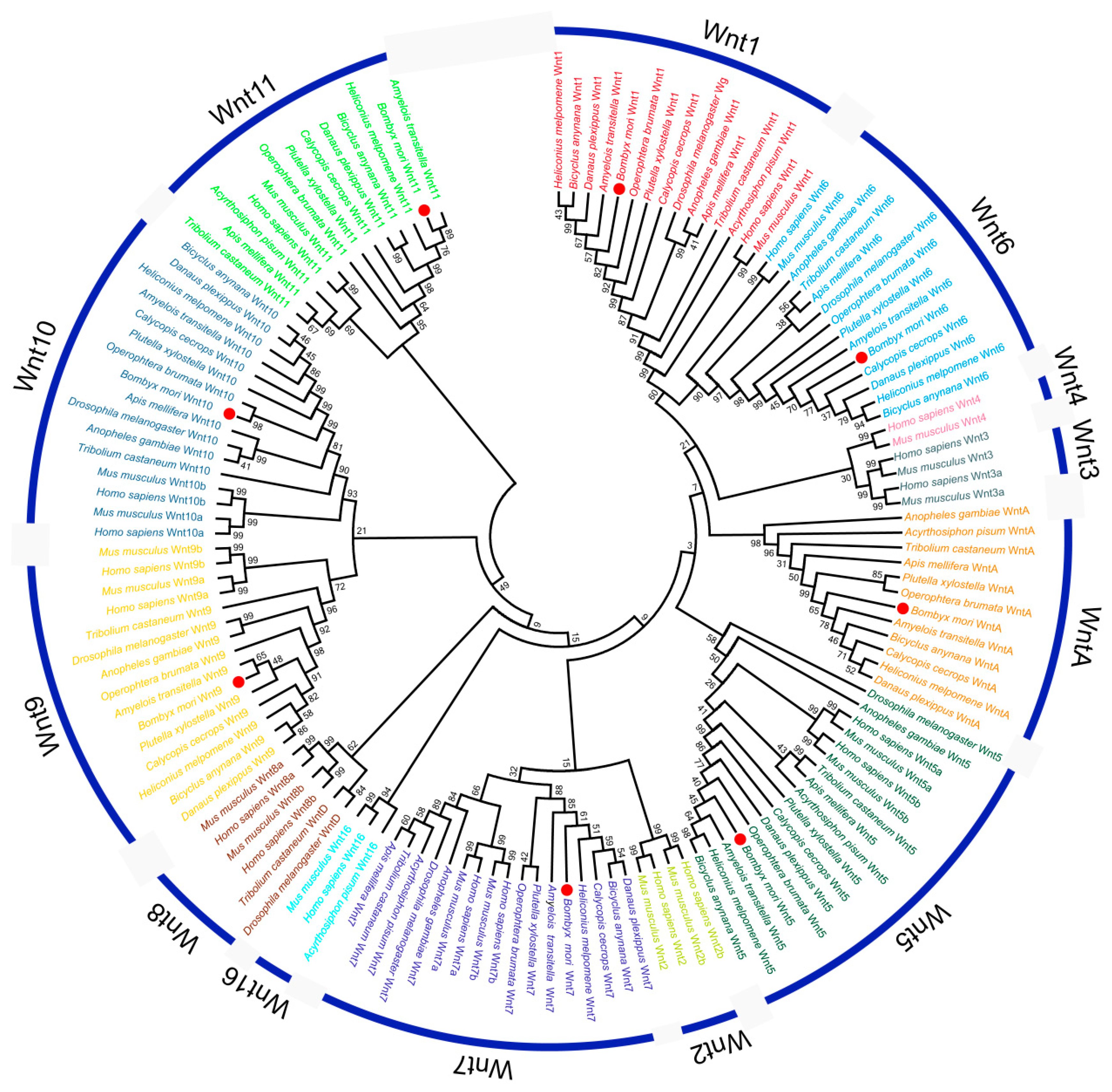
2.2. Phylogenetic Analysis of BmWnts
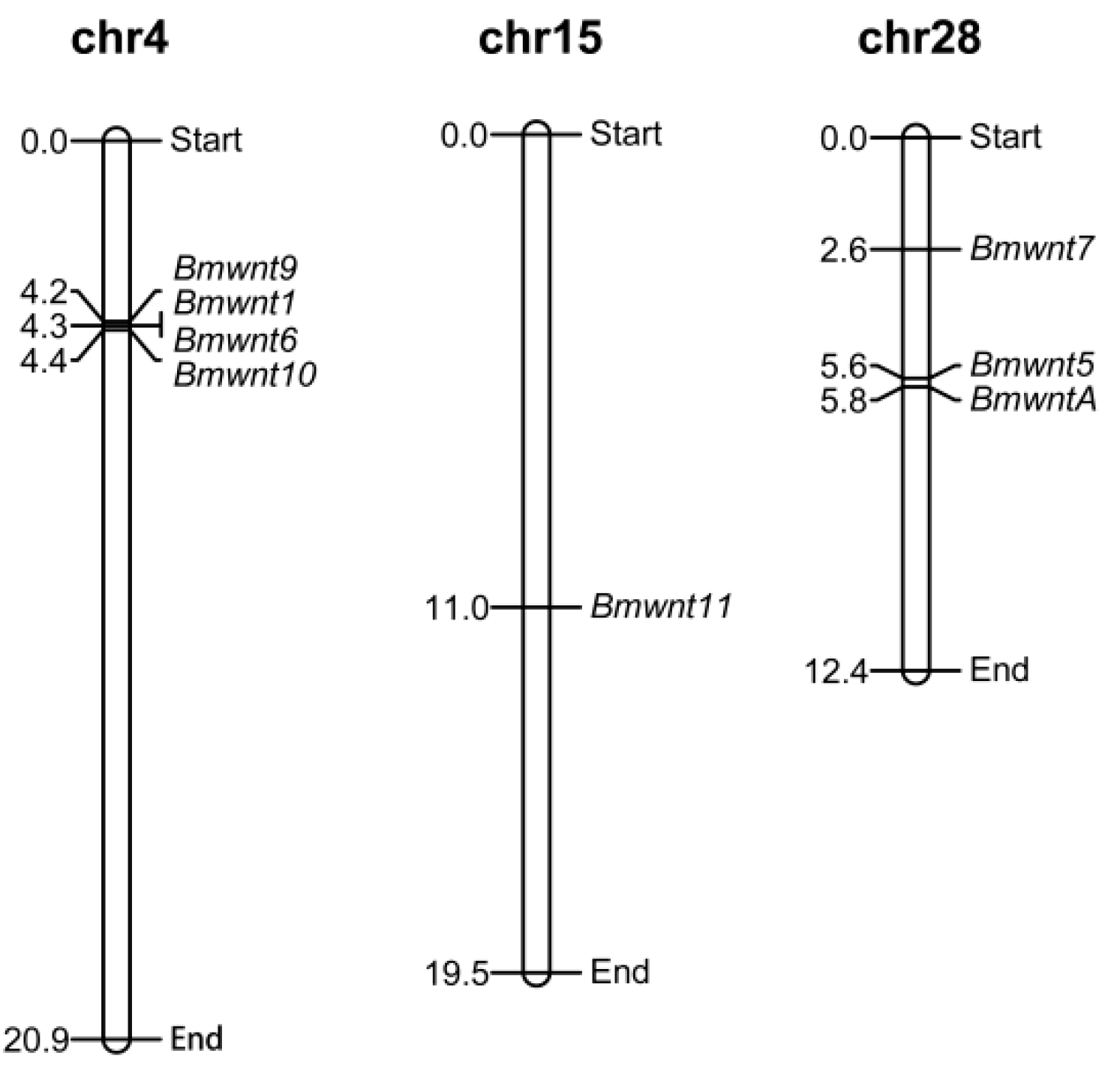
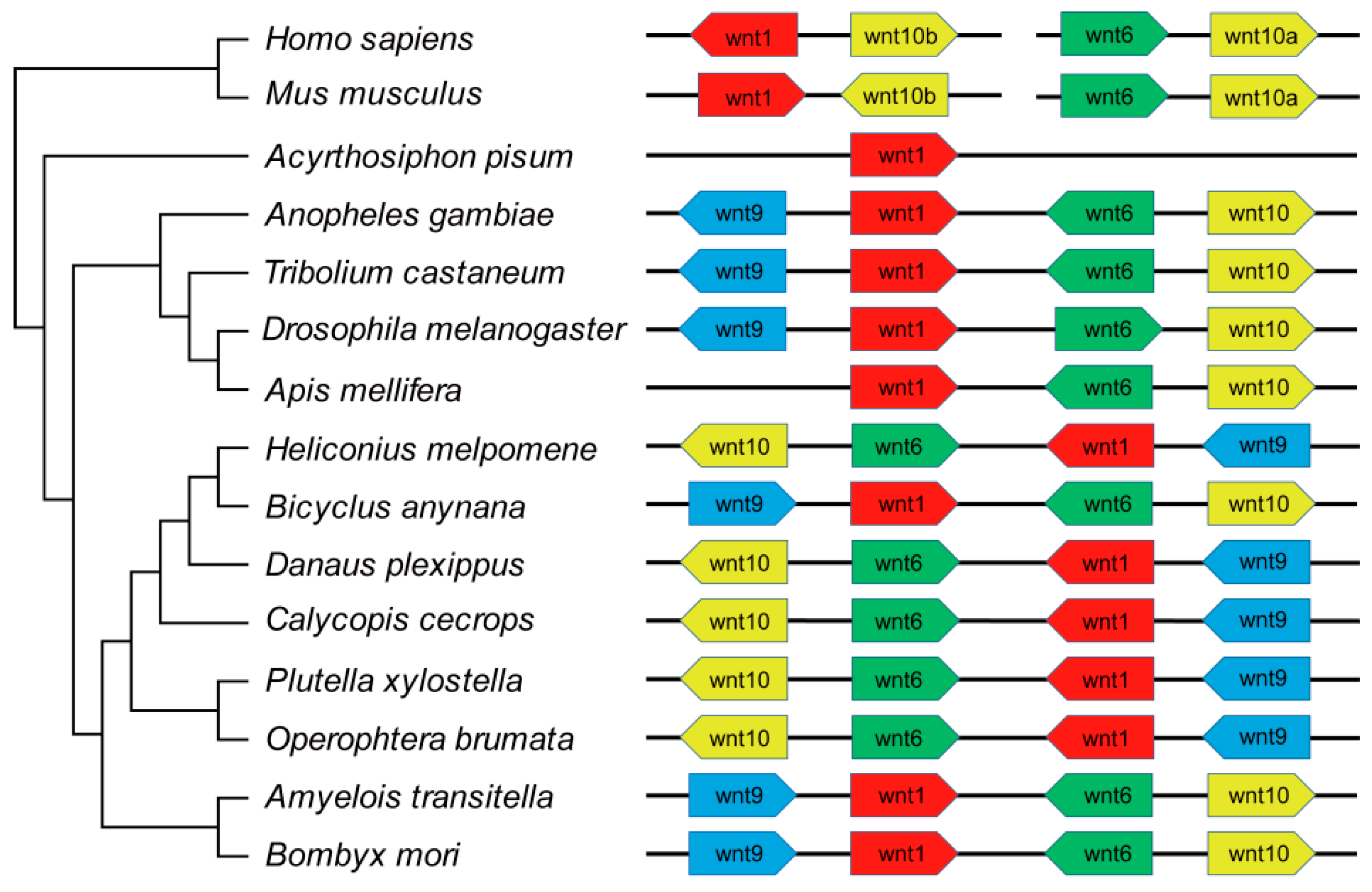
2.3. The Conserved Wnt Gene Cluster
2.4. Embryonic Expression Profile of the BmWnt Genes
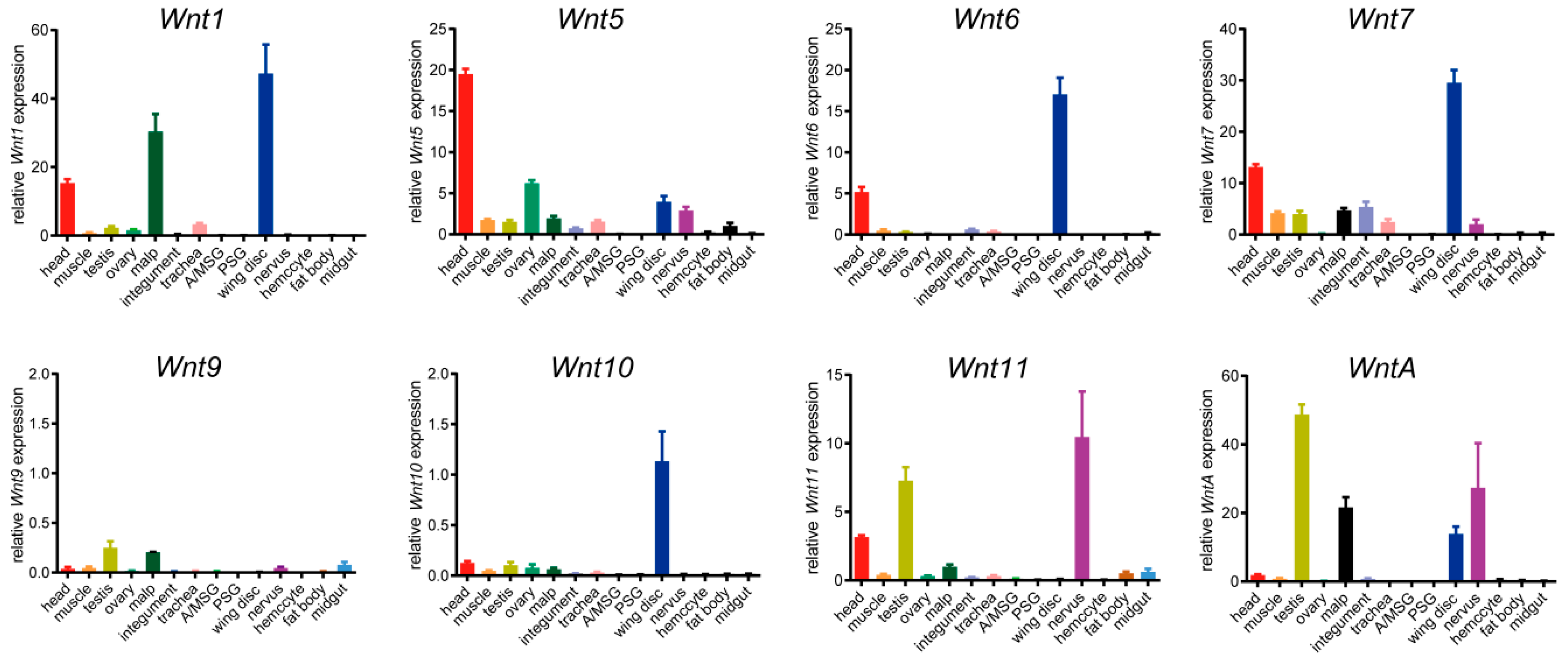
2.5. Different Tissue Expression Profiles of the BmWnt Genes
3. Discussion
4. Materials and Methods
4.1. Genome-Wide Identification of Wnt Proteins
4.2. Chromosomal Distribution and Protein Structure
4.3. Sequence Alignment and Phylogenetic Analysis
4.4. RNA Extraction and cDNA Synthesis
4.5. Real-Time Quantitative PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| PCP | planar cell polarity |
| Fzd | Frizzled |
| LEF1/TCF | T-cell factor/lymphoid enhancing factor |
| JNK | Jun N-terminal serine/threonine kinase |
| Rock | Rho-associated kinase |
| CamKI | Ca2+–calmodulin-dependent protein kinase II |
| PKC | protein kinase C |
| HMM | Hidden Markov Model |
| SMART | Simple Modular Architecture Research Tool |
| AEL | after egg laying |
References
- Nusse, R.; Varmus, H.E. Many tumors induced by the mouse mammary tumor virus contain a provirus integrated in the same region of the host genome. Cell 1982, 31, 99–109. [Google Scholar] [CrossRef]
- Rijsewijk, F.; Schuermann, M.; Wagenaar, E.; Parren, P.; Weigel, D.; Nusse, R. The Drosophila homolog of the mouse mammary oncogene int-1 is identical to the segment polarity gene wingless. Cell 1987, 50, 649–657. [Google Scholar] [CrossRef]
- Nusse, R.; Brown, A.; Papkoff, J.; Scambler, P.; Shackleford, G.; McMahon, A.; Moon, R.; Varmus, H. A new nomenclature for int-1 and related genes: The Wnt gene family. Cell 1991, 64, 231. [Google Scholar] [CrossRef]
- Loh, K.M.; van Amerongen, R.; Nusse, R. Generating Cellular Diversity and Spatial Form: Wnt Signaling and the Evolution of Multicellular Animals. Dev. Cell 2016, 38, 643–655. [Google Scholar] [CrossRef] [PubMed]
- Logan, C.Y.; Nusse, R. The Wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 2004, 20, 781–810. [Google Scholar] [CrossRef] [PubMed]
- Croce, J.C.; McClay, D.R. Evolution of the Wnt pathways. Methods Mol. Biol. 2008, 469, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Rao, T.P.; Kuhl, M. An updated overview on Wnt signaling pathways: A prelude for more. Circ. Res. 2010, 106, 1798–1806. [Google Scholar] [CrossRef] [PubMed]
- Van Amerongen, R.; Nusse, R. Towards an integrated view of Wnt signaling in development. Development 2009, 136, 3205–3214. [Google Scholar] [CrossRef] [PubMed]
- Semenov, M.V.; Habas, R.; Macdonald, B.T.; He, X. SnapShot: Noncanonical Wnt Signaling Pathways. Cell 2007, 131, 1378. [Google Scholar] [CrossRef] [PubMed]
- Rigo-Watermeier, T.; Kraft, B.; Ritthaler, M.; Wallkamm, V.; Holstein, T.; Wedlich, D. Functional conservation of Nematostella Wnts in canonical and noncanonical Wnt-signaling. Biol. Open 2012, 1, 43–51. [Google Scholar] [CrossRef] [PubMed]
- Andre, P.; Song, H.; Kim, W.; Kispert, A.; Yang, Y. Wnt5a and Wnt11 regulate mammalian anterior-posterior axis elongation. Development 2015, 142, 1516–1527. [Google Scholar] [CrossRef] [PubMed]
- Kuhl, M.; Sheldahl, L.C.; Park, M.; Miller, J.R.; Moon, R.T. The Wnt/Ca2+ pathway: A new vertebrate Wnt signaling pathway takes shape. Trends Genet. TIG 2000, 16, 279–283. [Google Scholar] [CrossRef]
- De, A. Wnt/Ca2+ signaling pathway: A brief overview. Acta Biochimica et Biophysica Sinica 2011, 43, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Adamska, M.; Larroux, C.; Adamski, M.; Green, K.; Lovas, E.; Koop, D.; Richards, G.S.; Zwafink, C.; Degnan, B.M. Structure and expression of conserved Wnt pathway components in the demosponge Amphimedon queenslandica. Evol. Dev. 2010, 12, 494–518. [Google Scholar] [CrossRef] [PubMed]
- Leininger, S.; Adamski, M.; Bergum, B.; Guder, C.; Liu, J.; Laplante, M.; Brate, J.; Hoffmann, F.; Fortunato, S.; Jordal, S.; et al. Developmental gene expression provides clues to relationships between sponge and eumetazoan body plans. Nat. Commun. 2014, 5, 3905. [Google Scholar] [CrossRef] [PubMed]
- Kusserow, A.; Pang, K.; Sturm, C.; Hrouda, M.; Lentfer, J.; Schmidt, H.A.; Technau, U.; von Haeseler, A.; Hobmayer, B.; Martindale, M.Q.; et al. Unexpected complexity of the Wnt gene family in a sea anemone. Nature 2005, 433, 156–160. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.N.; Pang, K.; Matus, D.Q.; Martindale, M.Q. A WNT of things to come: Evolution of Wnt signaling and polarity in cnidarians. Semin. Cell Dev. Biol. 2006, 17, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Lengfeld, T.; Watanabe, H.; Simakov, O.; Lindgens, D.; Gee, L.; Law, L.; Schmidt, H.A.; Ozbek, S.; Bode, H.; Holstein, T.W. Multiple Wnts are involved in Hydra organizer formation and regeneration. Dev. Biol. 2009, 330, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Prud’homme, B.; Lartillot, N.; Balavoine, G.; Adoutte, A.; Vervoort, M. Phylogenetic analysis of the Wnt gene family. Insights from lophotrochozoan members. Curr. Biol. CB 2002, 12, 1395. [Google Scholar] [CrossRef]
- Cho, S.J.; Valles, Y.; Giani, V.C., Jr.; Seaver, E.C.; Weisblat, D.A. Evolutionary dynamics of the wnt gene family: A lophotrochozoan perspective. Mol. Biol. Evol. 2010, 27, 1645–1658. [Google Scholar] [CrossRef] [PubMed]
- Garriock, R.J.; Warkman, A.S.; Meadows, S.M.; D’Agostino, S.; Krieg, P.A. Census of vertebrate Wnt genes: Isolation and developmental expression of Xenopus Wnt2, Wnt3, Wnt9a, Wnt9b, Wnt10a, and Wnt16. Dev. Dyn. Off. Pub. Am. Assoc. Anat. 2007, 236, 1249–1258. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, R.; Beermann, A.; Farzana, L.; Wittkopp, N.; Lutz, R.; Balavoine, G.; Brown, S.J.; Schroder, R. Tribolium Wnts: Evidence for a larger repertoire in insects with overlapping expression patterns that suggest multiple redundant functions in embryogenesis. Dev. Genes Evol. 2008, 218, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Murat, S.; Hopfen, C.; McGregor, A.P. The function and evolution of Wnt genes in arthropods. Arthropod. Struct. Dev. 2010, 39, 446–452. [Google Scholar] [CrossRef] [PubMed]
- Shigenobu, S.; Bickel, R.D.; Brisson, J.A.; Butts, T.; Chang, C.C.; Christiaens, O.; Davis, G.K.; Duncan, E.J.; Ferrier, D.E.; Iga, M.; et al. Comprehensive survey of developmental genes in the pea aphid, Acyrthosiphon pisum: Frequent lineage-specific duplications and losses of developmental genes. Insect Mol. Biol. 2010, 19 (Suppl. 2), 47–62. [Google Scholar] [CrossRef]
- Dearden, P.K.; Wilson, M.J.; Sablan, L.; Osborne, P.W.; Havler, M.; McNaughton, E.; Kimura, K.; Milshina, N.V.; Hasselmann, M.; Gempe, T.; et al. Patterns of conservation and change in honey bee developmental genes. Genome Res. 2006, 16, 1376–1384. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, J.; Banno, Y.; Mita, K.; Yamamoto, K.; Ando, T.; Fujiwara, H. Periodic Wnt1 expression in response to ecdysteroid generates twin-spot markings on caterpillars. Nat. Commun. 2013, 4, 1857. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Aslam, A.F.; Liu, X.; Li, M.; Huang, Y.; Tan, A. Functional analysis of Bombyx Wnt1 during embryogenesis using the CRISPR/Cas9 system. J. Insect Physiol. 2015, 79, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Edayoshi, M.; Yamaguchi, J.; Fujiwara, H. Protruding structures on caterpillars are controlled by ectopic wnt1 expression. PLoS ONE 2015, 10, e0121736. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed]
- Berntsson, R.P.; Smits, S.H.; Schmitt, L.; Slotboom, D.J.; Poolman, B. A structural classification of substrate-binding proteins. FEBS Lett. 2010, 584, 2606–2617. [Google Scholar] [CrossRef] [PubMed]
- Wiese, K.E.; Nusse, R.; van Amerongen, R. Wnt signalling: Conquering complexity. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed]
- Holstein, T.W. The evolution of the Wnt pathway. Cold Spring Harb. Perspect. Biol. 2012, 4, a007922. [Google Scholar] [CrossRef] [PubMed]
- Janssen, R.; Le Gouar, M.; Pechmann, M.; Poulin, F.; Bolognesi, R.; Schwager, E.E.; Hopfen, C.; Colbourne, J.K.; Budd, G.E.; Brown, S.J.; et al. Conservation, loss, and redeployment of Wnt ligands in protostomes: Implications for understanding the evolution of segment formation. BMC Evol. Biol 2010, 10, 374. [Google Scholar] [CrossRef] [PubMed]
- Pace, R.M.; Eskridge, P.C.; Grbic, M.; Nagy, L.M. Evidence for the plasticity of arthropod signal transduction pathways. Dev. Genes Evol. 2014, 224, 209–222. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.E. Molecular cloning of sequences from wingless, a segment polarity gene in Drosophila: The spatial distribution of a transcript in embryos. EMBO J. 1987, 6, 1765–1773. [Google Scholar] [CrossRef] [PubMed]
- Baker, N.E. Embryonic and imaginal requirements for wingless, a segment-polarity gene in Drosophila. Dev. Biol. 1988, 125, 96–108. [Google Scholar] [CrossRef]
- Carroll, L.M.N.S. Conservation of wingless patterning functions in the short-germ embryos of Tribolium castaneum. Nature 1994, 367, 460–463. [Google Scholar]
- Dhawan, S.; Gopinathan, K.P. Spatio-temporal expression of wnt-1 during embryonic-, wing- and silkgland development in Bombyx mori. Gene Expr. Patterns GEP 2003, 3, 559–570. [Google Scholar] [CrossRef]
- Buratovich, M.A.; Anderson, S.; Gieseler, K.; Pradel, J.; Wilder, E.L. DWnt-4 and Wingless have distinct activities in the Drosophila dorsal epidermis. Dev. Genes Evol. 2000, 210, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Gieseler, K.; Graba, Y.; Mariol, M.C.; Wilder, E.L.; Martinez-Arias, A.; Lemaire, P.; Pradel, J. Antagonist activity of DWnt-4 and wingless in the Drosophila embryonic ventral ectoderm and in heterologous Xenopus assays. Mech. Dev. 1999, 85, 123–131. [Google Scholar] [CrossRef]
- Russell, J.; Gennissen, A.; Nusse, R. Isolation and expression of two novel Wnt/wingless gene homologues in Drosophila. Development 1992, 115, 475–485. [Google Scholar] [PubMed]
- Llimargas, M.; Lawrence, P.A. Seven Wnt homologues in Drosophila: A case study of the developing tracheae. Proc. Natl. Acad. Sci. USA 2001, 98, 14487–14492. [Google Scholar] [CrossRef] [PubMed]
- Fradkin, L.G.; Noordermeer, J.N.; Nusse, R. The Drosophila Wnt protein DWnt-3 is a secreted glycoprotein localized on the axon tracts of the embryonic CNS. Dev. Biol. 1995, 168, 202–213. [Google Scholar] [CrossRef] [PubMed]
- Wouda, R.R.; Bansraj, M.R.; de Jong, A.W.; Noordermeer, J.N.; Fradkin, L.G. Src family kinases are required for WNT5 signaling through the Derailed/RYK receptor in the Drosophila embryonic central nervous system. Development 2008, 135, 2277–2287. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, S.; McKinnon, R.D.; Kokel, M.; Thomas, J.B. Wnt-mediated axon guidance via the Drosophila Derailed receptor. Nature 2003, 422, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Inaki, M.; Yoshikawa, S.; Thomas, J.B.; Aburatani, H.; Nose, A. Wnt4 is a local repulsive cue that determines synaptic target specificity. Curr. Biol. CB 2007, 17, 1574–1579. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Zhang, X.; Yuan, J.; Zhang, X.; Li, F.; Xiang, J. Wnt gene family members and their expression profiling in Litopenaeus vannamei. Fish Shellfish Immunol. 2018, 77, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Doumpas, N.; Jekely, G.; Teleman, A.A. Wnt6 is required for maxillary palp formation in Drosophila. BMC Biol. 2013, 11, 104. [Google Scholar] [CrossRef] [PubMed]
- Koshikawa, S.; Giorgianni, M.W.; Vaccaro, K.; Kassner, V.A.; Yoder, J.H.; Werner, T.; Carroll, S.B. Gain of cis-regulatory activities underlies novel domains of wingless gene expression in Drosophila. Proc. Natl. Acad. Sci. USA 2015, 112, 7524–7529. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.; Reed, R.D. Wnt signaling underlies evolution and development of the butterfly wing pattern symmetry systems. Dev. Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Kozopas, K.M.; Samos, C.H.; Nusse, R. DWnt-2, a Drosophila Wnt gene required for the development of the male reproductive tract, specifies a sexually dimorphic cell fate. Genes Dev. 1998, 12, 1155–1165. [Google Scholar] [CrossRef] [PubMed]
- DeFalco, T.; Camara, N.; Le Bras, S.; Van Doren, M. Nonautonomous sex determination controls sexually dimorphic development of the Drosophila gonad. Dev. Cell 2008, 14, 275–286. [Google Scholar] [CrossRef] [PubMed]
- Cohen, E.D.; Mariol, M.C.; Wallace, R.M.; Weyers, J.; Kamberov, Y.G.; Pradel, J.; Wilder, E.L. DWnt4 regulates cell movement and focal adhesion kinase during Drosophila ovarian morphogenesis. Dev. Cell 2002, 2, 437–448. [Google Scholar] [CrossRef]
- Duan, J.; Li, R.; Cheng, D.; Fan, W.; Zha, X.; Cheng, T.; Wu, Y.; Wang, J.; Mita, K.; Xiang, Z.; et al. SilkDB v2.0: A platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res. 2010, 38, D453–D456. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, G.; Schroeder, A.J.; Goodman, J.L.; Strelets, V.B.; Crosby, M.A.; Thurmond, J.; Emmert, D.B.; Gelbart, W.M.; FlyBase, C. FlyBase: Introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res. 2015, 43, D690–D697. [Google Scholar] [CrossRef] [PubMed]
- Zhan, S.; Merlin, C.; Boore, J.L.; Reppert, S.M. The monarch butterfly genome yields insights into long-distance migration. Cell 2011, 147, 1171–1185. [Google Scholar] [CrossRef] [PubMed]
- Heliconius Genome, C. Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 2012, 487, 94–98. [Google Scholar] [CrossRef]
- Giraldo-Calderon, G.I.; Emrich, S.J.; MacCallum, R.M.; Maslen, G.; Dialynas, E.; Topalis, P.; Ho, N.; Gesing, S.; VectorBase, C.; Madey, G.; et al. VectorBase: An updated bioinformatics resource for invertebrate vectors and other organisms related with human diseases. Nucleic Acids Res. 2015, 43, D707–D713. [Google Scholar] [CrossRef] [PubMed]
- Legeai, F.; Shigenobu, S.; Gauthier, J.P.; Colbourne, J.; Rispe, C.; Collin, O.; Richards, S.; Wilson, A.C.; Murphy, T.; Tagu, D. AphidBase: A centralized bioinformatic resource for annotation of the pea aphid genome. Insect Mol. Biol. 2010, 19 (Suppl. 2), 5–12. [Google Scholar] [CrossRef]
- Wang, L.; Wang, S.; Li, Y.; Paradesi, M.S.; Brown, S.J. BeetleBase: The model organism database for Tribolium castaneum. Nucleic Acids Res. 2007, 35, D476–D479. [Google Scholar] [CrossRef] [PubMed]
- Jouraku, A.; Yamamoto, K.; Kuwazaki, S.; Urio, M.; Suetsugu, Y.; Narukawa, J.; Miyamoto, K.; Kurita, K.; Kanamori, H.; Katayose, Y.; et al. KONAGAbase: A genomic and transcriptomic database for the diamondback moth, Plutella xylostella. BMC Genomics 2013, 14, 464. [Google Scholar] [CrossRef] [PubMed]
- Elsik, C.G.; Tayal, A.; Diesh, C.M.; Unni, D.R.; Emery, M.L.; Nguyen, H.N.; Hagen, D.E. Hymenoptera Genome Database: Integrating genome annotations in HymenopteraMine. Nucleic Acids Res. 2016, 44, D793–D800. [Google Scholar] [CrossRef] [PubMed]
- Challi, R.J.; Kumar, S.; Dasmahapatra, K.K.; Jiggins, C.D.; Blaxter, M. Lepbase: The Lepidopteran genome database. bioRxiv 2016. Available online: https://www.biorxiv.org/content/10.1101/056994v1.abstract (accessed on 6 June 2016). [CrossRef]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


| GeneName | GeneID | Scaff_ID | Probe ID |
|---|---|---|---|
| BGIBMGA006146 | BmWnt1 | nscaf2847: 4290086..4300264: + | sw07406 |
| BGIBMGA013783 | BmWnt5 | nscaf3097: 574395..598709: + | sw10651 |
| BGIBMGA006004 | BmWnt6 | nscaf2847: 4311541..4316713: − | sw10550 |
| BGIBMGA013981 | BmWnt7 | nscaf3099: 2430055..2453230: + | none |
| BGIBMGA006145 | BmWnt9 | nscaf2847: 4227852..4228750: + | sw17884 |
| BGIBMGA006147 BGIBMGA006148 | BmWnt10 | nscaf2847: 4406381..4410443: + | sw21982 |
| BGIBMGA007580 | BmWnt11 | nscaf2887: 11667287..1177617: + | sw15093 |
| BGIBMGA013787 | BmWntA | nscaf3097: 771657..781653: + | sw13710 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ding, X.; Liu, J.; Zheng, L.; Song, J.; Li, N.; Hu, H.; Tong, X.; Dai, F. Genome-Wide Identification and Expression Profiling of Wnt Family Genes in the Silkworm, Bombyx mori. Int. J. Mol. Sci. 2019, 20, 1221. https://doi.org/10.3390/ijms20051221
Ding X, Liu J, Zheng L, Song J, Li N, Hu H, Tong X, Dai F. Genome-Wide Identification and Expression Profiling of Wnt Family Genes in the Silkworm, Bombyx mori. International Journal of Molecular Sciences. 2019; 20(5):1221. https://doi.org/10.3390/ijms20051221
Chicago/Turabian StyleDing, Xin, Junxia Liu, Lu Zheng, Jiangbo Song, Niannian Li, Hai Hu, Xiaoling Tong, and Fangyin Dai. 2019. "Genome-Wide Identification and Expression Profiling of Wnt Family Genes in the Silkworm, Bombyx mori" International Journal of Molecular Sciences 20, no. 5: 1221. https://doi.org/10.3390/ijms20051221
APA StyleDing, X., Liu, J., Zheng, L., Song, J., Li, N., Hu, H., Tong, X., & Dai, F. (2019). Genome-Wide Identification and Expression Profiling of Wnt Family Genes in the Silkworm, Bombyx mori. International Journal of Molecular Sciences, 20(5), 1221. https://doi.org/10.3390/ijms20051221





