Ectopic Expression of a R2R3-MYB Transcription Factor Gene LjaMYB12 from Lonicera japonica Increases Flavonoid Accumulation in Arabidopsis thaliana
Abstract
1. Introduction
2. Results
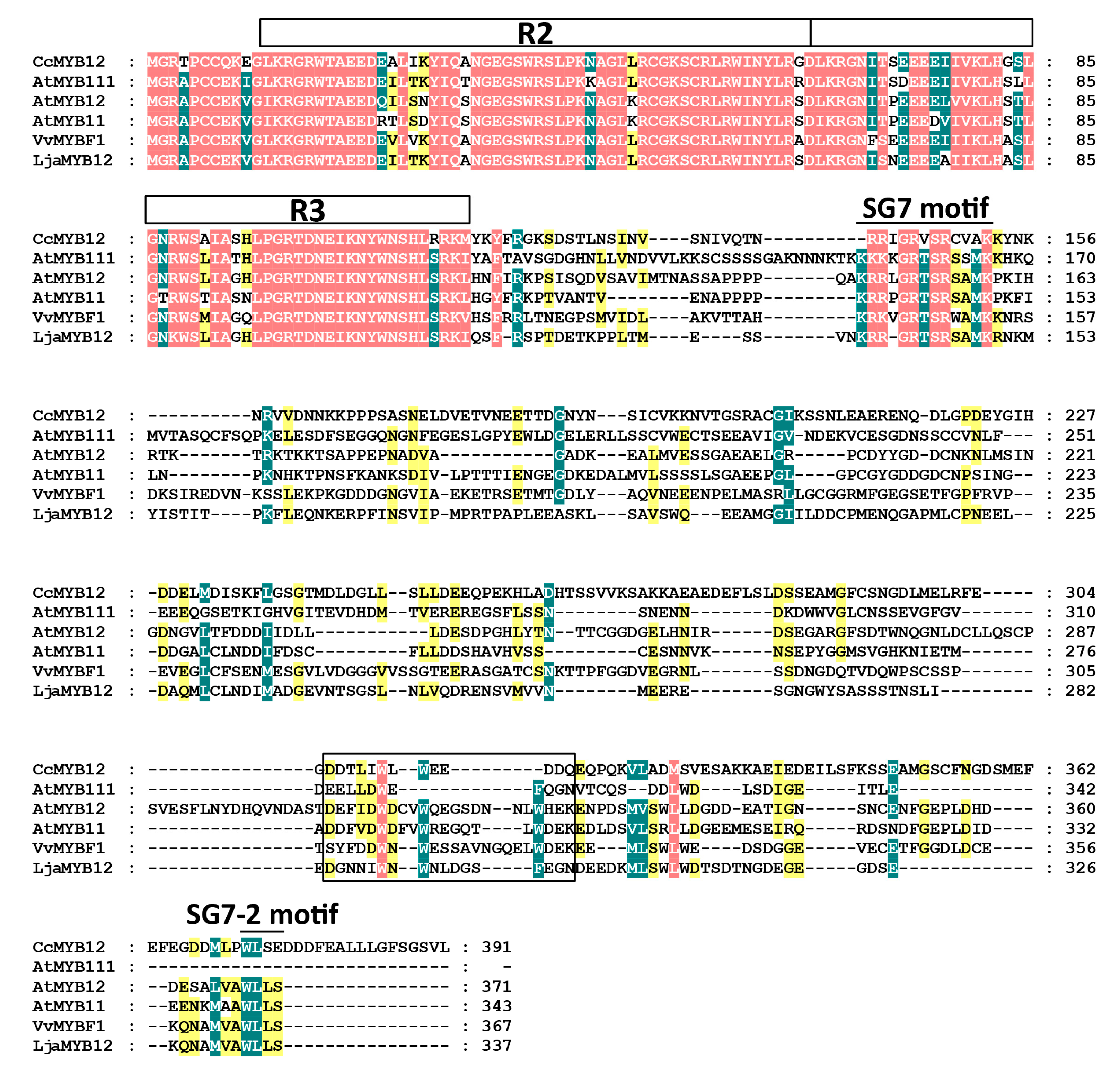
2.1. LjaMYB12 Is a R2R3-MYB Transcription Factor in L. japonica
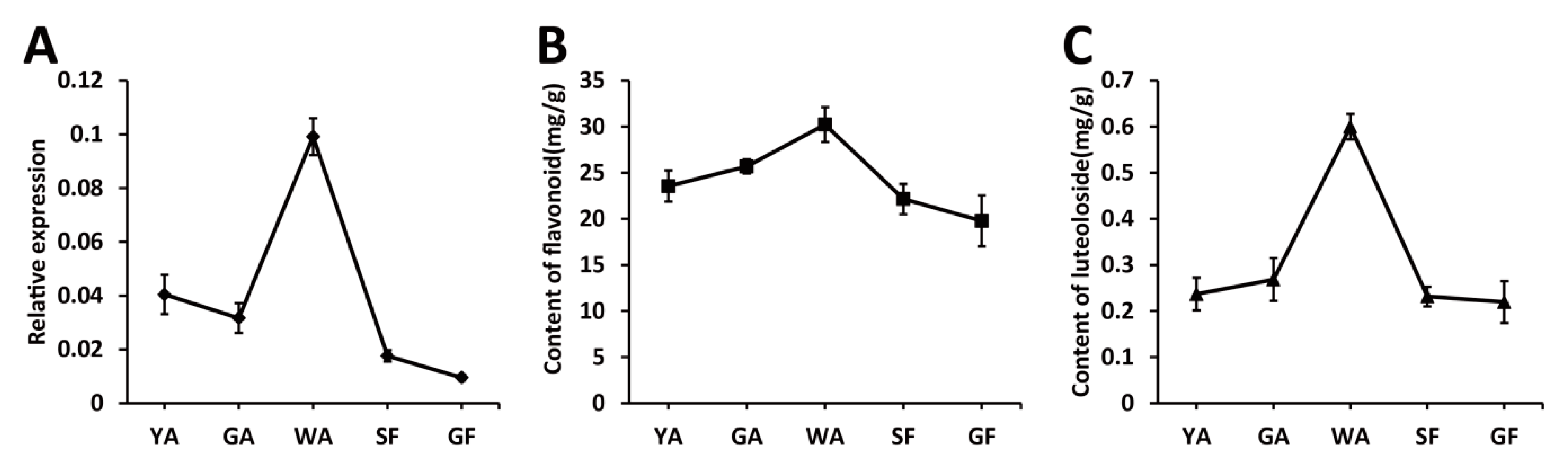
2.2. The Transcriptional Level of LjaMYB12 Is Proportional to the Total Flavonoid Content during the Development of L. Japonica Flowers
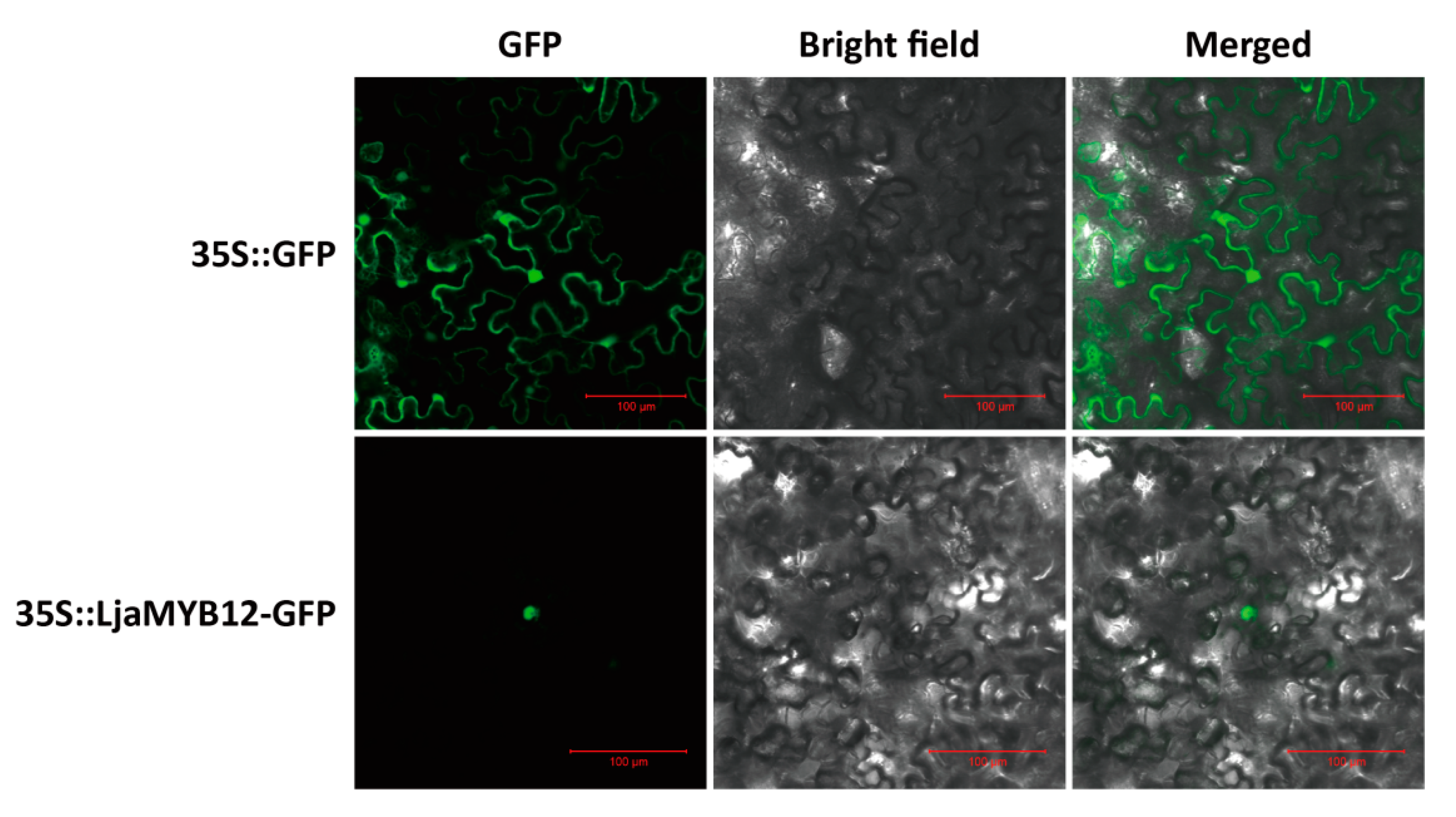
2.3. LjaMYB12 Localizes to the Nucleus
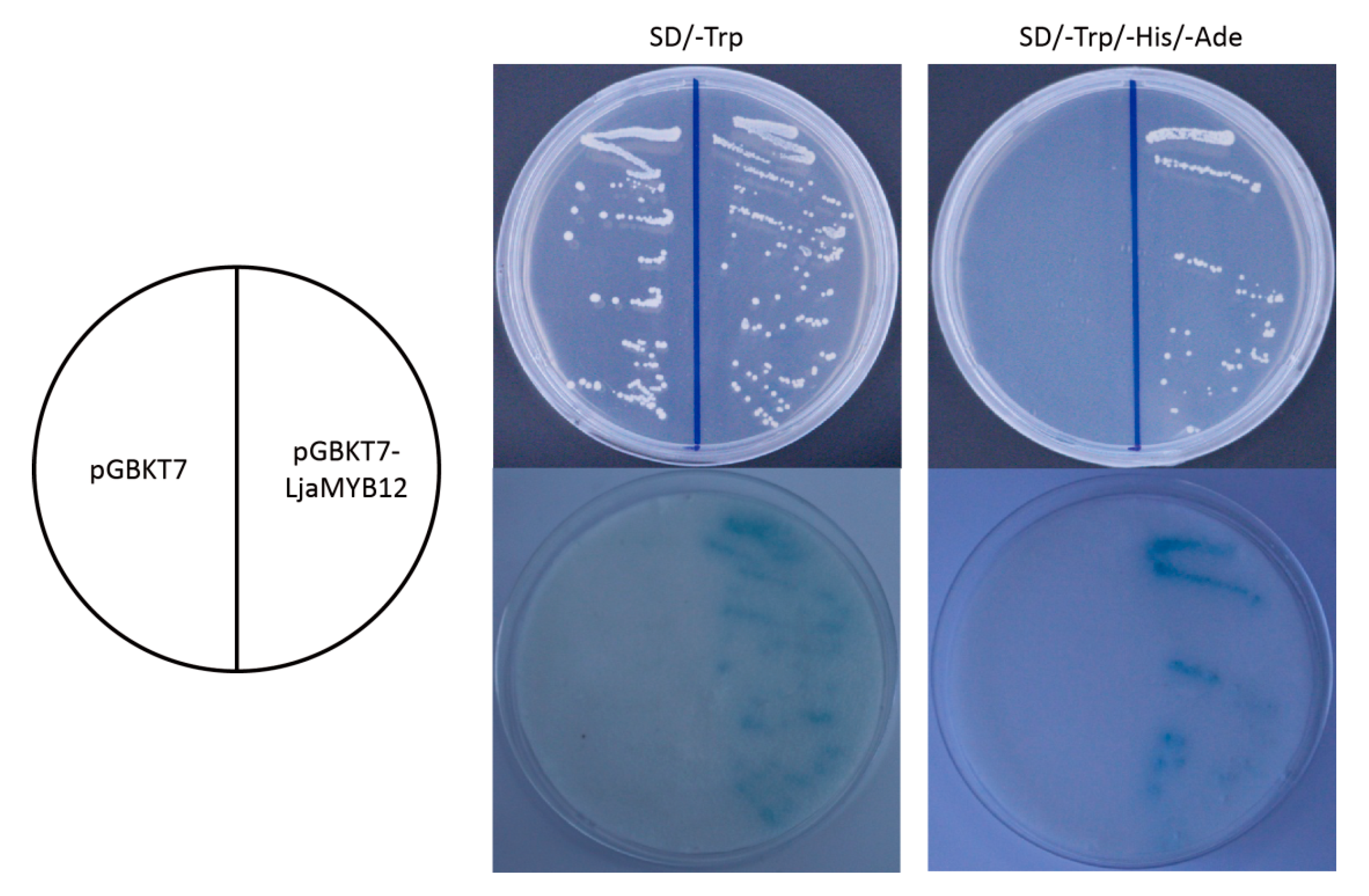
2.4. LjaMYB12 Has Transactivation Activity
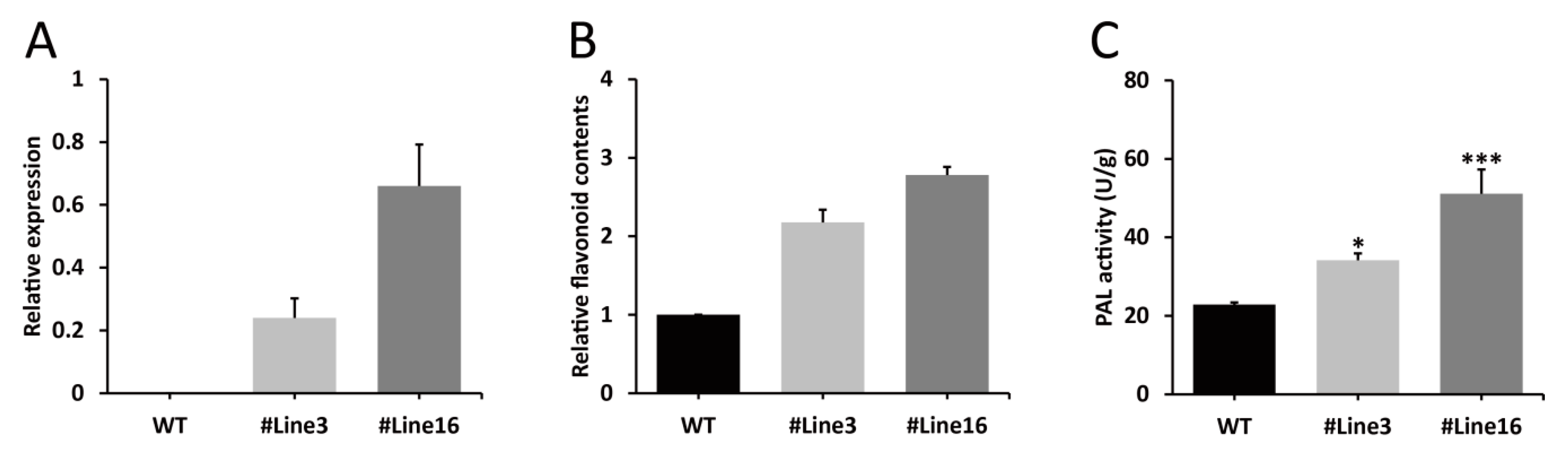
2.5. LjaMYB12 Overexpression Increases PAL Activity and Flavonoid Content in Transgenic Arabidopsis
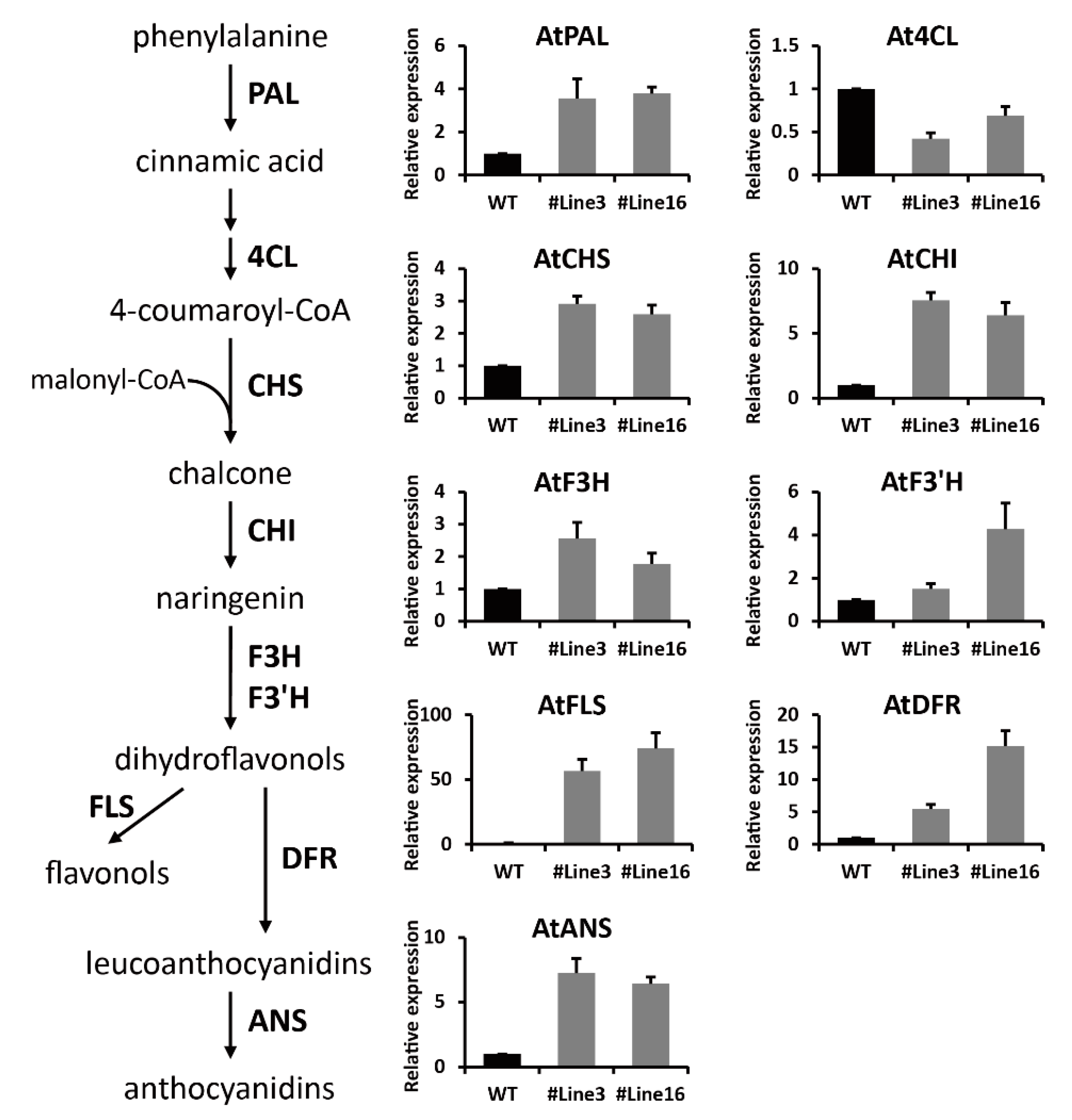
2.6. LjaMYB12 Promotes Transcription of a Range of Flavonoid Biosynthetic Genes in Transgenic Arabidopsis
3. Discussion
4. Materials and Methods
4.1. RNA Isolation and cDNA Synthesis of L. japonica
4.2. Isolation and Characterization of LjaMYB12
4.3. Subcellular Localization Assay
4.4. Transactivation Activity of LjaMYB12 in Yeast
4.5. Arabidopsis Transformation
4.6. PAL Activity Assay
4.7. Quantification of Flavonoid and Luteoloside
4.8. Quantitative Real-Time Reverse Transcriptional PCR (qRT-PCR)
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| bHLH | basic Helix-Loop-Helix |
| WDR | WD Repeats |
| qRT-PCR | quantitative real-time Reverse Transcriptional PCR |
| HPLC | High Performance Liquid Chromatography |
| ORF | Open Reading Frame |
| GFP | Green Fluorescent Protein |
| PAL | Phenylalanine Ammonia-Lyase |
| 4CL | 4-Coumarate:CoA Ligase |
| C4H | Cinnamate 4-Hydroxylase |
| CHS | Chalcone Synthase |
| CHI | Chalcone Isomerase |
| F3H | Flavanone 3-Hydroxylase |
| F3′H | Flavonoid 3′-Hydroxylase |
| FLS | Flavonol Synthase |
| DFR | Dihydroflavonol 4-Reductase |
| ANS | Anthocyanidin Synthase |
| RT | Flavonol-3-glucosiderhamnosyltransferase |
| GT | Flavonol-3-glucosyltransferase; |
| C3H | p-Coumaroyl ester 3-Hydroxylase |
| HCT | Cinnamoyl CoA shikimate/quinate Transferase |
| HQT | Hydroxycinnamoyl CoA Quinate Transferase. |
References
- Shang, X.; Pan, H.; Li, M.; Miao, X.; Ding, H. Lonicera japonica thunb.: Ethnopharmacology, phytochemistry and pharmacology of an important traditional chinese medicine. J. Ethnopharmacol. 2011, 138, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Kim, J.S.; Kim, H.P.; Lee, J.H.; Kang, S.S. Phenolic constituents from the flower buds of Lonicera japonica and their 5-lipoxygenase inhibitory activities. Food Chem. 2010, 120, 134–139. [Google Scholar] [CrossRef]
- Yoo, H.J.; Kang, H.J.; Song, Y.S.; Park, E.H.; Lim, C.J. Anti-angiogenic, antinociceptive and anti-inflammatory activities of Lonicera japonica extract. J. Pharm. Pharmacol. 2008, 60, 779–786. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.C.; Du, J.; But, P.P.; Deng, X.L.; Zhang, Y.W.; Ooi, V.E.; Xu, H.X.; Lee, S.H.; Lee, S.F. Antiviral Chinese medicinal herbs against respiratory syncytial virus. J. Ethnopharmacol. 2002, 79, 205–211. [Google Scholar] [CrossRef]
- Leung, H.W.; Hour, M.J.; Chang, W.T.; Wu, Y.C.; Lai, M.Y.; Wang, M.Y.; Lee, H.Z. P38-associated pathway involvement in apoptosis induced by photodynamic therapy with Lonicera japonica in human lung squamous carcinoma CH27 cells. Food Chem. Toxicol. 2008, 46, 3389–3400. [Google Scholar] [CrossRef] [PubMed]
- Schlotzhauer, W.S.; Pair, S.D.; Horvat, R.J. Volatile constituents from the flowers of Japanese honeysuckle (Lonicera japonica). J. Agr. Food Chem. 1996, 44, 206–209. [Google Scholar] [CrossRef]
- Chai, X.Y.; Li, S.L.; Li, P. Quality evaluation of Flos Lonicerae through a simultaneous determination of seven saponins by HPLC with ELSD. J. Chromatogr. A 2005, 1070, 43–48. [Google Scholar] [CrossRef]
- Chen, J.; Li, S.L.; Li, P.; Song, Y.; Chai, X.Y.; Ma, D.Y. Qualitative and quantitative analysis of active flavonoids in Flos Lonicerae by capillary zone electrophoresis coupled with solid-phase extraction. J. Sep. Sci. 2005, 28, 365–372. [Google Scholar] [CrossRef]
- Lu, H.T.; Jiang, Y.; Chen, F. Application of preparative high-speed counter-current chromatography for separation of chlorogenic acid from Flos Lonicerae. J. Chromatogr. A 2004, 1026, 185–190. [Google Scholar] [CrossRef]
- Zhu, Y.P. The Pharmacopoeia of the People’s Republic of China, 2010; China Medical Science Press: Beijing, China, 2010. [Google Scholar]
- Pourcel, L.; Routaboul, J.M.; Cheynier, V.; Lepiniec, L.; Debeaujon, I. Flavonoid oxidation in plants: From biochemical properties to physiological functions. Trends Plant Sci. 2007, 12, 29–36. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Lepiniec, L.; Debeaujon, I.; Routaboul, J.M.; Baudry, A.; Pourcel, L.; Nesi, N.; Caboche, M. Genetics and biochemistry of seed flavonoids. Annu. Rev. Plant Biol. 2006, 57, 405–430. [Google Scholar] [CrossRef] [PubMed]
- Saito, K.; Yonekura-Sakakibara, K.; Nakabayashi, R.; Higashi, Y.; Yamazaki, M.; Tohge, T.; Fernie, A.R. The flavonoid biosynthetic pathway in Arabidopsis: Structural and genetic diversity. Plant Physiol. Biochem. 2013, 72, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Broun, P. Transcriptional control of flavonoid biosynthesis: A complex network of conserved regulators involved in multiple aspects of differentiation in Arabidopsis. Curr. Opin. Plant Biol. 2005, 8, 272–279. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Gao, L.; Wang, H.; Chen, X.; Wang, Y.; Yang, H.; Wei, C.; Wan, X.; Xia, T. The R2R3-MYB, bHLH, WD40, and related transcription factors in flavonoid biosynthesis. Funct. Integr. Genomics 2013, 13, 75–98. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Q.; Fang, X.; Wu, X.M.; Mao, Y.B.; Wang, L.J.; Chen, X.Y. Transcriptional regulation of plant secondary metabolism. J. Integr. Plant Biol. 2012, 54, 703–712. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Osbourn, A.; Ma, P. MYB transcription factors as regulators of phenylpropanoid metabolism in plants. Mol. Plant 2015, 8, 689–708. [Google Scholar] [CrossRef] [PubMed]
- Klempnauer, K.H.; Gonda, T.J.; Bishop, J.M. Nucleotide sequence of the retroviral leukemia gene v-myb and its cellular progenitor c-myb: The architecture of a transduced oncogene. Cell 1982, 31, 453–463. [Google Scholar] [CrossRef]
- Rosinski, J.A.; Atchley, W.R. Molecular evolution of the Myb family of transcription factors: Evidence for polyphyletic origin. J. Mol. Evol. 1998, 46, 74–83. [Google Scholar] [CrossRef]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Gu, J.; Chopra, S.; Gu, X.; Peterson, T. Ordered origin of the typical two- and three-repeat Myb genes. Gene 2004, 326, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, O.; Nahal, H.; Foong, J.; Provart, N.J.; Campbell, M.M. Expansion and diversification of the Populus R2R3–MYB family of transcription factors. Plant Physiol. 2009, 149, 981–993. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Zhao, M.; Leavitt, J.M.; Lloyd, A.M. Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J. 2008, 53, 814–827. [Google Scholar] [CrossRef] [PubMed]
- Nesi, N.; Jond, C.; Debeaujon, I.; Caboche, M.; Lepiniec, L. The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 2001, 13, 2099–2114. [Google Scholar] [CrossRef]
- Mehrtens, F.; Kranz, H.; Bednarek, P.; Weisshaar, B. The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol. 2005, 138, 1083–1096. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Ishihara, H.; Huep, G.; Barsch, A.; Mehrtens, F.; Niehaus, K.; Weisshaar, B. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J. 2007, 50, 660–677. [Google Scholar] [CrossRef] [PubMed]
- Blanco, E.; Sabetta, W.; Danzi, D.; Negro, D.; Passeri, V.; De Lisi, A.; Paolocci, F.; Sonnante, G. Isolation and characterization of the flavonol regulator CcMYB12 from the globe artichoke [Cynara cardunculus var. scolymus (L.) Fiori]. Front. Plant Sci. 2018, 9, 941. [Google Scholar] [CrossRef]
- Luo, J.; Butelli, E.; Hill, L.; Parr, A.; Niggeweg, R.; Bailey, P.; Weisshaar, B.; Martin, C. AtMYB12 regulates caffeoyl quinic acid and flavonol synthesis in tomato: Expression in fruit results in very high levels of both types of polyphenol. Plant J. 2008, 56, 316–326. [Google Scholar] [CrossRef]
- Pandey, A.; Misra, P.; Khan, M.P.; Swarnkar, G.; Tewari, M.C.; Bhambhani, S.; Trivedi, R.; Chattopadhyay, N.; Trivedi, P.K. Co-expression of Arabidopsis transcription factor, AtMYB12, and soybean isoflavone synthase, GmIFS1, genes in tobacco leads to enhanced biosynthesis of isoflavones and flavonols resulting in osteoprotective activity. Plant Biotechnol. J. 2014, 12, 69–80. [Google Scholar] [CrossRef]
- Li, Y.; Kim, J.I.; Pysh, L.; Chapple, C. Four isoforms of Arabidopsis 4-coumarate: CoA ligase have overlapping yet distinct roles in phenylpropanoid metabolism. Plant Physiol. 2015, 169, 2409–2421. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Lee, C.; Zhou, J.; McCarthy, R.L.; Ye, Z.H. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 2008, 20, 2763–2782. [Google Scholar] [CrossRef] [PubMed]
- Schwinn, K.; Venail, J.; Shang, Y.; Mackay, S.; Alm, V.; Butelli, E.; Oyama, R.; Bailey, P.; Davies, K.; Martin, C. A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 2006, 18, 831–851. [Google Scholar] [CrossRef] [PubMed]
- Espley, R.V.; Hellens, R.P.; Putterill, J.; Stevenson, D.E.; Kutty-Amma, S.; Allan, A.C. Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J. 2007, 49, 414–427. [Google Scholar] [CrossRef] [PubMed]
- Quattrocchio, F.; Wing, J.F.; van der Woude, K.; Mol, J.N.; Koes, R. Analysis of bHLH and MYB domain proteins: Species-specific regulatory differences are caused by divergent evolution of target anthocyanin genes. Plant J. 1998, 13, 475–488. [Google Scholar] [CrossRef] [PubMed]
- Quattrocchio, F.; Wing, J.; van der Woude, K.; Souer, E.; de Vetten, N.; Mol, J.; Koes, R. Molecular analysis of the anthocyanin2 gene of petunia and its role in the evolution of flower color. Plant Cell 1999, 11, 1433–1444. [Google Scholar] [CrossRef] [PubMed]
- Lin-Wang, K.; Bolitho, K.; Grafton, K.; Kortstee, A.; Karunairetnam, S.; McGhie, T.K.; Espley, R.V.; Hellens, R.P.; Allan, A.C. An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biol. 2010, 10, 50. [Google Scholar] [CrossRef]
- Nakatsuka, T.; Saito, M.; Yamada, E.; Fujita, K.; Kakizaki, Y.; Nishihara, M. Isolation and characterization of GtMYBP3 and GtMYBP4, orthologues of R2R3-MYB transcription factors that regulate early flavonoid biosynthesis, in gentian flowers. J. Exp. Bot. 2012, 63, 6505–6517. [Google Scholar] [CrossRef]
- Czemmel, S.; Stracke, R.; Weisshaar, B.; Cordon, N.; Harris, N.N.; Walker, A.R.; Robinson, S.P.; Bogs, J. The grapevine R2R3-MYB transcription factor VvMYBF1 regulates flavonol synthesis in developing grape berries. Plant Physiol. 2009, 151, 1513–1530. [Google Scholar] [CrossRef]
- Yun, C.S.; Yamamoto, T.; Nozawa, A.; Tozawa, Y. Expression of parsley flavone synthase I establishes the flavone biosynthetic pathway in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 2008, 72, 968–973. [Google Scholar] [CrossRef]
- Qi, X.; Yu, X.; Xu, D.; Fang, H.; Dong, K.; Li, W.; Liang, C. Identification and analysis of CYP450 genes from transcriptome of Lonicera japonica and expression analysis of chlorogenic acid biosynthesis related CYP450s. Peer J. 2017, 5, e3781. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Zhang, H.; Gao, S.; Lercher, M.J.; Chen, W.H.; Hu, S. Evolview v2: An online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 2016, 44, W236–W241. [Google Scholar] [CrossRef] [PubMed]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 2006, 1, 2019–2025. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Liu, G.; Tang, N.; Li, Z. Transcriptome analysis reveals molecular signatures of luteoloside accumulation in senescing leaves of Lonicera macranthoides. Int. J. Mol. Sci. 2018, 19, 1012. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
| Genes | Forward Primer (5′–3′) | Reverse Primer (5′–3′) |
|---|---|---|
| Gene Cloning | ||
| LjaMYB12 | ATGGGGAGAGCACCATGCT | CTAAGAGAGCAGCCATGCAAC |
| Subcellular Localization Assay | ||
| LjaMYB12 | GGGGTACCATGGGGAGAGCACCATGCT | CGGGATCCAGAGAGCAGCCATGCAACC |
| EGFP | CGGGATCCATGGTGAGCAAGGGCGAG | ACGCGTCGACTTACTTGTACAGCTCGTCATGC |
| Transactivation Analysis | ||
| LjaMYB12 | GGAATTCCATATGATGGTGAGCAAGGGCGAG | ACGCGTCGACCTAAGAGAGCAGCCATGCAAC |
| Arabidopsis Transformation | ||
| LjaMYB12 | CGGGATCCATGGGGAGAGCACCATGCT | ACGCGTCGACCTAAGAGAGCAGCCATGCAAC |
| qRT-PCR | ||
| LjaMYB12 | CTTTGAAGGGAATGATGAGG | TTGCTTCTCGCTGTCTCC |
| LjActin | TGCTGGATTCTGGTGATGGT | ATTTCCCGCTCTGCTGTG |
| AtPAL | ATCGAAGTGATCCGTTACGC | ACTCCGATTGGTGTTCCTTG |
| At4CL | CGCAAACCCTTTCTTCACTC | ACTCCGTCGTCGTTTTGAAG |
| AtCHS | TGAGAACCATGTGCTTCAGG | CAGATGCATGTGACGTTTCC |
| AtCHI | TTTGTACCGTCCGTCAAGTC | CAATGACGGTGAAGATCACG |
| AtF3H | TCAGATCGTTGAGGCTTGTG | ATGTCGAAACGGAGCTTGTC |
| AtF3′H | CCTCCACCTCCGACTAGGGT | TGCTCGGCCACGGATTTA |
| AtFLS | CAACATTCCGAGGTCCAACG | TCTTCGTCGGGATCGCTTAG |
| AtDFR | GTCGGTCCATTCATCACAAC | TGAGCGTTGCATAAGTCGTC |
| AtANS | TCAAGAAAGCCGGAGAAGAG | TTGTCCACTCGCGTTGTTAG |
| AtACT2 | ACCCGATGGGCAAGTCATC | CGAGGGCTGGAACAAGACTTC |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qi, X.; Fang, H.; Chen, Z.; Liu, Z.; Yu, X.; Liang, C. Ectopic Expression of a R2R3-MYB Transcription Factor Gene LjaMYB12 from Lonicera japonica Increases Flavonoid Accumulation in Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 4494. https://doi.org/10.3390/ijms20184494
Qi X, Fang H, Chen Z, Liu Z, Yu X, Liang C. Ectopic Expression of a R2R3-MYB Transcription Factor Gene LjaMYB12 from Lonicera japonica Increases Flavonoid Accumulation in Arabidopsis thaliana. International Journal of Molecular Sciences. 2019; 20(18):4494. https://doi.org/10.3390/ijms20184494
Chicago/Turabian StyleQi, Xiwu, Hailing Fang, Zequn Chen, Zhiqi Liu, Xu Yu, and Chengyuan Liang. 2019. "Ectopic Expression of a R2R3-MYB Transcription Factor Gene LjaMYB12 from Lonicera japonica Increases Flavonoid Accumulation in Arabidopsis thaliana" International Journal of Molecular Sciences 20, no. 18: 4494. https://doi.org/10.3390/ijms20184494
APA StyleQi, X., Fang, H., Chen, Z., Liu, Z., Yu, X., & Liang, C. (2019). Ectopic Expression of a R2R3-MYB Transcription Factor Gene LjaMYB12 from Lonicera japonica Increases Flavonoid Accumulation in Arabidopsis thaliana. International Journal of Molecular Sciences, 20(18), 4494. https://doi.org/10.3390/ijms20184494





